Abstract
The chronologic localization of Mycoplasma hyopneumoniae was examined by in situ hybridization in experimentally infected pigs for a period of 35 days after intratracheal inoculation. M. hyopneumoniae DNA was detected in bronchial and bronchiolar epithelial cells from infected pigs at 7, 14, 21, and 28 days postinoculation (DPI) and in alveolar and interstitial macrophages and type I pneumocytes from infected pigs at 14, 21, 28, and 35 DPI. Strong hybridization signals for M. hyopneumoniae were detected mainly at the luminal surface of bronchial and bronchiolar lining epithelial cells. When a hybridization signal was detected at the luminal surface of bronchial and bronchiolar lining epithelial cells, a given bronchus or bronchiole also exhibited peribronchiolar lymphoid cuffing. These observations suggested that the presence of M. hyopneumoniae in different tissues could be due to a difference in the duration of the infection.
Mycoplasma hyopneumoniae is the primary agent of enzootic pig pneumonia,14,20 a chronic respiratory disease endemic to pig farms that is characterized by high morbidity rates and low mortality rates.24 The clinical signs associated with M. hyopneumoniae infection include sporadic, dry, and nonproductive cough. Histologic lesions in the acute stage of the disease include loss of respiratory cilia, exfoliation of ciliated cells, and accumulation of neutrophils and macrophages in lumina and around airways. As the disease progresses, prominent lesions consist of extensive monocytic cuffs with occasional lymphoid germinal center formation around the airways and vessels.4,12,14
In situ hybridization and immunohistochemistry are potentially useful techniques for studying the pathogenesis of infectious agents and can be combined with histologic preparation of tissues to allow the simultaneous morphologic study lesions.6–9 However, immunohistochemistry is of limited use in the study of M. hyopneumoniae infection because a specific monoclonal antibody for M. hyopneumoniae is not yet available. M. hyopneumoniae antigen, however, has been detected in airway epithelium by immunohistochemistry using a polyclonal antibody.11 The success of these immunohistochemistry techniques is limited because M. hyopneumoniae shares antigenic determinants with two other porcine mycoplasmas, M. flocculare and M. hyorhinis.3,5 Furthermore, the nonpathogenic M. flocculare is commonly found in swine lung.3,5 M. hyopneumoniae-specific DNA probes are therefore a potentially valuable tool for the detection of M. hyopneumoniae nucleic acids by in situ hybridization of formalin-fixed, paraffin-embedded lung tissue. The objectives of this study were to determine the temporal distribution of M. hyopneumoniae and the cell types affected in pigs experimentally infected with M. hyopneumoniae. By employing in situ hybridization protocols a better understanding of the pathogenesis of mycoplasmal infection was achieved.
A field isolate of M. hyopneumoniae strain SNU98703 identified by the disc growth inhibition test using antiserum against M. hyopneumoniae ATCC 25934 (strain J)25 and polymerase chain reaction (PCR)16 was used for the experimental infection. M. hyopneumoniae was grown in Friis medium and harvested at pH 6.9. The mycoplasmal suspensions were centrifuged (20,000 × g) and washed three times in 0.25 M NaCl. To avoid undesirable contaminants, the culture medium was treated as previously described.23
Twenty-five Landrace–Duroc crossbred pigs 13 days of age were randomly allocated to infected or control groups. All pigs were serologically negative for M. hyopneumoniae, porcine reproductive and respiratory syndrome virus, and swine influenza virus by enzyme-linked immunosorbent assay (IDEXX Laboratories, Westbrook, ME). A tissue homogenate containing a derivative of M. hyopneumoniae strain SNU98703 (105 color changing units/ml) was administered intratracheally to 15 pigs in the infected group at a dilution of 1:100 in 10 ml of mycoplasmal Friis medium as previously described.25 Ten control pigs were exposed in the same manner to Friis medium and were maintained individually in stainless steel isolators. Three infected and two control pigs were euthanatized at 7, 14, 21, 28, and 35 days postinoculation (DPI). Tissues were collected from each pig at necropsy. The methods used were previously approved by the Institutional Animal Care and Use Committee, Seoul National University.
M. hyopneumoniae ATCC 25934 (strain J) was grown and harvested in Friis medium (pH adjusted to 6.9).13 The mycoplasma suspension was centrifuged (20,000 × g) and washed three times in 0.25 M NaCl. Genomic DNA was purified using the Rapidprep microgenomic DNA isolation kit (Pharmacia Biotech, Uppsala, Sweden) following the manufacturer's instructions. A 520-base pair DNA fragment was used as a probe.16 The forward and reverse primers were 5′-GTGTATCAAAATTGCCAATC-3′ (nucleotides 851–870) and 5′-TCCCATAACCTTGTCTTCAG-3′ (nucleotides 1351–1370), respectively.16 PCR was carried out as previously described.16
PCR products were purified using a 30-kd cutoff membrane ultrafiltration filter. The nucleotide sequences of the purified PCR products were determined by use of BigDye chemistry with the ABI Prism Sequencer (Applied Biosystems, Foster City, CA). Sequences of the purified PCR products were determined before PCR products were labeled by random priming with digoxigenin-dUTP (Boehringer Mannheim, Indianapolis, IN) according to the manufacturer's instructions. The probe for porcine circovirus 2 (PCV-2) was used as a negative probe.8 In situ hybridization was carried out as previously described.18 Positive tissue controls for M. hyopneumoniae were taken from lung of naturally infected pigs.18
Coughing was observed in all M. hyopneumoniae-infected pigs beginning at 10–14 DPI but was not observed in pigs in the uninfected control group. At necropsy, no visible gross lesions were seen in infected pigs at 7 DPI. Well-demarcated areas of dark red to purple firm parenchyma were seen in the cranial, middle, accessory, and caudal lung lobes of the infected pigs at 14, 21, and 28 DPI. The extent and distribution of lesions varied in each infected pig. Typical white mucoid exudate was observed in the bronchioles, bronchi, and trachea of the infected animals. There were no obvious signs of pleuritis or pericarditis. At 35 DPI, infected pigs presented minimal mycoplasmal lesions.
