Abstract
Polypeptide growth factors, including epidermal growth factor (EGF), play a central role in regulating hepatocyte growth both in vivo and in primary culture. To characterize EGF gene expression in the pathogenesis of regenerative cirrhotic fibrosis, we employed biotinylated antisense oligonucleotide probes to localize hepatic mRNA transcripts in situ. In control tissue and regenerative hepatic nodules, EGF receptor (EGFR) mRNA transcripts were expressed constitutively. In contrast, oligonucleotide probes targeting the human EGF coding region showed that EGF transcription was extremely low in control liver but was highly elevated and localized to regenerative hepatic nodules and bile duct epithelia of cirrhotic liver. To determine whether EGF mRNA accumulation accompanied a comparable increase in the EGF peptide, we performed immunohistochemistry using an antibody specific for the nonprocessed peptide aminoterminus. We observed that positive localized EGF staining paralleled its mRNA transcript. These results indicate that EGF upregulation is a characteristic of cirrhotic liver disease and suggest that persistent de novo ligand synthesis and its signaling contribute to an autocrine-mediated hepatocyte proliferation within the regenerative nodule.
L
Hepatocytes express the tyrosine kinase EGF receptor (EGFR) and quickly respond to exogenous EGF in vivo (Milani et al. 1991) and in primary culture (Block et al. 1996; Ismail et al. 1991). EGFR signaling activates a specific group of signal transducers and activators of transcription (STATS) (Ruff-Jamison et al. 1993), increases transcription of a characteristic set of early growth response genes (Kruijer et al. 1986), and effects the mobilization of an intrinsically low rate of hepatic DNA replication. Previously, we have shown that in the first 30 min of rodent regenerative liver growth induced by two-thirds hepatectomy, EGF gene transcription is rapidly upregulated (G0-G1) but diminishes below baseline values before the first wave of cell division (G1-S) (Mullhaupt et al. 1994). These results and others (Fausto et al. 1995) suggest that hepatic EGF gene activation constitutes an autocrine mechanism regulating the hepatocyte pre-replicative growth program. In this study we investigated whether increased EGF expression is associated with hepatic regenerative nodules in late-stage cirrhosis. For these studies, we generated highly labeled antisense probes via direct incorporation of biotinylated thymidine phosphoramidites into the oligonucleotide synthesis. Using these probes in in situ hybridization, we show comparable levels of EGFR gene expression in control and cirrhotic liver sections. In contrast, EGF mRNA expression (as assayed by in situ hybridization followed by tyramide deposition-mediated signal amplification) is rare in control tissue but can be detected at high levels in regenerative nodules and associated bile duct epithelium of cirrhotic liver. To determine whether up-regulation of EGF transcripts is accompanied by an increase in intrahepatic EGF peptide levels, we performed parallel immunohistochemical localization with antiserum specific for the EGF pre-pro-peptide. Positive immunostaining of the EGF precursor was similar to EGF mRNA expression. These findings are consistent with the notion that EGF expression comprises a key hepatocyte growth regulatory mechanism and indicate that persistent ligand synthesis and EGFR signaling contribute to the autocrine dysregulation of regenerative nodules in cirrhotic liver disease.
Materials and Methods
Tissue Specimens
Liver specimens from three transplant patients with cirrhotic liver disease and control specimens (livers rejected for transplantation) were obtained at surgery according to UCSF policies of the Committee on Human Research at UCSF, in compliance with NIH guidelines. Tissue pieces were immediately placed in ice-cold 4% formaldehyde (freshly prepared from paraformaldehyde), dissolved in Dulbecco's Mg2+/Ca2+-free PBS, and were fixed for 24 hr at 4C. Tissues were dehydrated in increasing ethanol concentrations, cleared in xylenes, and embedded in paraffin by standard methods. Sections (5-μm) were collected on positively charged microscope slides (Superfrost Plus; Fisher Scientific, Pittsburgh, PA).
