Abstract
Glanzmann thrombasthenia (GT) is an inherited, intrinsic platelet defect characterized by a quantitative or qualitative change in the platelet glycoprotein complex IIb-IIIa (integrin αIIbβ3). The subunits are encoded by separate genes and both subunits must be expressed for a stable complex to form on the platelet surface; therefore, a defect in either gene can result in GT.
Keywords
Dr. Eduard Glanzmann, a Swiss pediatrician, first described Glanzmann thrombasthenia (GT) in his patients in 1918; however, the association of a deficiency of the platelet glycoprotein complex IIb–IIIa with GT was not made until 1974.
14–
16
In the decade that followed, this glycoprotein complex was recognized as being the receptor that mediated platelet aggregation and was termed the fibrinogen receptor. In the 1990s, it was recognized that the fibrinogen receptor of platelets was an integrin and was designated as αIIbβ3 to comply with the nomenclature for that system. It was recognized at this time that the subunits were encoded by separate genes and that both subunits were required for a stable complex to form on the platelet surface. This disease has been well documented at the clinical and molecular levels in human beings and dogs
2,
3,
7,
13
(
GT was recently described in two horses with histories of chronic, intermittent epistaxis that was unrelated to exercise. 12 Horse No. 1 was a 7-year-old thoroughbred cross gelding located in Hatfield, Hertfordshire, England, while horse No. 2 was a 4-year-old quarter horse mare located in Auburn, Alabama. These animals had normal coagulation screening test results, normal platelet numbers, and normal von Willebrand factor antigen levels. Clot retraction and platelet aggregation responses were markedly impaired. Flow cytometric studies indicated a reduction in the αIIbβ3 integrin on platelet surfaces. Based on these findings, both horses were diagnosed with GT.
The purpose of this study was to determine the cDNA sequences encoding normal equine αIIb and β3 and compare them with established human and canine cDNA sequences and to cDNA sequences determined in both affected horses.
Platelet-rich plasma was obtained by centrifuging 100–200 ml of EDTA-anticoagulated whole blood as previously described. 5 Platelets were counted and concentrated into pellets containing 5 × 109 platelets per pellet. Total RNA was harvested from platelet pellets using the Micro to Midi Total RNA Purification System (Invitrogen Corporation, Carlsbad, CA). First strand cDNA synthesis was performed using the Superscript™ First-Strand Synthesis System for reverse transcription-polymerase chain reaction (RT-PCR) using oligo(dT) primers (Invitrogen Corporation).
Genomic DNA was harvested from EDTA-anticoagulated whole blood using the QIAamp® DNA Blood Mini Kit (Qiagen, Inc., Valencia, CA). Most primers were designed based on homologous regions of canine (GenBank accession nos. AF153316 and AF116270) and human DNA sequences for αIIb and β3. 8, 17 Normal equine cDNA sequence was used to design some of the primers.
DNA segments were amplified by PCR in an overlapping fashion using normal equine cDNA or genomic DNA as templates initially (Tables 1 and 2). The experiments were then repeated using cDNA or genomic DNA isolated from horses 1 and 2. In selected experiments, genomic DNA from relatives of horse 2 (dam, sire, and two half-siblings) were also evaluated. Amplification products were separated via electrophoresis on 1.5% agarose gels. DNA was extracted from target bands using the QIAquick Gel Extraction Kit (Qiagen, Inc.). Harvested bands were sequenced directly by a laboratory using an ABI 3100 Genetic Analyzer. Nucleotide and amino acid sequences from different species were compared using the AlignX program of the VectorNTI suite (Informax Invitrogen Life Science Software, Frederick, MD).
