Abstract
MUC13 is a newly identified transmembrane mucin. Although MUC13 is known to be overexpressed in ovarian and gastric cancers, limited information is available regarding the expression of MUC13 in metastatic colon cancer. Herein, we investigated the expression profile of MUC13 in colon cancer using a novel anti-MUC13 monoclonal antibody (MAb, clone ppz0020) by immunohistochemical (IHC) analysis. A cohort of colon cancer samples and tissue microarrays containing adjacent normal, non-metastatic colon cancer, metastatic colon cancer, and liver metastasis tissues was used in this study to investigate the expression pattern of MUC13. IHC analysis revealed significantly higher (p<0.001) MUC13 expression in non-metastatic colon cancer samples compared with faint or very low expression in adjacent normal tissues. Interestingly, metastatic colon cancer and liver metastasis tissue samples demonstrated significantly (p<0.05) higher cytoplasmic and nuclear MUC13 expression compared with non-metastatic colon cancer and adjacent normal colon samples. Moreover, cytoplasmic and nuclear MUC13 expression correlated with larger and poorly differentiated tumors. Four of six tested colon cancer cell lines also expressed MUC13 at RNA and protein levels. These studies demonstrate a significant increase in MUC13 expression in metastatic colon cancer and suggest a correlation between aberrant MUC13 localization (cytoplasmic and nuclear expression) and metastatic colon cancer.
Worldwide, colorectal cancer is increasing in many areas with an estimated 1.2 million new colorectal cancer cases and 609,000 deaths in 2010 (Jemal et al. 2010). In the United States, colorectal cancer remains the third most common and third most lethal cancer in both men and women (Siegel et al. 2011). Colon cancer patients have a 5-year survival rate of up to 90% if diagnosed at an early stage; however, the 5-year survival rate declines to less than 10% when cancer is diagnosed with metastasis to distant organs (Siegel et al. 2011). Therefore, understanding the molecular events leading to metastatic colon cancer is highly desirable.
Mucins, high molecular weight glycoproteins predominantly expressed at the epithelial surface of tissues, provide protection under normal physiological conditions (McAuley et al. 2007; Cone 2009). However, aberrant mucin expression has been associated with breast (Workman et al. 2009), pancreatic (Singh et al. 2004), and lung (Gao et al. 2009) cancers. MUC13 is a newly identified transmembrane mucin expressed in the small intestine, large intestine, gastric mucosa, and trachea under normal physiological conditions (Williams et al. 2001). MUC13 consists of two distinct subunits, an extracellular α subunit (consisting of the mucin tandem repeat domain, one epidermal growth factor [EGF]-like domain, and a portion of the sperm enterokinase agarine [SEA] module) and a β subunit (consisting of the remaining SEA module, two EGF-like domains, the transmembrane domain, and the cytoplasmic tail) (Maher et al. 2011). The mucin repeat domain (the hallmark of mucins) contains a series of serine/threonine amino acids that provide the scaffold for O- and N-linked glycosylation (Williams et al. 2001). During protein processing, MUC13 is predicted to be cleaved into α and β subunits (within the SEA domain), and the subunits non-covalently associate at the cell membrane until shedding is initiated via environmental signals involved in outside-to-inside signaling that may alter cell behavior (Maher et al. 2011).
MUC13 has been shown to be overexpressed in gastrointestinal and ovarian cancers (Shimamura et al. 2005; Walsh et al. 2007; Chauhan et al. 2009). In reference to colon cancer, reports regarding the expression levels of MUC13 have been variable. For instance, in the first published report of MUC13, Williams et al. (2001) reported downregulation of MUC13 in a small number of colorectal cancers compared with adjacent normal tissues. Subsequently, Packer et al. (2004) also showed downregulation of MUC13 mRNA in colorectal cancer tissues compared with adjacent normal tissues. In contrast to these studies, Walsh et al. (2007) later reported the overexpression of MUC13 in the membrane and cytoplasm of colon adenocarcinomas using a polyclonal anti-MUC13 antibody. In the present study, we sought to resolve the discrepancies regarding MUC13 expression in colon cancer using a novel monoclonal antibody (ppz0020) (Shimamura et al. 2005). We examined four groups of tissues: (1) adjacent normal colon, (2) non-metastatic colon cancer, (3) metastatic colon cancer, and (4) the corresponding liver metastasis. In addition, we also analyzed colon cancer cell lines to determine the expression of MUC13 in colon cancer cell line models. Herein, we show that the expression of MUC13 significantly increases in colon cancer tissues and is aberrantly localized in metastatic colon cancer.
Materials and Methods
Tissue Specimens and Cell Culture
Tissue microarrays (TMAs) containing colon cancer tissues of various grades and adjacent normal tissues from eight patients were procured from AccuMax (catalog number A203 [VI]; ISU Abxis Co., Ltd., San Diego, CA). To facilitate accurate comparison of MUC13 expression between non-metastatic and metastatic colon cancer, the 10 tissue spots in this array that were from patients with metastatic colon cancer were excluded from analysis. The final number of colon cancer tissue spots suitable for analysis was 76. None of the 8 adjacent normal tissue spots were from patients with metastatic colon cancer. To examine MUC13 expression in metastatic colon cancer, additional TMAs (catalog numbers A203 [III] and A203 [IV]) were obtained from AccuMax. These arrays contained three types of tissue from each patient: normal colon, metastatic colon cancer, and liver metastasis tissue samples (tissue spots suitable for analysis were as follows: adjacent normal, n=27; metastatic colon cancer, n=56; and liver metastasis, n=49). In addition, archived colon cancer samples (n=60) were obtained from Sanford Health. The use of archived clinical samples in this study was approved by the institutional review board (IRB) at Sanford Research/USD, Sioux Falls, South Dakota. Furthermore, six colon cancer cell lines (LoVo, HT-29, T-84, SW620, SW480, and SW48) were used to determine MUC13 expression in colon cancer cell lines. Each cell line was grown in ATCC-specified cell culture media supplemented with 10% FBS (Hyclone Laboratories; Logan, UT) and 1% sodium pyruvate.
