Abstract
Prosaposin, the precursor of four lysosomal cofactors required for the hydrolysis of sphingolipids, is transported to the lysosomes via the alternative receptor, sortilin. In this study, we identified a specific domain of 17 amino acids within the C terminus of prosaposin involved in binding to this sorting receptor. We generated six prosaposin deletion constructs and examined the effect of truncation by coimmunoprecipitation and con-focal microscopy. The experiments revealed that the first half of the prosaposin C terminus (aa 524–540), containing a saposin-like motif, was required and necessary to bind sortilin and to transport it to the lysosomes. Based on this result, we introduced twelve site-directed point mutations within the first half of the C terminus. Although the interaction of prosaposin with sortilin was pH dependent, the mutation of hydrophilic amino acids that usually modulate pH-dependent protein interactions did not affect the binding of prosaposin to sortilin. Conversely, a tryptophan (W530) and two cysteines (C528 and C536) were essential for its interaction with sortilin and for its transport to the lysosomes. In conclusion, our investigation demonstrates that a saposin-like motif within the first half of the prosaposin C terminus contains the sortilin recognition site.
M
An independent report also demonstrated the occurrence of prosaposin/pro-cathepsin D complexes and suggested that they are formed spontaneously after the folding of the two precursors in the endoplasmic reticulum and transported in tandem to lysosomes and to the extracellular space (Gopalakrishnan et al. 2004). Our laboratory findings showed that the lysosomal trafficking of prosaposin is dependent on the D domain along with its highly conserved C-terminal region (Zhao and Morales 2000). More recently, we have demonstrated that prosaposin can be targeted to the endosomes and lysosomes by the alternative trafficking receptor, sortilin (Lefrancois et al. 2003). Sortilin is a 95- to 100-kDa membrane protein expressed in several tissues, such as brain, testis, and spinal cord (Petersen et al. 1997). Sortilin consists of a short cytoplasmic domain containing a sorting signal, a single transmembrane domain, and a luminal segment composed of a single VPS10 domain (Petersen et al. 1997; Nielsen et al. 2001). The VPS10 motif is also found in the VPS10p receptor, the sorting receptor implicated in the transport of carboxypeptidase Y to the yeast lysosomal vacuole (Marcusson et al. 1994). More recently, sortilin has been implicated as an alternative receptor to the MPR for some soluble hydrolases (Ni and Morales 2006; Canuel et al. 2008).
Prosaposin is the precursor of four heat-stable SAPs termed saposins A, B, C, and D (Morimoto et al. 1988,1989; Stastny et al. 1992; Hiraiwa et al. 1993). Prosaposin contains four central and two peripheral domains. The four central domains are delimited by three linker regions and correspond to the functional saposins that are also referred to as B-type domains (Patthy 1991; Kishimoto et al. 1992). Subsequent to prosaposin's lysosomal transport, the linker regions of the central domains are proteolytically cleaved to liberate saposins into the lumen of the lysosomes (O'Brien and Kishimoto 1991). The lysosomal hydrolysis of sphingolipids with short oligosaccharide side chains requires the presence of these SAPs (Sandhoff et al. 1998). The peripheral domains, also referred to as A-type domains, are situated at the N and C termini of prosaposin (Patthy 1991). While the B-type domains are composed of 80 amino acid residues, the A-type domains are composed of 34 amino acid residues corresponding to a saposin-like structure (Patthy 1991; Kishimoto et al. 1992). Deletion analysis revealed that unlike the N terminus behavior, removal of the C terminus abolished the trafficking of prosaposin to the lysosomes (Zhao and Morales 2000; Canuel et al. 2009). Furthermore, chimeric albumin proteins linked to saposin D and the C terminus of prosaposin reached the lysosomes (Zhao and Morales 2000; Canuel et al. 2009).
In this study, we have identified the critical domain within the first half of the C terminus of prosaposin (aa 524–540) that is responsible for the binding to sortilin and required for the targeting of prosaposin to the lysosomes.
Materials and Methods
Reagents and Constructs
The wild-type prosaposin cDNA construct linked to a myc tag was produced in our laboratory (Zhao and Morales 2000). The sortilin-myc construct was a gift from Dr. C. Petersen (University of Aarhus, Aarhus, Denmark). The pcDNA 3.1A/Myc-His vector was from Invitrogen (Carlsbad, CA). All of the primers and Escherichia coli DH-5α-competent cells were from Bio S and T (Montreal, QC, Canada). All restriction enzymes were from New England Biolabs (Pickering, ON, Canada). T4 DNA ligase was from Roche (Laval, QC, Canada). DMEM, FBS, and trypsin were from Invitrogen. Protease inhibitor cocktail was from Roche. ProLong Gold antifade reagent, 4',6-diamidino-2-phenylindole (DAPI), and Hoechst 33342 stain were from Invitrogen. The pGEM-T Easy Vector System II was from Promega (Madison, WI). StrataClone Ultra PCR cloning kit, QuikChange II site-directed mutagenesis kit, and QuikChange Lightning site-directed mutagenesis kit were from Stratagene (La Jolla, CA). Long Range PCR kit, Taq PCR kit, QIAquick gel extraction kit, QIAprep Spin mini-prep kit, Hispeed Plasmid mini-kit, and PolyFect transfection reagent were all from Qiagen (Mississauga, ON, Canada). Protein A Sepharose 4 Fast Flow, ECL Plus Western blotting detection reagents, and Amersham Hyperfilm MP autoradiography films were from GE Healthcare (Piscataway, NJ). MG-132 (Z-Leu-Leu-Leu-al), citric acid, and Na2HPO4 were from Sigma (Oakville, ON, Canada).
Antibodies
The rabbit polyclonal anti-prosaposin antibody was generated in our laboratory (Lefrancois et al. 2003). Monoclonal anti-c-myc antibody and goat anti-mouse IgG antibodies conjugated to horseradish peroxidase (HRP) were from Sigma. Rabbit polyclonal anti-sortilin antibodies were purchased from Alomone Labs (Jerusalem, Israel). Goat anti-rabbit IgG antibodies conjugated to HRP was from Santa Cruz Biotechnology (Santa Cruz, CA). Monoclonal anti-LAMP-1 antibody was from BD Biosciences (Mississauga, ON, Canada). Monoclonal anti-Golgin 97 antibody and goat anti-mouse IgG antibody conjugated to Alexa 594 were from Invitrogen. Chicken polyclonal anti-myc antibody, fluorescein-labeled goat anti-chicken IgY antibody, and BlokHen were from Aves Labs (Tigard, OR).
