Abstract
Although apoptosis can be readily assessed in vitro with a variety of techniques, the detection of apoptosis in the in vivo setting poses a much more difficult proposition. Apoptosis in an organism is followed almost inevitably by rapid clearance of dying cells via phagocytosis, thus limiting the ability to analyze apoptosis in vivo using classical techniques. To address this issue, we developed a method to enhance in vivo apoptosis detection using pretreatment with chloroquine, an inhibitor of macrophage activity, in Swiss albino mice. This technique resulted in a significant increase in the accumulation of apoptotic cells induced by 5-fluorouracil, as detected by propidium iodide staining in solid and ascitic forms of Ehrlich ascitic tumors and in bone marrow cells. We further validated our technique using DNA fragmentation and endonuclease assays. Our results demonstrated that chloroquine pretreatment can significantly enhance accumulation of apoptotic cells in organisms, and we envision combining this method with modern imaging techniques to optimize in vivo detection of apoptosis.
Introduction
In recent years, efforts in cancer research have focused on understanding the molecular mechanisms that regulate apoptosis and tumor cell proliferation [1,2]. Classically, apoptotic cell death is defined as programmed cell death resulting in certain morphological and biochemical characteristics that can be distinguished from other forms of cell death [3]. The detection of apoptosis in patient biopsies can be used as an important tool to assess the prognosis of a disease or response to a therapy [2,4]. Apoptosis can be readily studied in vitro by various techniques, including those based on morphology (via chromatin condensation or DNA fragmentation), cysteine protease activity (as assessed by poly[ADP-ribose] polymerase [PARP] or lamin cleavage), flow cytometry (as measured by annexin V binding or the sub-G1 peak), or DNA fragmentation (as assayed by electrophoresis and terminal deoxyribonucleotidyl transferase [TdT]-mediated dUTP nick end labeling [TUNEL]) [5–8]. However, the study and detection of apoptosis in an organism is much more difficult because apoptotic cells are rapidly engulfed by macrophages [9–13]. Although this rapid clearance of apoptotic cells may help suppress inflammation and modulate the immune response [14], it severely reduces the number of apoptotic cells that are available for analysis at any given time, thus limiting the study of apoptosis in vivo.
We hypothesized that inhibition of phagocytosis would result in enrichment of the apoptotic cells and enhance our ability to detect apoptosis in vivo. Such a technique would be a very useful tool for future studies investigating the efficacy of various clinical therapies. For this purpose, we used chloroquine, a common antimalarial drug that selectively accumulates in lysosomes and blocks lysosome function completely [15,16]. Chloroquine is a weak base that alkalinizes lysosomes and thereby inhibits enzyme release [17–19]. In addition, chloroquine has been demonstrated to cause time- and dose-dependent inhibition of phagocytosis in a variety of cells, such as rat peritoneal macrophage and human monocytes and neutrophils [15–20], which are primarily responsible for clearance of apoptotic cells. In this report we show that chloroquine pretreatment enhanced our capacity to detect 5-fluorouracil (5-FU, a thymidylate synthase inhibitor)-induced apoptosis [21] in Ehrlich ascitic carcinoma (EAC) solid and ascitic tumors and in bone marrow cells.
Materials and Methods
Reagents
Ehrlich ascitic carcinoma cells were obtained from the Cancer Research Institute (Mumbai, India) and were maintained by serial propagation in Swiss albino mice. Chloroquine and 5-FU were purchased from Sigma (St. Louis, MO).
Animals and Tumor Propagation
Male Swiss albino mice (8-12 weeks old), maintained in an institutional breeding colony (Indian Institute of Science, Bangalore, India), were used for all the experiments described. Animals were kept under a 12-hr light/12-hr dark regimen and were given food and water ad libitum. One million ascitic EAC cells were transplanted via subcutaneous and intraperitoneal injection to obtain solid and ascitic tumors, respectively.
Experimental Design
Animals were treated with chloroquine (20 mg/kg body weight) every 8 hr for 3 days. At the end of this period, they received a single intraperitoneal injection of 5-FU (200 mg/kg body weight). Sixteen hours after the 5-FU injection, mice were then sacrificed by cervical dislocation. Additional groups of animals received only chloroquine, 5-FU, or no treatment for comparison with the group treated with both chloroquine and 5-FU.
