Abstract
Immunocytochemistry (IHC) and immunofluorescence (IF) offer crucial diagnostic insights in clinical settings. Most IF assays are performed using fluorophore-conjugated antibodies, but these fluorophores can be subject to issues, such as photobleaching and autofluorescence, that result in lower signal-to-noise ratios (SNR). Gold nanoparticles provide greater signal stability and scatter light well, making them easily separable from the background of biological tissues and improving SNR. In this study, we sought to determine the labeling efficiency, signal quality, and image artifacts of 2.2, 10, and 40 nm diameter gold nanoparticle probes conjugated to antibodies in IF applications on fixed, permeabilized cells in comparison to traditional fluorophores. Overall, micrographs of nanoparticle labels had higher SNR due to lower background signal, and punctate appearances as compared with the continuously distributed signal of the immunofluorescent label. Signal-to-noise ratios varied with nanoparticle diameter, and signal fidelity was worse for keratin versus epithelial growth factor receptor (EGFR). Labeling of EGFR was successful using both extracellular and intracellular epitopes, while poor labeling of keratin 19 with 10 nm diameter nanoparticles was improved by pretreatment with heat and sonication, suggesting hindrance of nanoparticle labels within the fixed, permeabilized cell.
Keywords
Introduction
Immunohistochemistry (IHC) and immunofluorescence (IF) are longstanding diagnostic tools, offering crucial insights into the spatial distribution and expression patterns of biomolecules within tissues.1–3 Most commonly applied in immunostaining practices due to the high selectivity of antibodies to their respective antigen proteins, IHC and IF have many applications in diagnostic pathology and basic research due to the relative simplicity of the assay.4,5 These staining techniques can be run as a direct or indirect method, where the labeled antibodies are bound to the primary target or a primary antibody specific for the epitope of interest, respectively. 6 As with any such test, false positives and cases of ambiguous staining occur. Indirect methods lower false positives in IHC and IF assays through washing steps to remove unbound antibodies and multiple secondary antibodies bound to the primary, leading to higher signal to noise. 6
Fluorescent organic dyes serve as labels in imaging assays due to their simplicity and safety as compared with radioisotopes, as well as their photostability compared with fluorescent proteins. 7 However, photobleaching and autofluorescence lower the signal strength and stability of fluorescent dyes over time, presenting potential issues during time-lapse studies and less accurate quantitative analysis of fluorescence signals.5,8–10 In addition, autofluorescence endogenous to cells and tissues or derived from fixatives also contributes to false positives and higher background, especially for imaging of widely distributed molecular targets within a labeled specimen.5,11
Nanoparticles are well-suited for biological imaging assays due to their tunable surface chemistry and distinctive optical (permittivity), magnetic (permeability), and electric (conductivity) characteristics. 12 Nanoparticle labels allow for a greater signal stability in imaging spatial biology assays as well, demonstrating less signal variation as compared with fluorescent counterparts.13–15 Gold nanoparticles strongly scatter visible light and are detectable by darkfield microscopy, 16 making them easily separable from the low background scattering of biological tissue.17–20 The stability and strength of nanoparticle scattering signals allow for efficient detection and monitoring of nanoparticle labeling in a time-lapse setting and have demonstrated utility in live cell imaging applications. 12 A critical issue in nanoparticle labeling applications is the potential impact of nanoparticle size on tissue penetration and cellular uptake. While larger, brighter nanoparticles may encounter challenges in efficiently penetrating tissues and accessing cellular compartments, smaller nanoparticle-conjugates may label targets in subresolution clusters, enhancing signal to noise. 21 In addition, while most organic dyes and fluorophores demonstrate signal increase in a linear fashion as binding increases, closely packed nanoparticles exhibit nearfield effects and aggregation that may impact scattering in a nonlinear fashion. Antigen retrieval combined with PEGylation, 22 and/or sonoporation 23 improves nanoparticle penetration and select binding to target molecules. Customizable nanoparticle size and shape may also lower the probability of undesirable nanoparticle aggregation or nonspecific binding in IHC and IF, 19 a phenomenon shown to be thermodynamically and entropically favorable due electrostatic interactions. 24 Apart from false signals produced by nonspecific aggregation, directed nanoparticle aggregation at the binding site combines the advantages of better penetration of smaller nanoparticles with higher signal from the aggregate versus single nanoparticles. Several nanoparticle labeling methods use this approach. 25
Based on previous successful immunogold labeling results, and on the diffusion and optical characteristics of gold nanoparticles and conventional fluorophores for molecular labeling, we hypothesized that nanoparticle-antibody conjugates would effectively label extracellular and intracellular epitopes, but with lower specificity and higher off-target/false signal using large-diameter nanoparticles and for extended intracellular targets. In this study, we sought to determine the labeling efficiency, signal quality, and image artifacts of 2.2, 10, and 40 nm diameter gold nanoparticle probes conjugated to antibodies in IF applications on fixed, permeabilized cells in comparison to traditional fluorophores across different subcellular locations. Labeling comparisons were performed with antibodies for three different targets within human gingival epithelial cells: an extracellular epitope of epithelial growth factor receptor (EGFR) to demonstrate extracellular performance; an intracellular EGFR epitope to demonstrate intracellular performance with a subplasma membrane target; and keratin 19 for intracellular targets deep into the cytoplasm (anti-keratin). Findings indicate that for extracellular and subplasma membrane epitopes, but not extended intracellular targets, gold nanoparticle-conjugated antibodies provide superior labeling performance to conventional fluorophore-conjugated antibodies. Understanding the advantages and limitations of both nanoparticle and fluorophore-based approaches allows researchers to make informed decisions in selecting the most appropriate tools for their specific imaging assays. Labeling and imaging applications of nanoparticles require a critical evaluation of their advantages and drawbacks in comparison to traditional fluorophores. Subplasma membrane intracellular regions are more favorable for nanoparticle labeling than the bulk cytoplasm containing extended cytoskeletal structures.
Methods
Immunofluorescence
Immortalized human gingival epithelial (Gie-No3B11; T0049, Applied Biological Materials) cells 26 were cultured in 60 mm recessed glass bottom petri dishes (MatTek) for approximately 30 hr at 37C in a CO2 incubator before being fixed. Dishes used for intracellular anti-EGFR IF were fixed using 500 µl of −20C-chilled 100% ACS Grade Methanol (Lab Alley Essential Chemicals), for 5 min at room temperature; all other experiments were performed using 500 µl of 3.7% paraformaldehyde (Thermo Scientific) for 20 min at room temperature. Dishes were rinsed twice with 1× PBS, then permeabilized as follows: 30 min at room temperature with 0.1% Triton-X (Acros Organics) in PBS for anti-keratin 19 dishes; 5 min at room temperature with 0.1% Triton-X in PBS for intracellular anti-EGFR dishes; no permeabilization for extracellular anti-EGFR dishes. All dishes were blocked overnight at 4C in a solution containing 10% normal goat serum (NGS; Abcam), 1% bovine serum albumin (BSA; Sigma), and 0.1% Triton-X in PBS, then rinsed twice with a rinse buffer of 1% NGS, 1% BSA, and 0.1% Triton-X in PBS. Anti-keratin 19 primary antibody (ab192643; Abcam) 27 was diluted 1:200, intracellular Anti-EGFR primary antibody (ab193244, Abcam) 28 and extracellular Anti-EGFR primary antibody (NBP2-52671, Novus Biologicals) 29 were diluted 1:100, each with rinse buffer. All primary antibodies used were rabbit anti-human monoclonal antibodies; intracellular EGFR and keratin 19 antibodies were conjugated with Alex Fluor 488. Dishes were incubated with 500 µl of diluted primary antibody for 90 min at room temperature with agitation in a dark box, then rinsed twice with rinse buffer.