Truncation Constructs
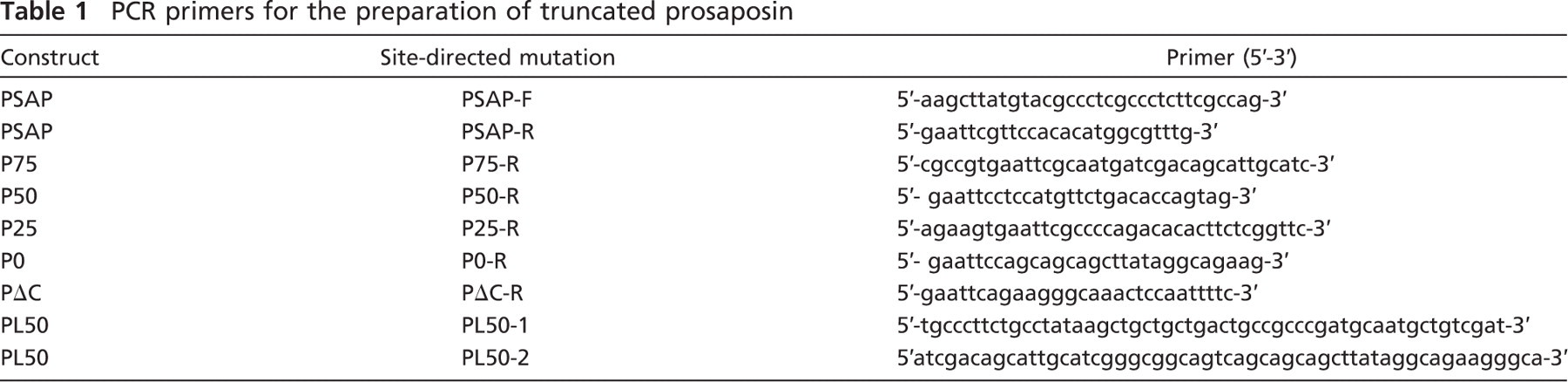
For sequential deletion of the C terminus, a prosaposin cDNA was amplified by PCR with the primers PSAP-F (forward) and PSAP-R (reverse) (Table 1). Subsequently, a HindIII restriction site was added into the 5’ end of PSAP-F, and an EcoRI restriction site was added into the 5’ end of PSAP-R. The purified PCR product was subcloned into the pGEM-T vector and transformed into DH-5α-competent cells. The resulting prosaposin TA clone was purified with a QIAprep Miniprep kit and digested with HindIII and EcoRI restriction enzymes. After it was purified with a QIAquick gel extraction kit, the wild-type prosaposin cDNA was ligated with the pcDNA 3.1A vector previously digested with the HindIII and EcoRI enzymes. After the second round of transformation and purification, the resulting wild-type prosaposin-myc construct was harvested and designated PSAP-WT (Figures 1A and 2A).
In the same manner, we obtained the first truncated construct, P-75, where 25% of the DNA sequence encoding the A-type domain of the C terminus was deleted from the 3’ end. This truncated prosaposin contains 75% of the C terminus, lacking aa 552–557. To achieve this deletion, a pair of primers, PSAP-F and P75-R (Table 1), was used for PCR amplification (Figure 2A).
PCR primers for the preparation of truncated prosaposin
The second truncated construct, P-50, lacked aa 541–557, representing 50% of the DNA sequence encoding the C terminus (Figure 2A). The primers PSAP-F and P50-R (Table 1) were used to attain the truncation.
The third construct, P-25, lacked 75% of the C-terminal end (aa 532–557). The primers for the amplification of P-25 were PSAP-F and P25-R (Table 1).
The fourth truncated construct, P-0, lacked the entire A-type domain, excluding the linker region (aa 524–557). The primers used in the PCR amplification were PSAP-F and P0-R (Table 1).
The final truncated construct, P-ΔC, lacked the whole C terminus (aa 518–557), including the linker region connecting this domain to saposin D. The primers for P-ΔC amplification were PSAP and PΔC-R (Table 1). All of the above-described constructs were produced following the same protocol used for the generation of PSAP-WT (Figure 2A).
Additionally, we engineered a truncated construct which we named P-L50. In contrast to P-50, in which 50% of the DNA sequence encoding the A-type domain was deleted from the 3’ end, P-L50 lacked the first half of the A-type domain of C terminus (aa 524–540). That is, 50% of the DNA sequence was removed from the 5’ end of the DNA segment encoding the A-type domain on the C terminus (Figure 2A). To generate P-L50, we used a QuikChange Lightning site-directed mutagenesis kit and deleted half of the DNA sequence of the A-type domain. The pair of primers for P-L50 amplification was designed with QuikChange primer design software. The primers for P-L50 mutagenesis, PL50-1 and PL50-2, were complementary to each other (Table 1).
Site-directed Mutagenesis
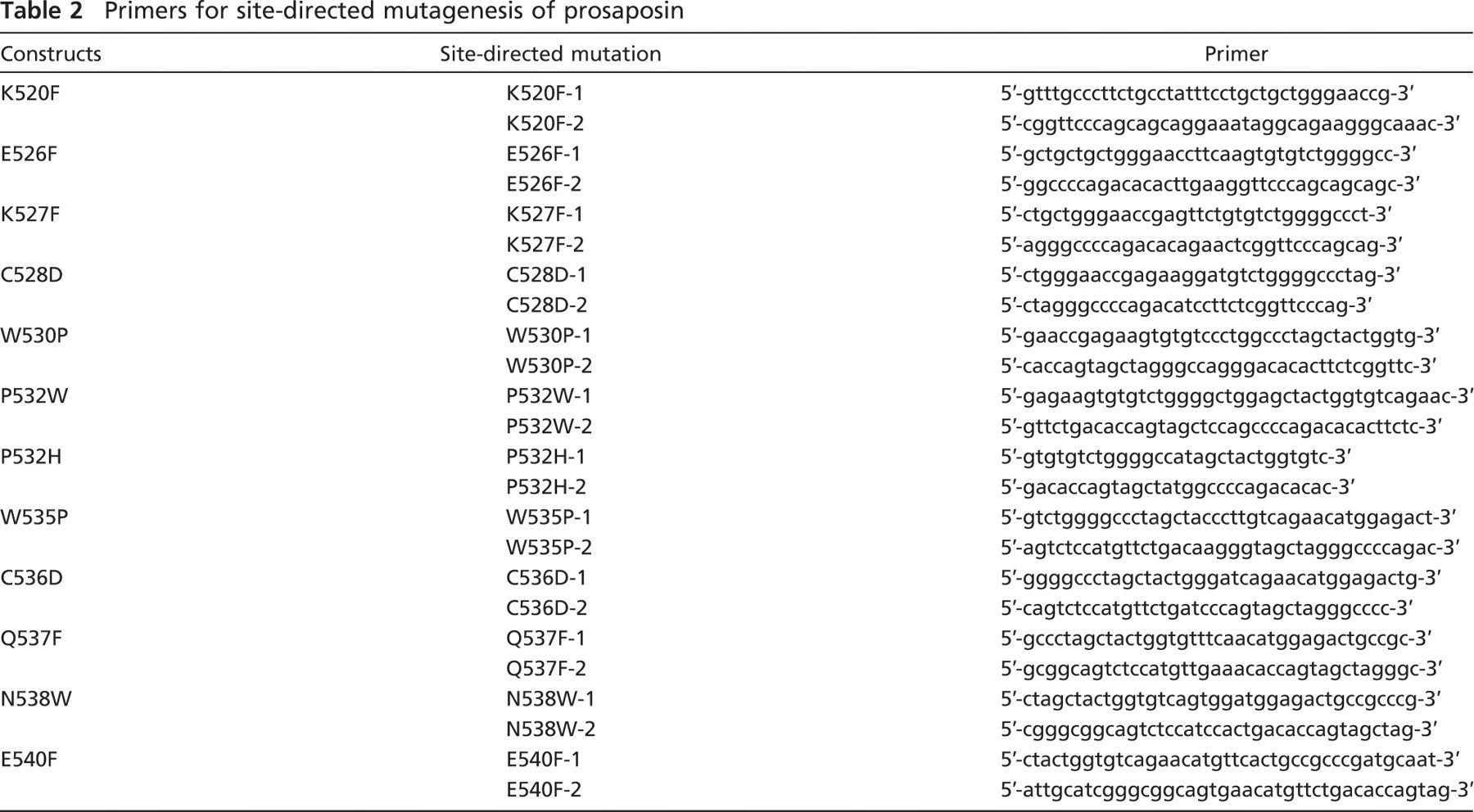
Twelve site-directed mutations in the C terminus of prosaposin were generated by either the QuikChange II site-directed mutagenesis or the QuikChange Lightning site-directed mutagenesis kit. For each mutation, one of the amino acids within the first half of the C terminus was changed to the most disfavored substitute amino acid (Betts and Russell 2003). The site-directed mutations were K520F, E526F, K527F, C528D, W530P, P532W, P532H, W535P, C536D, Q537F, N538W, and E540F. The pairs of primers were designed with QuikChange primer design software. In each pair, the primers were complementary to each other (Table 2).