Immunohistochemistry
Slides were processed for immunohistochemistry (IHC) analysis using a polymer-based MACH4 IHC kit (Biocare Medical; Concord, CA) according to the manufacturer’s recommendation (unless indicated, all reagents were purchased from Biocare Medical). In brief, tissues were heated at 60C for 30 min, deparaffinized in SlideBrite, and rehydrated with graded ethanols. Tissues were incubated in Peroxidazed solution to quench endogenous peroxide activity, and antigen retrieval was performed in a Biocare Decloaking Chamber (125C for 30 sec) while submerged in 1× Diva solution. Tissue sections were incubated with Sniper for 10 min to block nonspecific binding sites. Tissues were washed with Tris-buffered saline containing 0.1% Tween-20 (TBS-T) and then incubated with anti-MUC13 MAb (clone ppz0020) (1:1000) (Shimamura et al. 2005) in Davinci Green Diluent for 30 min. After washing with TBS-T, the tissues were incubated with the mouse probe for 30 min followed by incubation with the horseradish peroxidase–polymer for 30 min. The reaction color was developed by incubating tissue sections with 3,3′-diaminobenzidine reagent (DAB) solution for 2 min. Tissue sections were counterstained with hematoxylin solution, washed, dehydrated, and mounted with Ecomount mounting media. To confirm that immunostaining was specific, a negative control (no primary antibody control) was also included, and these samples had no detectable staining (data not shown).
Immunohistochemical Analysis
All slides were viewed under an Olympus BX 41 microscope (Olympus Corporation; Center Valley, PA). Quantitative analysis of immunoreactivity was done by calculating the composite score, which is a function of the percentage of cancer cells positively immunostained multiplied by the MUC13 staining intensity (range, 0–16). Percentage of cancer cells positively stained for MUC13 was scored on the scale of 0 to 4 as follows: 0 for less than 5%, 1 for 5–25%, 2 for 26–50%, 3 for 51–75%, and 4 for >75% positively stained cancer cells. Immunostaining intensity was also graded on a 0 to 4 scale as follows: 0 for no immunostaining, +1 for faint, +2 for weak, +3 for high, and +4 for very high staining. The mean composite score (MCS) of MUC13 staining was calculated based on total immunoreactivity or immunoreactivity localized to the membrane, cytoplasm, or nucleus as scored by independent pathologists (MDK, DWL, and TB). In most cases, the independent evaluations were consistent between samples; however, in cases of discrepancies, consensus was arrived by reevaluation by MDK. This type of semi-quantitative analysis (i.e., MCS), which takes into account both intensity of staining as well as the extent of tumor cells stained, is a well-recognized and accepted analytical method that has been widely used for the quantification of protein expression obtained from IHC.
Total RNA Isolation and RT-PCR Analysis
Total RNA was isolated from colon cancer cell lines SW48, SW480, SW620, T-84, LoVo, and HT-29 by using RNeasy Mini kit (Qiagen; Valencia, CA). RNA samples (2 µg) were reversed transcribed into cDNAs from their respective cell lines using the High Capacity cDNA kit (Applied Biosystems; Foster City, CA) as per the manufacturer’s instructions. The resulting cDNA samples were subjected to PCR amplification using 10 pmol MUC13 and glyceraldehyde 3-phosphate dehydrogenase (GAPDH)-specific forward and reverse primers (MUC13: CCTTCGGTGTGATTATTATGGC [forward], GCATCTGGCTGTCTCTGGA [reverse]; GAPDH: TGAAG GTCGGAGTCAACGGATTTGGT [forward], CATGTGG GCCATGAGGTCCACCAC [reverse]) in a final volume of 50 µl containing 250 µmol/L dNTP, 1.5 mmol/L MgCl2, and 1.25 units of Taq DNA polymerase (MasterMix; Eppendorf, Westbury, NY). A PCR thermocycler (BioRad; Hercules, CA) was used with an initial denaturation at 94C for 4 min followed by 30 cycles of 95C for 1 min, 58C for 1 min, and 72C for 2 min. PCR products were resolved on a 2% agarose gel.
Real-time PCR
Total RNA was isolated from colon cancer cell lines SW48, SW480, SW620, T-84, LoVo, and HT-29 by using RNeasy Mini kit (Qiagen). RNA samples (2 µg) were reversed transcribed into cDNAs from their respective cell lines using the High Capacity cDNA kit (Applied Biosystems) as per the manufacturer’s instructions. The SYBR Green RT PCR kit was used to amplify DNA using MUC13 primers (MUC13: CCTTCGGTGTGATTATTATGGC [forward], GCATCTGGCTGTCTCTGGA [reverse]) as per the manufacturer’s instructions (Applied Biosystems). GAPDH-specific primers were used for the normalization (GAPDH: TGAAGGTCGGAGTCAACGGATTTGGT [forward], CATGTGGGCCATGAGGTCCACCAC [reverse]). A Real Time PCR thermocycler (Applied Biosystems 7500) was used with an initial denaturation at 95C for 10 min followed by 40 cycles of 95C for 15 sec and 60C for 1 min. The relative quantity (RQ) value was calculated based on the ∆∆Ct method using the Gene Expression Macro provided by Bio-Rad (Bio-Rad;Hercules, CA).
Immunofluorescence Confocal Microscopy
Colon cancer cell lines were subjected to immunofluorescence confocal microscopy as described earlier (Chauhan et al. 2009). Briefly, colon cancer cells were grown in chamber slides in the ATCC-specified media at 37C overnight. Cells were washed with Hank’s buffer containing 0.1 M HEPES media and washed with phosphate-buffered saline (PBS) containing 0.05% Tween-20. Cells were fixed with 70% ice-cold methanol for 2 min and blocked with 10% goat serum. Cells were incubated with anti-MUC13 MAb (ppz0020) for 1 hr at room temperature. Cells were then washed with PBS-T (3×, 5 min each) and incubated in FITC-conjugated goat anti-mouse secondary antibody for 1 hr. Cells were washed and incubated in DAPI (1 µg/ml) for 2 min and mounted using Vectashield mounting medium (Vector Laboratories; Burlingame, CA). Immunostaining was observed and photographed under an Olympus Fluoview FV1000 confocal laser microscope (Olympus Corporation). Photomicrographs of MUC13 staining were taken using the green channel.
Statistical Analysis
Mean composite scores between adjacent normal and cancer tissues were compared using analysis of variance (ANOVA) and two-tailed Student’s t-tests with unequal variance. For clinical samples, median values of the continuous measures were compared between MUC13 expression groups using a non-parametric Kruskal-Wallis test, and categorical distributions were compared using Fisher’s exact test. Pairwise comparisons were performed if the overall test suggested differences between the three groups. p<0.05 was considered statistically significant. All analyses were performed using SAS 9.1 (SAS Institute; Cary, NC) by one of the authors (SEP).