Four secondary antibodies were used, all goat anti-rabbit antibodies, conjugated with TRITC (ab6718; Abcam) 30 , and 40, 10, and 2.2 nm gold nanoparticles (CAS11, Nanopartz), 31 respectively. The fluorophore-conjugated secondary antibody was diluted 1:100 and the nanoparticle-conjugated secondary antibodies were diluted 1:20 in rinse buffer. Dishes were incubated with 500 µl of diluted secondary antibody for 90 min at room temperature with agitation in a dark box, then rinsed twice with rinse buffer. Cells were not cleared after staining. Dishes were then filled with 500 µl of 1× PBS, coverslipped, and imaged on an Eclipse TE300 Inverted Microscope (Nikon) using a 40× oil immersion lens and oil immersion condenser and Photometrics X1 camera (Thermo Scientific) with PVCamTest software version 3.4.183 (Teledyne Photometrics).
Sonication and Heat Application
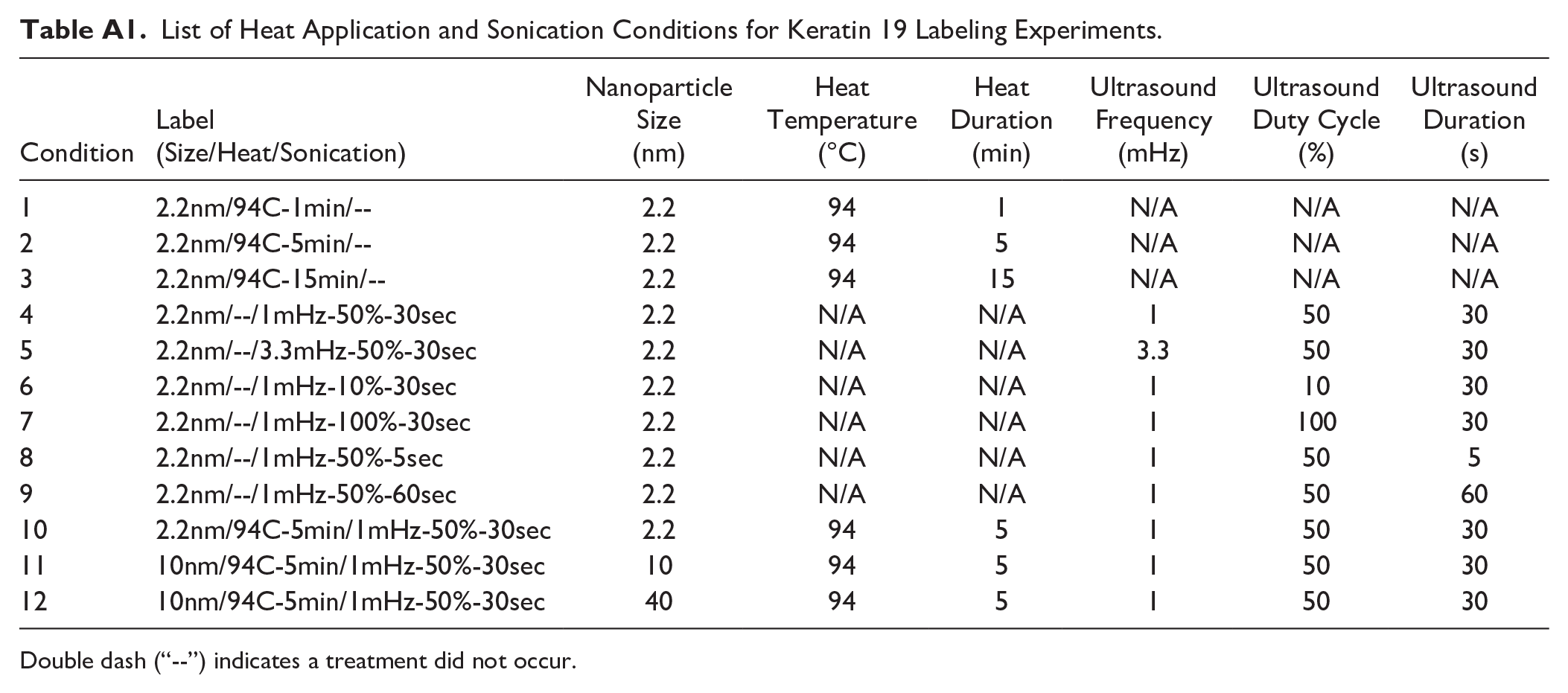
Anti-keratin 19 dishes underwent heat application and sonication tests to determine impacts on nanoparticle labeling. Nine conditions outlined in Appendix Table A1 were tested following permeabilization but before blocking as mentioned above using 2.2 nm gold nanoparticle-conjugated goat anti-rabbit secondary antibody. A combined pretreatment (designated 2.2nm/94C-5min/1mHz-50%-30sec, Table A1) was then performed using levels of heat application and sonication that performed well as single pretreatments, on the 2.2 nm secondary antibodies. The 10 pretreatment conditions were then evaluated by determining the SNR from darkfield microscopy and the best overall pretreatment condition was performed using the 10 and 40 nm secondary antibodies (combined pretreatments designated as 10nm/94C-5min/1mHz-50%-30sec, and 40nm/94C-5min/1mHz-50%-30sec, respectively).
Heat application was performed by submerging the sample dish in a 1× PBS-filled beaker heated to 94C. 32 Plates remained submerged at temperature for durations listed in Table A1 before proceeding with blocking as stated above. Sonication was performed by submerging the sample dish in a 1× PBS-filled beaker at room temperature. Dishes were seated on top of ultrasound-resistant material, Aptflex F28, to minimize impact of indirect ultrasound and the ultrasound wand was partially submerged in the beaker as well, held 6.7 cm from the glass bottom of the sample dish, as previously studied.23,33 All dishes were processed at an intensity of 1 W/cm2 and frequency, duty cycle, and duration were varied depending on condition as listed in Table A1, then followed up with blocking and staining protocol above. When both techniques were performed, heat application preceded sonication.
Post-stain Rinsing and Sonication
Conditions 2.2nm/94C-5min/1mHz-50%-30sec, 10nm/94C-5min/1mHz-50%-30sec, and 40nm/94C-5min/1mHz-50%-30sec from above were then rinsed an additional 3 times with 1 ml 1× PBS and agitated for 30 sec before decanting each rinse and reimaged to ensure unbound nanoparticles were removed. These dishes were then imaged, and underwent sonication in a room temperature water bath for 1 min followed by additional imaging as stated above.
Exposure Time
To test photobleaching effects on the conjugated tags, anti-keratin 19 dishes were exposed to light from their imaging condition for multiple timepoints and reimaged: 0 sec (initial exposure), 10 sec, 30 sec, 1 min, 2 min, 5 min, 10 min, 20 min, and 30 min. Primary fluorophores, secondary fluorophores, and secondary nanoparticles were all subjected to the same time and continuous exposure conditions. Specimens were stored for 1 week at 4C and imaged again with the same microscope and acquisition settings.
