Abstract
We investigated the frequencies of abnormalities involving either chromosome 1, 16, 18, or 21 in failed-fertilized human oocytes. Although abnormalities involving chromosome 16 showed an age-dependent increase, results for the other chromosomes did not show statistically significant differences among the three age groups, <35 years, 35–39 years, and >39 years. The scoring of four chromosomes is likely to underestimate the true rate of aneuploid cells. Therefore, for a pilot study investigating a more-comprehensive analysis of oocytes and their corresponding first polar bodies, we developed a novel eight-probe chromosome enumeration scheme using fluorescence in situ hybridization and spectral imaging analysis.
C
Using fluorescence in situ hybridization (FISH), we investigated the frequency of aneuploidy and chromatid predivision for chromosomes 1, 16, 18, and 21 in 273 failed-fertilized oocytes from 95 patients, stratified by age (<35 years, 35–39 years, and >39 years; age range 26.1 to 42.2 years). Oocytes were prepared as previously described (Racowsky et al. 1992).
Our probes were labeled with four different fluorochromes so that they could be identified and scored easily using the fluorescence microscope (Figures 1A–1D). The DNA probes for chromosomes 1, 18, and 21 were labeled by random priming using a BioPrime kit (Invitrogen; Gaithersburg, MD) to incorporate digoxigenin-11-dUTP (Roche Molecular Biochemicals; Indianapolis, IN), fluorescein-12-dUTP (FITC; Roche Molecular Biochemicals), or Spectrum Orange-dUTP (Vysis; Downers Grove, IL). A cloned 1.77-kb EcoRI fragment of human satellite III DNA specific for chromosome 1 (pUC1.77) (Fung et al. 2001) was labeled with Spectrum Orange. The probe specific for satellite II DNA of chromosome 16 was prepared from clone pHUR195 (Fung et al. 2001) and labeled with FITC as well as digoxigenin. The single-copy DNA probe specific for chromosome 21 was selected from the Genethon/CEPH yeast artificial chromosome (YAC) library (YAC 141G6) and labeled with FITC as previously described (Fung et al. 2001). The DNA probe specific for chromosome 18 (CEP18), labeled with Spectrum Aqua, was obtained from the manufacturer (Vysis).
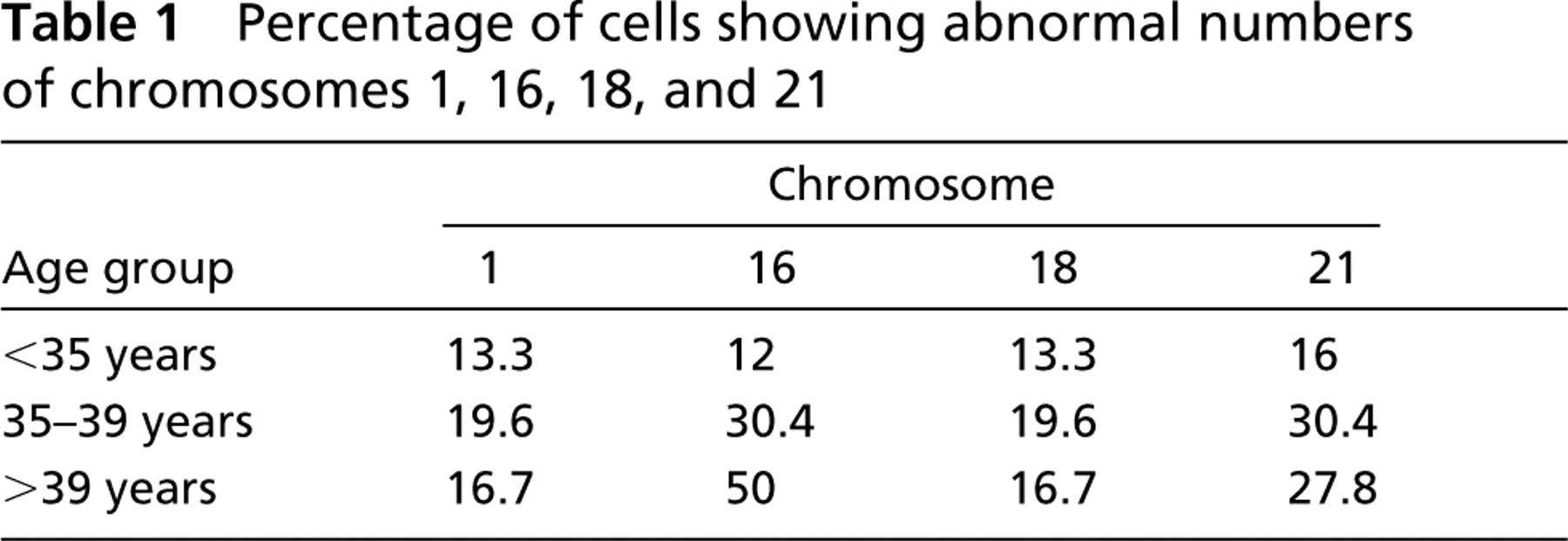
The results demonstrated an overall high rate of aneuploidy among failed-fertilized eggs, in agreement with reports from other laboratories (Martini et al. 1997; Pellestor et al. 2003). However, published reports had suggested a disproportionally high rate of aneuploid cells with abnormalities related to the number of copies of chromosome 1 (Martini et al. 1997; Bahçe et al. 1999). In our study, the rate of cells showing aneusomy involving chromosome 1 was 15.8%, and it was neither age dependent nor significantly different from that for chromosomes 16 (22.5%), 18 (16.3%), and 21 (22.7%) (t-test, p=0.241). Table 1 summarizes the chromosome-specific rates of aneuploid cells for each of the three age groups. Only chromosome 16 exhibited a significant age-dependent increase in aneuploidy (12.0%, 30.4%, and 50.0% for <35, 35–39, and >39 years, respectively; p=0.0009) (Table 1).