At the time of sacrifice, ascitic tumor was aspirated by using a 20-gauge needle, and the solid tumors were removed surgically. To obtain single-cell suspensions from solid tumors, a small piece of tumor was minced into pieces and incubated with trypsin-EDTA (0.25%, 53 mM) in PBS for 1 hr at 37°C. Bone marrow cells were collected from all the animals by surgically removing the femur and tibia, which were then flushed with ice cold sterile PBS using a 26-gauge needle-fitted syringe. Samples were then processed for propidium iodide (PI) staining, DNA fragmentation assays, and endonuclease assays, as described below.
Propidium Iodide Staining
Cells were rewashed in PBS, fixed in 70% ethanol, stained with 1% PI with RNase, and viewed under a fluorescent microscope. For each sample, triplicates of 200 cells were counted, and the number of apoptotic cells were scored.
DNA Fragmentation Assay
After treatment, the cells were washed with PBS, suspended in cytoplasm extraction buffer (10 mM Tris, 150 mM NaCl, 5 mM MgCl2, and 0.5% Triton X), incubated on ice for 30 min, and then centrifuged at 14,000 rpm at 4°C. The resultant pellet was discarded, and the supernatant was incubated with RNase (0.2 mg/mL) for 30 min at 55°C and then with Proteinase K (0.1 mg/mL) for 2 hr at 37°C. DNA was extracted by treatment of the supernatant with phenol/chloroform (1:1) and then precipitated with isopropanol. Immediately after addition of isopropanol, the high molecular weight DNA was removed with the help of a sterile glass capillary to enrich the fragmented DNA. Samples were then incubated overnight at −80°C. DNA was then precipitated by centrifugation at 14,000 rpm at 4°C for 15 min. The DNA pellet was air-dried and resuspended in 40 μL of TE buffer (10 mM Tris-HCl, pH 8.0, 1 mM EDTA). Total DNA was resolved on a 1.8% agarose gel, containing 0.3 μg of ethidium bromide in Tris-borate EDTA buffer. DNA was visualized under an ultraviolet (UV) transilluminator.
Endonuclease Assay
About 107 cells were disrupted in 1 mL of STEM-1 buffer (200 mM NaCl, 10 mM Tris-HCl [pH 7.4], 10 mM MgCl2, 2.5 mM EGTA, 0.5 mM DTT, and 0.2% Triton X-100). Samples were then incubated at 4°C for 15 min. Nuclei were collected after centrifugation at 200 rpm for 5 min at 4°C, resuspended, and incubated for 3 hr at 4°C in STEM-2 buffer (200 mM NaCl, 10 mM Tris-HCl [pH 7.4], 10 mM MgCl2, 2.5 mM EGTA, and 0.5 mM DTT). The supernatant containing nuclear endonuclease was collected after centrifugation at 13,000 rpm for 15 min. Protein concentration was determined by Lowry's method [22]. Endonuclease activity was assayed by using supercoiled plasmid bluescript DNA as a substrate in the presence of 2 mM CaCl2 and MgCl2. The reaction products were separated by electrophoresis on a 0.8% agarose gel.
Statistics
Results are presented as the mean ± SE of at least three experiments. The Student's t test was used to assess the statistical significance of difference. A significant level threshold of p ≤ .05 was used in this study.
Results
Baseline Detection of Apoptosis in EAC Tumors and Bone Marrow Tissue After 5-FU Treatment
To determine the baseline for apoptosis detection, we assessed DNA fragmentation using PI staining, gel electrophoresis, and an endonuclease assay in ascitic tumors, solid tumors, and bone marrow cells from mice that received either no treatment or 5-FU. Control samples showed that only 1.3–2.9% of cells were undergoing apoptosis (Table 1). After a single injection of 5-FU, PI staining revealed a nonsignificant increase in apoptotic cells when compared to controls. Representative microscopy images from each sample are demonstrated in Figure 1, showing few apoptotic cells (as demarcated by arrows) in either control samples or those treated with 5-FU alone. This was also confirmed by DNA fragmentation patterns upon agarose gel electrophoresis. No DNA fragmentation was visualized in any control samples, but a very faint DNA ladder was detectable in samples treated with 5-FU alone (Figure 2A). Assays of endonuclease activity also demonstrated comparable results. No endonuclease activity was demonstrated in control samples and in 5-FU-treated bone marrow, but slight endonuclease activity was revealed in 5-FU-treated tumor samples (Figure 2B).