DNA Oligonucleotide Probes
Oligonucleotides were synthesized with an ABI 390 synthesizer. Biotin-dT was incorporated by programming the addition of 5′-dimethoxytrityloxy-5-N((4tbutylbenzoyl)-biotinyl)-aminohexyl) 3′acryimido) 2′deoxy-U-3′-2-cyanoethyl-N,N-diisopropyl)-phosphoramidite (Glen Research; Sterling, VA). After column elution, oligos were deprotected overnight at 58C, dried in vacuo, resuspended in H2O, adjusted to 50% deionized formamide (pH 7.0), heated to 65C for 5 min, and fractionated by electrophoresis on a 15% acrylamide/Bis (30:1), 7 M urea, 1 × TBE gel. Bands were identified by UV shadowing, excised, and eluted in H2O, precipitated twice with 3 volumes of ethanol, and stored in H2O at −20C. Oligo sequences (bT, biotinylated thymidine) were selected from the GeneBank (NCBI) database: human EGF antisense: bTTACAAAGCACTGTbTGTCCCAATTTGGGTbTT (4325-4295) and AbTGTAGCCAACAACACAGbTTGCATGCATACTTGbTC (3459-3425); human EGF sense probe bTAATGGAGCAAGCTbTTCATATGCCCTCCTA (3915-3945). The human EGF receptor antisense, GGAGCGCTGCCCCGGCCGTCCGG(bT)3 (197–220) and sense, CCGGGACGGCCGGGGCAGCGCTC(bT)3 (476-499), were purchased from Research Genetics (Huntsville, AL).
In Situ Localization of EGFR mRNA Using Biotin-labeled Oligonucleotide Probes Detected by ABC-Peroxidase
Sections were dewaxed with Safeclear (CMS; Houston, TX), rehydrated in ethanol, washed with PBS, and treated with 0.2 N HCl for 20 min, followed by 1.5% H2O2 for 30 min, then by 0.3% Triton X-100 for 15 min, and incubated with 1 μg/ml proteinase K (Amresco; Solon, OH) for 30 min at 37C. After washing with 0.1 M triethanolamine buffer, pH 8.0, sections were acetylated for 10 min in 0.25% acetic anhydride in triethanolamine buffer, incubated with 40% formamide in 1 × SSC at 37C for 2 hr, and air-dried. One pmole biotinylated probe was diluted in 100 μl hybridization buffer containing 40% formamide, 1 × SSC, 10 × Denhardt's solution, 0.5% SDS, 0.5% sodium pyrophosphate, yeast tRNA (500 μg/ml) in 10 mM Tris buffer, pH 7.4, denatured at 85C, chilled on ice and applied to the sections, and hybridized in a humidified chamber at 40C. After 12 hr the sections were washed at room temperature (RT) (5 ml/ slide) with 2 × SSC, followed by 1 × SSC and 0.25 × SSC for 30 min each at 37C. Sections were briefly washed in TBS (10 mM Tris, pH 7.6, containing 500 mM NaCl), blocked with 4% BSA, 0.5% teleostean gelatin, 0.1% Tween-20 in TBS for 30 min, and incubated with ABC peroxidase (Vector; Burlingame, CA) for 30 min, followed by three-min washes in blocking buffer and three-min washes in TBS. Peroxidase activity was revealed with DAB substrate (5 min), purchased from Qual-Tek Laboratories (Santa Barbara, CA). The sections were washed with several changes of distilled water, counterstained lightly with Carazzi's hematoxylin, dehydrated with alcohol and xylenes, and coverslipped.
Localization of EGF Expression by In Situ Hybridization Using Biotin-labeled Oligonucleotide Probes Detected by Tyramide-mediated Signal Amplification
The conditions of pretreatment, hybridization, and posthybridization washes for EGF mRNA were essentially as described above, with the following modifications. Hybridization washes were followed by a brief wash in TBS and sections were blocked with 4% BSA, 0.5% teleostean gelatin, 0.1% Tween-20 in TBS for 30 min. Sections were then incubated with ABC-peroxidase (Vector) for 30 min, followed by three 5-min washes in blocking buffer, and were incubated with biotinylated tyramide (Dako; Carpinteria, CA) for 15 min, followed by ABC-peroxidase for 15 min. After three 5-min washes with blocking buffer and three 5-min min washes in TBS, peroxidase activity was revealed with DAB substrate for 5 min. Sections were then washed with several changes of distilled water, counterstained lightly with Carazzi's hematoxylin, dehydrated with alcohol and xylenes, and coverslipped. For all hybridizations, a set of controls was used to establish hybridization specificity: (a) sense control probes did not yield signal above background; (b) omission of the biotin-labeled antisense probes resulted in no signal; and (c) omission of ABC-peroxidase resulted in no signal, indicating the specificity of the signal amplification procedure.
