Abstract
In the clinical setting, routine identification of the main types of tissue amyloid deposits, light-chain amyloid (AL) and serum amyloid A (AA), is based on histochemical staining; rarer types of amyloid require mass spectrometry analysis. Raman spectroscopic imaging is an analytical tool, which can be used to chemically map, and thus characterize, the molecular composition of fluid and solid tissue. In this proof-of-concept study, we tested the feasibility of applying Raman spectroscopy combined with artificial intelligence to detect and characterize amyloid deposits in unstained frozen tissue sections from kidney biopsies with pathologic diagnosis of AL and AA amyloidosis and control biopsies with no amyloidosis (NA). Raman hyperspectral images, mapped in a 2D grid-like fashion over the tissue sections, were obtained. Three machine learning–assisted analysis models of the hyperspectral images could accurately distinguish AL (types λ and κ), AA, and NA 93–100% of the time. Although very preliminary, these findings illustrate the potential of Raman spectroscopy as a technique to identify, and possibly, subtype renal amyloidosis.
Introduction
Amyloidosis is an uncommon systemic disease, and the kidney is one of the most frequently involved organs. Diagnosis requires tissue biopsy, where amyloid deposits appear positive with Congo red stain and display apple-green birefringence when examined under polarized light. Although various forms of renal amyloidosis have been characterized,1,2 the most frequent types of amyloid detected in kidney biopsies are immunoglobulin light-chain amyloidosis (AL), arising from plasma cell dyscrasia, and amyloidosis derived from serum amyloid A (AA). 3 AL and AA amyloid deposits can be identified by immunohistochemical staining for kappa and lambda light chain and for amyloid A, respectively. Recently, laser microdissection/mass spectrometry has been used to detect other, rare types of amyloid, identifying 92–95% of known amyloid types with objective, semiquantitative measures of protein composition.1,3,4 However, clinical implementation of mass spectrometry–based characterization of tissue amyloid deposits is resource- and time-intensive and available only in very few specialized labs. Raman spectroscopic imaging, which uses inelastic light scattering to probe vibrational modes of molecules, enables multiplexed chemical fingerprinting at a specific location in unstained biological samples.5–7 With spatially resolved spectroscopic information, Raman spectroscopy can generate hyperspectral images that offer equivalent scale and quality to traditional tests, including immunochemical and histochemical staining.8,9 Recently, several studies, using various artificial intelligence tools for analysis, have proposed automated classification and assessment of Raman spectroscopic images, which may aid pathologic diagnoses.8,10
Owing to its ability to detect changes in protein secondary structure with exquisite molecular specificity and sensitivity, this technique is well suited to examine functional and pathological amyloid fibril formation, employing discrete, point-based spectroscopic measurements. Raman spectroscopy has been used to analyze structural features of different types of amyloid, such as α-synuclein amyloid, 11 β-amyloid, 12 and apolipoprotein C-III, 13 suggesting that this technology could be used to investigate experimental models of amyloidogenesis and could have potential future applications in the clinical diagnosis of amyloidosis.