Primers and polymerase chain reaction (PCR) conditions used to amplify equine αIIb cDNA.∗
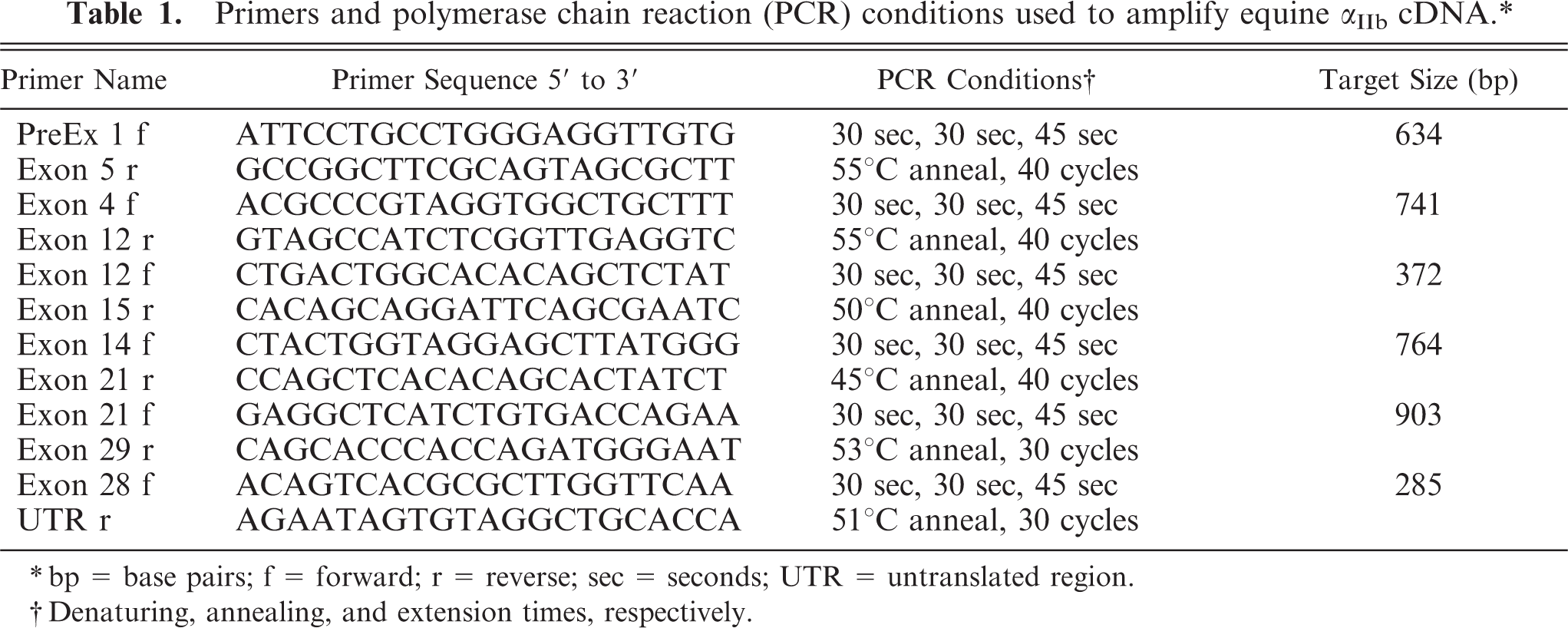
∗bp = base pairs; f = forward; r = reverse; sec = seconds; UTR = untranslated region.
†Denaturing, annealing, and extension times, respectively.
Primers and polymerase chain reaction (PCR) conditions used to amplify equine β 3 cDNA.∗

∗bp = base pairs; f = forward; r = reverse; sec = seconds; UTR = untranslated region.
†Denaturing, annealing, and extension times, respectively.