Immunohistochemical Detection of Pro-EGF Peptide
Antiserum to the human EGF pre-pro-peptide amino acids 1-21 was a kind gift of Dr. B Mroczkowski and has been characterized elsewhere (Mroczkowski and Reich 1993). Five-μm sections were collected on microscope slides (Superfrost Plus; Fisher Scientific) and, after deparaffinization and rehydration, antigen retrieval was performed by micro-waving in 10 mM citrate buffer, pH 6.0. Nonspecific antibody binding was prevented by preincubation with blocking buffer: 2% BSA, 0.5% teleostean fish skin gelatin (Sigma Chemicals; St Louis, MO), 0.1% Tween-20, and 500 mM NaCl in Tris, pH 7.6, for 30 min at RT. Sections were incubated overnight at RT in 100 μl blocking buffer with a 1:125 antibody dilution. After five washes in blocking buffer, bound antibody was detected with biotinylated goat anti-rabbit IgG, followed by ABC-peroxidase conjugate (Vector). Peroxidase activity was developed with DAB substrate (5 min) and sections counterstained with hematoxylin before mounting. Omission of the first antibody resulted in no signal detection in either control or cirrhotic liver sections.
Results
Localization of EGF Receptor mRNA in Control and Cirrhotic Human Liver
To localize the cellular distribution of EGFR gene expression in human liver, we used an antisense EGFR biotinylated oligonucleotide probe (Radinsky et al. 1993) that was complementary to a segment of the receptor mRNA coding region. As shown in Figure 1A, after stringent washing and color development, hybridization of the EGFR antisense probe to its target transcript could be clearly resolved in control hepatic sections, in which staining displayed a cell specific pattern restricted to hepatocytes. Kupffer and endothelial cells did not contain detectable levels of the EGFR transcript. In the individual lobules, EGFR mRNA expression was nonuniformly distributed, showing a peripheral bias (Figure 1A). To verify the specificity of the hybridization/detection procedure, parallel control sections were hybridized with a sense-control biotinylated EGFR probe, which did not produce detectable staining (Figure 1B).
We next compared the EGFR mRNA expression pattern observed in control tissue to sections obtained from advanced cirrhosis. Here, EGFR mRNA staining was evident throughout the cirrhotic tissue (Figure 1C), displaying a higher intensity bias in the smaller nodules and an increased concentration at the nodular periphery (Figure 1C). Moreover, positive signals were evident throughout the highly proliferative bile duct epithelium but were absent from the internodular fibrotic matrix. In all cases, the EGFR mRNA signal was restricted to the cytoplasm (Figure 1D).
Localization of EGF mRNA in Control and Cirrhotic Liver Using Biotin-labeled Oligoprobes Followed by Tyramide-catalyzed Signal Amplification
In previous studies we have demonstrated that a low constitutive level of EGF expression was detectable in rodent liver and was rapidly upregulated in the immediate-early phase of regenerative liver growth (Mullhaupt et al. 1994). To examine whether an increase in EGF expression could also be correlated with the regenerative nodular expansion of human cirrhosis, we compared the cellular distribution of EGF gene transcripts in control and diseased liver sections. To target hepatic EGF mRNA transcripts, we designed biotinylated oligonucleotides complementary to the coding region of the processed mature EGF peptide. With the use of two different biotin-labeled antisense oligonucleotide probes, staining for the EGF mRNA was essentially negative in control liver (not shown) but could be identified at a low albeit significant level in cirrhotic sections when ABC-peroxidase, followed by DAB substrate alone was used for detection (Figure 2A). To increase the detection sensitivity of the biotin oligo-mRNA hybrids, we applied a tyramide-deposition mediated signal amplification procedure which affords a 100- to 1000-fold signal increase (Adams 1992; Speel et al. 1999). With this method, hepatocytes remained negative for EGF mRNA in the control samples, although weak staining could be discerned in isolated bile duct cells (Figure 2B). In contrast, with the tyramide deposition-mediated signal amplification procedure, EGF mRNA accumulation was clearly evident in the cytoplasm of the cirrhotic hepatocytes (Figures 2C and 2D). EGF staining was highly localized to regenerative nodules and the presumptive proliferative bile ductules (Figure 2D). Staining appeared most concentrated at the nodule periphery. In cirrhotic liver, the EGF mRNA signal was more intense in the newly forming bile epithelium (Figure 2D). No EGF mRNA staining was detected in the fibroblasts of the internodular matrix (Figures 2C and 2D). Similar results were obtained with both EGF antisense oligoprobes. To verify the hybridization pattern, parallel sections were hybridized to an EGF sense control probe and were negative (not shown).