Cell Culture
COS-7 cells were cultured in DMEM supplemented with 10% FBS, 5% penicillin and streptomycin, and
Sortilin–Prosaposin pH-dependent Binding Assay
COS-7 cells were cotransfected with sortilin-myc and wild-type prosaposin-myc constructs. Twenty-four hr after transfection, the cells were harvested and lysed in lysis buffer [0.5% NP-40 and protease inhibitor cocktail in citric acid/Na2HPO4 buffer (pH 4.0, 5.0, 5.5, 6.0, 6.5, or 7.0)]. The supernatants were incubated with protein A Sepharose beads coupled to anti-prosaposin antibody at 4C overnight. The beads were washed three times with washing buffer [50 mM citric acid/Na2HPO4 buffer, 0.15 M NaCl (pH 4.0, 5.0, 5.5, 6.0, 6.5, or 7.0)] and boiled in 3 × SDS-PAGE loading buffer. After proteins were separated with a 10% SDS-PAGE gel, proteins were transferred to a nitrocellulose membrane and immunoblotted with anti-sortilin antibody at a 1:2500 dilution. Anti-rabbit secondary antibody coupled to HRP was used at a 1:10,000 dilution. The experiment was performed three times.
Sortilin–Prosaposin Binding Assay
COS-7 cells were cotransfected with the sortilin-myc plasmid and one of the following constructs: PSAP-WT, P-70, P-50, P-25, P-0, P-ΔC, P-L50, K520F, E526F, K527F, C528D, W530P, P532W, P532H, W535P, C536D, Q537F, N538W, E540F, or pcDNA3.1A empty vector. Twenty four hr after transfection, the cells were harvested and lysed [50 mM Tris-HCl, 0.15M NaCl, 0.5% NP-40, and protease inhibitor cocktail (pH 6.0)]. The cellular supernatants were incubated with protein A Sepharose beads coupled to anti-sortilin antibody at 4C overnight. The resulting protein–bead complexes were washed three times with washing buffer [50 mM Tris–HCl, 0.15M NaCl (pH6.0)] and boiled in 3 × SDS–PAGE loading buffer. Proteins were separated on a 10% SDS–PAGE gel, transferred to a nitrocellulose membrane, and immunoblotted with anti-myc antibody diluted 1:5000. Anti-mouse secondary antibodies coupled to HRP were used at a dilution of 1:10,000. The desired protein bands were detected with ECL Plus Western blotting detection reagents. The experiment was performed three times.
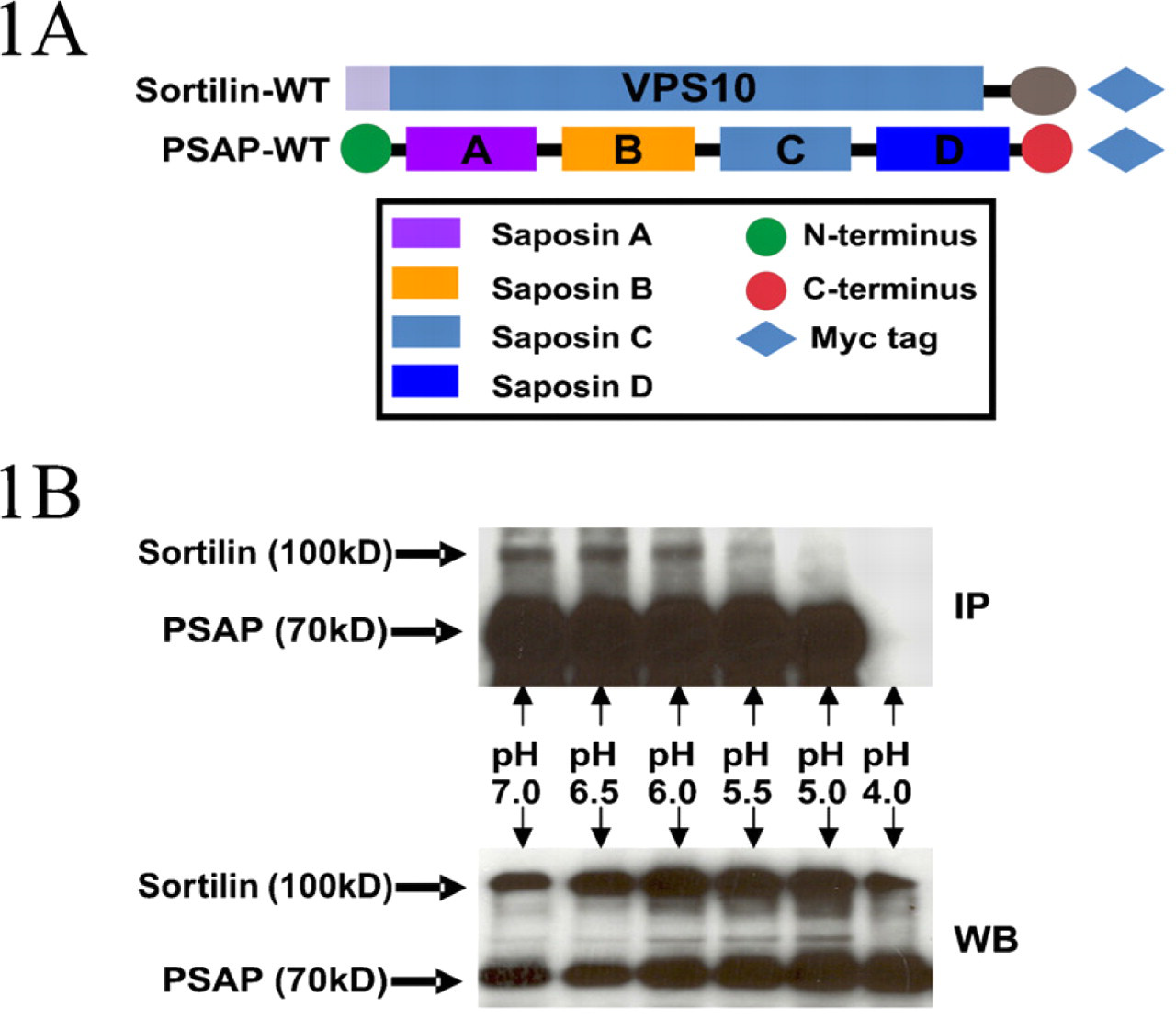
Effect of pH on the interaction of prosaposin and sortilin. (
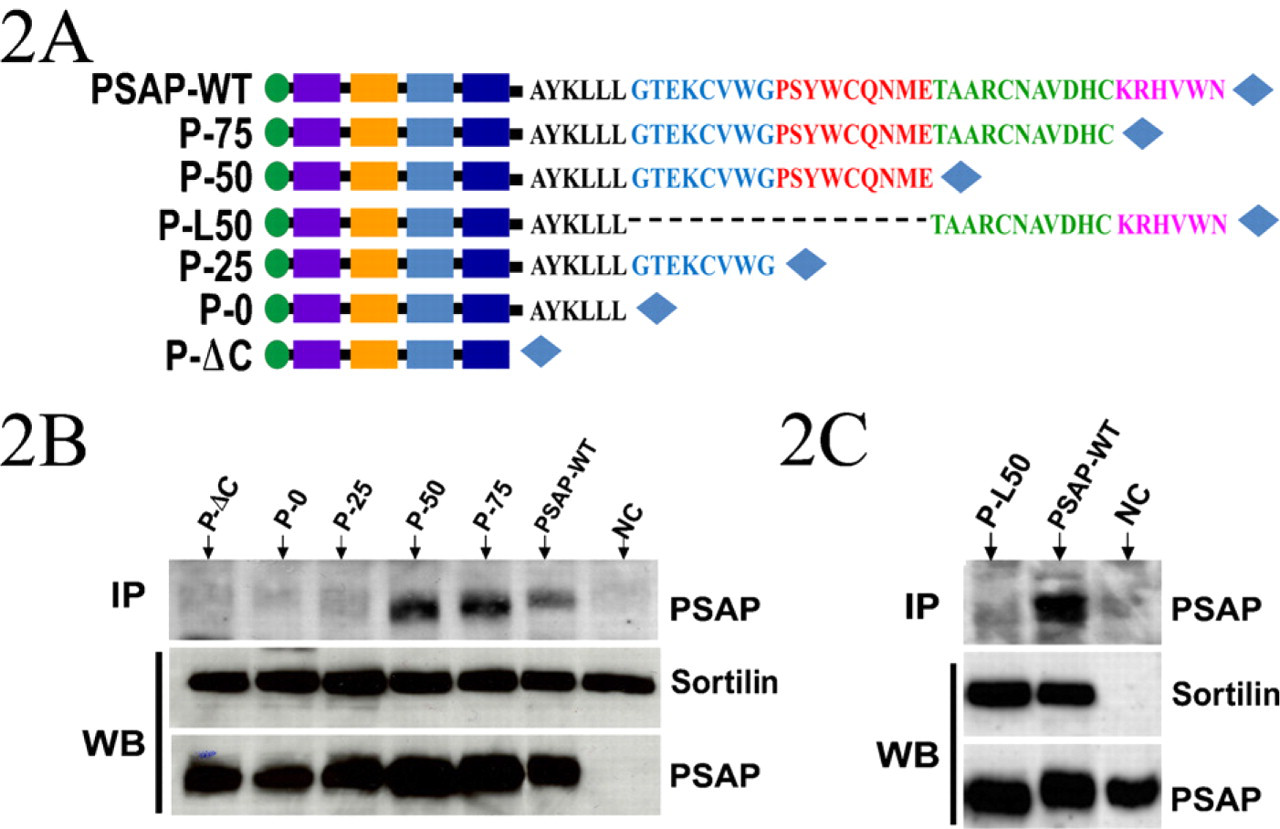
Effects of sequential truncations of prosaposin C terminus. (
Immunofluorescent Confocal Microscopy
COS-7 cells were seeded onto cover slips in 6-well plates and cultured for 24 hr. The cells were then transfected with PSAP-WT, P-70, P-50, P-25, P-0, P-ΔC, P-L50, K520F, E526F, K527F, C528D, W530P, P532W, P532H, W535P, C536D, Q537F, N538W, E540F, or pcDNA3.1A empty vector. After 24 hr, the cells were washed with 1x PBS, fixed for 10 min in 4% paraformaldehyde, and then permeabilized with 0.25% Triton X-100 for 5 min. Subsequently, the cells were incubated for 1 hr with 3% goat serum, BlokHen diluted 1:10, and 0.1% Triton X-100. The incubation of double primary antibodies was performed at a 1:200 dilution overnight at 4C with chicken anti-myc antibody and mouse anti-LAMP-1 or anti-Golgin 97 antibody. The cells were washed with PBS and incubated with mixed secondary antibodies diluted to 1:400 (i.e., goat anti-mouse IgG antibody conjugated to Alexa 594 and fluorescein-labeled goat anti-chicken IgY antibody) for 1 hr at room temperature. Then the cells were washed again with PBS, and the nuclei were counterstained with 300 μl of Hoechst 33342 (0.007 μg/μl) stain. Finally, the cells were rinsed with PBS and water and mounted onto slides using ProLong Gold antifade reagent with DAPI. The immunofluorescent staining was visualized with an LSM 510 META confocal microscope from Carl Zeiss (Toronto, ON, Canada).