In this proof-of-concept study, we explore Raman spectroscopic imaging coupled with machine learning (ML) to identify amyloid in renal biopsy tissue and to discriminate between AA and AL deposits through 2D maps of hyperspectral Raman data. Despite the subtlety of the explicit differences in spectral features between each type, trained ML decision models, particularly, a fully connected neural network, enabled accurate subtyping based on hyperspectral Raman imaging of amyloid tissues. This study of Raman spectroscopic imaging holds significance, as it is, to the best of our knowledge, the first report on Raman spectroscopy-based identification of renal amyloidosis. Our ML analysis of hyperspectral Raman images shows this technique’s potential for subtyping amyloid deposits.
Materials and Methods
A schematic illustration of the experimental design is provided in Fig. 1. Briefly, consecutive tissue sections were prepared for Raman measurement and pathological evaluation of amyloidosis for validation. Raster-scanned Raman measurements of tissue sections were processed and analyzed to localize and subtype renal amyloid deposits through multiple ML approaches.
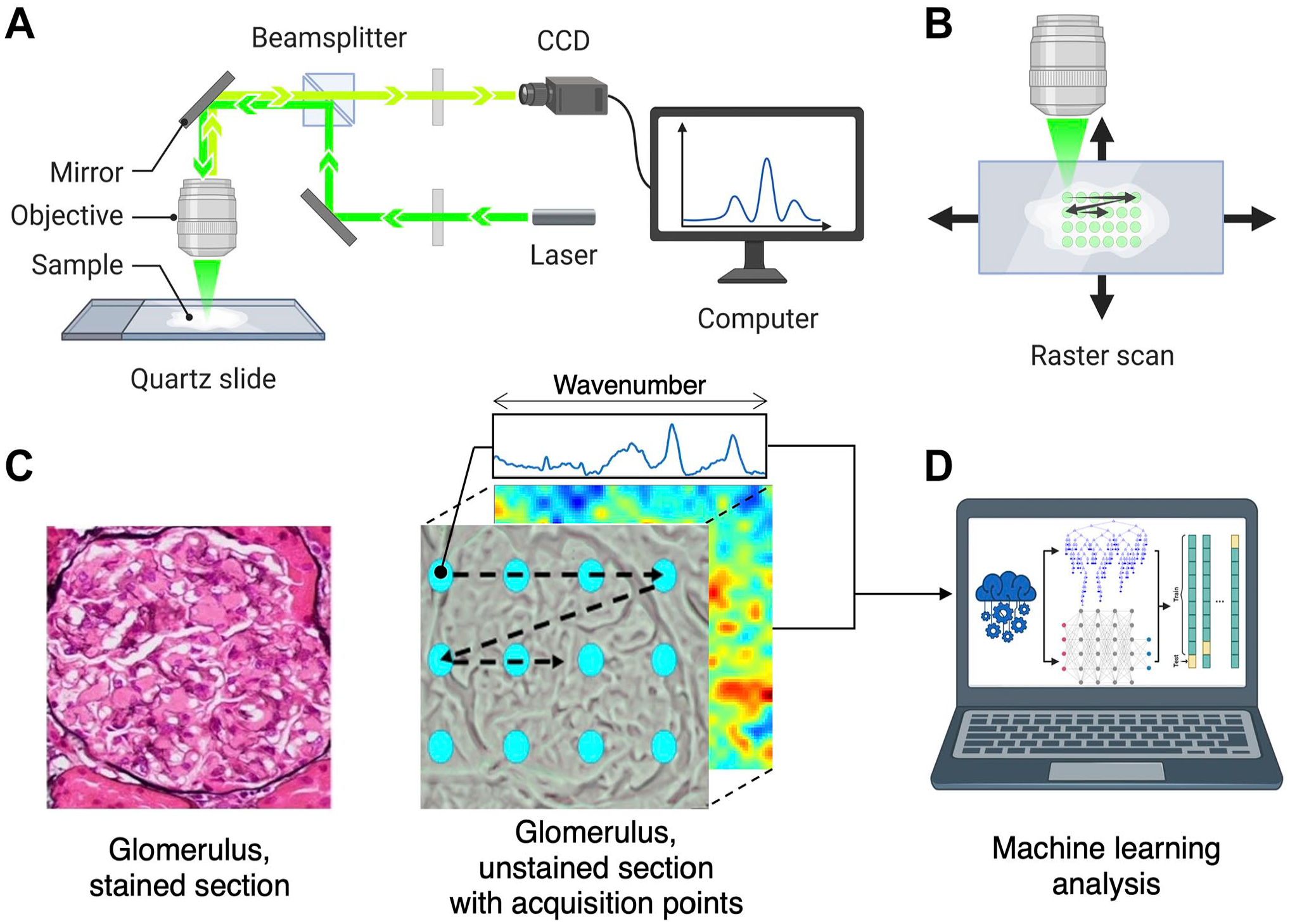
Overview of the Raman spectroscopic imaging of unstained frozen sections of kidney biopsy tissues. (A) Schematic illustration of Raman spectroscopy. (B) Raster scanning of Raman spectroscopy on a tissue section. (C) Kidney glomerulus with amyloid deposits in a section from paraffin-embedded tissue stained with PAS-MS silver stain and unstained frozen section including a glomerulus with each acquisition point corresponding to an individual Raman spectrum reconstructed into a 2D image. (D) Machine learning analysis of Raman hyperspectral tissue imaging. (Created with BioRender.com). Abbreviations: PAS-MS, periodic acid Schiff-methenamine silver; CCD, charge-coupled device.
Kidney Biopsies
Remnant de-identified tissues from kidney biopsies collected for diagnostic purposes were used for this study, including 4 AL (3 lambda-positive, 1 kappa-positive), 4 AA, and 5 non-amyloid cases (2 diabetic glomerulopathy, 1 focal segmental glomerulosclerosis, 1 immunoglobulin (Ig) A nephropathy, 1 fibrillary glomerulonephritis). The study was approved by an institutional review board (IRB00090103). The biopsies were processed for standard light microscopy and stained with hematoxylin–eosin (H&E), periodic acid Schiff (PAS), PAS-methenamine silver (PASMS), trichrome, Congo red, immunohistochemical staining for amyloid A, and kappa and lambda light chains. Immunofluorescence for IgG, IgA, IgM, kappa, and lambda light chains, along with complement (C3 and C1q), was performed on frozen tissue, and electron microscopy was performed on glutaraldehyde fixed tissue. A diagnosis of amyloidosis was made based on positive Congo red staining with the characteristic green birefringence under polarized light, the fibrillary ultrastructure detected by electron microscopy, and the type of amyloid was further characterized as amyloid A or amyloid L (kappa- or lambda-positive) based on the immunohistochemical and immunofluorescence stains (Fig. 2).