Detection of the EGF Pre-pro-peptide in Human Cirrhotic Liver
We next carried out immunohistochemical localization to determine whether the regenerative parenchyma produced detectable levels of the EGF peptide. Because the hepatocyte EGF receptor can serve as an uptake module for processed EGF (see Discussion), we used an antiserum raised against the pro-peptide region, amino acids 1-21 (Mroczkowski and Reich 1993), which discriminates the precursor from the endocytosed mature EGF peptide. No immunoreactivity could be detected in control liver samples (not shown). However, the regenerative nodules in cirrhotic liver sections stained positive with the pro-peptide antiserum (Figure 3). Remarkably EGF propeptide staining appeared nonuniform across the lobules, where it was seen with a distinct variation in cell-to-cell intensity. Fibroblasts in the adjoining internodular fibrotic matrix were negative. A less intense staining was consistently seen in the presumptive proliferative hepatic ductules, which in many cases exhibited a slight luminal bias (Figure 3).
Discussion
In this work we show that the polypeptide mitogen EGF and its cell surface receptor are expressed differentially in control and cirrhotic human liver. In normal tissue, receptor transcripts appear dispersed throughout the parenchyma, whereas its cognate ligand, i.e., EGF, remains at nondetectable levels. In contrast, in advanced cirrhosis, a continuous receptor synthesis is accompanied by EGF expression that is localized to hepatocytes and proliferative bile epithelia. Furthermore, with an EGF precursor-specific antiserum, our results show that nascent peptide synthesis parallels its mRNA throughout the regenerative tissue. Collectively, these results reveal that EGF and its receptor are synthesized coordinately with nodule expansion and suggest that this co-expression of a mitogenic cell surface receptor and its associated binding ligand may consititute an autocrine growth mechanism of cirrhotic liver disease.
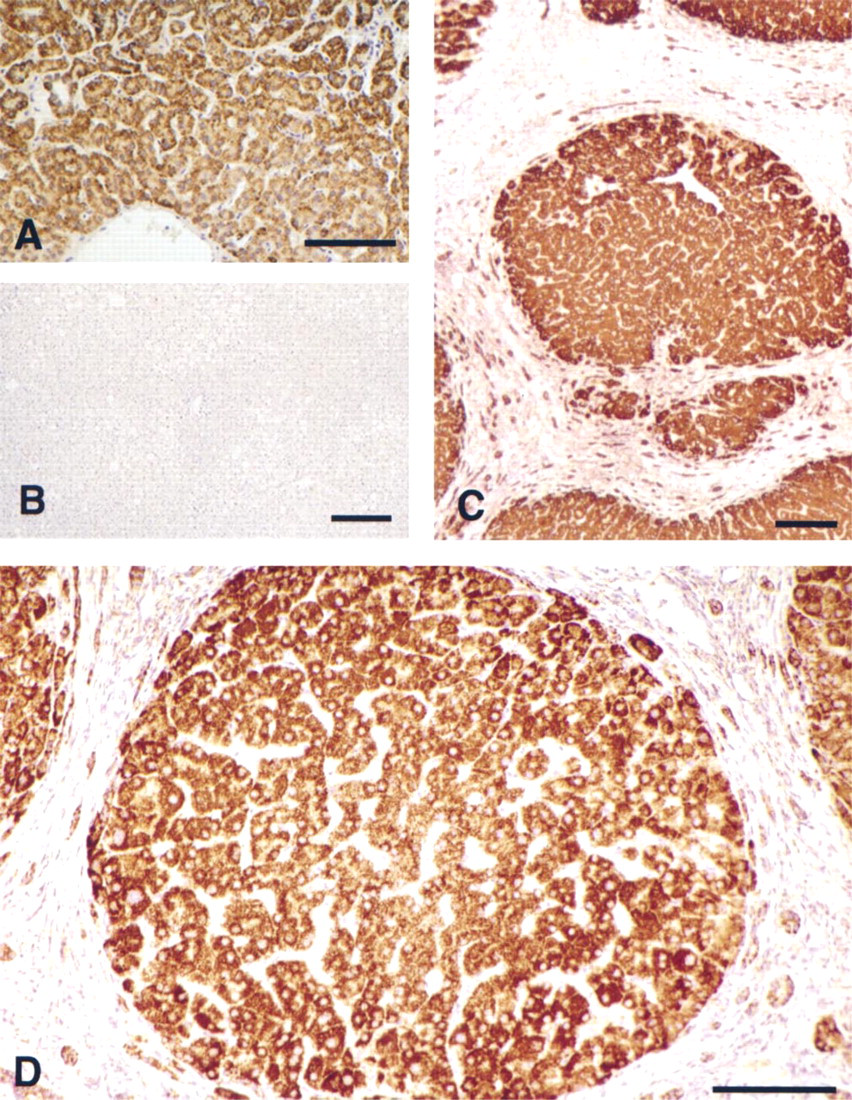
EGFR mRNA expression in control and cirrhotic liver. A human biotinylated EGFR antisense oligo targeted to the receptor coding region was hybridized to liver sections and detected with ABC-peroxidase. Peroxidase activity was revealed with DAB substrate. (
A number of studies have shown that both quiescent and proliferative hepatocytes maintain significant levels of the transmembrane EGF receptor, although its functional role in liver homeostasis has remained controversial (Chabot et al. 1986). In normal tissue, EGFR exhibits a high affinity for EGF and is believed to operate primarily as a mechanism to effect the blood-to-bile clearance of its serum binding ligands (St Hilaire et al. 1983; Burwen et al. 1984; Wollenberg et al. 1989). On the other hand, in the early phases of liver injury or after resection, EGF surface receptor numbers decrease and acquire a lowered ligand affinity, which is postulated to mediate signaling of hepatocyte cell division (Earp and O'Keefe 1981; Wollenberg et al. 1989; Oguey et al. 1992). This apparent inverse relationship among receptor mass reduction, acquisition of a lowered binding affinity, and hepatocyte growth induction is not fully understood. However, it remains possible that endogenous ligand occupation may mask true receptor numbers (thus affecting measured binding constants). Furthermore, because EGFR comprises a heteromeric complex (Schlessinger 1989), rapid compositional changes (i.e., cErb2,3,4) may transmodulate receptor specificity and redefine the downstream signaling pathways with direct effects on hepatic cell replication.