Primers for site-directed mutagenesis of prosaposin
MG-132 Treatment
COS-7 cells were transfected with one of the following constructs: W530P, P532W, P532H, W535P, PSAP-WT, or pcDNA3.1A empty vector, and 24 hr later the cells were incubated in DMEM containing 10 μM MG-132 for 6 hr to inhibit the proteasome degradation pathway. The cell lysates were subjected to Western blotting. The experiment was performed three times.
Statistical Analysis
The results of confocal microscopy were statistically analyzed using SPSS version 16.0 software. The number of vesicles stained by the LAMP-1 antibody (red) and the number of overlaid vesicles (yellow) stained by both anti-LAMP-1 and anti-myc antibodies in transfected cells were counted. The counts were done in twenty cells from a minimum of three different experiments for each mutation. Subsequently, the percentage of overlaid structures for each construct was calculated. A Welch t-test or a one-way ANOVA test was performed to compare the effect of truncations and mutations in the C terminus of prosaposin with wild-type prosaposin (PSAP-WT).
Results
Effect of pH on the Interaction of Prosaposin and Sortilin
Ligand–receptor binding and dissociation are pH-dependant processes (Mellman et al. 1986). While mild acidic compartments, such as the lumens of the trans-Golgi network (TGN) (pH 6.0), favor the binding of soluble hydrolases to the MPRs, the strong acidic compartments of the endosomes and lysosomes (pH 5.0 or lower) induce their release from the MPRs (Coffey and De Duve 1968; van Weert et al. 1995; Schindler et al. 1996; Demaurex et al. 1998). To test the hypothesis that the binding of prosaposin to sortilin is pH dependent, we performed an in vivo pH-dependent assay. Since the amounts of endogenous prosaposin and sortilin in the COS-7 cells are insufficient for coimmunoprecipitation assays, the cells were cotransfected with wild-type prosaposin and sortilin constructs (Figure 1A). The transfected cells were homogenized in lysis buffer at pH levels of 4.0, 5.0, 5.5, 6.0, 6.5, or 7.0. All samples subjected to immunoprecipitation contained equal concentration of lysate, and the washing buffer for each sample had the same pH as the initial lysis buffer, to keep the sample in a stable pH condition. The complexes pulled down by anti-prosaposin antibody were resolved with a 10% acrylamide gel. After being transferred to a nitrocellulose membrane, the samples were immunoblotted for prosaposin and sortilin by using anti-prosaposin and anti-sortilin antibodies, respectively (Figure 1B). The results showed that the presence and/or intensity of the sortilin bands varied at different pH levels. The strongest sortilin bands were observed at pH 6.0, 6.5, and 7.0, whereas a substantial decrease in intensity was detected at pH 5.5, and no sortilin band was observed at pH 5.0.