Statistical Analysis
Colocalization analysis was completed in Fiji (ImageJ) using the “Colocalization Threshold” and “Coloc 2” plugin with standard thresholds. Micrographs of each secondary antibody were colocalized with the corresponding primary antibody micrograph. Colocalization plots were created in which fluorescence and scattering signal were compared for every signal-containing pixel in the micrographs. The background threshold was set at 50 on an 8-bit (0–255) scale. Several colocalization parameters were then computed, ignoring pixels beneath the background threshold levels for the two channels. Pearson’s and Spearman’s correlation coefficients simply describe the strength of linear and rank-order correlations between the two channels’ signal levels within colocalized pixels. Manders colocalization coefficient m1 describes the normalized percentage overlap of the primary with the secondary antibody (fluorescence and nanoparticle) signals, while m2 describes the inverse situation of secondary antibody signal overlap with primary. 34 The Kendall Tau-b rank correlation parameter is a nonparametric coefficient measure of association of two overlapping datasets, that is closer to 1 when there are more pixel pairs with signal intensities from the two channels in rank order, as opposed to having opposing signal trends (discordant pixel pairs). 35
Signal-to-noise ratios (SNR) were calculated within Fiji using a standardized area to determine the mean signal and standard deviation of background noise of at least 15 distinct regions for each condition tested. SNRs were then averaged to assess each condition. ANOVA and post hoc pairwise comparisons were performed in SPSS Statistics (IBM) using the “One-Way ANOVA,” “Univariate General Linear Model,” and “Independent Sample Nonparametric Test” methods.
Results
Labeling of Keratin 19
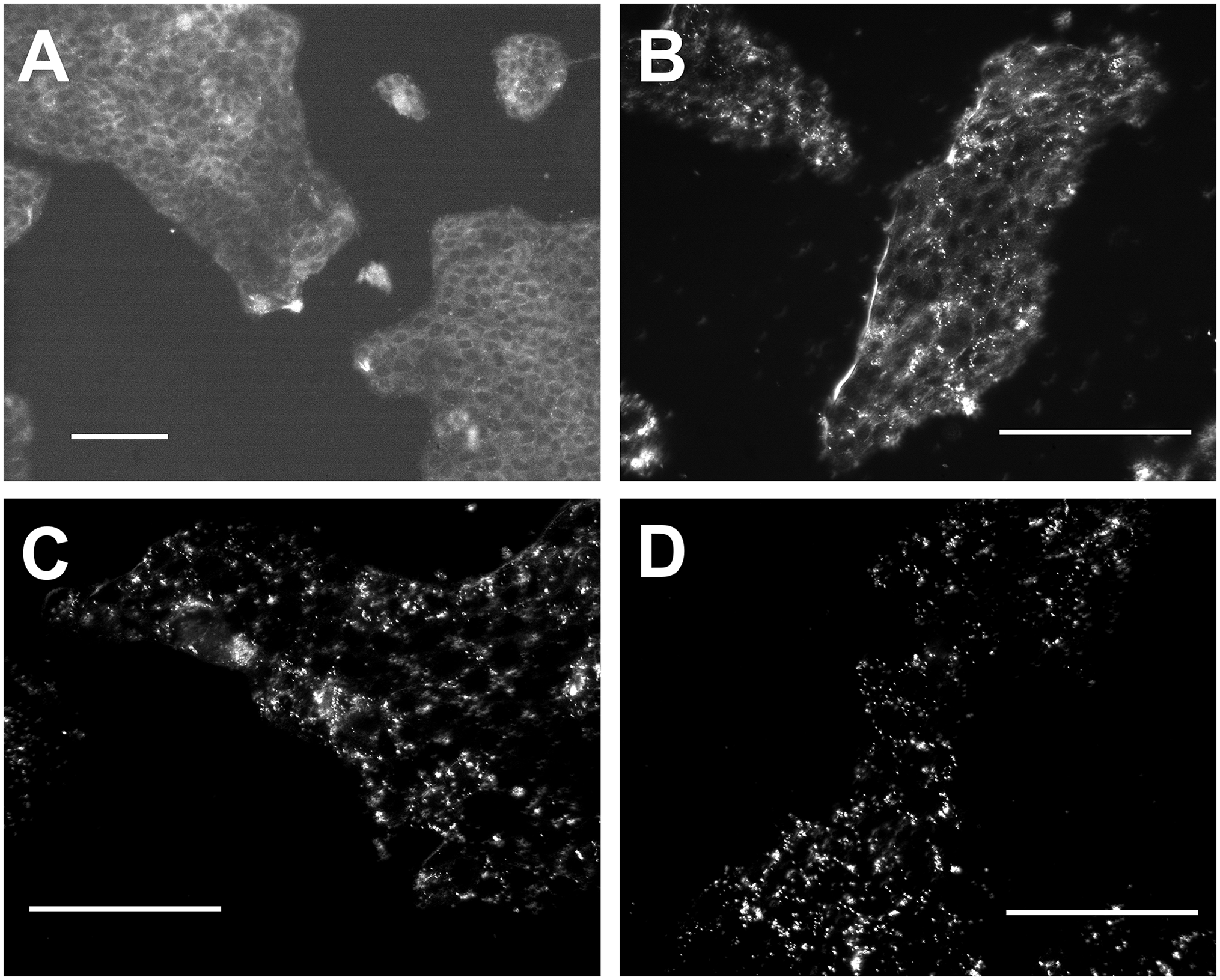
Bright, punctate nanoparticle immunolabeling of keratin 19 in gingival epithelial cells contrasted with continuously distributed filament-like fluorescent label signal with regional variation in brightness. Representative fluorescence and darkfield scattering micrographs revealed labeling and signal intensity differences between goat anti-rabbit IgG secondary antibodies conjugated to TRITC (Fig. 1A), 2.2 nm (Fig. 1B), 10 nm (Fig. 1C) and 40 nm (Fig. 1D) diameter gold nanospheres. The TRITC label produced a lower SNR (30 ± 1 arbitrary units or AU) versus 2.2 nm (164 ± 30 AU), 10 nm (55 ± 18 AU), and 40 nm (162 ± 61 AU) diameter nanosphere labels. In comparison, darkfield scattering signal from the cells not exposed to nanoparticles was 25 ± 11 AU. Label type significantly affected SNR (one-factor ANOVA, F = 24, p<0.001). Specifically, 2.2 nm and 40 nm diameter gold nanoparticle conjugates produced SNR higher than background scattering of the cells (Tukey tests, p<0.001) and higher than the TRITC SNR (p<0.001). Both fluorophore and nanoparticle labels localized to cortical and cytoplasmic compartments, but the TRITC label was continuous and predominantly cortical, whereas the nanoparticle labels were punctate and predominantly cytoplasmic, and rarely located within the image regions containing cell nuclei.