Our present studies are in agreement with previous work that has shown relatively high receptor transcript levels occurring throughout the control hepatic parenchyma. However, comparison of these results to the staining pattern in cirrhotic sections indicates that receptor transcript density actually increases in localized regions of the diseased tissue, suggesting receptor gene upregulation. Because the fibrotic/parenchymal tissue ratio is relatively high in advanced cirrhosis, final receptor levels, when normalized to accumulated tissue mass, may be decreased overall.
As early as 1973 it was noted that EGF administration to normal liver transiently increases the hepatic labeling index and, more recently, has been shown to directly activate receptor-dependent STAT signaling pathways in situ (Bucher et al. 1977; Ruff-Jamison et al. 1993). Although such studies have indicated a mitogenic role for EGF in quiescent tissue, efficient and continual initiation of cell division appears to depend on tissue alterations that produce partial hepatic destruction. These studies have led to the theory that growth factor signaling of hepatic cell division is ineffective unless it is preceded by an as yet unknown upstream priming event (Webber et al. 1994).
Attempts to correlate changes in circulating pools of EGF with liver growth induction have thus far proved equivocal, and only a limited effect on liver growth has been reported after sialoadenectomy (Lambotte et al. 1997). To determine whether ligand signals may arise from a direct (autocrine) hepatic route, we previously reported that a low steady-state level of EGF expression could indeed be demonstrated in rodent liver (Mullhaupt et al. 1994) and that ligand expression rapidly increased over 10-fold in the immediate-early (G0-G1) time period after two-thirds HPX, returning to basal levels before cell replication (Mullhaupt et al. 1994). Such findings indicated a direct role for EGF in the early stages of the regenerative response, because previous data have shown that TGF-α (which also binds EGFR) is increased only after the first wave of regenerative cell division (Fausto et al. 1995). Further recent evidence using a rodent bile duct ligation model (Napoli et al. 1997) has suggested a role for EGF in cirrhotic disease. In this setting, progressive fibrosis and nodule formation were accompanied by increased bFGF, HGF, and a 25-fold increase in EGF mRNA at Day 21 post ligation. Interestingly, in this model and others (Polimeno et al. 1995; Harada et al. 1996) TGF-α increases were not observed. This suggests that early signaling pathways are persistent in chronic cirrhosis and may abrogate the sequential downstream pathways observed in the two-thirds HPX model (Fausto 1994).