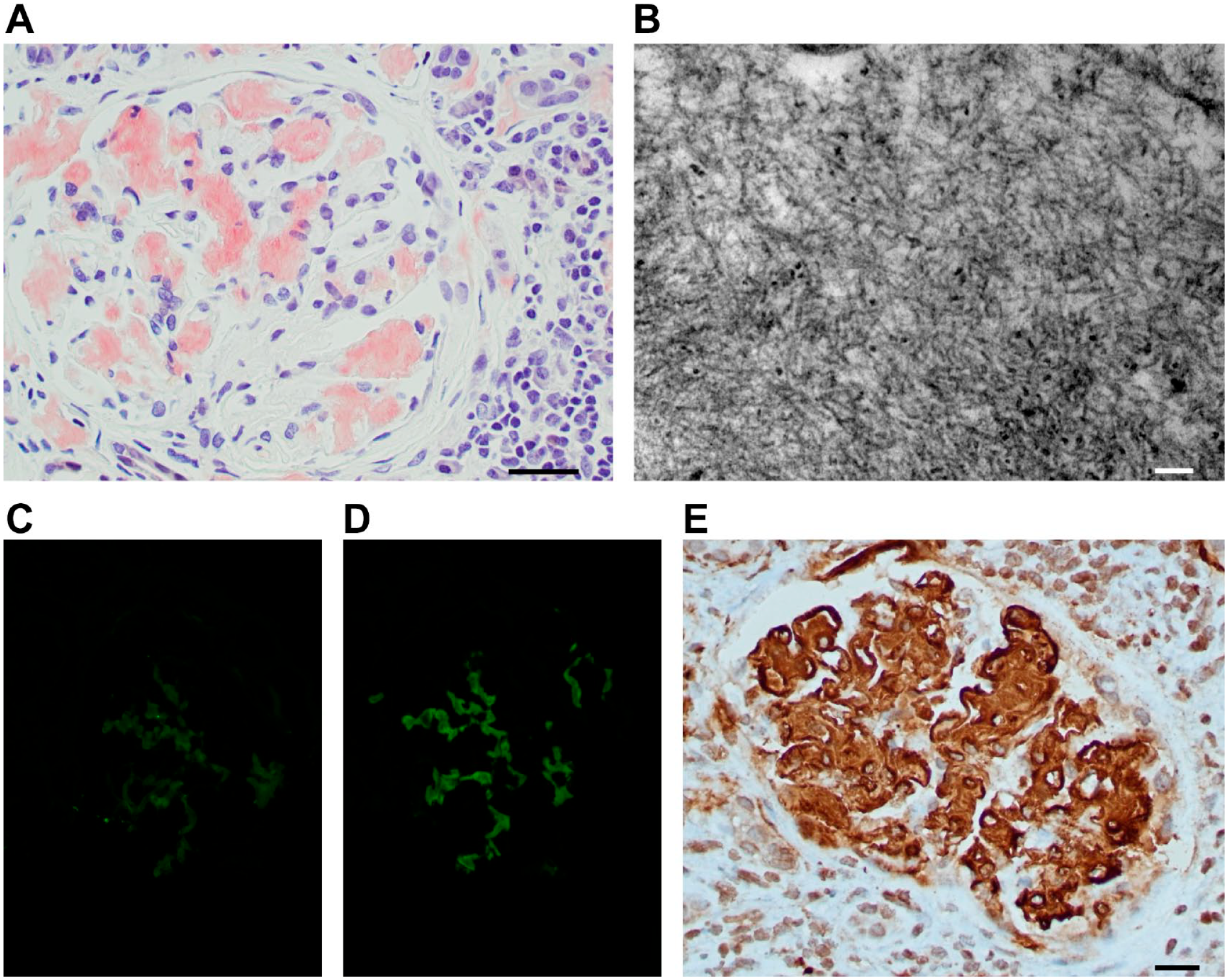
Histological features of amyloid deposits. (A) Congo red-positive glomerular amyloid deposits (magnification 400×); (B) electron microscopy image showing the characteristic amyloid substructure with randomly oriented fibrils; (C) negative glomerular immunofluorescence stain for kappa light chain in a case of light-chain amyloid (magnification 400×); (D) positive glomerular immunofluorescence stain for lambda (magnification 400×); (E) immunohistochemical stain for amyloid A is positive in a glomerulus with amyloid A deposits (magnification 400×). Scale bars A and E = 50 µm; bars B = 100 nm.
Raman Spectroscopy
From each case of the 14 biopsies, 1 to 3 unstained sections were prepared on quartz microscope slides to minimize spectral interference with the biochemical fingerprint of the tissue sample. Raman spectra were obtained from serial sections of frozen tissue cores measuring from 200 to 600 µm in width and from 2000 to 6000 µm in length. Each section ranged between 200 and 400 µm in width and over 1000 µm in length. Glomerular or arterial wall regions within each tissue section, with radii approximately 100–200 µm, were identified by pathologists for Raman measurements. Localization of amyloid deposits was confirmed by evaluating in parallel two adjacent tissue sections. One unstained section was placed on a quartz slide for Raman hyperspectral imaging, and the other was placed on a glass slide and stained by Congo red to verify the position of amyloid deposits.
Hyperspectral Raman data of tissue sections were collected using a confocal Raman microscope (Horiba Jobin Yvon-XploRA PLUS, Horiba France SAS, Longjumeau, France) adapted from a published design (Fig. 1A), as previously described. 14 We collected Raman measurements of kidney tissue placed on each slide by localizing amyloid with a 100× objective (MPlan N, Olympus Life Science, Center Valley, PA, US). Illumination with a laser of 532-nm wavelength was projected at every 5-µm interval along both x- and y-axes within each tissue section, with spatial resolution achieved by a 100-μm confocal pinhole. The tissue sections were placed on a computer-assisted motorized stage with a resolution of 0.5 μm to enable translation in both x- and y-directions for spectral mapping. Each image comprised a total of 625 acquisition points and raster scanned in a grid distribution to obtain a 2D spectral map, covering 72 × 72 µm2 of tissue (Fig. 1B). Mapping areas were aimed at selected glomerular and arterial wall regions within tissue, defined as “amyloid positive” or “amyloid negative” by Congo red staining on serial tissue sections. Raman spectroscopic signals representing the biochemical properties of scanned kidney tissue were collected at each acquisition point with a 1200 lines/mm grating and recorded through a thermoelectrically cooled charge-coupled device (CCD) detector (1024 × 256, Syncerity, Horiba Scientific) (Fig. 1C). The wavenumber of Raman spectra focused on the Raman biological fingerprint region, ranging from 800 to 1800 cm−1, to identify peaks corresponding to different types of amyloidosis.
Data Processing and Analysis
The collected digital raw Raman spectra were corrected by removing cosmic ray artifacts, denoising with Savitzky–Golay filters, and discriminating outliers and saturated signals by using Hotelling’s T2 versus Q residuals test. The spectra were then processed based on background signal followed by locally weighted smoothing. 15 The data preprocessing procedure was conducted by utilizing MATLAB 2021b (MathWorks, Inc., Natick, MA).
ML-based classification is an algorithmic approach that learns from the data to predict pertinent classes based on the learned decision model (Fig. 1D). ML techniques serve as a powerful tool to unravel the complex Raman spectra of biological samples and identify their molecular properties. 10 To classify amyloid types based on their Raman spectral mappings, we used three different supervised ML algorithms, including k-nearest neighbors (kNN), extreme gradient boosting (XGBoost), and multilayer perceptron (MLP).