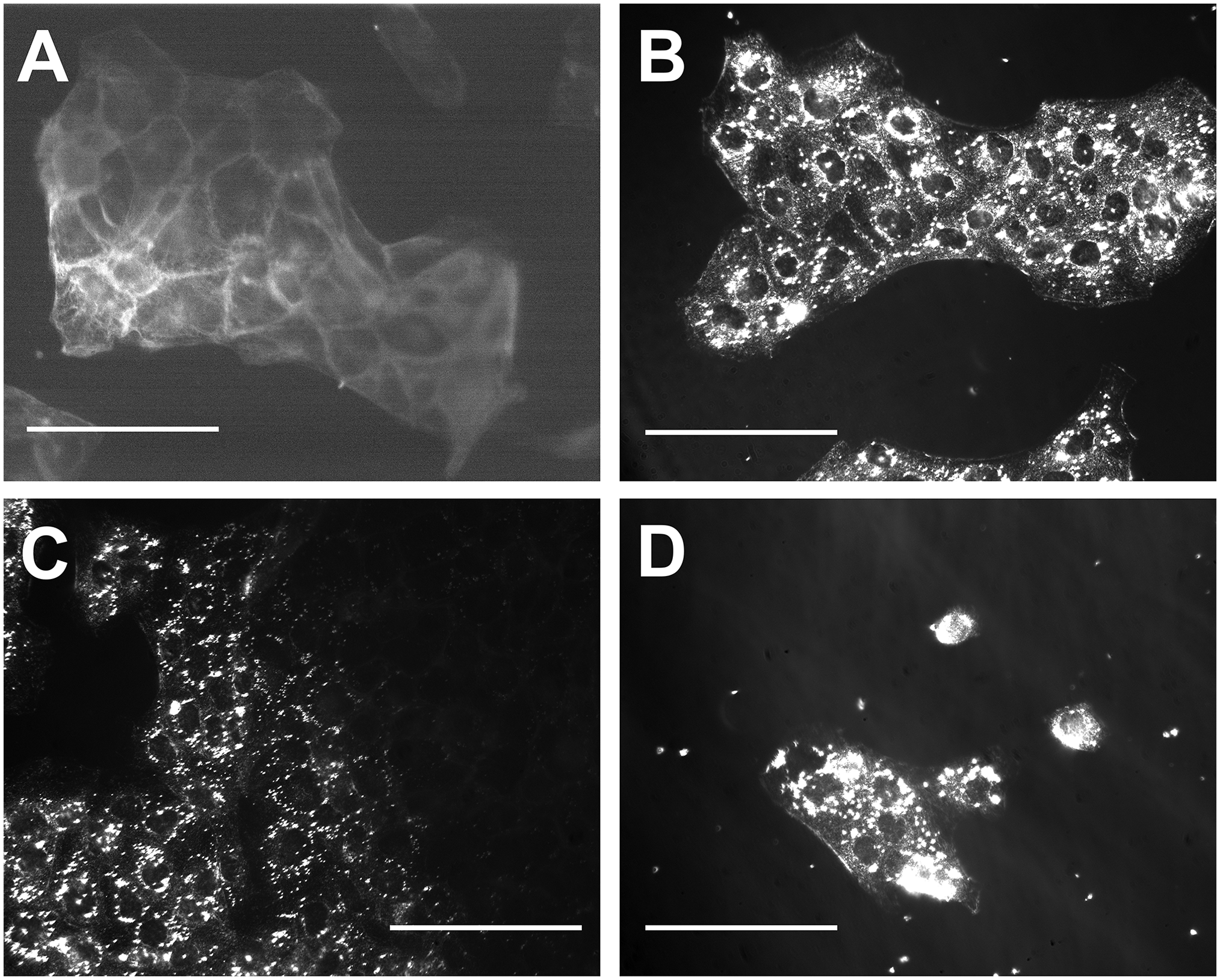
Micrographs of fixed human gingival epithelial cells incubated with a keratin 19 primary antibody. Secondary antibody labels were (A) TRITC-tagged, and conjugated with (B) 2.2 nm, (C) 10 nm, and (D) 40 nm diameter gold nanospheres. All fixed cells were imaged using (A) epifluorescence or (B–D) darkfield microscopy at constant exposure with a double oil immersion 40× magnification objective. Scale bars are indicated. Scale bars indicate 50 µm length.
Signal from imaging tags did not diminish after direct exposure to excitation/darkfield illumination for durations of 10 sec to 30 min, but nanoparticle labels were essentially unchanged after 1 week of slide storage in a refrigerator, whereas the TRITC label intensity was much lower after this interval (data not shown). Colocalization analysis determined a partial spatial overlap of fluorescence and nanoparticle signal (Fig. 2). However, the TRITC-tagged secondary antibody displayed the highest Pearson’s (one-factor ANOVA, F = 9.6, p<0.001) and Manders m2 (F = 3.4, p<0.05) in colocalizing with the AlexaFluor 488-tagged primary antibody. Specifically, TRITC had a higher m2 coefficient than 40 nm conjugates (Tukey test, p<0.05), and a higher Pearson’s coefficient than 2.2 nm (p<0.05) and 10 nm conjugates (p<0.01). Spearman’s, m1, and Kendall Tau-b colocalization parameters were not significantly different between groups.
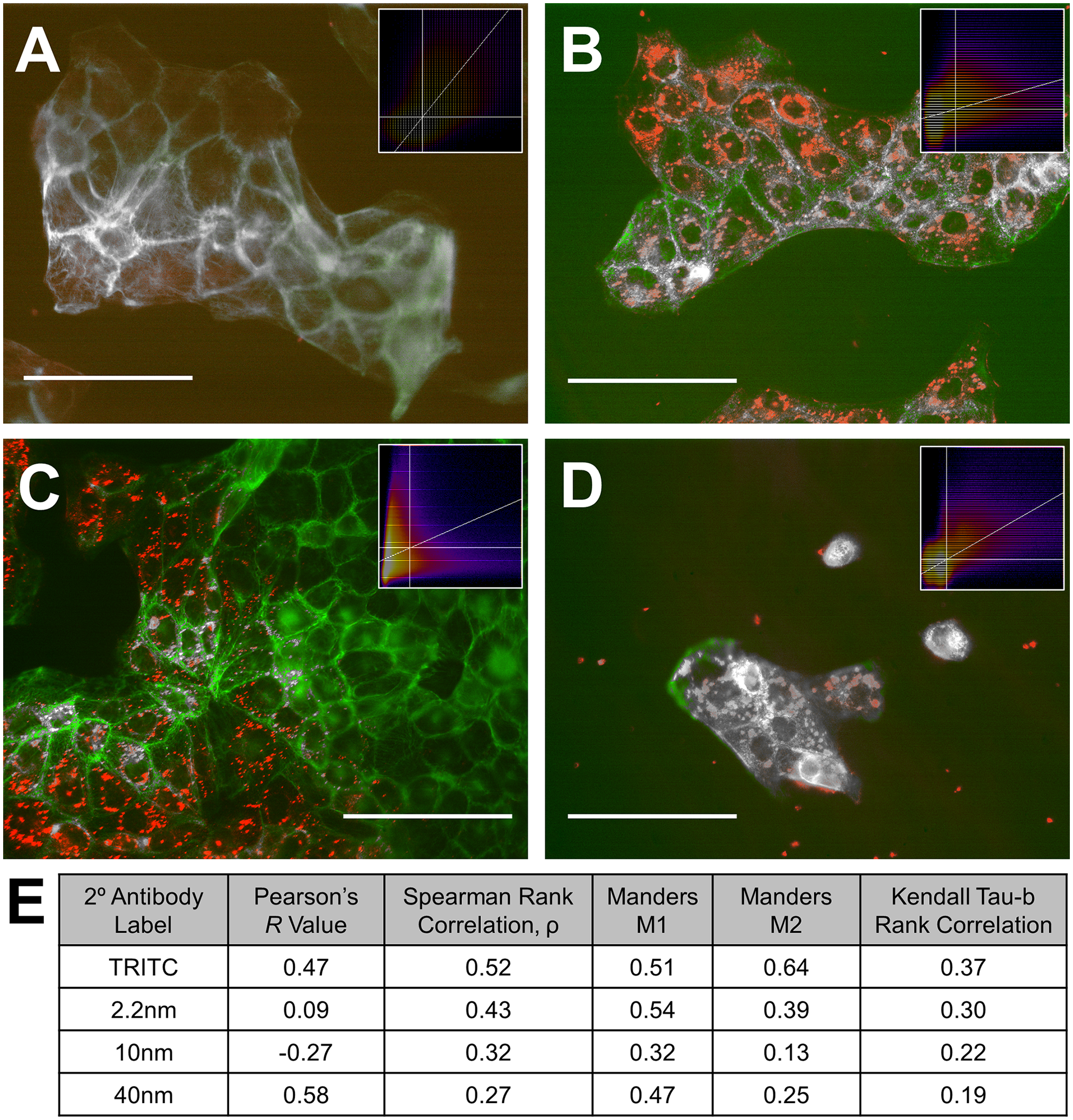
Colocalization of fixed human gingival epithelial cells incubated with keratin-19 primary antibody. Secondary antibody labels were (A) TRITC-tagged, and conjugated with (B) 2.2 nm, (C) 10 nm, and (D) 40 nm diameter gold nanospheres. Secondary antibodies are denoted in red, primary antibody in green, fully colocalized regions in grey. Inlaid: 2D intensity histogram with secondary antibody intensity y-axis and primary antibody intensity x-axis. Auto-assigned thresholds are shown by white lines and the color of each bin indicates the number of pixels with those specific intensity combinations with yellow indicating highest pixel count and dark blue indicating lowest (E) shows a table containing statistical colocalization values for each secondary antibody. Scale bars indicate 50 µm length.