The primary focus of our present study was to investigate whether EGF gene expression could be identified in human liver and whether it may exhibit up-regulation in human regenerative liver disease. As we have shown here, in contrast to rodent liver, human liver EGF transcripts are virtually undetectable in control liver, with only occasional weak staining in the bile epithelium. In contrast, advanced cirrhotic tissue showed staining for EGF mRNA, which was most prominent at the nodule periphery and notable within presumptive proliferative bile duct cells. Fibroblasts throughout the accumulated internodular matrix were devoid of EGF expression.
The novel identification of human hepatic EGF mRNA expression in cirrhotic liver reported here is probably a result of technical improvements that have enabled the detection of low transcript levels. First, to minimize RNA degradation, tissue obtained at the time of surgery was immediately quenched in fixative, thus minimizing nucleolytic activity that can hinder retrospective analysis of archival samples (Komminoth et al. 1994), particularly when low transcript levels are targeted. Second, uniformly labeled DNA probes were produced by incorporating biotin-dT during the oligo synthesis, thus eliminating variations in specific activity that frequently occur in post-synthesis labeling procedures (Bloch 1993). Third, oligo probes were designed to incorporate biotin-dT within a 15-base interval, which minimizes side-chain steric hindrance after hybridization and eliminates terminal instability which can occur with bT-tailed probes. Finally, to increase sensitivity, we employed a post-hybridization signal amplification procedure in which biotinylated tyramide deposition is targeted directly to the oligo-mRNA hybrids, yielding a 100- to 1000-fold increase in sensitivity over standard methods (Adams 1992; Speel et al. 1999).
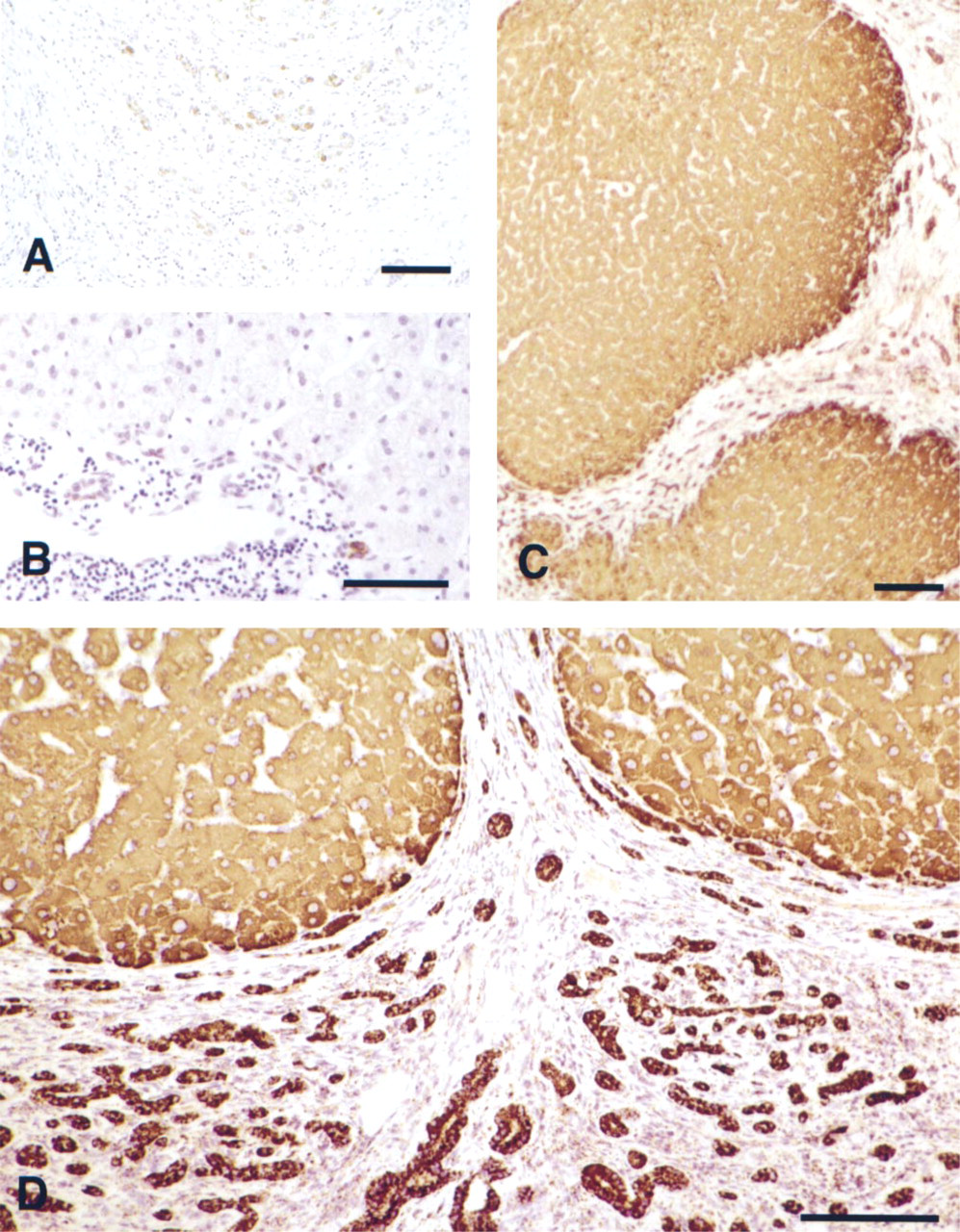