The kNN algorithm is one of the most fundamental classifiers that arranges data sets with respect to the geometric distance between each datapoint. We measured the Euclidean distance between each Raman spectra and predicted the class among its five nearest.
XGBoost is an advanced tree boosting-based algorithm, making predictions based on an ensemble of decision trees that learn patterns and features of the data set. The learned model classifies with respect to the consensus vote from independent decision trees, intended to minimize individual error and control overfitting. We designed a classification model with 100 decision trees at a learning rate of 0.3.
MLP is the simplest fully connected neural network that constitutes most of highly complex deep learning algorithms. The model parameters we used were logistic activation with Adam optimizer for a maximum of 2000 iterations until convergence.
Results
Raman Spectra of Frozen Tissues Reveal Regions Affected by AL and AA
Multiple hyperspectral Raman images were collected from each frozen section. Characterization of tissue as containing AA, AL, or NA was based on spectral features unique to each type that are observed in the region between 800 and 1800 cm−1 (Fig. 3).
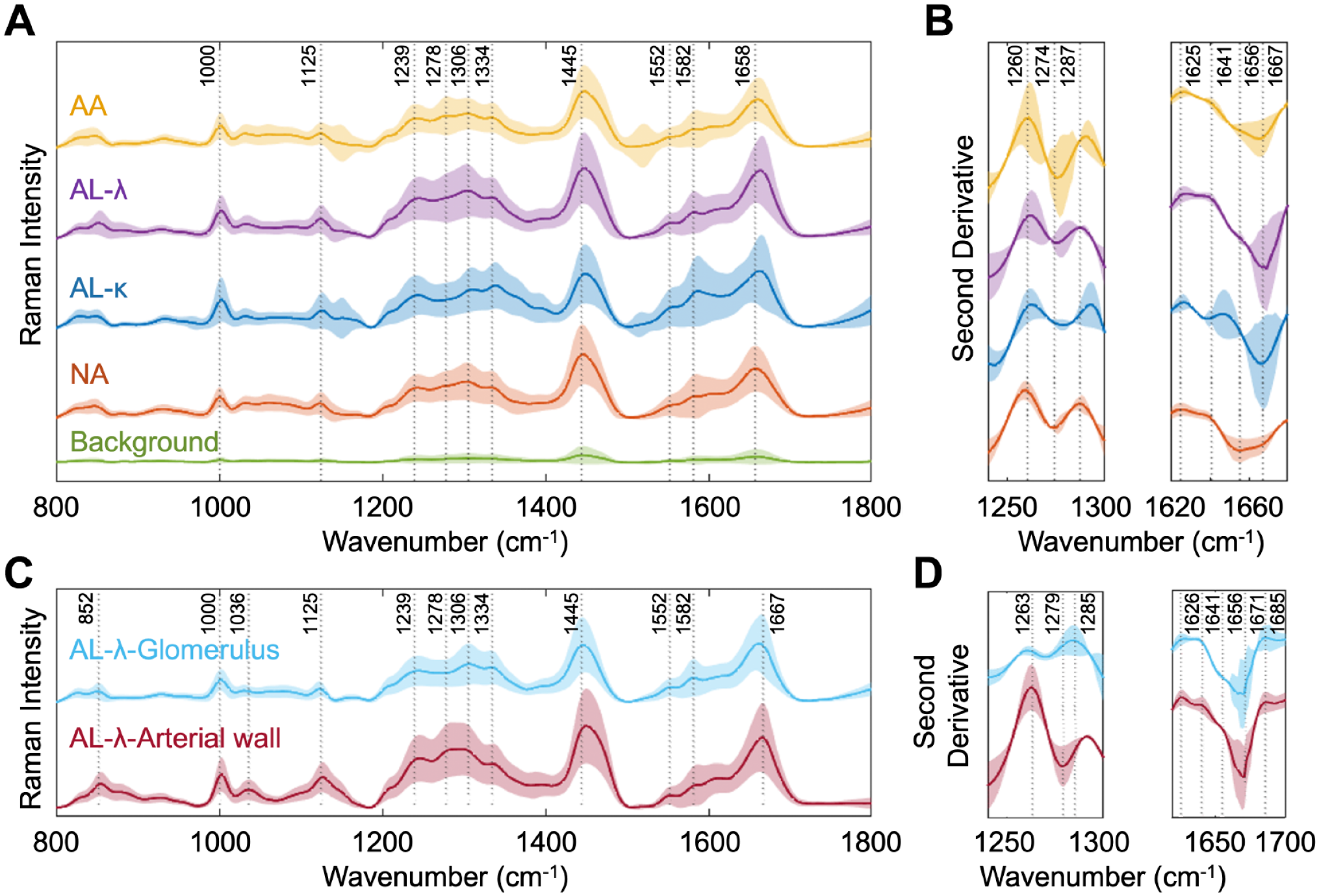
Raman spectra of different types of renal amyloidosis and deposition sites. (A) Baseline corrected and smoothed average Raman spectra of different amyloid tissue types with 1 standard deviation shaded. (B) Second derivative analysis of average Raman spectra with 1 standard deviation shaded. (C) Baseline corrected and smoothed average Raman spectra of AL within glomerulus and arterial wall within unstained frozen tissue with 1 standard deviation shaded. (D) Second derivative analysis of average Raman spectra of AL tissue with 1 standard deviation shaded. Abbreviations: AL, light-chain amyloid; AA, amyloid A; NA, no amyloidosis.