Effects of Sonication and Heat Application as Pretreatment
Sonication and heating during antibody application improved the colocalization of 10 conjugated antibody signals with fluorophore-antibody signal labeling keratin 19 intermediate filaments. Representative darkfield scattering micrographs depict labeling and signal intensity differences between different pretreatment conditions before incubation goat anti-rabbit IgG secondary antibodies conjugated to 2.2 nm (Fig. 3A–C), 10 nm (Fig. 3D) and 40 nm (Fig. 3E) diameter gold nanospheres. For 2.2 nm diameter nanospheres, the optimized heat application (Fig. 3A), sonication (Fig. 3B), and combined pretreatment (Fig. 3C) produced no improvement compared with the no-pretreatment option (181 ± 16 AU, 175 ± 25 AU, and 220 ± 76 AU, respectively, vs. 164 ± 30 AU with no pretreatment, in Fig. 2B). However, the combined pretreatment method enhanced SNR for the 10 nm (186 ± 11 AU vs. 55 ± 18 AU, p<0.001, t-test, Figs. 3D vs. 2C) diameter gold nanospheres. All treated nanoparticle-labeled cells produced significantly higher SNR than the primary antibody-negative control sample (one-factor ANOVA, F = 14.4, p<0.001), but there were no significant differences in SNR between the treatment conditions. The 2.2 nm conjugated secondary antibody colocalized poorly with the AlexaFluor 488-tagged primary antibody via several colocalization statistics, while the 10 nm and 40 nm diameter gold nanoparticles showed increased colocalization as compared with respective no-pretreatment options (Figs. 3F vs. 2E).
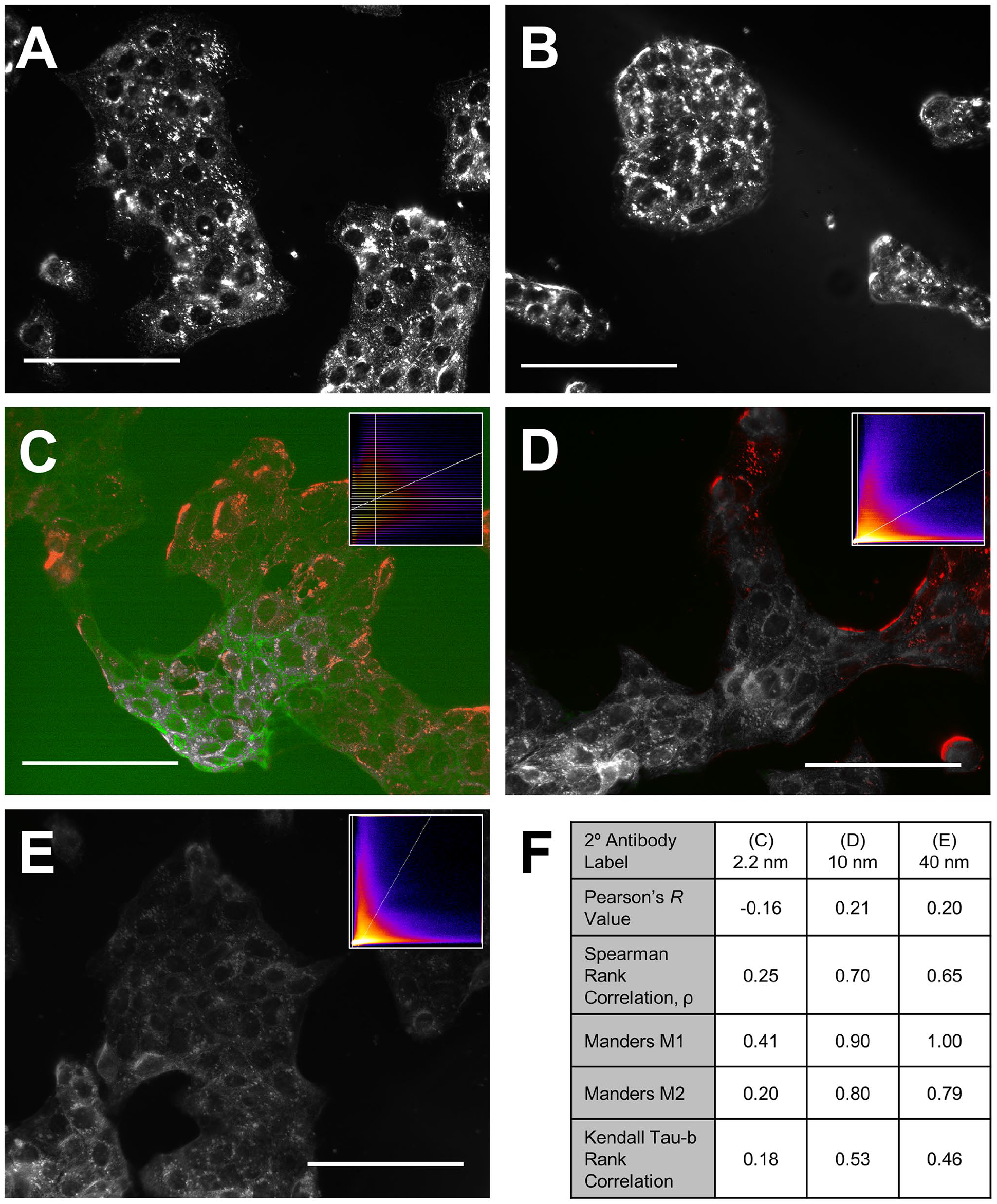
Micrographs and colocalization of fixed human gingival epithelial cells incubated with keratin-19 primary antibody. Secondary antibody labels were conjugated with (A-C) 2.2 nm, (D) 10 nm, and (E) 40 nm diameter gold nanospheres. For colocalization (C-E) secondary antibodies are denoted in red, primary antibody in green, fully colocalized regions in grey. Inlaid: 2D intensity histograms with secondary antibody intensity y-axis and primary antibody intensity x-axis. Auto-assigned thresholds are shown by white lines and the color of each bin indicates the number of pixels with those specific intensity combinations with yellow indicating highest pixel count and dark blue indicating lowest. (F) shows a table containing statistical colocalization values for each secondary antibody. Scale bars indicate 50 µm length.