Results
MUC13 Is Overexpressed in Colon Cancer
Immunohistochemical analysis of colon cancer TMA containing adjacent normal and non-metastatic colon cancer samples showed very faint MUC13 expression in adjacent normal tissues (n=8; Fig. 1A and Table 1) and significantly higher MUC13 expression in various grades of non-metastatic colon cancer samples (n=76; Fig. 1B–D and Table 1). MUC13 expression was evaluated and analyzed in terms of MCS (as described in Materials and Methods). The MCS of MUC13 expression was significantly (p<0.001) higher in colon cancer tissues (MCS = 14.2) compared with adjacent normal tissues (MCS = 5) (Table 1). The MCS was significantly higher in well, moderately, and poorly differentiated grades of non-metastatic colon cancer compared with adjacent normal tissue (Fig. 1A–D). In addition, very high (4+) to high (3+) MUC13 staining intensity was observed in 97.3% of the colon cancer tissues (74 of 76 samples) compared with 0% of the adjacent normal tissues (0 of 8 tissues) (Table 1).
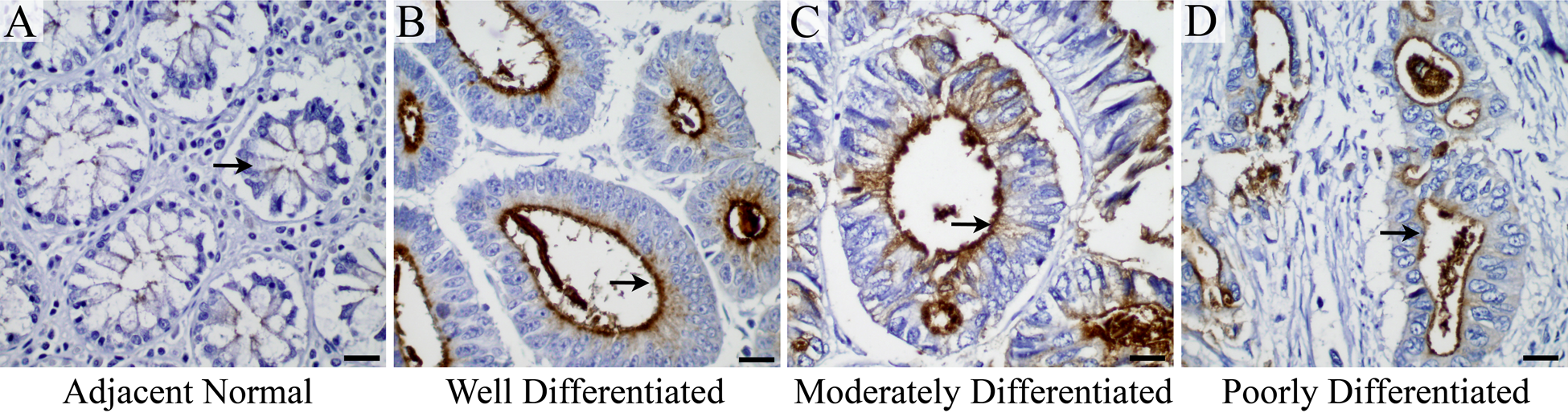
(A–D) MUC13 is overexpressed in colon cancer: MUC13 expression (brown color, representative areas indicated by the black arrows) is detected in (A) adjacent normal, (B) well-differentiated, (C) moderately differentiated, and (D) poorly differentiated colon cancer. Scale bars = 20 µm.
Immunohistochemical Analysis of MUC13 in Adjacent Normal and Various Grades of Colon Cancer
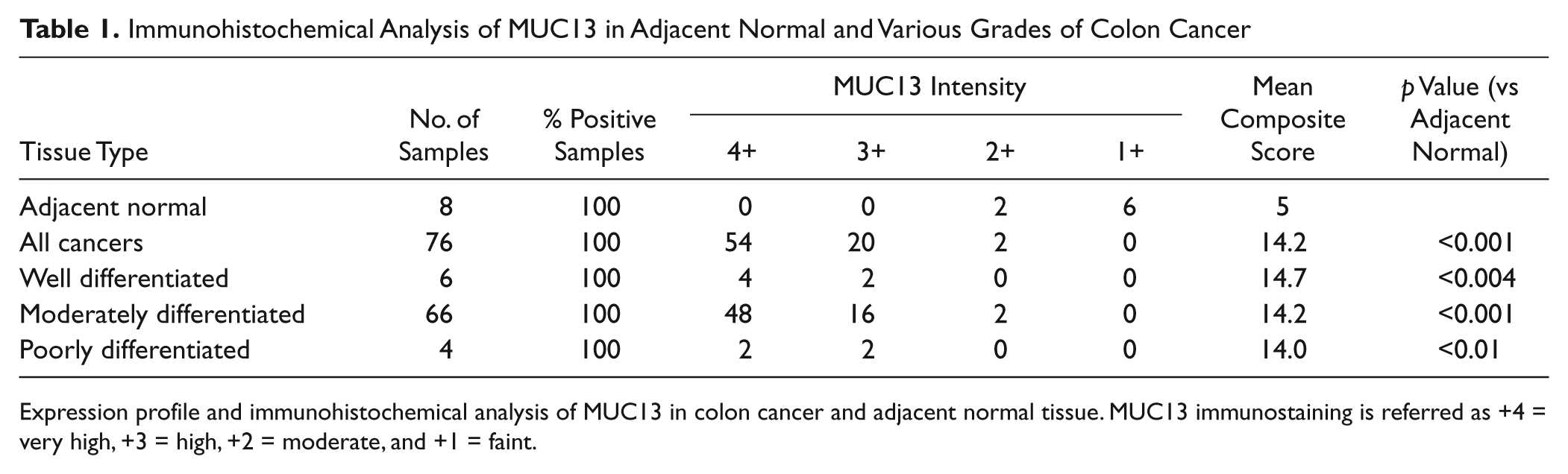
Expression profile and immunohistochemical analysis of MUC13 in colon cancer and adjacent normal tissue. MUC13 immunostaining is referred as +4 = very high, +3 = high, +2 = moderate, and +1 = faint.