Normal equine cDNA sequence encoding αIIb had 82% nucleotide identity with human and 87% with canine sequences. At the amino acid level, percent identities were 82 and 81%, respectively. The cDNA encoding β3 in horses had 92% nucleotide identity with human and 93% with canine sequences. Amino acid identity for β3 was 95% when compared with both human and canine sequences. Within the four calcium-binding domains of αIIb, equine cDNA exhibited 86–88% identity with human nucleotide sequence. Within the four calcium-binding domains, amino acid identity between equine and human ranged from 83 to 91%. When equine nucleotide sequences were compared with canine sequences within the four calcium-binding domains, nucleotide identities ranged from 83 to 91%, and amino acid identities ranged from 75 to 100%. cDNA sequences for equine αIIb and β3 have been submitted to GenBank (GenBank accession nos. AY322154 and AY999064).
cDNA sequences encoding β3 in horses 1 and 2 with GT were identical to sequences obtained from a normal horse. cDNA sequences encoding αIIb were also identical except for a single guanine to cytosine (CGG to CCG) substitution in codon 41 in exon 2 (Fig. 1A, B). This change would result in the substitution of a proline for an arginine in a highly conserved region of the encoded protein.
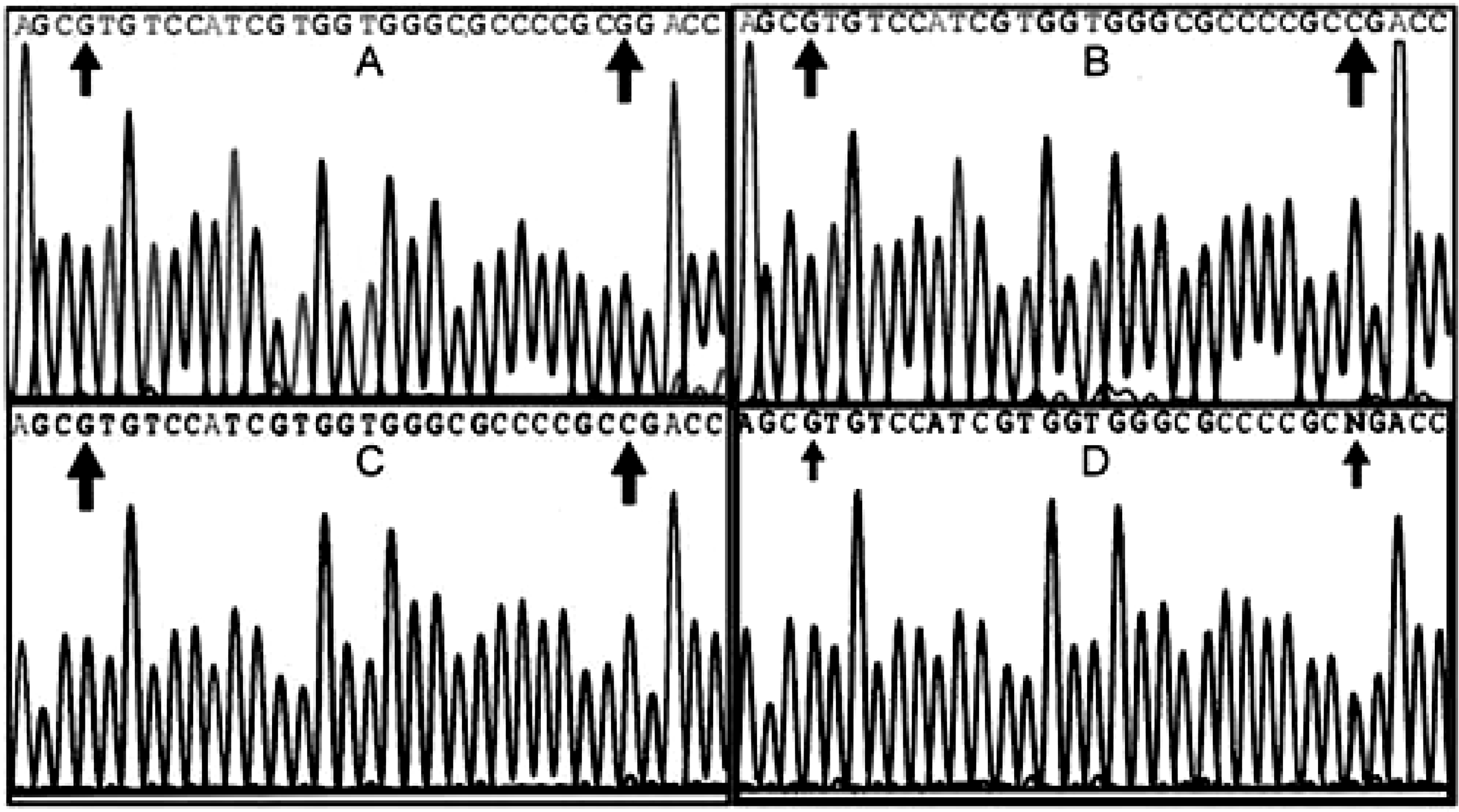
Exon 2 was evaluated by PCR amplification and sequencing using as template genomic DNA from 10 normal horses, from GT horses 1 and 2, and from the sire, dam, and two half-siblings of horse 2 that shared the same dam. Genomic DNA sequencing results indicated that horse 1 was homozygous for the single guanine to cytosine base change, while horse 2 was heterozygous for the base change (Fig. 1C, D). The genomic DNA sequences encoding exon 2 of 10 normal horses and the dam and two siblings of horse 2 were all identical. The sire of horse 2 was heterozygous for the base change. The sire, dam, and siblings were clinically normal.