Intracellular Labeling of EGFR
Nanoparticle immunolabeling of EGFR on an intracellular epitope produced bright, punctate labeling with low background, compared with a much higher background for immunofluorescence labels. Representative fluorescence and darkfield scattering micrographs depict labeling and signal intensity differences between goat anti-rabbit IgG secondary antibodies conjugated to TRITC (Fig. 4A), 2.2 nm (Fig. 4B), 10 nm (Fig. 4C) and 40 nm (Fig. 4D) diameter gold nanospheres. The labeling type significantly affected SNR for intracellular EGFR (one-factor ANOVA, F = 54.2, p<0.001). The TRITC label produced a lower SNR (54 ± 11 AU) versus 10 nm (137 ± 24 AU), and 40 nm (192 ± 26 AU) diameter nanosphere labels (Tukey tests, p<0.001 vs. TRITC), but not 2.2 nm nanoparticle conjugates (90 ± 17 AU, p=0.12 vs. TRITC). TRITC labels displayed minimal EGFR clusters with strong autofluorescence visible within the image, whereas all nanoparticle label images displayed punctate signal with dim darkfield background from cellular structures. The relative intensity of darkfield background was lower with higher nanosphere diameter due to a higher intensity label signal.
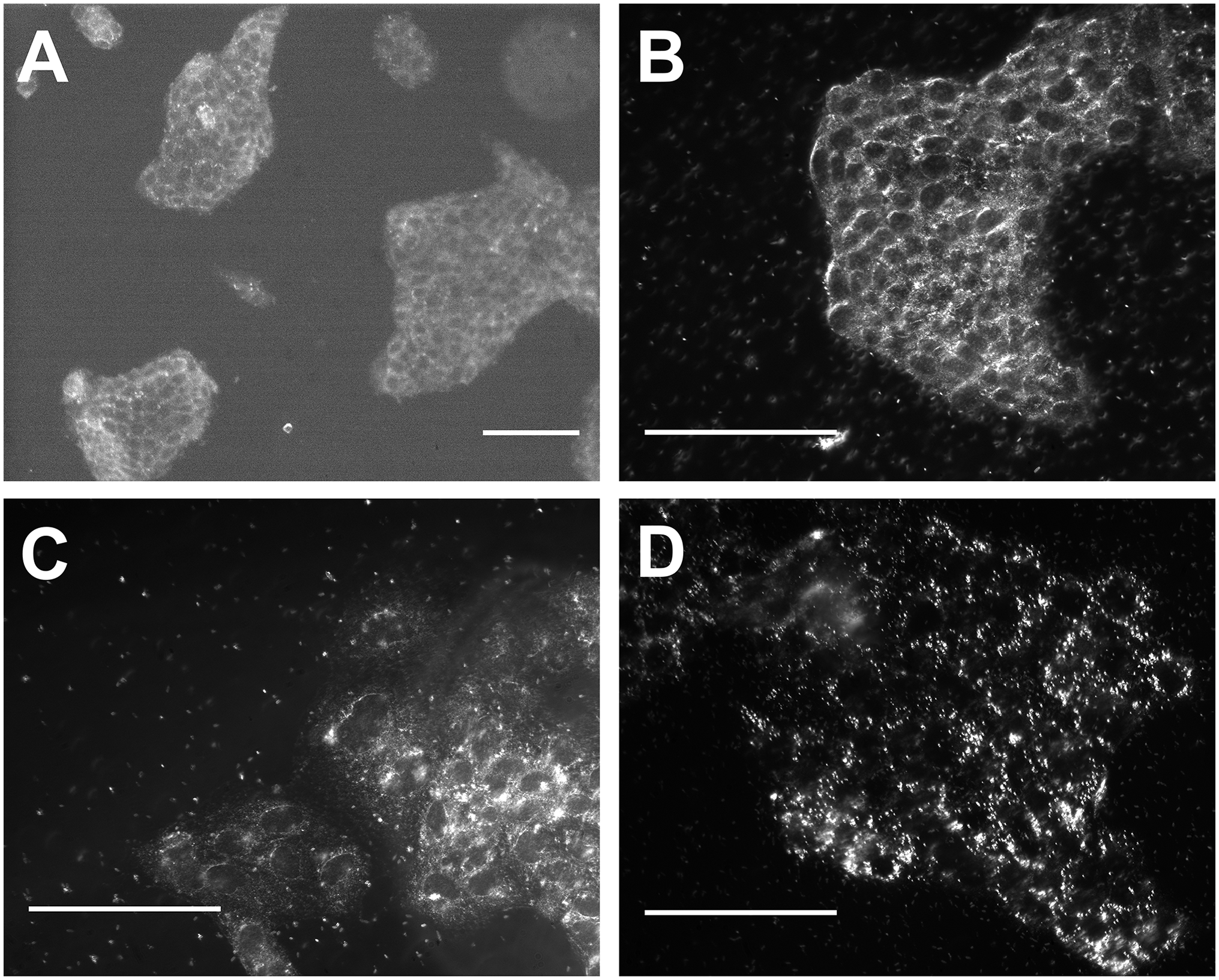
Micrographs of fixed human gingival epithelial cells incubated with an intracellular EGFR primary antibody. Secondary antibody labels were (A) TRITC-tagged, and conjugated with (B) 2.2 nm, (C) 10 nm, and (D) 40 nm diameter gold nanospheres. All fixed cells were imaged using (A) epifluorescence with an LWD 20× magnification objective or (B–D) darkfield microscopy at constant exposure with a double oil immersion 40× magnification objective. Scale bars indicate 50 µm length.
Extracellular Labeling of EGFR
Nanoparticle immunolabeling of EGFR on an extracellular epitope, residue C225, in gingival epithelial cells produced bright, punctate labeling with high SNR, a stark contrast with the primarily cellular background of immunofluorescence labels. Representative fluorescence and darkfield scattering micrographs depict labeling and signal intensity differences between goat anti-rabbit IgG secondary antibodies conjugated to TRITC (Fig. 5A), 2.2 nm (Fig. 5B), 10 nm (Fig. 5C), and 40 nm (Fig. 5D) diameter gold nanospheres. Specifically, the SNR was higher for all nanoparticle conjugates versus TRITC and darkfield imaging of primary antibody-negative controls (one-factor ANOVA, F = 21.0, p<0.001). The TRITC label produced a lower SNR (58 ± 16 AU) versus 2.2 nm (151 ± 47 AU), 10 nm (183 ± 20 AU), and 40 nm (165 ± 44 AU) diameter nanosphere labels. The SNRs of all nanoparticle labeling conditions were not significantly different. TRITC labels displayed minimal EGFR clusters with predominantly cellular morphology visible within the image, whereas all nanoparticles displayed punctate signal and dim darkfield background from cellular structures, becoming negligible in the 40 nm diameter nanoparticle labeling condition.