Effects of Sequential Truncations of the Prosaposin C Terminus
To identify the specific region within the prosaposin C terminus that binds sortilin, we have generated several truncated prosaposin constructs (Figure 2A). Before engineering these constructs, we analyzed the predicted secondary structure of the C terminus, using EMBOSS Garnier algorithm (Garnier et al. 1978). The EMBOSS output file predicted the existence of two α-helices within the C terminus. The first helix is localized to the linker region between aa 518 and 523, and the second helix is between aa 540 and 550. To generate the first construct (P-75) and to avoid disruption of the predicted helixes, we deleted the C-terminal region immediately after C551. In the P-50 construct the deleted sequence spans aa 541 to 557. In this case, the second helix (E540 to H550) was eliminated. In P-25, the deletion spans aa 531 to 557, resulting in the elimination of two cysteine residues. The P-0 construct lacks the entire A-type domain (aa 524–557). The final construct, P-ΔC, is a truncated prosaposin lacking the entire C terminus and the linker that connects this region to saposin D (aa 518–557).
COS-7 cells were cotransfected with sortilin and each of the prosaposin constructs described above. The cells were homogenized in lysis buffer (pH 6.0) and subjected to immunoprecipitation. The complexes were pulled down with anti-sortilin antibody and resolved with a 10% acrylamide gel. Immunoblotting with anti-myc antibody showed that sortilin pulled down the wild-type prosaposin PSAP-WT and truncated constructs P75 and P50, while it failed to precipitate P-ΔC (Figure 2B). P-L50, a prosaposin construct lacking the first half of A-type domain, was not pulled down by sortilin (Figure 2C).
To determine if the truncations affected the transport of prosaposin to the lysosomes, COS-7 cells transfected with each prosaposin construct were examined by confocal microscopy after immunofluorescence labeling. Both the wild-type and the truncated prosaposin constructs appeared green while the TGN and lysosomes were stained red with anti-Golgin 97 antibody or anti-LAMP1 antibody, respectively. The nuclei were stained blue with Hoechst 33342. The results demonstrated that wild-type prosaposin (PSAP-WT) exhibited Golgi and vesicular staining and overlaid with both anti-Golgin 97 (data not shown) and anti-LAMP-1 staining (Figure 3A). The distribution of P75 and P50 was similar to that of PSAP-WT (Figures 3B–3C). However, P-ΔC, P0, and P-25 localized only in the perinuclear region and overlaid with anti-Golgin 97 (data not shown) but not with anti-LAMP-1 staining (Figures 3D–3G). Statistical analysis showed that the percentage of overlaid vesicles of P-ΔC, P-0, P-25, and P-L50 significantly decreased compared with that of PSAP-WT (p<0.01), while P-50 and P-75 did not (p>0.05) (Figure 4A).
Effects of Point Mutations in the C Terminus of Prosaposin
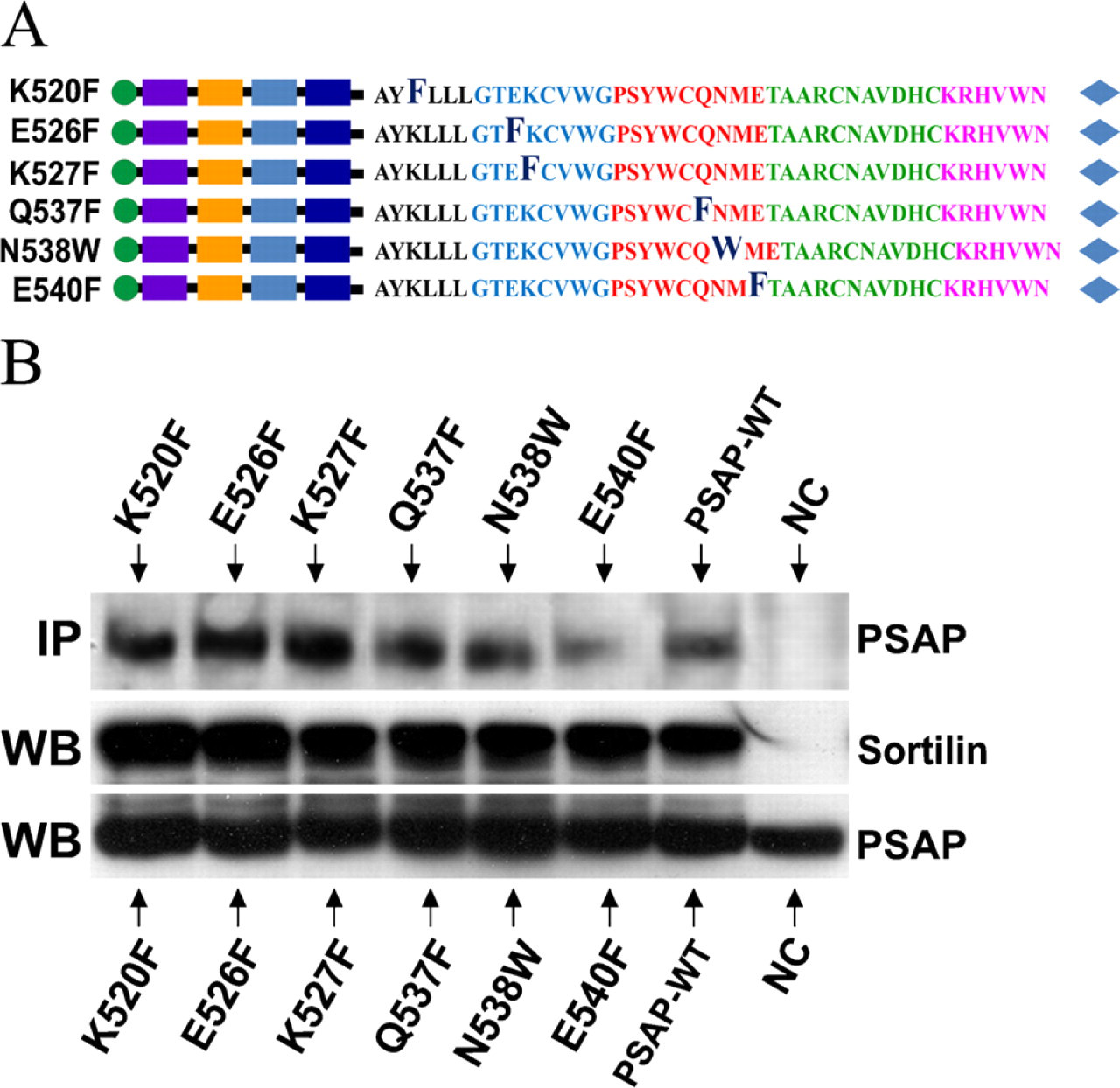
In this experiment we introduced six point mutations to hydrophilic amino acid residues in the first half of the C terminus of prosaposin, bearing in mind that those residues might play key roles in the pH-dependent binding of prosaposin to sortilin. The point mutations were directed to several conserved polar amino acids in the first half of the A-type domain (Zhao et al. 1998) (Figure 5A).