We observed spectral features unique to AA, AL, and NA types due to their protein conformational changes, appearing across amide I, II, and III regions (Fig. 3A). In the amide I region, dominated by the C=O stretch of protein secondary structure, 1658 cm−1 peaks are observed, while AL types are shifted to a higher frequency. Compared to AA and NA spectra, AL spectra exhibit subtle peaks at 1552 cm−1, associated with amide II. 16 Multiple peaks, at 1239, 1278, and 1306 cm−1, are observed in amide III region. For the average AL spectrum, distinctive peaks are observed at 1582 cm−1, which can be attributed to aromatic acids including phenylalanine and tryptophan. 5 Our findings are consistent with previous studies on Raman spectroscopy performed on amyloid.16,17 A second derivative analysis further revealed spectral differences between amyloid types (Fig. 3B).
We then examined the spectral difference based on amyloid infiltration sites (Fig. 3C and D). The differences between the averaged spectra collected from glomeruli and arterial walls of AL tissue samples showed features native to the structure.
While peaks unique to AL types observed in Fig. 3A appeared in measurements collected in both glomeruli and arterial walls, we still observed some site-specific spectral features (Fig. 3C). In the amide III region, a broad bandwidth was observed between 1278 and 1306 cm−1 peaks for the arterial wall sections, while glomeruli measurements have a peak around 1307 cm−1. Also, measurements collected at the arterial wall showed an elevated intensity at 1036 cm−1, associated with phenylalanine. 16 We further conducted a second derivative analysis in the amide I and III regions to identify spectral features due to the associated sites (Fig. 3D). At 1263 cm−1 peaks, attributed to lipids, Raman measurements of glomerular region were less shifted while those of arterial walls had an elevated intensity. Such a Raman peak shift, despite being collected from regions involving an identical amyloid subtype, indicates the potential for other contributing factors, such as intermolecular interactions, pressure, and more, as the deposits interact with a slightly different surrounding environment specific to the deposition site.11,13
ML-Based Classification Can Subtype Amyloidosis Based on Hyperspectral Raman Data of AL Lambda, AL Kappa, and AA