Micrographs of fixed human gingival epithelial cells incubated with an extracellular EGFR primary antibody targeting C225. Secondary antibody labels were (A) TRITC-tagged, and conjugated with (B) 2.2 nm, (C) 10 nm, and (D) 40 nm diameter gold nanospheres. All fixed cells were imaged using (A) epifluorescence with an LWD 20× magnification objective or (B–D) darkfield microscopy at constant exposure with a double oil immersion 40× magnification objective. Scale bars indicate 50 µm length.
Discussion
The labeling effectiveness of gold nanoparticle-conjugated antibodies for cellular epitopes depends on epitope presentation and location within or on the cell. Keratin 19 presented spatially extended epitopes deep within the cell throughout the cytoplasm, and nanoparticle conjugates labeled this cytoskeletal element poorly. In contrast, conjugates effectively labeled the intracellular epitope of EGFR, located on the cytoplasmic side of the cell membrane. As expected, the extracellular epitope of EGFR was also labeled well by the nanoparticle conjugates. Although 2.2, 10, and 40 nm diameter nanoparticles all produced high signal-to-noise, the 40 nm diameter nanoparticles produced the most intense optical signal, apparent from comparison of signal from each nanoparticle size to the background darkfield scattering from the bulk of the Gie-No3B11 cells. All nanoparticle labels produced higher signal-to-noise than fluorescent labels, which in turn already demonstrate higher SNR than gold standard IHC techniques such as peroxidase staining where SNR has historically been a low performance metric. 36 Nonspecific nanoparticle retention was most apparent in the keratin 19 labeling experiment and was comparatively lower in the EGFR labeling experiments. These data define conditions and targets for effective nanoparticle-antibody labeling, summarized in Fig. 6, and suggest that steric hindrance of target and/or nanoparticle aggregation primarily limit labeling of targets within the core or bulk of permeabilized cells.
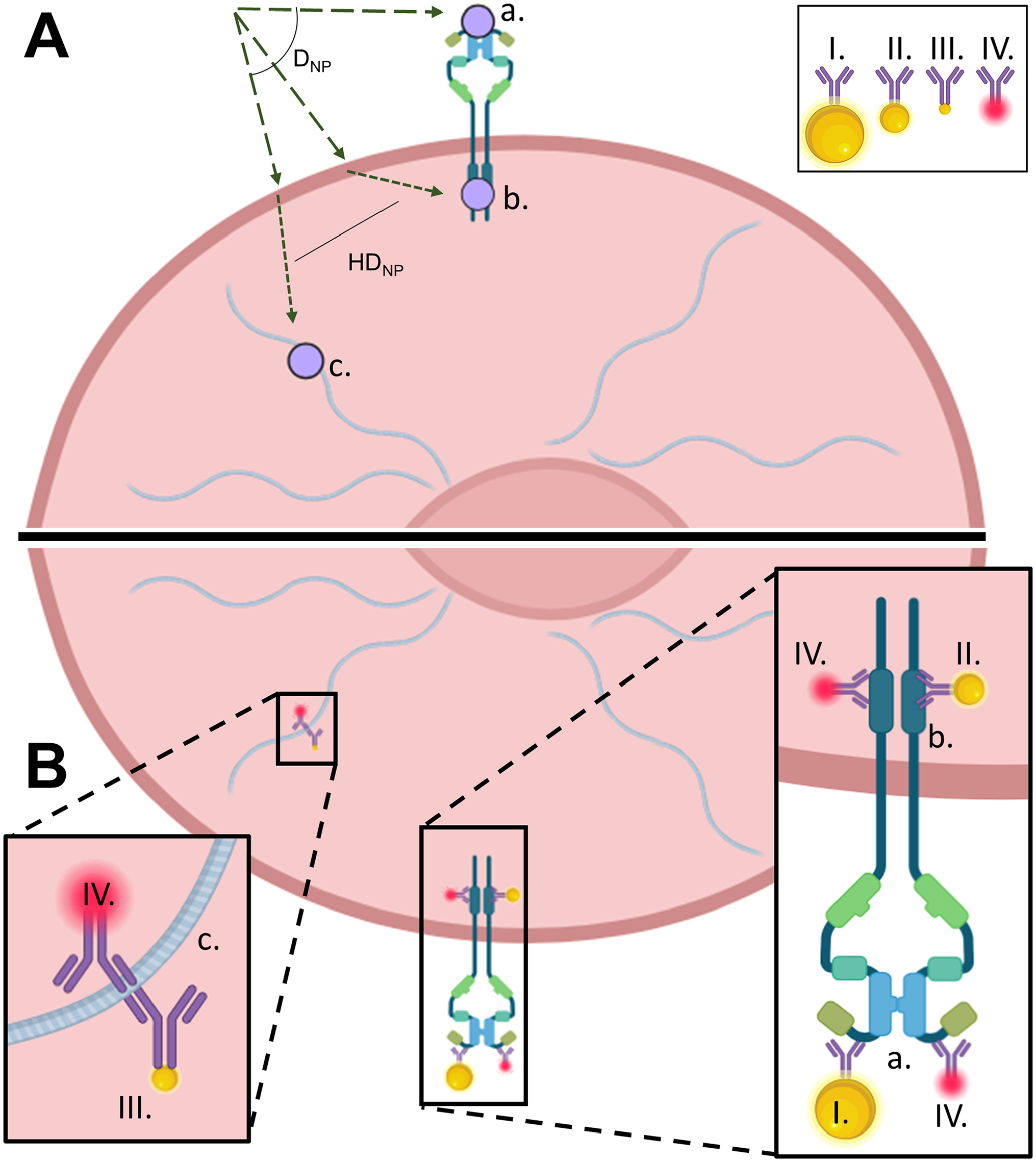
Schematic showing (A) diffusion path of antibody to (a.) extracellular EGFR, (b.) intracellular EGFR, and (c.) cytokeratin-19; and (B) binding sites with the bound secondary antibody attached. Antibody conjugates are denoted in the legend at diameters (I.) 40 nm, (II.) 10 nm, (III.) 2.2 nm, and (IV.) TRITC conjugated. TRITC displayed for molecular size comparison to nanoparticle labels. Primary antibodies not shown.