To determine the expression of MUC13 in metastatic colon cancer with lesions to the liver, MUC13 IHC analysis was performed on a set of tissues containing adjacent normal (n=27), primary tumor (n=56), and liver metastasis (n=49) samples from each patient (Fig. 2 and Table 2). In contrast to adjacent normal tissues collected from non-metastatic colon cancer patients (Fig. 1A), MUC13 expression was expressed at a higher level in adjacent normal samples obtained from patients with metastatic colon cancer (Fig. 2A). However, MUC13 expression was still significantly (p<0.001) higher in primary colon tumor samples (MCS = 16) and liver metastatic lesions (MCS = 16) compared with adjacent normal tissue from these patients (MCS = 12.9; Table 2). The normal-appearing liver parenchyma did not express MUC13 at detectable levels. Strong MUC13 staining (+4 and +3) was observed in 100% of primary tumors (56 of 56) and liver metastasis lesions (49 of 49; Table 2).
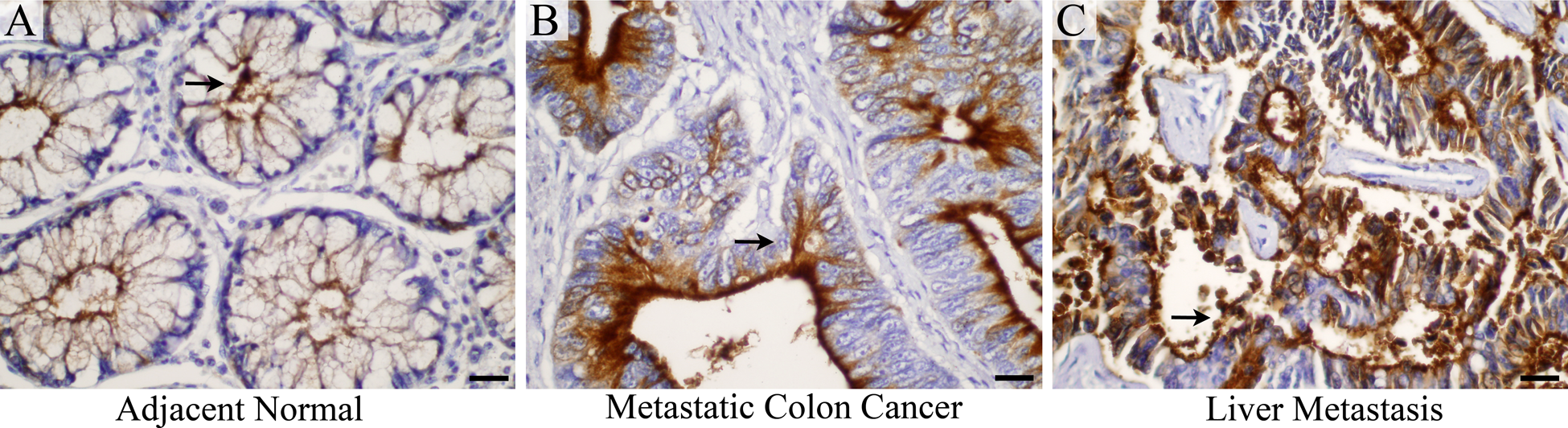
(A–C) MUC13 is overexpressed in metastatic colon cancer and liver metastasis: MUC13 expression (brown color, representative areas indicated by the black arrows) is shown in adjacent normal (A), metastatic colon cancer (B), and liver metastasis (C). Scale bars = 20 µm.
Immunohistochemical Analysis of MUC13 in Adjacent Normal and Metastatic Colon Cancer
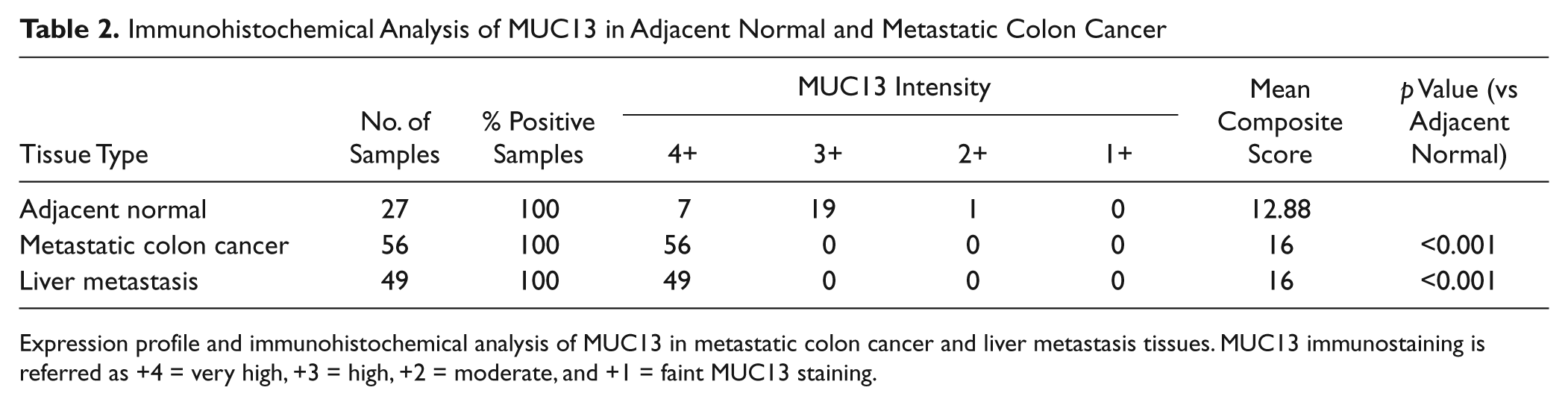
Expression profile and immunohistochemical analysis of MUC13 in metastatic colon cancer and liver metastasis tissues. MUC13 immunostaining is referred as +4 = very high, +3 = high, +2 = moderate, and +1 = faint MUC13 staining.
Aberrant Subcellular Localization of MUC13
Under normal physiological conditions, MUC13 is expressed at the apical surface of epithelial cells of the colon. Aberrant localization of MUC13 (i.e., expression on the basal surface, in the cytoplasm, or in the nucleus) may affect the cell’s ability to function appropriately. Hence, we sought to determine the subcellular localization of MUC13. In the adjacent normal tissues from patients with non-metastatic colon cancer, MUC13 localization was restricted to the apical surface of cells (membranous expression) and was not detected in the cytoplasm or nucleus (Fig. 3A–E, F). The non-metastatic colon cancer tissue samples predominantly showed membrane MUC13 expression; however, in addition to membrane staining, 23.7% of non-metastatic tissue samples also showed cytoplasmic and 10.5% showed nuclear staining (Fig. 3G). Although the expression of MUC13 in the cytoplasm and nucleus of non-metastatic colon cancer tissue samples was relatively faint, the localization pattern was clearly different compared with the adjacent normal tissue. Interestingly, metastatic colon cancer samples (both the primary colon tumor and the liver metastasis) had significantly higher levels of cytoplasmic and nuclear MUC13 staining (p<0.05) compared with adjacent normal tissue (Fig. 3A–E, F, colon cancer with liver metastasis). In metastatic colon cancer patients, both the primary colon tumor and the liver metastasis frequently exhibited aberrant MUC13 expression in the cytoplasm (89.3% and 85.7%, respectively) and the nucleus (64.3% and 69.4%, respectively; Fig. 3G). In addition, although the adjacent normal tissues of metastatic colon cancer with liver metastasis demonstrated relatively high membranous MUC13 expression, only 7.4% of samples exhibited cytoplasmic MUC13 expression, and no nuclear staining was observed. These data show that there is increased cytoplasmic and nuclear expression in metastatic colon cancer samples, suggesting a positive correlation of aberrant MUC13 localization with advanced metastatic stages of colon cancer.