Spatially resolved hyperspectral Raman data were taken to ML approaches to demonstrate classification capability for subtyping renal amyloidosis (Fig. 4). We employed three ML algorithms, kNN, XGBoost, and MLP models, to learn amyloid features and to differentiate biochemical features embedded in Raman spectra of AA, AL-λ, AL-κ, and NA tissues. We used Raman spectra collected from glomeruli in AL, AA, and NA tissues (6875, 7500, and 13,125 spectra, respectively), and from arterial walls in one AL case (2500 spectra), summing up a total of 30,000 spectra from 48 tissue mappings of glomerular or vascular regions. These were used for training ML models that were optimized by 10-fold cross-validation analysis. Briefly, all 30,000 Raman spectra were randomly split into 10 subsets. At each iteration, the decision model was trained with nine subsets and tested with the remaining one subset for the model to learn the data set and check prediction capability. Throughout multiple iterations, the model generalizes the data set and reduces decision variability.
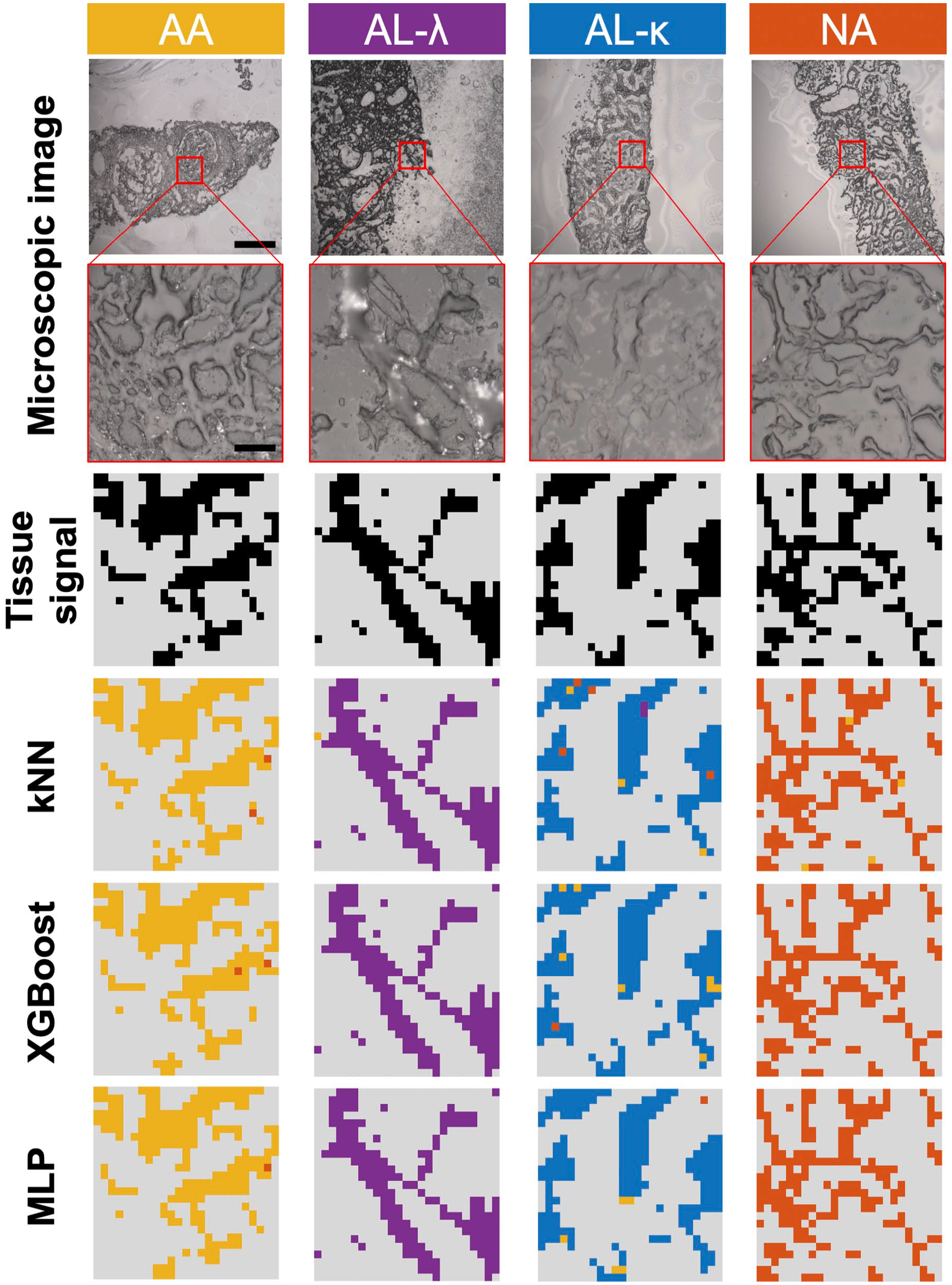

ML classification (kNN, XGBoost, and MLP) results of AA, AL λ and κ, and NA tissues. From top to bottom: Microscopic images of Raman imaging area (first row: scale bar = 2 µm, second row: scale bar = 20 µm); distribution of tissue signal (black) from background (gray) reconstructed based on hyperspectral Raman imaging; ML-based classification distribution within each amyloid type (AA [yellow], AL-λ (purple), AL-κ (blue), NA (orange), background(gray)). Abbreviations: ML, machine learning; kNN, k-nearest neighbors; XGBoost, extreme gradient boosting; MLP, multilayer perceptron; AA, amyloid A; AL, light-chain amyloid; NA, no amyloidosis.
Three independently trained ML models made predictions for each pixel, as shown in Fig. 4, where the decisions are color-coded as yellow, purple, blue, and orange for AA, AL-λ, AL-κ, and NA, respectively; signals dominated by background are represented as gray. Two-dimensional reconstructed images allowed localization of each spectrum within the tissue sample, represented per pixel within the image. ML classification images show subtyping results of tissue sections. All three decision models presented close representation of the tissue signal image with respect to the morphology of respective types.
For each ML classification image of tissue sections, the amyloid type was decided based on the majority class among the pixels within the image excluding background pixels. If no consensus was reached to any of the three types, such that none of the three types constituted over 50% of all predictions made for spectra of the tissue section, it remained unclassified. Within the classification maps based on trained ML models, we spotted several misclassified pixels with incorrect predictions on amyloid subtyping. However, most of them are located at the tissue periphery, where a relatively weak tissue signal mixed with background may have led to the incorrect decision.