EGF mRNA expression in normal and cirrhotic human liver. EGF mRNA was detected with a biotinylated EGF antisense probe targeted to the mature coding region. (
The observed derepression of the EGF gene is accompanied by appearance of the nascent EGF peptide, as detected by immunohistochemistry. EGF is synthesized as a 160-kD transmembrane precursor and is proteolytically processed at the extracellular surface, releasing a mature 6-kD exocrine peptide (Carpenter and Cohen 1979). Although the small EGF peptide is believed to mediate paracrine responsiveness, increasing evidence points to the involvement of larger partially processed forms of EGF and related peptides (Mroczkowski and Reich 1993; Thorne and Plowman 1994; Marechal et al. 1999) that function in localized mitogenic signaling. To discriminate between the mature peptide, which may be endocytosed from serum, and nascent hepatic synthesis, we employed an antiserum raised against the human EGF pre-pro-region, which is removed before extracellular release (Carpenter and Cohen 1979). Lack of immunostaining in control liver parenchyma indicates a virtual absence of precursor synthesis, with the exception of an occasional presence in bile epithelium. However, in concert with its gene transcript, positive staining of the newly synthesized pre-pro-EGF peptide was evident throughout regenerative nodules and bile duct epithelium.
Interestingly, considerable variation was seen within individual nodules, as well as a distinct mosaic distribution indicating considerable variations in synthesis or processing. Although circulating levels of the mature EGF peptide in advanced cirrhosis have not been determined, the present findings are consistent with the hypothesis that the EGF precursor or its partially processed and nonsoluble forms may contribute to regenerative juxtacrine signaling. In previous studies we have shown that in regenerating rodent liver (after two-thirds HPX), EGF accumulates primarily as a partially processed 60-kD product (Mullhaupt et al. 1994).