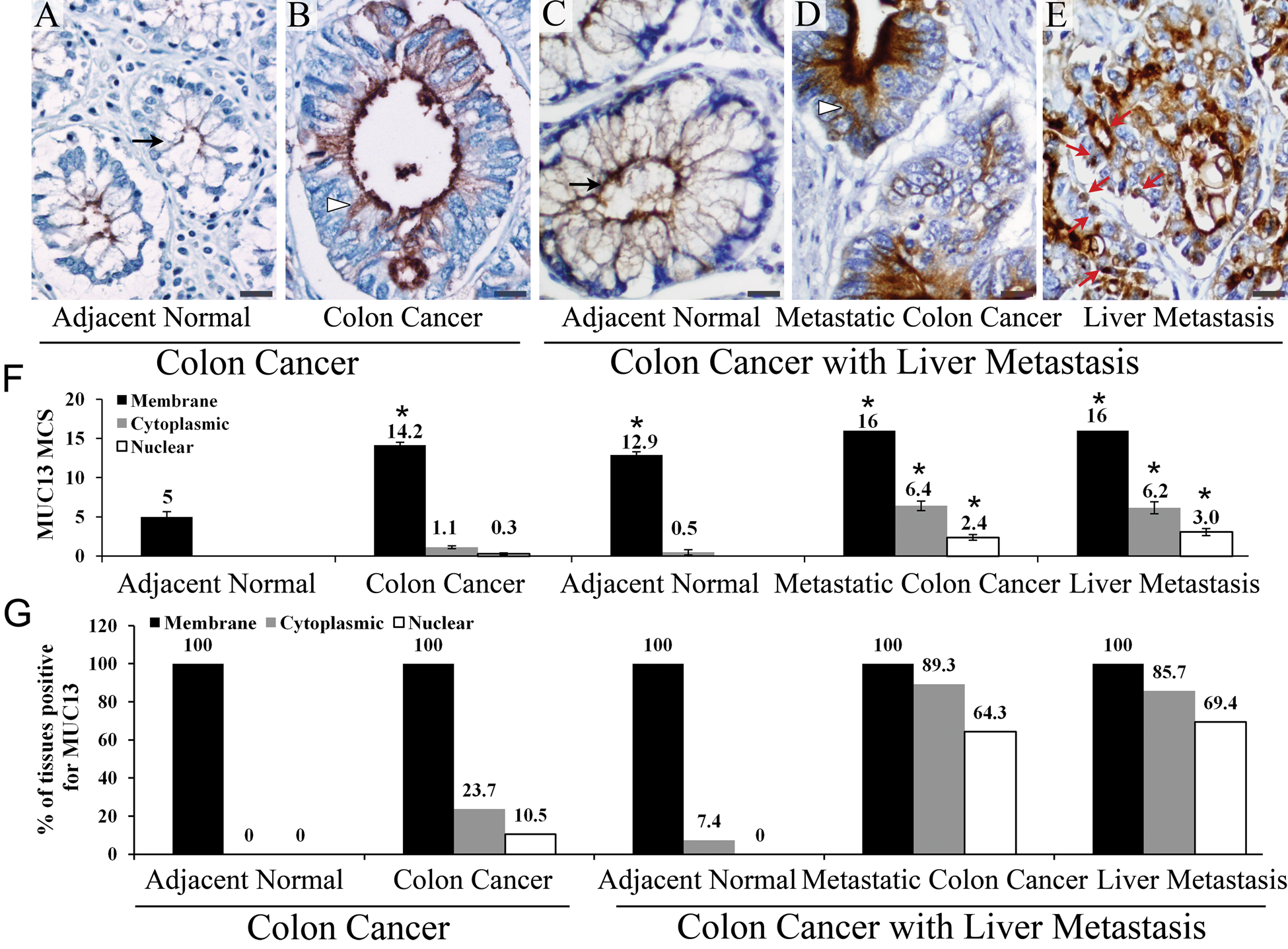
Comparative study of expression and localization of MUC13 in non-metastatic colon cancer and metastatic colon cancer with liver metastasis. (A–F) MUC13 expression and localization pattern: Membranous, cytoplasmic, and nuclear MUC13 localization is shown by black arrow, arrowhead, and a red arrow, respectively. *p<0.05. (G) Distribution of membranous, cytoplasmic, and nuclear MUC13 expression in tissues of non-metastatic colon and metastatic colon cancer with liver metastasis. Scale bars = 20 µm.
Association of MUC13 Subcellular Localization with Clinicopathological Features
To investigate the association between aberrant MUC13 subcellular localization and colon cancer progression, we analyzed the expression pattern of MUC13 in colon cancer tissue samples (n=60) obtained from Sanford Health. MUC13 localization was analyzed in three separate groups based on expression at the (1) membrane (M), (2) membrane plus cytoplasm (M + C), and (3) membrane plus cytoplasm plus nucleus (M + C + N). The clinicopathological parameters were compared among these three groups (Table 3). There was no significant correlation between the localization of MUC13 expression and the patient’s age, patient’s sex, and tumor’s depth of invasion. However, MUC13 subcellular localization significantly correlated with both tumor size and grades (Table 3). In pairwise comparisons for tumor size (>5 cm vs ≤5 cm), M + C + N and M + C groups had relatively larger tumors (>5 cm tumor) compared with the M group (p=0.005 and p=0.04, respectively). Similarly, for tumor differentiation (well vs moderately vs poorly differentiated), M + C + N and M + C groups had more poorly differentiated tumors compared with the M group (p=0.001 and p=0.005, respectively). In addition, although not statistically significant, 38.5% of samples with extensive lymph node involvement (N2) had nuclear staining, whereas only 13.8% of samples without nodal involvement (N0) were positive for nuclear MUC13 staining. Together, these data suggest a correlation between increased tumor volume and poor differentiation with aberrant MUC13 subcellular localization.