Poor labeling of keratin 19 by nanoparticle conjugates was independent of nanoparticle size and the application of heat and ultrasound during specimen incubation with the antibodies. Incomplete, punctate labeling of intermediate filaments was not improved by heat treatment that would be expected to retrieve a portion of the antigens made unavailable during fixation. 37 Nor did 2.2 nm diameter nanoparticle label keratin 19 better than 10 nm or 40 nm diameter gold, indicating hindered transport by small pore size was not an issue. Indeed, the 2.2 nm nanoparticle conjugate size compares favorably to that of the primary antibody, which did not have penetration issues. Despite the nanoparticle PEGylation, large patches of signal clustering occurred. While the clustered appearance is consistent with known cluster sizes of EGFR, especially in cancerous or immortalized cell lines, 38 it is not consistent with keratin targets in intermediate filaments. It is likely that nonspecific aggregation and/or crowded labeling of reaction-available keratin sites also occurred. The probability of nonspecific aggregation is enhanced by the rich protein, ionic and macromolecular environment in the permeabilized cell, in which ionic and Van der Waals interactions between nanoparticles, surfactant, and the molecular milieu overcame the steric repulsion of the polyethylene glycol layer on the nanoparticle surfaces. 39 Residual presence of primary antibody in subcellular compartments is also a likely cause of aggregation and darkfield signal flairs, a fact supported by the absence of nanoparticle aggregation in primary antibody-negative control dishes imaged and stock solution.
In contrast to labeling of keratin 19, nanoparticle conjugate labeling of both intracellular and extracellular epitopes on EGF receptors produced similar results, with brighter labeling signal from larger diameter nanoparticles. The consistent pattern of EGFR labeling matches the known distribution of EGFR on the cell membrane, 40 and also indirect immunofluorescence labels (Figs. 4A and 5A). Furthermore, the higher signal intensity of 40 nm versus 10 nm and 2.2 nm diameter nanoparticle conjugates in labeling intracellular EGFR reflects the 4 × 103- and 1 × 108-fold higher scattering cross-section ratio, respectively, of the largest nanoparticles tested here, and is easily distinguished by comparison to the relatively low background darkfield scattering signal of unlabeled portions of the cells. It is possible to recommend 40 nm diameter nanoparticle conjugates for labeling intracellular EGFR. While gold nanoparticles have often been used to label EGFR extracellularly, these findings suggest that intracellular labeling of many cell membrane receptors and membrane-associated proteins is possible with nanoparticle conjugates. Other studies have labeled intracellular and extracellular portions of membrane-associated proteins with nanoparticles, but typically utilize conjugated fluorescent labels for intracellular epitopes, while extracellular targets are more frequently targeted for purposes such as computed tomography (CT) contrast agents41,42 It should be noted that multiple specimens were reimaged after 2 weeks, after storage and also after additional sonication and heating. While the fluorescent labels were significantly degraded, nanoparticle signal remained bright and consistent with the initial pattern and extent of scattering signal, indicating that the nanoparticles were likely bound to primary antibodies rather than loosely trapped within pores of the permeabilized cells. The sensitivity of labeling signal to nanoparticle size was highest for intracellular EGFR binding. It is likely that all nanoparticle-antibody conjugates labeled EGFR effectively, so that signal depended on nanoparticle scattering cross-section, whereas keratin 19 was poorly labeled by aggregated nanoparticles.
The strengths and limitations of this study affect the conclusions able to be drawn from the data. All experiments used indirect immunolabeling with antibodies, and PEGylated spherical gold nanoparticles conjugates. The success of labeling EGFR and failure of labeling keratin 19 by nanoparticle methods should not be interpolated to other kinds of labeling molecules, or other compositions, sizes and shapes of nanoparticles. Nevertheless, the failure of accurate nanoparticle labeling of keratin filaments is consistent with the known propensity of nanoparticles to aggregate and/or to be sterically blocked in the complex environment of the permeabilized cell. 43 Immunogold labeling has a long history in ultrathin cell and tissue sections in transmission electron microscopy, 44 suggesting that such conjugates can traverse a small distance in fixed tissue to accurately label a target. Very few studies have attempted to label intracellular targets in eukaryotic cells with colloidal nanoparticles, likely due to issues with effective labeling. Although heat and sonication were not enough to consistently overcome these issues leading to improved labeling in fixed gingival epithelial cells, intracellular epitopes of EGFR on these cells were well labeled by nanoparticle conjugates, indicating that aggregation was not a problem for subplasma membrane labeling assays.
Overall, nanoparticles may not serve as substantial replacements for fluorophores in all use cases due to significant hindered diffusion across the cell (see HDNP, Fig. 6), nor do they serve well for labeling of highly abundant cellular components such as cytoskeleton due to poor distribution of the nanoparticle conjugates to the keratin filaments, even after PEGylation. In cases where extracellular or submembrane intracellular labeling assays are needed, nanoparticles offer several advantages over fluorophores including increased SNR, enhanced resolution without cellular background, and prolonged signal stability. These findings indicate nanoparticle labeling may best serve niche cases such as live cell imaging, allowing tracking of membrane-bound molecules such as EGFR and RAS with more dependable signal strength than fluorophores may offer. Nanoparticle labeling may also serve to provide quantitative information through subsequent epitope capture and examination through mass spectrometry or enzyme-linked immunosorbent assay (ELISA). At the same time, if aggregation issues can be mitigated in the future, nanoparticle-antibody conjugates could be used on a wide scale for labeling of any epitope in fixed, permeabilized cells and tissues.
Footnotes
Appendix
List of Heat Application and Sonication Conditions for Keratin 19 Labeling Experiments.
| Condition | Label |
Nanoparticle Size |
Heat Temperature |
Heat Duration |
Ultrasound Frequency |
Ultrasound Duty Cycle |
Ultrasound Duration |
|---|---|---|---|---|---|---|---|
| 1 | 2.2nm/94C-1min/-- | 2.2 | 94 | 1 | N/A | N/A | N/A |
| 2 | 2.2nm/94C-5min/-- | 2.2 | 94 | 5 | N/A | N/A | N/A |
| 3 | 2.2nm/94C-15min/-- | 2.2 | 94 | 15 | N/A | N/A | N/A |
| 4 | 2.2nm/--/1mHz-50%-30sec | 2.2 | N/A | N/A | 1 | 50 | 30 |
| 5 | 2.2nm/--/3.3mHz-50%-30sec | 2.2 | N/A | N/A | 3.3 | 50 | 30 |
| 6 | 2.2nm/--/1mHz-10%-30sec | 2.2 | N/A | N/A | 1 | 10 | 30 |
| 7 | 2.2nm/--/1mHz-100%-30sec | 2.2 | N/A | N/A | 1 | 100 | 30 |
| 8 | 2.2nm/--/1mHz-50%-5sec | 2.2 | N/A | N/A | 1 | 50 | 5 |
| 9 | 2.2nm/--/1mHz-50%-60sec | 2.2 | N/A | N/A | 1 | 50 | 60 |
| 10 | 2.2nm/94C-5min/1mHz-50%-30sec | 2.2 | 94 | 5 | 1 | 50 | 30 |
| 11 | 10nm/94C-5min/1mHz-50%-30sec | 10 | 94 | 5 | 1 | 50 | 30 |
| 12 | 10nm/94C-5min/1mHz-50%-30sec | 40 | 94 | 5 | 1 | 50 | 30 |
Double dash (“--”) indicates a treatment did not occur.
Acknowledgements
We thank Khanh Ly, PhD, Hanadi Alqosiri, PhD, and Elijah Feret, MS, at The Catholic University of America, for help with human Gie-No3B11 cell culture.
Competing Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
R.N.B. and C.B.R. conceptualized and designed experiments. R.N.B. performed experiments and analysis; C.B.R. and G.N. supervised data acquisition and C.B.R. supervised analysis. All authors contributed to interpretation of data. R.N.B. and C.B.R. wrote the manuscript and all authors were involved in revision. All authors have read and approved the final manuscript.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