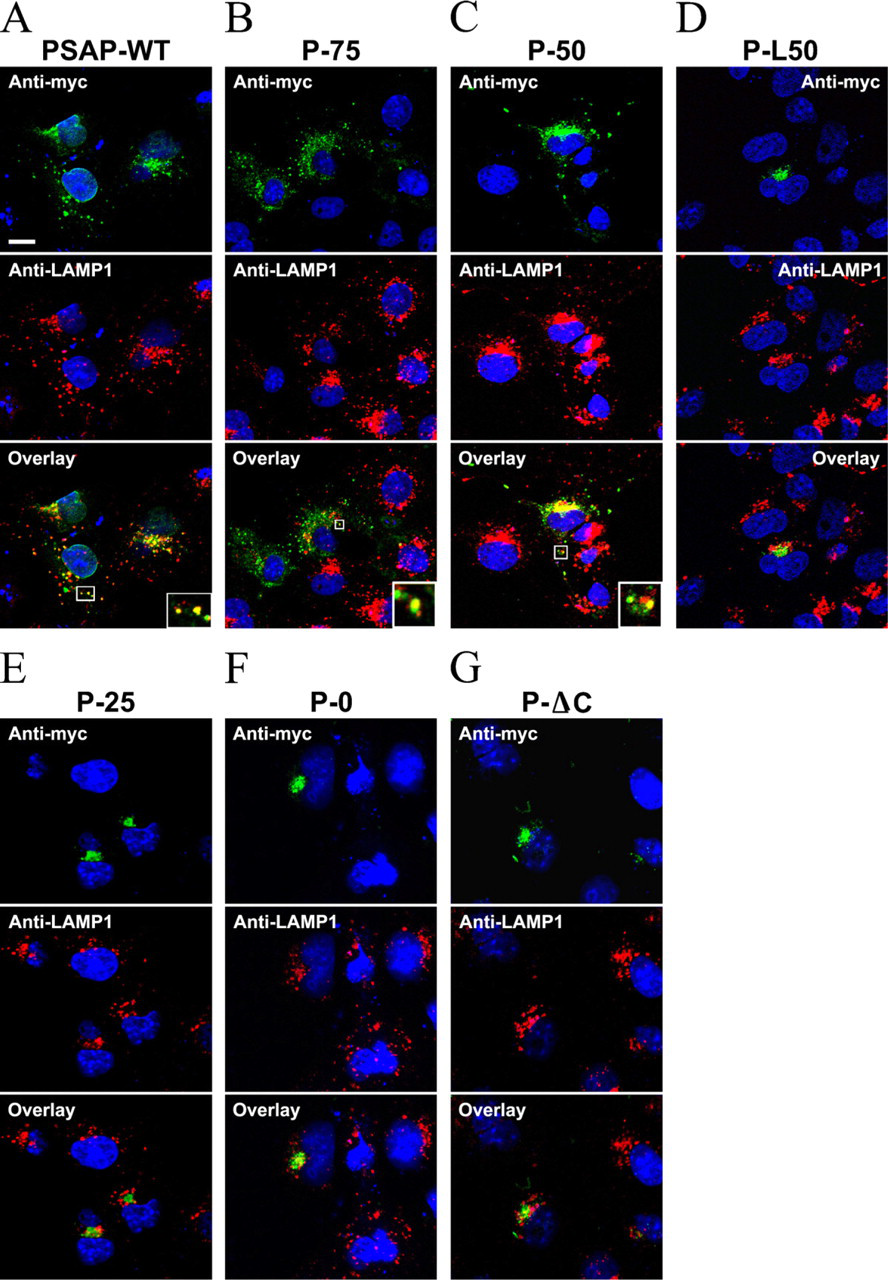
Lysosomal targeting of sequential truncations of prosaposin. The COS-7 cells transfected with the truncated constructs were examined by confocal microscopy. Truncated prosaposin was stained green with chicken anti-myc antibody. Lysosomes were stained red with anti-LAMP-1 antibody. Nuclei appear in blue. Anti-myc staining of cells transfected with PSAP-WT, P-75, and P-50 constructs labeled the perinuclear region and cytoplasmic vesicular structures and overlaid with anti-LAMP1 staining (
Each mutated probe (K520F, E526F, K527F, Q537F, N538W, and E540F) was cotransfected to COS-7 cells with the wild-type sortilin construct and subjected to immunoprecipitation. After being pulled down with anti-sortilin antibody, the protein complexes were immunoblotted with anti-myc antibody. All of the hydrophilic amino acids mutations (K520F, E526F, K527F, Q537F, N538W, and E540F) were pulled down by sortilin (Figure 5B).
To determine if the mutations affected the transport of prosaposin to the lysosomes, the cells transfected with mutated prosaposin constructs were immunostained with anti-myc, anti-Golgin 97, and/or anti-LAMP-1 antibodies, and the results were examined by confocal microscopy. The hydrophilic amino acid mutations K520F, E526F, K527F, Q537F, N538W, and E540F localized to vesicular structures and in the perinuclear region and overlaid with the anti-LAMP-1 (Figure 6) and anti-Golgin 97 (data not shown), respectively. One-way ANOVA testing showed that anti-myc/anti-LAMP-1-overlaid vesicles remained unchanged in those mutations compared with those of PSAP-WT (p>0.05) (Figure 4B).
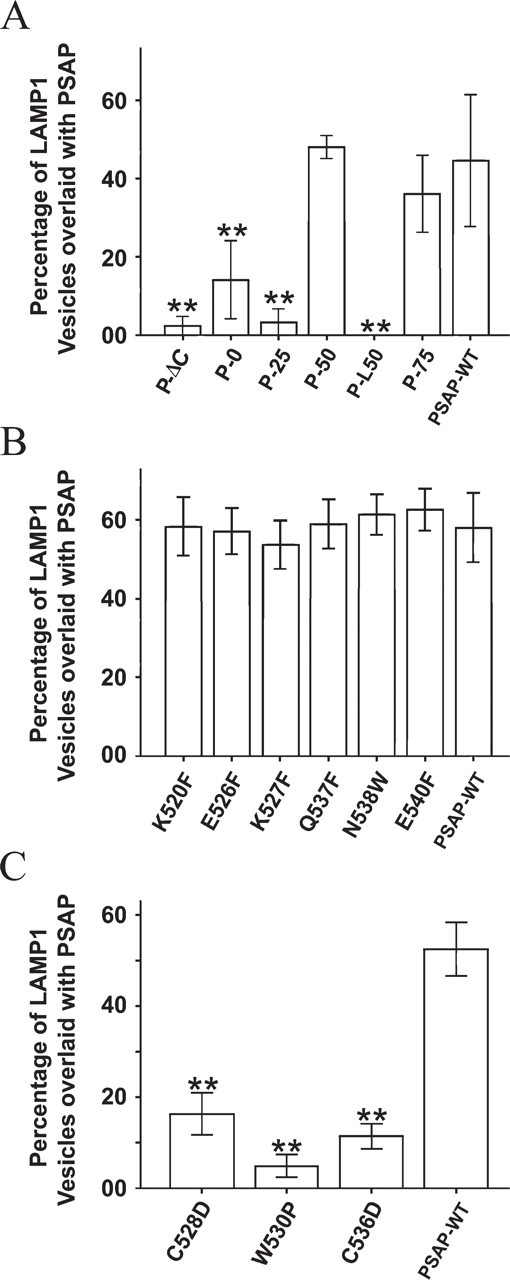
Lysosomal transport of prosaposin truncation/mutation constructs. The percentage of LAMP-1 vesicles overlaid with each prosaposin truncation/mutation construct was calculated. (
We also introduced six additional point mutations to hydrophobic residues in the first half of the A-type domain, C528D, W530P, P532W, P532H, W535P, and C536D (Figure 7A). In the case of the W530P, C528D, and C537D mutations, the anti-myc staining overlaid with anti-Golgin 97 (data not shown) but rarely with vesicular structures stained by anti-LAMP-1 (Figures 7B–7D). As expected, the percentage of anti-myc/anti-LAMP-1-overlaid vesicles was significantly lower than in PSAP-WT (p<0.01) (Figure 4C). Mutated constructs P532W, P532H, and W535P abolished the immunostaining of the protein carrying the mutations, and Western blot analysis failed to detect bands for any of the three mutations (Figure 7E). To examine whether these mutations resulted in the misfolding and degradation of prosaposin, we used MG-132 to inhibit the proteasome degradation pathway. Our results showed that MG-132 treatment allowed detection of bands for the three mutated prosaposins (Figure 7E).