Association of MUC13 Expression with Clinicopathological Parameters
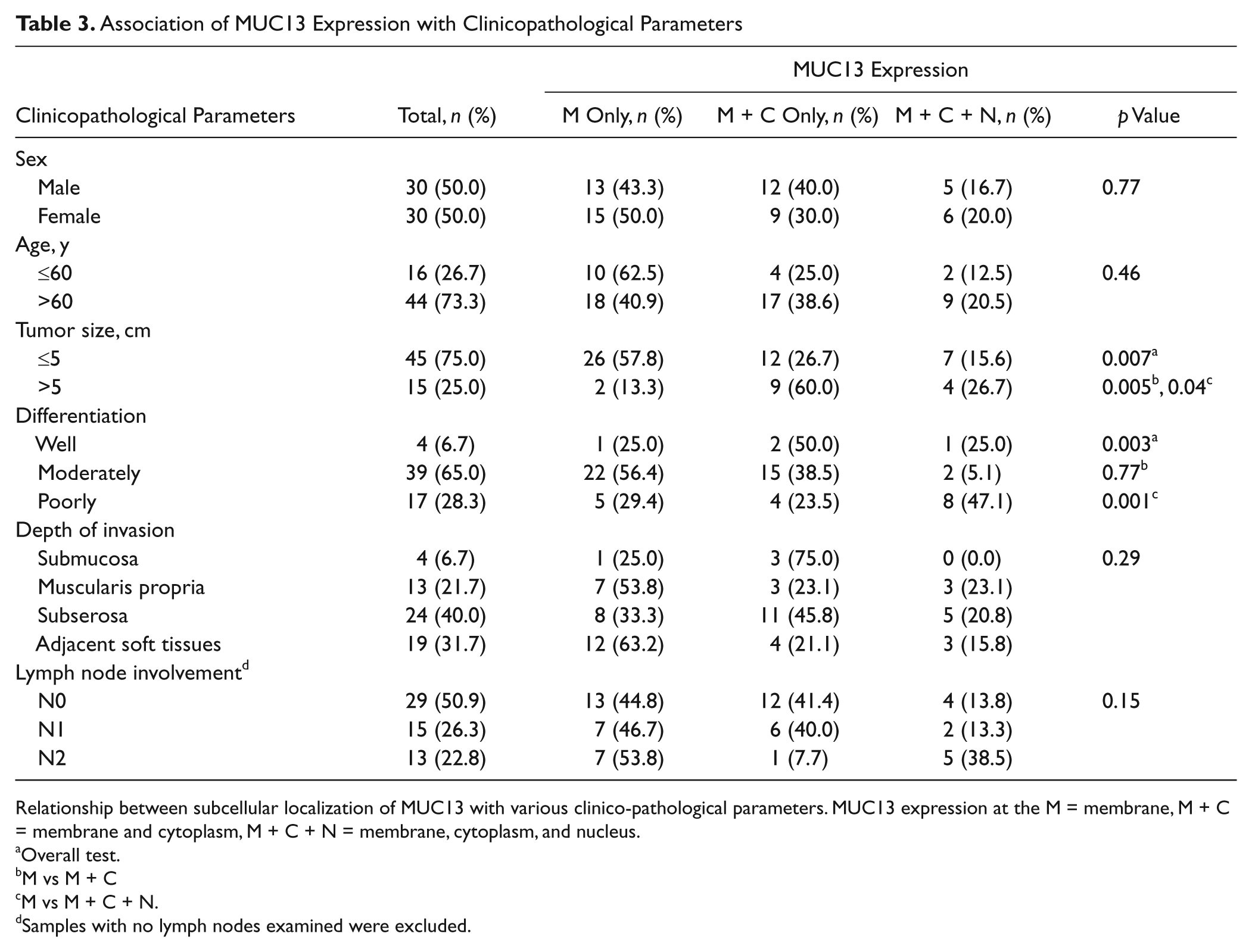
Relationship between subcellular localization of MUC13 with various clinico-pathological parameters. MUC13 expression at the M = membrane, M + C = membrane and cytoplasm, M + C + N = membrane, cytoplasm, and nucleus.
Overall test.
M vs M + C
M vs M + C + N.
Samples with no lymph nodes examined were excluded.
Expression of MUC13 in Colon Cancer Cell Lines
In addition to tumor samples, the MUC13 expression profile was also analyzed in six different colon cancer cell lines using end-point polymerase chain reaction (PCR), quantitative real-time PCR (qRT-PCR), and confocal immunofluorescence microscopy analyses. Of the six colon cancer cell lines tested, MUC13 RNA was not detectable or only faintly detectable in SW48 and SW480 colon cancer cell lines. SW620 cells showed high levels of MUC13 RNA, whereas LoVo, T-84, and HT-29 showed moderate MUC13 RNA levels (Fig. 4A). MUC13 mRNA expression in these cell lines by qRT-PCR was very similar to end-point PCR. The PCR results were confirmed by confocal immunofluorescence microscopy. MUC13 expression was undetectable in SW48 and SW480 cell lines, and moderate to high expression of MUC13 was detected in SW620, LoVo, T-84, and HT-29 colon cancer cell lines (Fig. 4B). At the subcellular localization level, MUC13 was predominantly localized in the membrane and cytoplasm of HT-29, LoVo, SW620, and T-84 colon cancer cell lines.