Amyloid subtyping results are summarized in Fig. 5 and Table 1. One to six images per patient, collected across various glomerular and vascular regions within each tissue section, were subject to classification. Subtyping is determined by the majority voting when more than 50% of all pixels are predicted to be the same amyloid type. Among the three ML decision models, MLP classification shows the best performance, correctly predicting all tissue types with minimal misclassified pixels. For one AA case, kNN failed to reach a consensus between AA and NA, resulting in a result of “unclassified,” and while XGBoost correctly classified the sample, slight differences between the two types were observed. Overall, the classification summary implies little to no misclassification across different amyloid types, demonstrating the specificity of ML model’s decision capability based on hyperspectral Raman imaging of tissues. Most misclassifications of amyloidogenic tissues result in an NA prediction; most of these misclassified pixels are concentrated on tissue periphery, as shown in Fig. 4. Thus, by localizing tissue signature, we can make a more accurate decision on subtyping with ML models.
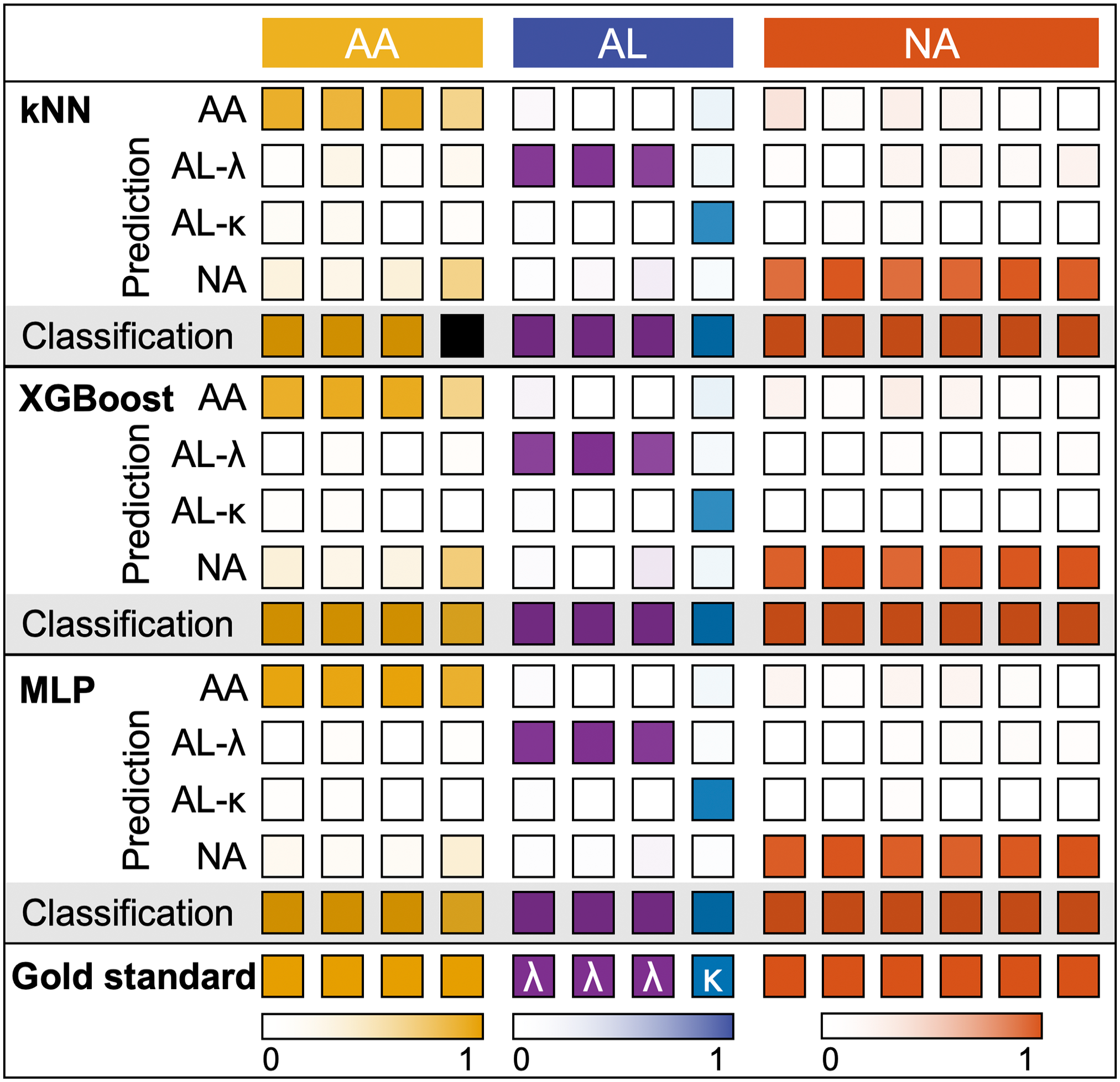
Summary of ML classification (kNN, XGBoost, and MLP) results of AA, AL λ and κ, and NA tissues. From top to bottom: prediction and subtyping results of kNN, XGBoost, and MLP decision models on hyperspectral Raman imaging of each tissue type. Classification decisions made based on the majority voting (classification threshold: >50%—AA (yellow), AL-λ (purple), AL-κ (blue), and NA (orange); unclassified: otherwise (black). Abbreviations: ML, machine learning; kNN, k-nearest neighbors; XGBoost, extreme gradient boosting; MLP, multilayer perceptron; AA, amyloid A; AL, light-chain amyloid; NA, no amyloidosis.
Summary of ML Classification (kNN, XGBoost, and MLP) Results of AA, AL λ and κ, and NA Tissues.
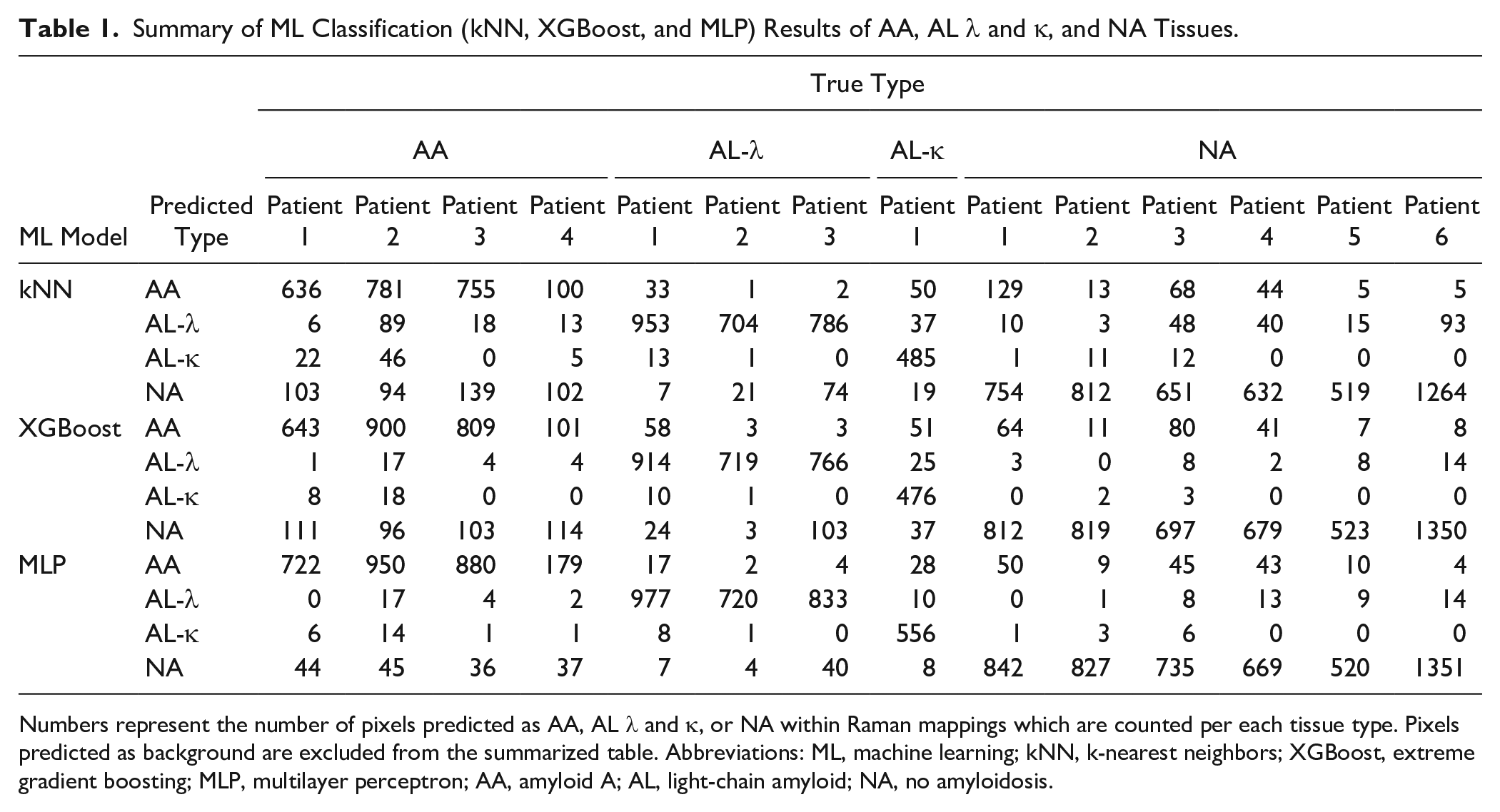
Numbers represent the number of pixels predicted as AA, AL λ and κ, or NA within Raman mappings which are counted per each tissue type. Pixels predicted as background are excluded from the summarized table. Abbreviations: ML, machine learning; kNN, k-nearest neighbors; XGBoost, extreme gradient boosting; MLP, multilayer perceptron; AA, amyloid A; AL, light-chain amyloid; NA, no amyloidosis.