Discussion
Sphingolipid activator proteins (SAPs) are soluble cofactors involved in the lysosomal degradation of glycosphingolipids (Schuette et al. 2001). SAPs include five activator proteins, the G
Effects of point mutations of hydrophilic residues in the interaction of prosaposin and sortilin. (
It is accepted that soluble lysosomal proteins are sorted within the TGN by a sorting receptor and that the ligand-receptor complex is targeted to the endosomes by accessory proteins that interact with the sorting receptor (Hirst and Robinson 1998; Rouille et al. 2000). Recently, our laboratory demonstrated that unlike the majority of soluble hydrolases, which are transported via the MPRs, prosaposin is translocated to endosomes via the alternative receptor sortilin (Lefrancois et al. 2003). Preliminary evidence suggests that the C terminus of prosaposin is required for the transport of prosaposin to the lysosomes (Zhao and Morales 2000). In a previous comparative study of the structure of human, mouse, and rat prosaposin, we found that the D domain and the C terminus were the most conserved regions among the three species, with more than 90% of sequence identity (Zhao et al. 1998). The high level of conservation suggests that this region carries a critical function. More recently, we found that the deletion of the C terminus abolished the transport of prosaposin to the lysosomes and that addition of this region to albumin induced the binding of the chimeric protein to sortilin, the penetration of the complex within detergent resistant micro-domains and the targeting of albumin to the lysosomes (Zhao and Morales 2000).
This observation led us to propose a model in which a specific domain within the C-terminal region of prosaposin interacts with the luminal domain of sortilin (Lefrancois et al. 2003). However, the precise nature of this interaction, as well as the intervening domains of these molecules, is still unknown. Interestingly, the C terminus of prosaposin contains a saposin-like motif significantly similar to the A-type domain found in the N terminus of the surfactant B associated protein (Patthy 1991) which has been implicated in the transport of this protein to the lamellar bodies of type II pneumocytes (Lin et al. 1996). Therefore, the elucidation of the recognition process between prosaposin and sortilin is important to understand the mechanism of this interaction.
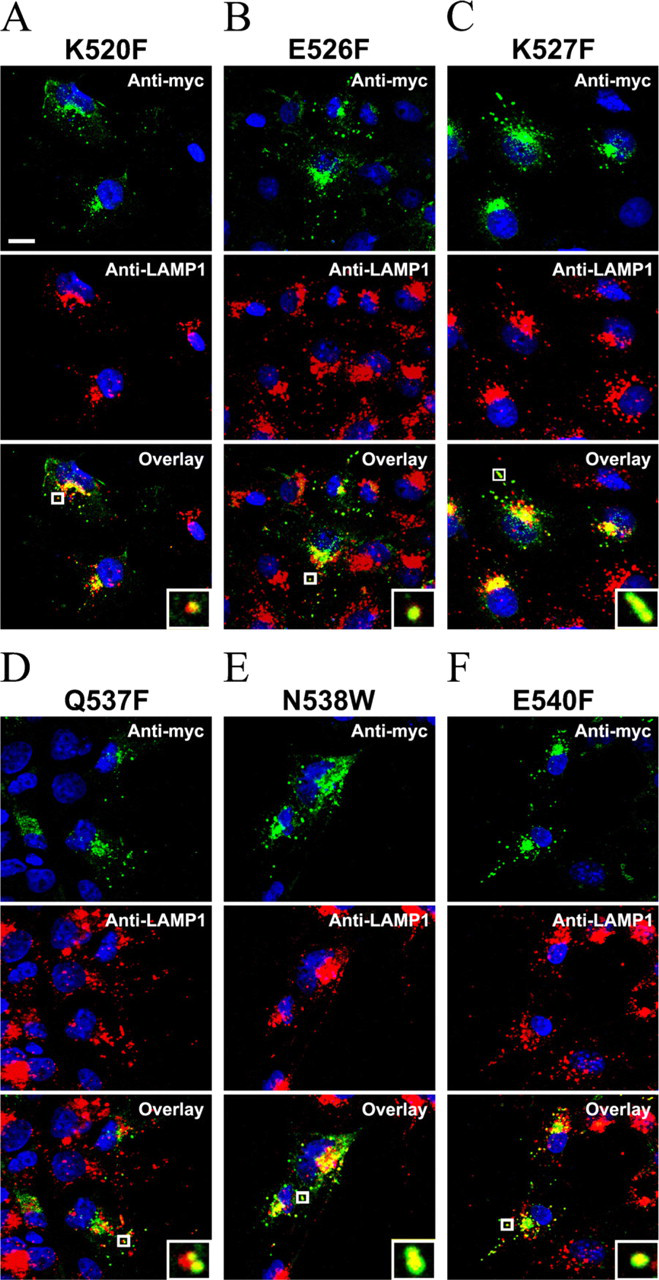
Lysosomal targeting of prosaposin point mutations on hydrophilic residues. The COS-7 cells transfected with the mutation constructs K520F, E526F, K527F, Q537F, N538W, and E540F were examined by confocal microscopy. Prosaposin mutation constructs were stained green, lysosomes stained red, and nuclei stained blue. Anti-myc staining of cells transfected with K520F, E526F, K527F, Q537F, N538W, and E540F constructs labeled the perinuclear Golgi region and cytoplasmic vesicular structures and overlaid with anti-LAMP1 staining (
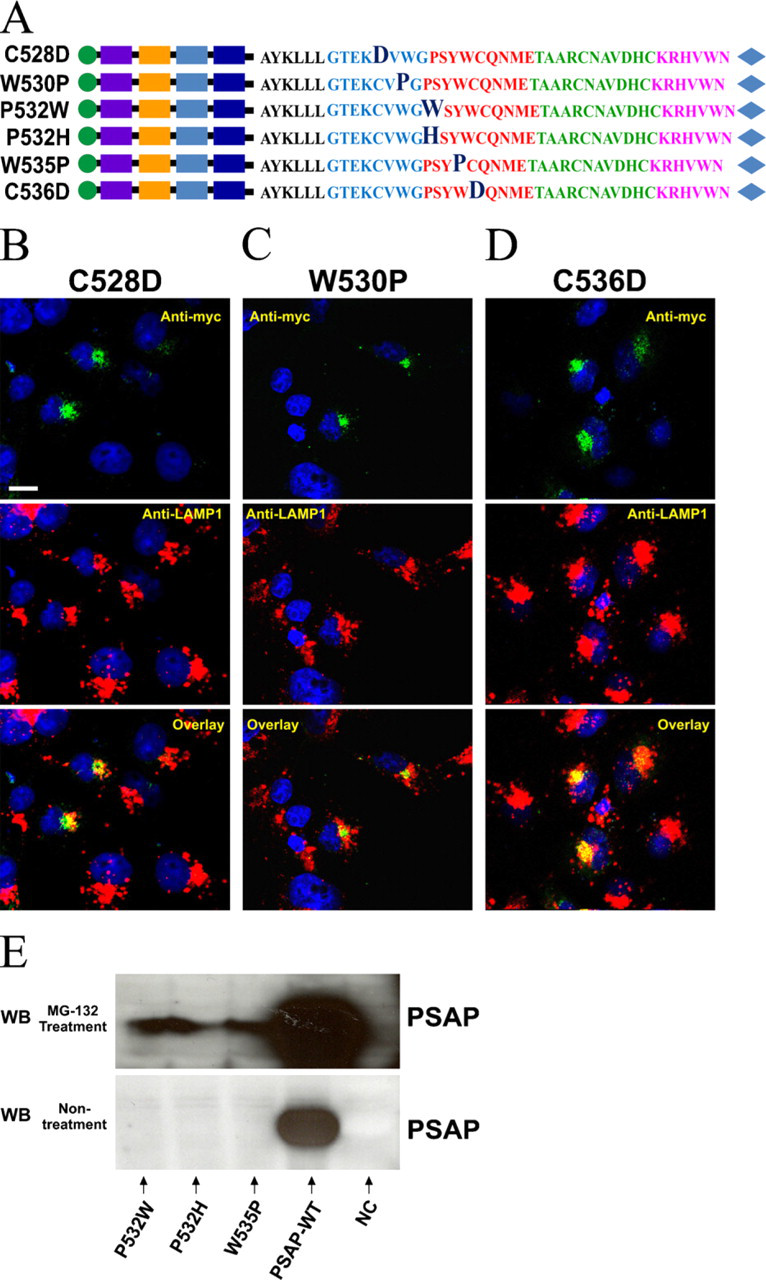
Effects of point mutations on hydrophobic residues in the first half of prosaposin C terminus. (
The objective of this investigation was to identify the specific domain within the C terminus of prosaposin implicated in its binding to sortilin. To accomplish this objective, six truncated prosaposin constructs were generated by deleting specific regions of the C terminus. The effect of these truncations was examined by con-focal microscopy. The results demonstrated that the elimination of the terminal half of the A-type domain (aa 541–557), which includes an α-helix, did not affect the transport of prosaposin to the lysosomes. However, the elimination of the first half of A-type domain (aa 524–540), which contains 17 amino acids located between the two α-helices, abolished the transport of prosaposin to the lysosomes. This result correlated with the immunoprecipitation assays which demonstrated that after the elimination of the first half of A-type domain, prosaposin could not be pulled down by sortilin.