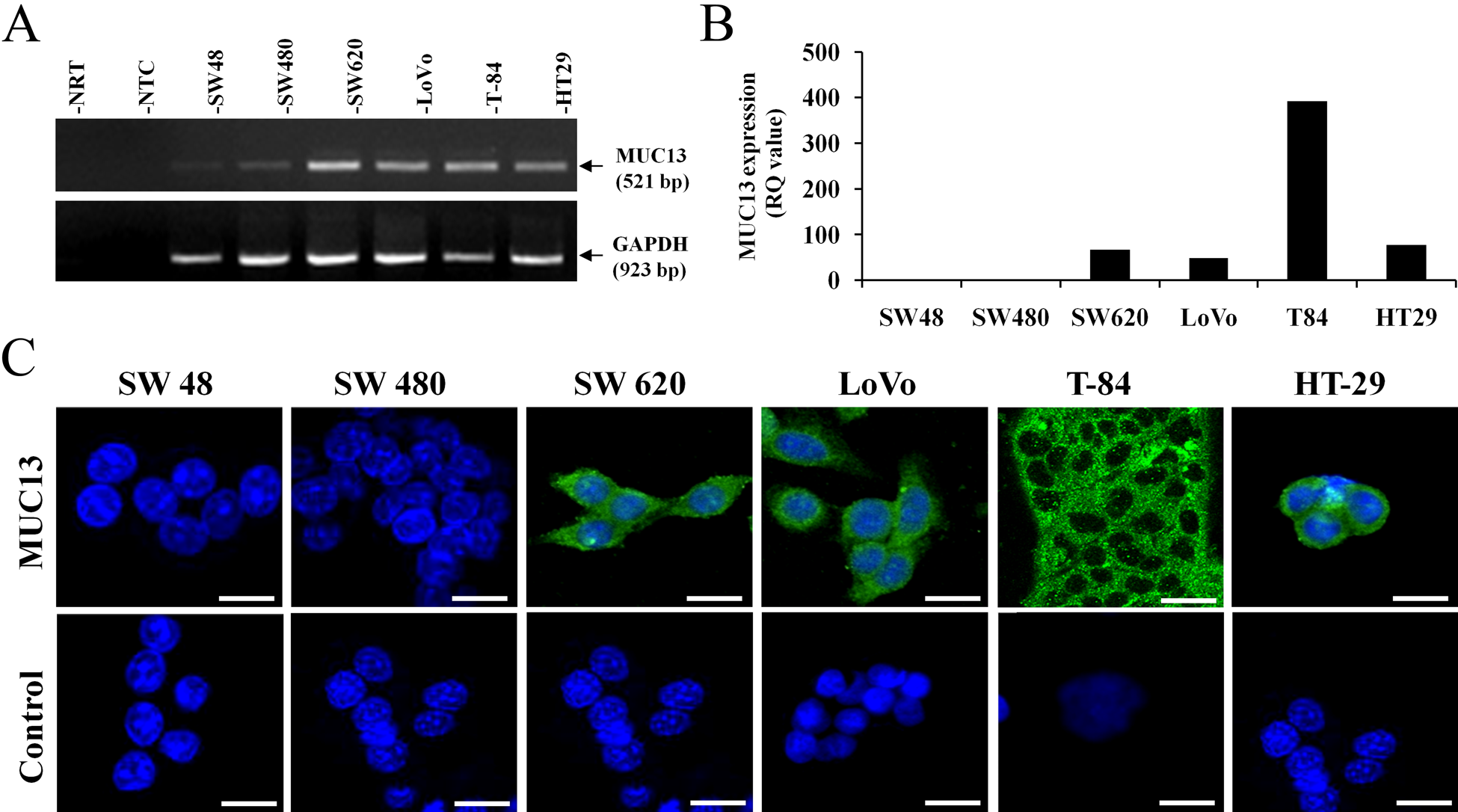
MUC13 expression in colon cancer cell lines. (A) End-point PCR analysis (NTC, non-template control; NRT, no reverse transcriptase control), (B) quantitative real-time PCR analysis (RQ, relative quantity), and (C) immunofluorescence confocal microscopy analysis. Scale bars = 20 µm.
Discussion
Colon cancer is still a leading cause of cancer-related deaths in the United States. The mortality rate due to advanced colon cancer has not significantly changed despite considerable progress in cancer research. Surgical resection of polyps and adjuvant therapy has improved patient survival when detected at the early disease stage. However, once colon cancer has metastasized to distant organs (e.g., liver, lung, and bones), the 5-year survival rate drops significantly, to approximately 10% (Siegel et al. 2011). Therefore, further understanding of the molecular events involved in metastasis is important for the development of more effective treatment strategies. In the present study, we have investigated the expression profile and localization patterns of the transmembrane mucin, MUC13, in a panel of non-metastatic and metastatic colon cancer with liver metastasis tissue samples by using a novel anti-MUC13 monoclonal antibody. Our study demonstrated that MUC13 is overexpressed in both metastatic and non-metastatic colon cancer tissues. In addition, MUC13 is aberrantly localized in the cytoplasm and nucleus in metastatic colon cancer samples (in both the primary colon tumor and in the liver metastasis).
In adjacent normal tissues of colon cancer, MUC13 localizes at the apical membrane, and its expression is fairly low. In contrast, MUC13 expression was significantly higher in various grades of colon cancer (well, moderately, and poorly differentiated). In non-metastatic colon cancer, MUC13 was predominantly localized at the cell membrane, with low levels of MUC13 expression in the cytoplasm. Furthermore, metastatic colon cancer with liver metastasis tissues had high MUC13 expression at the membrane, in the cytoplasm, and in the nucleus of the cancer cells. These observations suggest that the consistent increase in expression of MUC13 and its cytoplasmic/nuclear localization may play a role in the development and progression of colon cancer and its potential to metastasize at distant sites. This observation is in accordance with our findings from clinical tissues in which cytoplasmic and nuclear localization was associated with larger tumor size (>5 cm) and poorly differentiated stage of colon cancer. In a recent publication, Sheng et al. (2011) reported that MUC13 knockout mice exposed to dextran sulfate sodium (DSS) developed more severe colitis with an increased number of apoptotic cells. This report also showed that knockdown of MUC13 increased the sensitivity of human colon cancer cells to apoptosis from DSS, Actinomycin D, UV, and TRAIL exposure. In addition, MUC13 expression in colon biopsies revealed that increased MUC13 expression was associated with inflammatory conditions (Sheng et al. 2011). These findings support our results suggesting an association between MUC13 expression and the development and progression of colon cancer.
Surprisingly, adjacent normal tissues of metastatic colon cancer had high MUC13 expression, which is in contrast to the MUC13 expression in adjacent normal tissues of non-metastatic colon cancer. In addition, immunohistochemical analysis of MUC13 in diverticulosis (n=10) and dysplasia (n=10) tissue samples showed a moderate level of MUC13 expression (higher than adjacent normal from non- metastatic but lower than adjacent normal from metastatic colon cancer; data not shown). The difference in MUC13 expression between the adjacent normal tissues of these two groups of colon cancer samples suggests that MUC13 expression may be regulated by the adjoining malignancy, possibly through various signals from the tumor microenvironment, such as inflammation. Alternatively, the increased expression of MUC13 may be part of a “field effect” (Fearon and Vogelstein 1990). Development of colon cancer is a multistep process in which genetic changes occur in a stepwise fashion resulting in multiple acquired genetic changes that favor the proliferation and survival of a subset of cells that ultimately develop oncogenic phenotypes. Cells adjacent to the cancer, the so-called adjacent normal cells, also contain some of the genetic aberrations that are present in cancer cells and therefore are not truly normal. Our current observation of increased MUC13 in adjacent “normal” tissue from metastatic colon cancer patients suggests that overexpression of MUC13 may serve as an important indicator of metastasis.