Treatment Effects on the Percentage of Apoptotic Cells by PI Staining
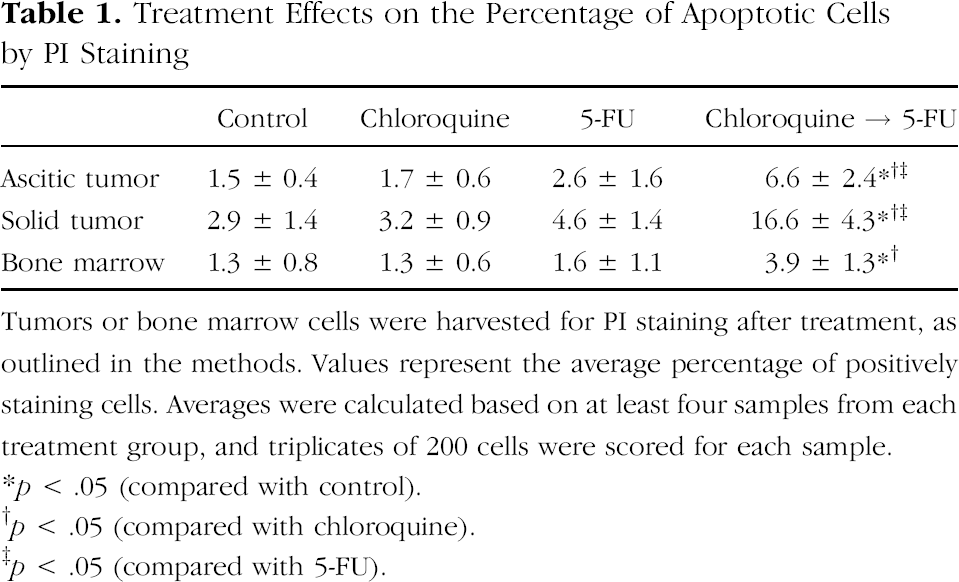
Tumors or bone marrow cells were harvested for PI staining after treatment, as outlined in the methods. Values represent the average percentage of positively staining cells. Averages were calculated based on at least four samples from each treatment group, and triplicates of 200 cells were scored for each sample.
p < .05 (compared with control).
p < .05 (compared with chloroquine).
p < .05 (compared with 5-FU).
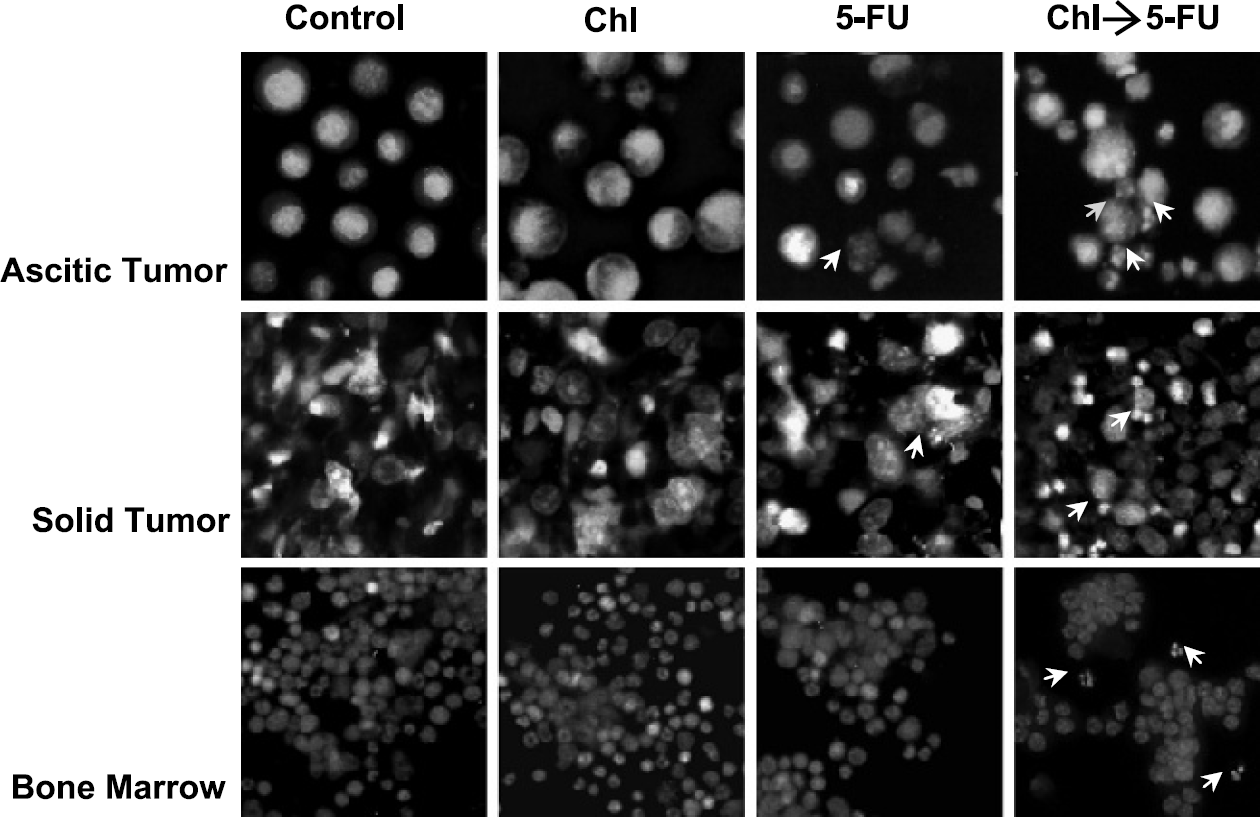
Representative microscopic fields demonstrating apoptotic cells, as detected by PI staining. Animals received either no treatment, only chloroquine (Chl), only 5-FU, or a combination of both, as described in the materials and methods section. Samples were stained with PI and viewed under a fluorescent microscope. Multiple fields were assessed within each sample, and a representative image is shown above. Apoptotic cells, as recognized by chromatin condensation and DNA fragmentation, are depicted with small white arrows. Few apoptotic cells were noted in control samples or those treated with chloroquine or 5-FU alone. However, within each field, several apoptotic cells were noted in samples that received chloroquine pretreatment followed by 5-FU.
Enrichment of Apoptosis Detection Using Chloroquine Pretreatment
Given the low levels of 5-FU-induced apoptosis visualized by fluorescent staining, DNA electrophoresis, and endonuclease assays, we suspected that our detection of apoptosis was being limited by phagocytosis of apoptotic cells by macrophages. To address this issue, we pretreated mice with chloroquine before