Because chromatid predivisions are considered to be a major mechanism leading to aneuploid oocytes (Pellestor et al. 2003), they were also recorded in our study of failed-fertilized eggs. The rate of chromatid predivisions for chromosome 1 was lower than for chromosome 18 (11.9% vs 25.4%; p=0.01) but was not significantly different from that for chromosomes 16 (16.4%) or 21 (19.1%). Moreover, in contrast to previous reports, we found that the rate of chromatid predivision involving chromosome 1 did not show any indication of age dependence. Because the size of our study cohort is in the broad range of the populations reported in the above-cited studies, it is unlikely that the observed discrepancies stem from problems related to cohort size. Other factors, such as ethnicity or life style (e.g., diet, caffeine consumption, exercise, smoking, drug use), should be taken into consideration.
Percentage of cells showing abnormal numbers of chromosomes 1, 16, 18, and 21
It is also likely that our FISH assay scoring only 4 of the 23 chromosome types found in human oocytes underestimates the true incidence of aneuploidy. One approach to determine the number and type of chromosomes in oocytes is based on the delineation of all chromosomes with chromosome-specific whole chromosome painting (WCP) probes. The chromosome-specific WCP probes were prepared by combining probes labeled with one or more of five reporter molecules so that each chromosome-specific WCP probe had a unique, identifiable spectral signature. The technology, termed “spectral karyotyping (SKY),” was developed initially to screen metaphase spreads and is used primarily in karyotype analysis of constitutional chromosome abnormalities and in cancer studies. We have applied the SKY technology in the past to demonstrate specific alterations in human and mouse tumors (Fung et al. 2000). In their elegant study, Sandalinas et al. (2002) demonstrated the application of SKY to the analysis of oocytes (Figures 1E–1G). However, because of overlap between hybridization domains, this combinatorial labeling and detection scheme works only with very well spread metaphase chromosomes. For chromosome enumeration in oocytes and polar bodies, as well as in interphase cell nuclei of other origins, the use of chromosome-specific DNA repeat or single-copy probes appears to be more accurate.
At present, no commercial test is available that scores more than five chromosomes in a single hybridization experiment. Consequently, investigators have applied repeated hybridization schemes to increase the number of chromosomes scored (Martini et al. 1997). We developed a test system based on nine-color FISH [including the 4',6-diamidino-2-phenylindole (DAPI) counterstain] and spectral imaging (SIm) analysis (Figures 1H–1J). We prepared uniquely labeled DNA probes for simultaneous scoring of chromosomes 13, 15, 16, 18, 21, 22, X, and Y. Next, we optimized cell pretreatment and FISH conditions to ensure that each DNA probe reached a hybridization efficiency of at least 90%, a margin acceptable for most IVF programs. Probes were tested initially on lymphocytes (Figures 1I and 1J) before being used in the analysis of oocytes and their corresponding first polar bodies (1PBs) (Figures 1K and 1L).
