Abstract
We report a case of a de novo complex chromosomal rearrangement among five chromosomes found in a clinically healthy woman. The only indication for chromosome analysis was a planned intracytoplasmatic sperm injection. Physical examination, including internal and external genitals, and ovaries and hormone status were normal. Banding cyto-genetics showed a rearrangement among chromosomes #3, #4, #7, #9, and #17. Twenty-four-color fluorescence in situ hybridization and multicolor banding were applied to characterize the translocations and breakpoints more precisely. This confirmed the involved chromosomes and revealed two breakpoints in chromosome #4. This six-breakpoint rearrangement [der(3)t(3;4), der(4)t(17;4;7), der(7)t(3;7), der(9)t(4;9), and der(17)t(9;17)] seemed to be balanced on a molecular cytogenetic level, although submicroscopic deletions or duplications close to the breakpoints cannot be excluded.
Keywords
C
We report here on a clinically healthy, 34-year-old woman. She was referred for genetic diagnosis because of unwanted childlessness. Moreover, she had a history of two early abortions. Gynecological examination, including external and internal genitals and ovaries, and hormone status, were normal. Her partner was diagnosed with oligoasthenoteratozoospermia I. Therefore, use of an artificial reproductive technology was planned. In preparation for intracytoplasmatic sperm injection (ICSI), chromosome analysis was performed on both partners. The male had a normal karyotype [46,XY], but the GTG-banding results of the female showed a hCCR involving five chromosomes (chromosomes #3, #4, #7, #9, and #17, see Figure 1). These aberration occurred de novo as karyotype analyses of her parents showed no cytogenetic abnormalities.
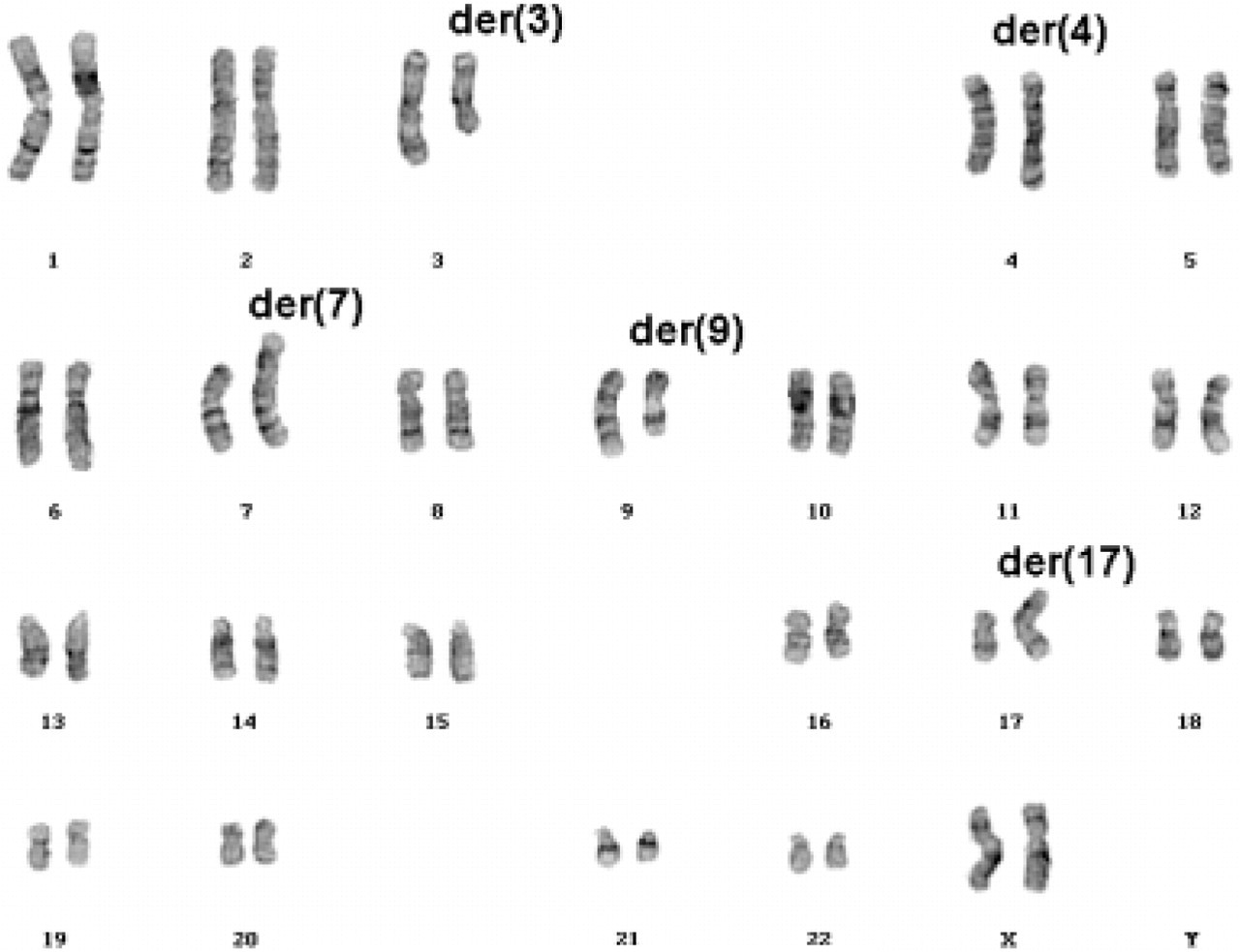
Cytogenetic banding analysis of the patient's karyotype showing five aberrant chromosomes (GTG banding).
Further characterization of this rearrangement was performed using molecular cytogenetic techniques. Twenty-four-color fluorescence in situ hybridization (FISH) according to Senger et al. (1998) indicated the following aberrations: del(3), der(4)t(17;4;7), der(7)t(3;7), der(9)t(4;9), and der(17)t(9;17) (see Figure 2). To substantiate from which chromosomal arm the translocated material was derived, arm-specific probes were also used. This revealed that the derivative chromosome 3 had no simple deletion but contained a subtle translocation of material of chromosome 4q. In the next step, multicolor banding (MCB) (Liehr et al. 2002) was applied to characterize the breakpoints more precisely and to define the orientation of the involved chromosomal material.
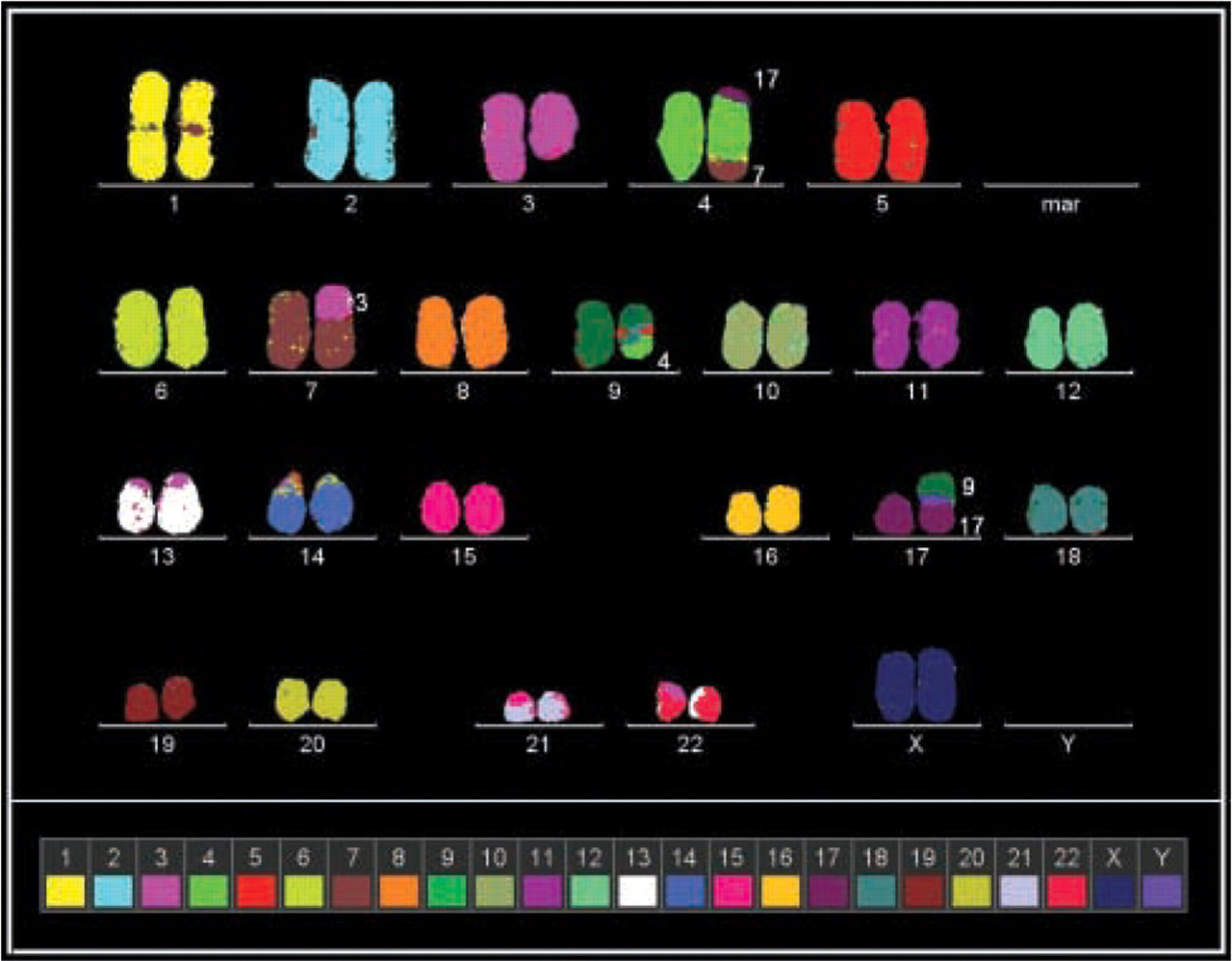