Based on the fact that most ligand-receptor interactions depend on small pH shifts (Mellman et al. 1986) we tested whether or not the binding of prosaposin to sortilin was also pH-dependent. While prosaposin bound to sortilin at a pH above 6.0, a substantial decrease in binding was detected at pH 5.5. No binding was detected at pH 5.0. This result indicates that the binding of prosaposin to sortilin is pH dependent and suggests that the pH of the TGN (pH 6.0) (Demaurex et al. 1998) might favor the association between prosaposin and sortilin. Our results also indicate that the dissociation of prosaposin and sortilin is pH dependent and suggest that the pH of endosomes (pH 5.5) (Coffey and De Duve 1968; van Weert et al. 1995) favors the dissociation between prosaposin and sortilin.
It is also accepted that hydrophilic amino acids found at the surface of water-soluble proteins, contribute to their solubility and form binding sites for charge molecules (Lodish et al. 2003). As a consequence, small shifts of cellular pH affect the charge of hydrophilic amino acids and modulate the binding properties of soluble proteins (Lodish et al. 2003). Prosaposin is a soluble globular protein. The sequence analysis of the C terminus revealed the presence of a saposin-like motif composed of two α-helices separated by several turns and our deletion experiments demonstrated that those turns are essential for the binding to sortilin. Thus, it is possible that small pH shifts produce conformational changes that affect the binding of this domain to sortilin. In fact, pH variations have been shown to induce conformational changes in all four saposins (Vaccaro et al. 1995). Thus, the stability of the tertiary structure of the first half of the saposin-like motif found within the prosaposin C terminus is probably affected by pH shifts. Consequently, it is tempting to speculate that the acidic pH of late endosomes induces a conformational change in the C terminus that favors the release of prosaposin from sortilin. Based on these facts, we replaced hydrophilic amino acids in the first half of the A-type domain to test the hypothesis that peripheral polar amino acids in the prosaposin C terminus may be essential for the lysosomal transport of prosaposin. To examine this hypothesis we generated six point mutations of conserved polar amino acid residues. However, the results demonstrated that none of these point mutations interfered with the sorting and transport of prosaposin to the lysosomes.
Banares-Hidalgo et al. (2008) found that the tryptophan residues within the saposin A-type domain of the SP-B precursor N terminus are responsible for the conformational change of SP-B precursor induced by low pH (Banares-Hidalgo et al. 2008). We wanted to know if tryptophan residue was important for the conformational structure of prosaposin C terminus. In addition, we also wanted to disclose the potential amino acid residues involved in the binding center of prosaposin-sortilin complex and in stabilizing the tertiary structure of prosaposin. Therefore, we introduced six point mutations to hydrophobic amino acid residues in the first half of the C terminus, which might be important for the trafficking of prosaposin.
Two of these mutations correspond to cysteines (C528D and C536D) that might form disulphide bonds. As expected, both mutations significantly decreased the transport of prosaposin to lysosomes, suggesting that they are essential for the stability of prosaposin structure and/or for the interaction of prosaposin with sortilin.
Additionally, we have introduced two mutations to proline residues (P532W and P532H), which are often found within tight turns of proteins, and two mutations to tryptophan residues (W530P and W535P), which are implicated in molecular recognition and stabilization of tertiary structure of proteins (Kay et al. 2000; Betts and Russell 2003). In the case of proline 532 we replaced it by the bulky amino acid residue tryptophan or by the positively charged histidine which is a small amino capable of inducing conformational changes (Betts and Russell 2003). Interestingly, the two proline mutations abolished the expression of prosaposin. However, after treating the transfected cells with MG-132, an inhibitor of the proteasome degradation pathway (Lee and Goldberg 1998), we visualized the presence of the native protein by Western blot analysis. This result suggests that the proline mutations resulted in the misfolding and rapid degradation of prosaposin. The mutations of the highly conserved tryptophan residues 530 and 535 in the first half of the C terminus were introduced while considering that this amino acid is the top ranking residue in hot spots of protein–protein binding (Ma et al. 2003). Tryptophan plays a unique role in the stability of protein tertiary structure and in protein–protein binding interactions due to its aromatic side chain and its bulky size (Samanta et al. 2000). While the W530P mutation blocked the lysosomal trafficking of prosaposin, the W535P mutation resulted in the rapid degradation of prosaposin. Consequently, our results suggest that W530 is likely involved in the stability of the tertiary structure of this domain and/or in the binding of prosaposin to sortilin. Recently, a study of the N terminus of the surfactant protein B (SP-B) precursor, which contains a saposin A-type domain, suggested that this region is susceptible to pH-dependent conformational changes and that the tryptophan residues within this region are responsible for a conformational change induced by low pH (Banares-Hidalgo et al. 2008). Taken together, it is possible that the prosaposin C terminus is also susceptible to pH-dependent conformational changes that favor or disfavor the binding of prosaposin to sortilin.
In conclusion, our findings suggested that the putative sortilin binding site in prosaposin is located within the first half of the A-type domain on the C terminus between aa 524 and 540. The tryptophan, proline, and cysteine residues in this region are critical for the stability and lysosomal trafficking of prosaposin. Our results demonstrated that the deletion of this region and the substitution of those critical hydrophobic residues abolished the binding of prosaposin to sortilin and the targeting of prosaposin to the lysosomes.
Footnotes
Acknowledgements
This research was supported by a CIHR operating grant to C.R.M.