Comparisons of the cDNA sequences that encode αIIb and β3 in humans and dogs have been reported. 9, 10 Horse cDNA sequence encoding αIIb was found to be more similar to dog (87%) than to human (82%), although when translated to the amino acid level, the percent identity with dog and human were very similar (81 and 82%, respectively). Murine monoclonal antibodies generated against human αIIb do not cross-react with either the canine or equine platelet integrin subunit (Boudreaux, personal observation). Horse cDNA sequences encoding β3 were more similar to human and dog sequences at both the nucleotide and amino acid level, with identities ranging from 92 to 95%. In spite of this similarity, monoclonal antibodies to human β3 that cross-react with the canine subunit do not recognize horse β3 (Boudreaux, personal observation). The reason for this discrepancy is not known but may be related to the conformation the β3 subunit achieves when it is complexed with αIIb on the platelet surface.
GT has been well characterized at the functional, biochemical, and molecular level in humans and dogs. In dogs, two different mutations, both in the gene encoding αIIb, have been reported.
1,
11
According to the human GT database, maintained by the Mount Sinai School of Medicine, over 60 mutations have been documented in the gene encoding αIIb and over 40 mutations have been documented in the gene encoding β3 (
The base change documented in exon 2 of horses 1 and 2 would be predicted to result in the change of encoded amino acid 41 from an arginine to a proline. The marked difference in structure and charge of proline compared with arginine, as well as the presence of two adjacent prolines at positions 40 and 41, would be predicted to result in marked instability of the beta-propeller region of the encoded protein. A similar missense mutation in exon 2 of the gene encoding αIIb has been described in humans. 19 In this case, a single nucleotide change resulted in the substitution of a proline for a leucine at amino acid position 55. Experiments demonstrated that the mutation resulted in severe impairment of expression of the αIIbβ3 complex on the surface of transfected COS7 cells. The authors hypothesized that the substituted proline caused an aberrant conformation in the encoded protein that prevented association of αIIb with β3, with ultimate lack of expression of the complex on the surface of cells.
This is the first characterization of cDNA sequences encoding platelet αIIb and β3 in normal horses and in two horses with GT. Future studies will be aimed at identifying the second mutation in the apparently compound heterozygote horse 2. Horses that have platelet-type hemorrhage, particularly epistaxis, that do not have thrombocytopenia, vasculitis, or von Willebrand disease, are candidates for the diagnosis of GT. Molecular-based screening assays, based on molecular defects identified in horses with GT, will greatly facilitate the identification of GT in horses with otherwise unexplained platelet-type bleeding.
Footnotes
Acknowledgements
We thank Dr. J. Taintor, Dr. J. Schumacher, Mrs. D. Czerkawski, E. Whatley, S. Spencer, A. Hall, D. Shiver, A. Mitchell, and K. Worley for their help with this project.