The ML amyloid subtype classification results illustrated in Fig. 5 are summarized in detail in Table 1, with the number of Raman spectra listed in order corresponding to the prediction results. Each number in the table represents the number of pixels predicted as corresponding amyloid types based on their Raman spectra. Among all 30,000 spectra used for classification, predictions made as background are excluded in the table. For all amyloid types, the MLP model delivered the highest number of accurate predictions, showing its superior prediction performance among the three ML models.
Discussion
In this proof-of-concept study, we describe, for the first time, the application of hyperspectral Raman imaging to identify and subtype amyloidosis in human kidney tissue. We show that hyperspectral Raman imaging powered with ML analysis can detect amyloid deposits in tissue sections and correctly discriminate between AA and AL (types λ and κ) amyloid, which are the most common types of amyloidosis involving the kidney.
Raman spectroscopy can be performed with relatively simple, inexpensive instrumentation that requires minimal sample preparation. For these reasons, this technique appears attractive for a broader use in biomedical fields, and in the past few years, it has received increasing attention as a novel tool with possible clinical applications, for example, in the examination of biofluids and tissue18,19 and in the evaluation of cancerous lesions.14,20,21
Before beginning this study, we had recently successfully employed this technique to identify the nature of crystal deposits in kidney biopsies, 19 prompting us to explore its further use in this type of small tissue specimen. Based on previous reports of Raman analysis of amyloid protein structural characteristics,11–13 we tested Raman spectroscopy to discriminate AA and AL (types λ and κ) amyloidosis from non-amylogenic kidney tissues and identify each type through spectroscopic mapping. In addition to molecular distinctions based on subtypes, we also observed subtle changes specific to amyloid infiltration sites, demonstrated by the difference in peak shifts of amyloid deposits in glomeruli and arterial walls. Such anatomical differences may influence the molecular structure of amyloid fibrils and their interactions with surrounding molecules, despite an identical amyloid subtype, which can be uncovered by Raman spectroscopy. Specifically, we observed changes in the amide III region, which showed a broad bandwidth between 1278 and 1306 cm−1 in the arterial wall sites, while the measurements from the glomeruli exhibited a peak around 1307 cm−1. Also, we spotted an increased intensity at 1036 cm−1, attributed to phenylalanine, from the measurements in the arterial wall region. According to a second derivative analysis, we found that the Raman measurements of the glomerular region exhibited a lower shift at 1263 cm−1 peaks, associated with lipids, whereas measurements from the arterial walls showed an elevated intensity.
We recognize the limitations of this proof-of-concept study obtained in a small number of specimens, which will further require validation on a large number of samples, including other types of amyloid proteins. Currently, identification of unusual types of amyloid is not possible by histochemistry alone, requiring more sophisticated tools. Mass spectrometry has become the assay of choice for these cases and can identify several non-AA and non-AL amyloid proteins such as leukocyte chemotactic factor 2 (ALECT2), fibrinogen α-chain (AFib), gelsolin (AGel), apolipoprotein (AI, AII, AIV, CII), and lysozyme (ALys). 1 The performance of Raman spectroscopy for identification of these rare types of amyloid remains to be determined and is an interesting area for investigation of ML-powered hyperspectral Raman imaging, since there are no previous reports of such application. As for the current scope of the study, it is premature to conclude whether deposition site will influence classification performance due to the small sample size we tested. However, additional renal amyloid cases can be examined using Raman spectroscopy coupled with ML analysis to further explore the potential application of this method in the classification of this rare disease. While amyloidosis inveterately suffers from scarce, precious sample availability, a label-free and non-destructive Raman spectroscopic approach will enable a reproducible and reusable procedure to supplement the current diagnostic pipeline.
In conclusion, our findings show the potential of Raman spectroscopy coupled with ML algorithms to identify amyloid subtypes, although further study will be necessary to evaluate its suitability and reliability for wider use in the diagnostic assessment of amyloid tissue deposits.
Footnotes
Competing Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
All authors have contributed to this article as follows: conceptualization of this study (IB, CJS and SMB), kidney biopsies and sample preparation (CJS and SMB), Raman experiments (JHK and CZ), Raman data processing and analysis (JHK), pathological evaluation (SMB), study supervision (IB and SMB), drafting manuscript and designing figures (JHK), and all authors reviewed, provided critical feedback, and contributed to the manuscript.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: IB, CZ, and JHK report financial support was provided by National Institute of General Medical Sciences (DP2GM128198) and by National Institute of Biomedical Imaging and Bioengineering (2-P41-EB015871-31).
Ethics Approval
The study was approved by an institutional review board (IRB00090103).
Data Availability
All data are included in the manuscript.