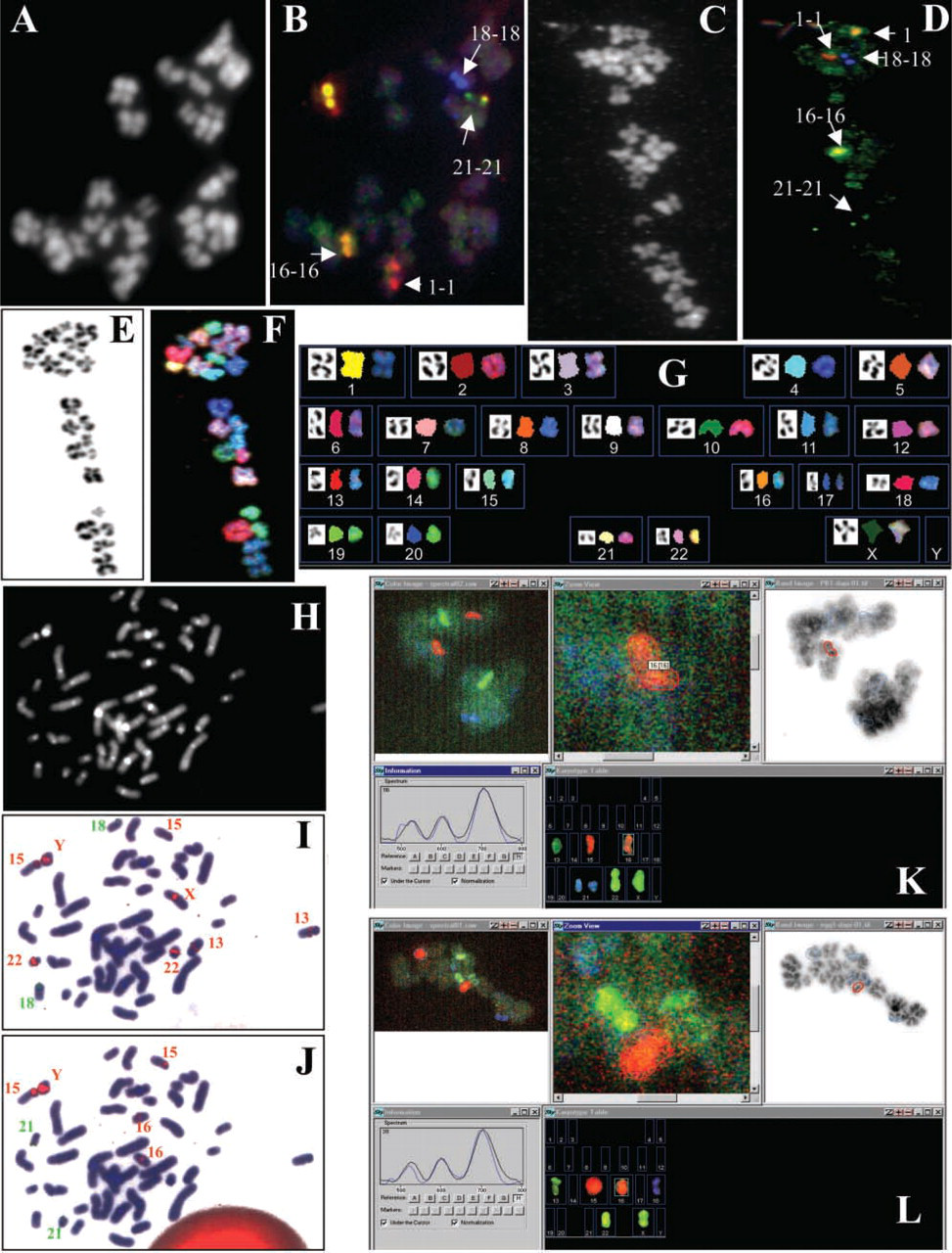
In situ hybridization results. FISH analysis of human failed-fertilized metaphase II eggs using specific probes for chromosomes 1 (red signals), 16 (yellow signals), 18 (blue signals), and 21 (green signals). Hybridization of the four-probe set to a spread prepared from a normal oocyte
We used an SIm-based approach to detect bound probes after FISH. SIm combines fluorescence spectroscopy and digital imaging for the analysis of FISH signals. A typical system is composed of a fluorescence microscope equipped with an interferometer and a charge-coupled device camera plus powerful computer software to perform rapid Fourier spectroscopy. Such systems are commercially available from Applied Spectral Imaging (Carlsbad, CA).
Immature, never-inseminated, frozen oocytes were matured in vitro and fixed. Oocytes and 1PBs were fixed separately so that they could be analyzed independently. The rate of in vitro maturation of the oocytes was low. Less than 50% (42/85) of immature eggs extruded a 1PB. In total, 85 eggs were needed to obtain 29 polar bodies and their respective oocytes. These specimens were analyzed with the eight-probe set. Of those 29 polar bodies, four did not have a corresponding oocyte properly fixed or analyzable. In total, there were 25 analyzable pairs of 1PBs and oocytes.
The screen dumps in Figures 1K and 1L show the typical steps during the analysis. Counterclockwise, shown from the top left, are the pseudo-RGB display of the spectral image, automated selection of the region of interest (here a close-up of a chromosome 16–specific signal), the overlay of regions of interest with the inverted DAPI image, and the karyotype table. The small image at lower left shows a comparison of prerecorded “pure dye” spectra (black line) with the fluorescence spectrum under the cursor (blue line), which is used to check the accuracy of the automated signal classification.
The concordance between the eggs and 1PBs was found to be 92%, thus proving that the probes, the slide pretreatment, and the detection protocol were ready for PGD applications. One example is shown in Figures 1K and 1L, where the 1PB (Figure 1K) contained two chromosomes 21 (i.e., disomy 21) whereas the corresponding oocyte carried no chromosome 21 (i.e., nullisomy 21). These results are very reliable, because the single-copy probe for chromosome 21 was the only green fluorescent probe in our set.
Footnotes
Acknowledgements
Supported in part by US National Institutes of Health Grant 1-R44-HD-44313 and by Brigham and Women's Hospital Ob/Gyn IVF Discretionary Funds (CR).
We gratefully acknowledge the assistance of the embryology team in the IVF laboratory at Brigham and Women's Hospital and wish to thank the large number of anonymous patients for donating oocytes for research. All experiments were carried out according to the Declaration of Helsinki and conducted following protocols approved by the institutions' Institutional Review Boards for the Protection of Human Subjects.