The 24-color FISH result (pseudocolor representations) of this highly complex karyotype showed the following aberrations: del(3), der(4)t(17;4;7), der(7)t(3;7), der(9)t(4;9), and der(17)t(9;17). Images were captured with the ISIS3 digital FISH imaging system (MetaSystems; Altlussheim, Germany) using a PCO VC45 CCD camera (PCO; Kehl, Germany) on an Axioplan 2 microscope (Zeiss; Jena, Germany) equipped with suitable filter combinations (DAPI/FITC/SpectrumOrange/TexasRed/Cyanine 5/Cyanine 5.5).
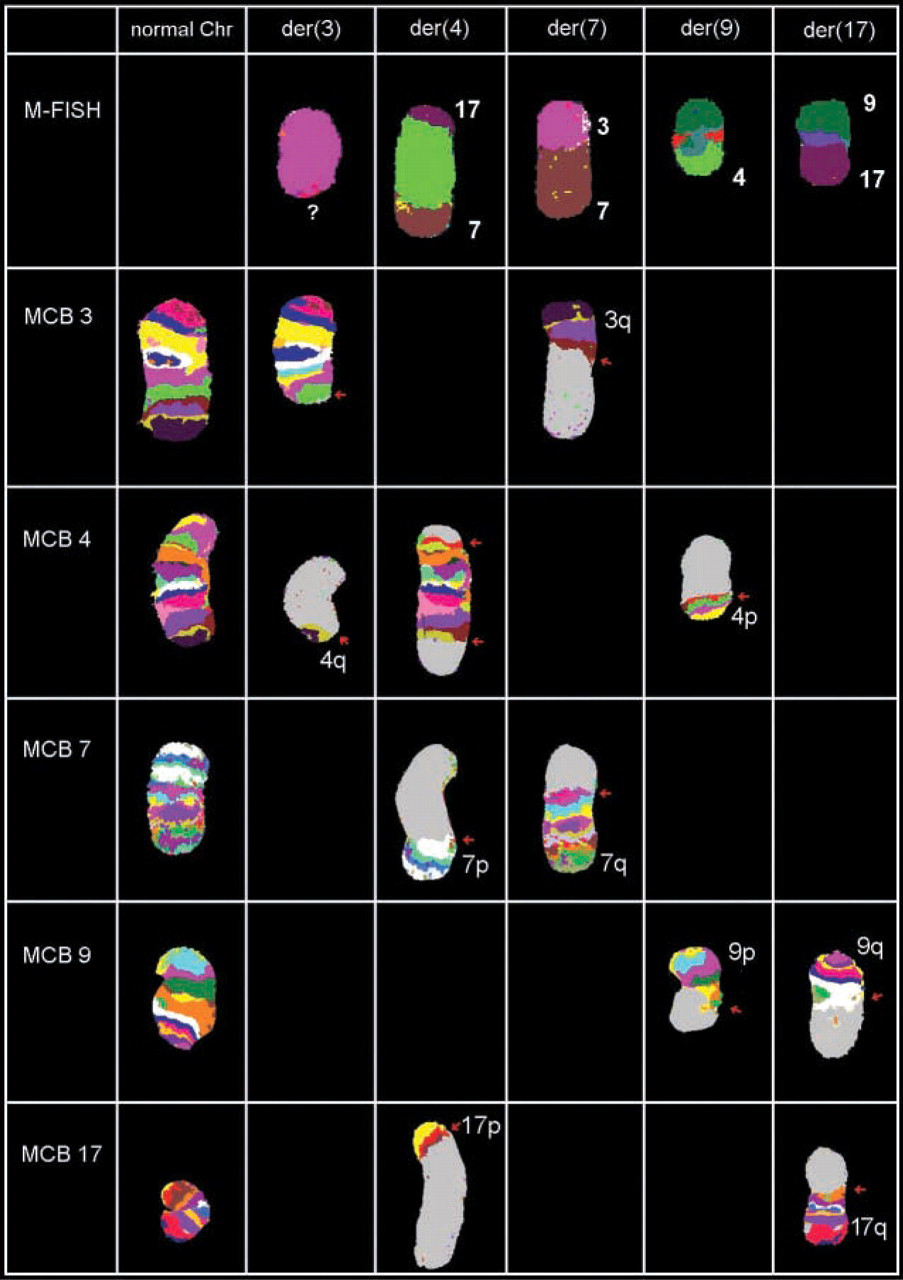
Summary of MCB results of the five derivative chromosomes. The hCCR consisted of a total of six breaks (breakpoints marked with little red arrows) and seemed to be balanced on a molecular cytogenetic level. Use of arm-specific probes revealed that the derivative chromosome 3 had no simple deletion but contained a subtle translocation of material of chromosome 4, which could be confirmed by pcp4q (not shown) and MCB 4. For patient's karyotype, see text.
By this comprehensive analysis, the karyotype could be established as follows:
ish 46,XX,der(3)t(3;4)(3pter→3q22::4q34~35→4qter), der(4)t(17;4;7)(17pter→17p13~12::4p14→4q34~35::7p13~12→7pter), der(7)t(3;7)(3qter→3q22::7p13~12→7qter), der(9)t(4;9)(4pter→4p14::9q13→9pter), der(17)t(9;17)(9qter→9q13::17p13~12→17qter).
The rearrangement contained in summary six breakpoints derived from five chromosomes and seemed to be balanced on a molecular cytogenetic level. The hybridization results of 24-color FISH and MCB for all five aberrant chromosomes are depicted in Figure 3.
Despite the knowledge of this hCCR result, one course of ICSI was performed but without success.
As mentioned above, there are few cases of reported hCCR involving five or more chromosomes. The number of descriptions increased with the introduction of new techniques (Astbury et al. 2004), which also improved detection of intrachromosomal rearrangements (Weise et al. 2003), but they remain rare findings. In our case, we detected in summary six breakpoints, all not located in known heterochromatic regions (such as 1q12 or centromeric regions). Thus, we speculate that all six breaks did not affect important genes or, more precisely, gene functions; however, this suspicion cannot be proven with the applied molecular cytogenetics approaches. Additionally, submicroscopic deletions or duplications close to the breakpoints could be excluded only by sequencing of the breakpoints.
The infertility of the couple, especially the history of two abortions, can easily be explained by the complexity of the hCCR itself and the extremely low probability of a cytogenetically balanced or normal pregnancy (Madan et al. 1997; Siffroi et al. 1997). In such cases, the ICSI method also reaches its limits (Siffroi et al. 1997), and careful genetic counseling of affected couples is required.
The lesson learned from this case is that, in preparation for ICSI, chromosome analysis should always be performed on both partners.