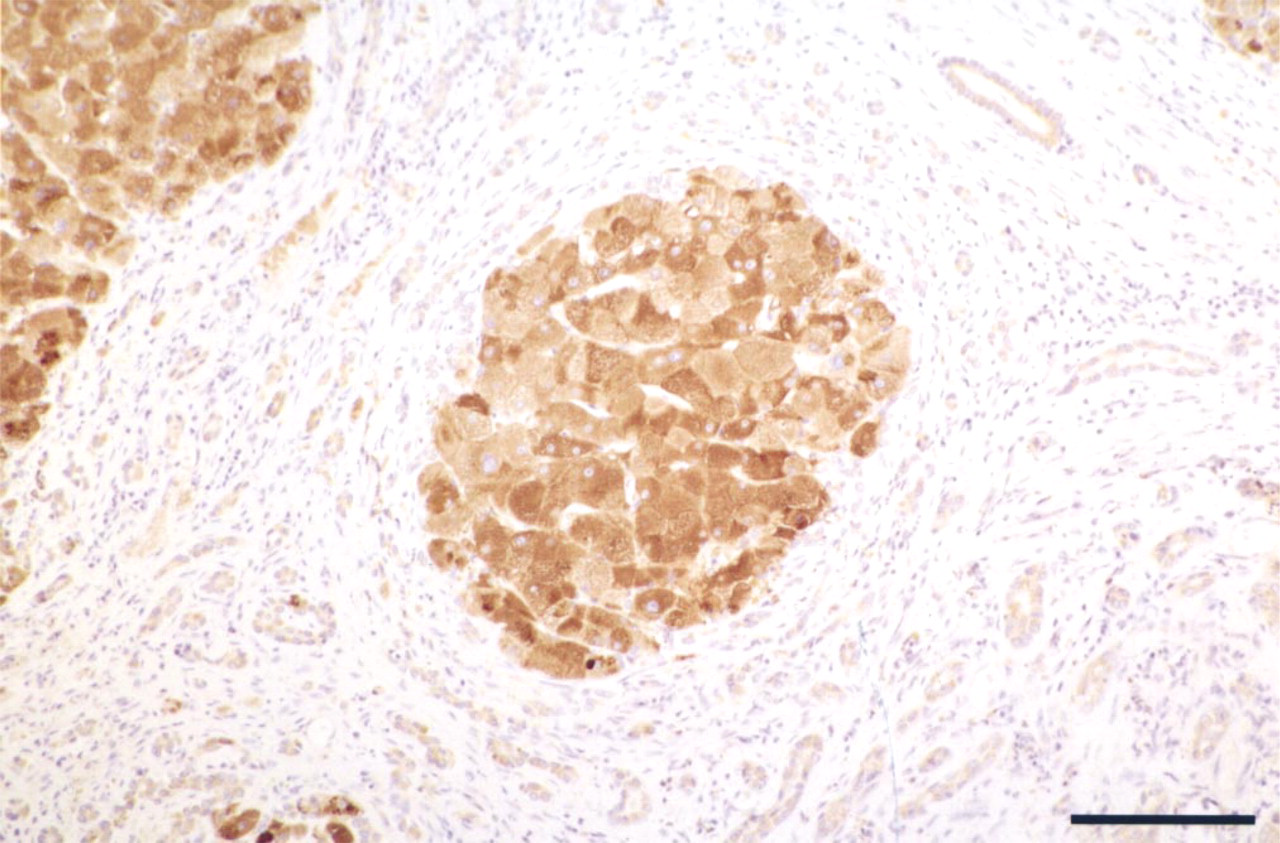
EGF pre-pro-peptide localization in a human cirrhotic liver. Immunohistochemistry was carried out using an antiserum raised against a synthetic human EGF precursor N-terminal pre-pro-peptide (amino acids 1–21). Immunostaining displays a distinct variation in cell-to-cell juxtaposition within the nodule. Note the weaker signal in bile ductules. Bar = 100 μm.
Although the long-term effects of continual hepatic EGF production remain unknown, recent studies have shown that EGFR upregulation is common in hepatic carcinogenesis (DeCicco et al. 1996,1997) and is frequently observed in carcinomas of the breast and prostate (Glynne-Jones et al. 1996; De Jong et al. 1998). Because EGF autoregulates the synthesis of its own receptor (Clark et al. 1985) and increases expression of other EGFR binding ligands (Barnard et al. 1994), such findings support the probability that EGF deregulation constitutes an early upstream event that facilitates growth factor signal amplification and thus contributes to increases in hepatic cell division during the course of chronic toxicity or injury.
Footnotes
Acknowledgements
Supported in part by a Department of Veterans Affairs Merit Review Grant and NIH grant PO1-AR39448 (LGK).
We thank Dr B. Mroczkowski (Agouron Pharmaceutical; La Jolla, CA) for kindly supplying the EGF N-terminal antiserum and Dr T. Wright and Ms T. Ryan (UCSF) for their help in obtaining tissue samples. We are grateful to members of the Liver Center for helpful comments and to Dr B. Sommers (Glen Research) for advice on oligo design.