administering 5-FU, as described in materials and methods. Chloroquine treatment itself did not result in any increase in apoptosis compared to control in any of our assays (Table 1, Figure 1, Figure 2A and B). However, the administration of chloroquine before 5-FU resulted in substantial increases in apoptosis detection by PI staining compared to groups receiving chloroquine alone (Figure 1). In the presence of chloroquine, 5-FU resulted in 6.6% apoptosis in ascitic tumors (compared to 1.7% with chloroquine alone, p = .02), 16.6% apoptosis in solid tumors (compared to 3.2%, p < .01), and 3.9% apoptosis in bone marrow (compared to 1.3%, p = .07) (Table 1). Compared to 5-FU treatment alone, the addition of chloroquine also improved the detection of 5-FU-induced apoptosis in ascitic EAC tumors (from 2.6% to 6.6%, p = .01), solid EAC tumors (4.6% to 16.6%, p < .01), and bone marrow cells (1.6% to 3.9%, p = .05) (Table 1). Additionally, compared to groups treated with chloroquine alone, there was markedly augmented DNA fragmentation in groups receiving both chloroquine and 5-FU, as revealed on gel electrophoresis (Figure 2A). Endonuclease activity was also substantially amplified in groups treated with chloroquine and 5-FU versus those receiving chloroquine alone (Figure 2B), as demonstrated by enlargement of the band of low molecular weight DNA in tumor samples and the appearance of a smear of lower molecular weight DNA in bone marrow samples. Taken together, these results demonstrate that although chloroquine alone does not cause an increase in apoptosis, chloroquine pretreatment will significantly improve the detection of apoptosis triggered by another agent, such as 5-FU.