MUC13 is a newly identified transmembrane protein, and its role in cancer has only recently been explored in epithelial cancers. High levels of MUC13 expression have been shown in ovarian cancer (Chauhan et al. 2009) and the intestinal type of gastric cancer (Shimamura et al. 2005). Reports regarding the expression levels of MUC13 in colon cancer have varied. In the first published report of MUC13, Williams et al. (2001) reported a decrease of MUC13, which was supported by Packer et al. (2004). Walsh et al. (2007) later reported the overexpression of MUC13 in the membrane and cytoplasm of colon adenocarcinomas using a polyclonal anti-MUC13 antibody with no reference to MUC13 expression in the nucleus. Our current study sought to resolve the discrepancies regarding MUC13 expression in non-metastatic and metastatic colon cancer using a novel monoclonal antibody (Shimamura et al. 2005). We observed a clear increase in MUC13 expression in a large cohort of colon cancer samples compared with adjacent normal colon tissues, supporting the findings published by Walsh et al. (2007). In addition, we report increased aberrant expression of MUC13 (cytoplasmic and nuclear) in metastatic colon cancer with liver metastasis. Previously published work from our laboratory shows that MUC13 can be involved in modulating the expression and activation of several proteins, including HER2, PAK1, and S-100A4 (Chauhan et al. 2009, 2012). These proteins are known to be involved in pathways that stimulate cell migration/metastasis, suggesting that MUC13 could directly play a role in the ability of cancer cells to metastasize. In addition, the extracellular domain of MUC13 is predicted to interfere with adhesion to both adjacent cells and the extracellular matrix, which could cause increased migration/metastasis of MUC13-overexpressing cells.
We also investigated MUC13 expression in colon cancer cell lines in vitro. Of the six colon cancer cell lines tested, high MUC13 expression was observed in four cell lines (HT-29, T-84, LoVo, and SW620). The localization of MUC13 was predominantly membranous and cytoplasmic. In contrast, faint or undetectable MUC13 expression was observed in SW480 and SW48 cell lines at the mRNA and protein levels. Interestingly, MUC13 expression was different between SW620 and SW480 cell lines, which are paired colon cancer cell lines. SW480 (non-metastatic cells) were obtained from a patient, and 1 year later, SW620 (metastatic cells) were obtained from a lymph node metastatic lesion from the same patient (Leibovitz et al. 1976). PCR and immunofluorescence analysis clearly indicate that MUC13 is only faintly expressed in the non-metastatic colon cancer cell line (SW480) and expressed at a high level in the metastatic colon cancer cell line (SW620), which is in accordance with our results (higher and more aberrant expression of MUC13 in metastatic colon cancer). These data further indicate that MUC13 may be a contributing factor for colon carcinogenesis and metastasis. We are currently investigating the potential pathways involved in colon cancer carcinogenesis and metastasis using SW620 and SW480 paired cell line culture models.
In conclusion, our immunohistochemical analysis revealed differential patterns of MUC13 expression and localization in adjacent normal, colon cancer, and liver metastasis tissues as summarized in Figure 5. In addition to being a potential biomarker for colon cancer, the high expression level of MUC13 in colon cancer tissue may identify MUC13 as a potential target for antibody-guided therapy in the future. Further studies are warranted to determine the functional roles of MUC13, including its translocation into the cell nucleus and effects on cell proliferation and metastasis. In addition, a study with a large cohort of samples is needed for exploration of MUC13 in colon cancer carcinogenesis, metastasis, and survival.
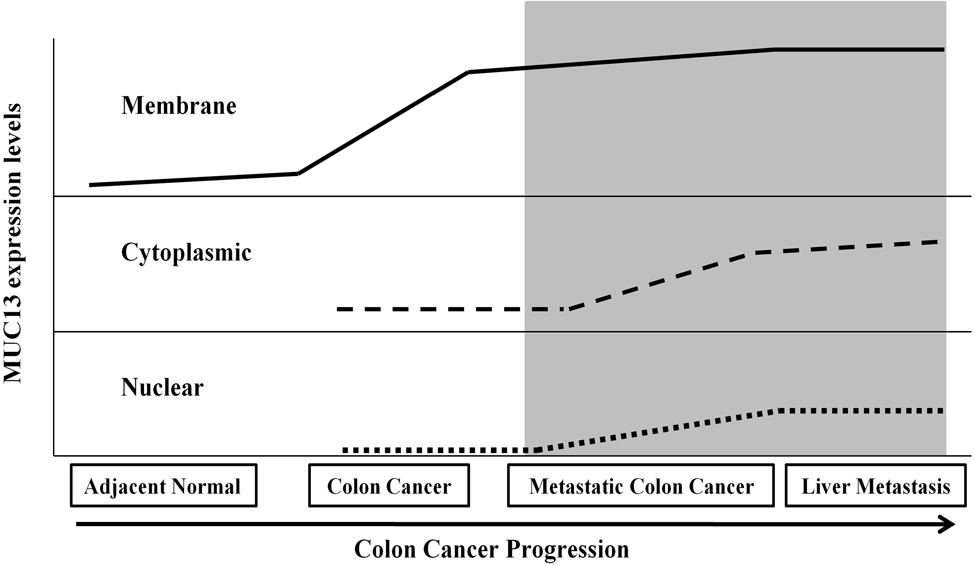
A summarized schematic representation of expression and localization of MUC13 with various stages of colon cancer progression and metastasis. The shaded area represents high MUC13 expression and its localization in the membrane, cytoplasm, and nucleus in advanced stage colon cancer (metastatic colon cancer and liver metastasis).
Footnotes
Acknowledgements
The authors gratefully acknowledge Cathy Christopherson for editorial assistance. The authors also thankfully acknowledge Sanford Research/USD Core facilities, which are supported by NIH R01 CA142736 to SCC, P20 RR17662, and P20RR024219 COBRE grants.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was partially supported by grants from PC073887, Governor’s Cancer 2010, and NIH R01 CA142736.