Discussion
In this report, we have introduced a novel technique for improving the detection of apoptosis in vivo using chloroquine pretreatment. Given the increasing evidence that dysregulation of apoptosis plays a critical role in oncogenesis [1,2,4,23], in certain autoimmune diseases [24], and in various degenerative diseases [25], the detection of apoptosis in these pathologic settings is crucial. While novel advances are being made within the arena of noninvasive apoptotic imaging [26,27], the evaluation of therapeutic agent efficacy using these techniques will fundamentally be limited by phagocytosis of apoptotic cells. We believe that the use of chloroquine pretreatment will reduce this limitation, and we envision combining our method with modern imaging techniques to optimize the in vivo detection of apoptosis.
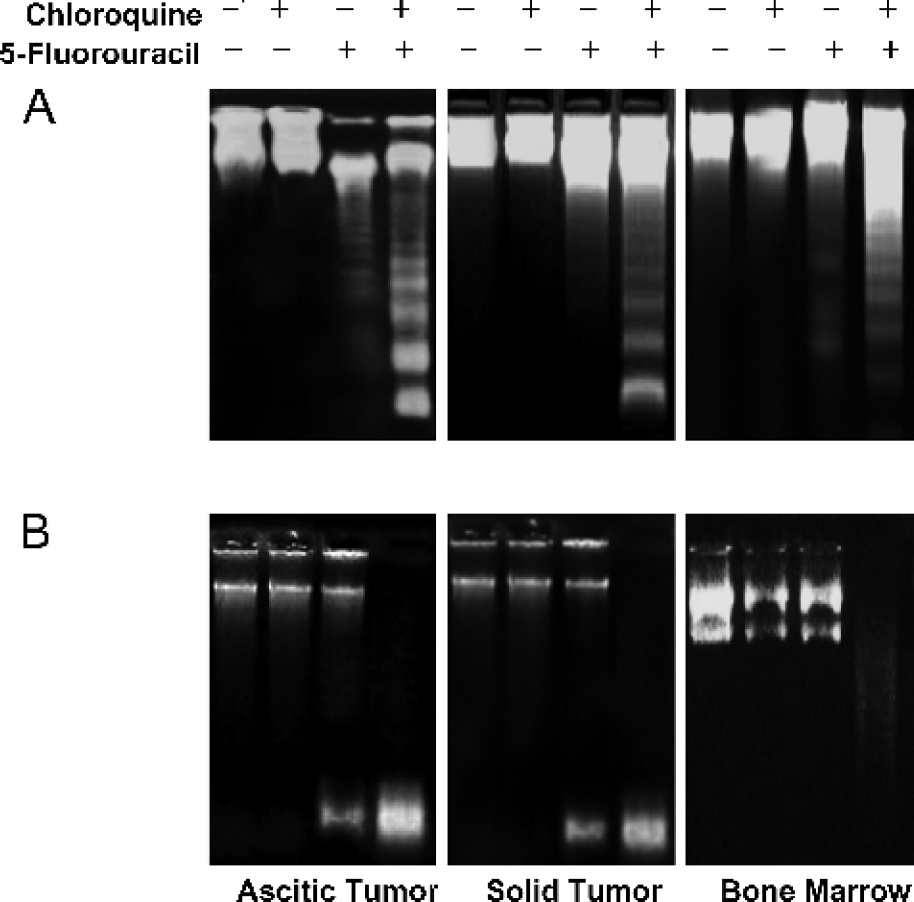
Effects of treatment on apoptosis detection using DNA fragmentation and endonuclease assays. (A) DNA samples from each treatment group were resolved on a 1.8% agarose gel and visualized under a UV transilluminator. The presence of apoptosis was detected by the fragmentation of DNA from a sample, as depicted by the appearance of a “DNA ladder”. Intense DNA ladders resulted from treatment with chloroquine and 5-FU, and faint DNA ladders were viewed in response to 5-FU alone. (B) Endonuclease activity from each treatment group was assessed, as described in materials and methods. Endonuclease activity is reflected by the appearance of low molecular weight DNA resulting from endonuclease cleavage of a plasmid substrate. Treatment with chloroquine and 5-FU amplified endonuclease assay substantially in tumor samples and to a lesser degree in bone marrow samples.
Footnotes
Acknowledgments
We thank P.V. Balasubramanyam for technical help, the animal facility at the Indian Institute of Science for mice, and the Department of Biotechnology (Government of India) for providing a postdoctoral fellowship to M.K.N.