Microscopic examination revealed typical lesions consisting of peribronchial, peribronchiolar, and perivascular lymphoid hyperplasia in all M. hyopneumoniae-inoculated pigs. Peribronchial, peribronchiolar, and perivascular lymphoid nodules were mild at 7 DPI and significantly more severe at 28 DPI. Large mononuclear cells, polymorphonuclear leukocytes, and edema fluid accumulated in alveoli and lumina of airways. Thickened interalveolar septa were accompanied by regions of alveolar collapse at 14, 21, and 28 DPI. Neither macroscopic or microscopic lesions were observed in the uninfected control pigs.
Results of the in situ hybridization experiments are summarized in Table 1. The morphology of host cells was preserved despite the relatively high temperatures and chemical treatment required in the procedure. A very close cell-to-cell correlation among serial sections from each lung sample was confirmed by in situ hybridization. The signal intensity varied within and between anatomical structures in sections of a single animal and also between pigs. When lung tissues from M. hyopneumoniae-infected pigs at 7 DPI were hybridized with the nonradioactive digoxigenin-labeled DNA probe, bronchial and bronchiolar epithelial cells showed positive hybridization signals. A strong hybridization signal was detected mainly at the luminal surface of bronchial and bronchiolar lining epithelial cells (Fig. 1), whereas no hybridization signal was seen in the cytoplasm of these cells. Application of in situ hybridization to sections of lung from all infected pigs necropsied at 14, 21, and 28 DPI revealed. M. hyopneumoniae DNA at the luminal surface of bronchial and bronchiolar epithelial cells, alveolar and interstitial macrophages, and type I pneumocytes (Fig. 2). Positive hybridization signal was especially intense at bronchial and bronchiolar epithelial cells, whereas hybridization of alveolar and interstitial macrophages and type I pneumocytes was sparse. When hybridization signal was detected at the luminal surface of bronchial and bronchiolar lining epithelial cells, a given bronchus or bronchiole also exhibited peribronchiolar lymphoid cuffing. Hybridization signals, however, were not seen in these peribronchiolar lymphoid cuffs. Considerably fewer bronchial and bronchiolar epithelial cells in sections from the infected pigs had M. hyopneumoniae DNA at 35 DPI. Hybridization signal was not detected in alveolar and interstitial macrophages or type I pneumocytes from infected pigs at 7 DPI.
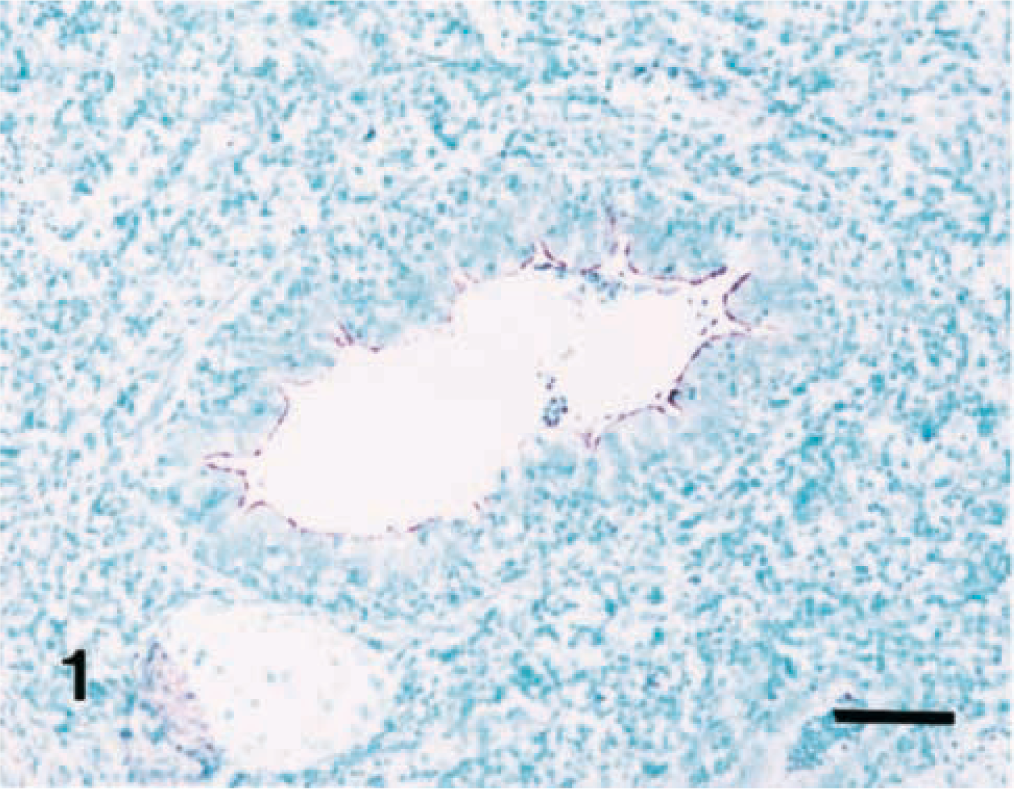
Lung; pig. Experimental infection with Mycoplasma hyopneumoniae, 14 days postinoculation. M. hyopneumoniae DNA was detected on the luminal surface of bronchiolar epithelial cells. In situ hybridization, nitroblue tetrazolium/5-bromocresyl-3-indolylphosphate, methyl green counterstain. Bar = 55 μm.
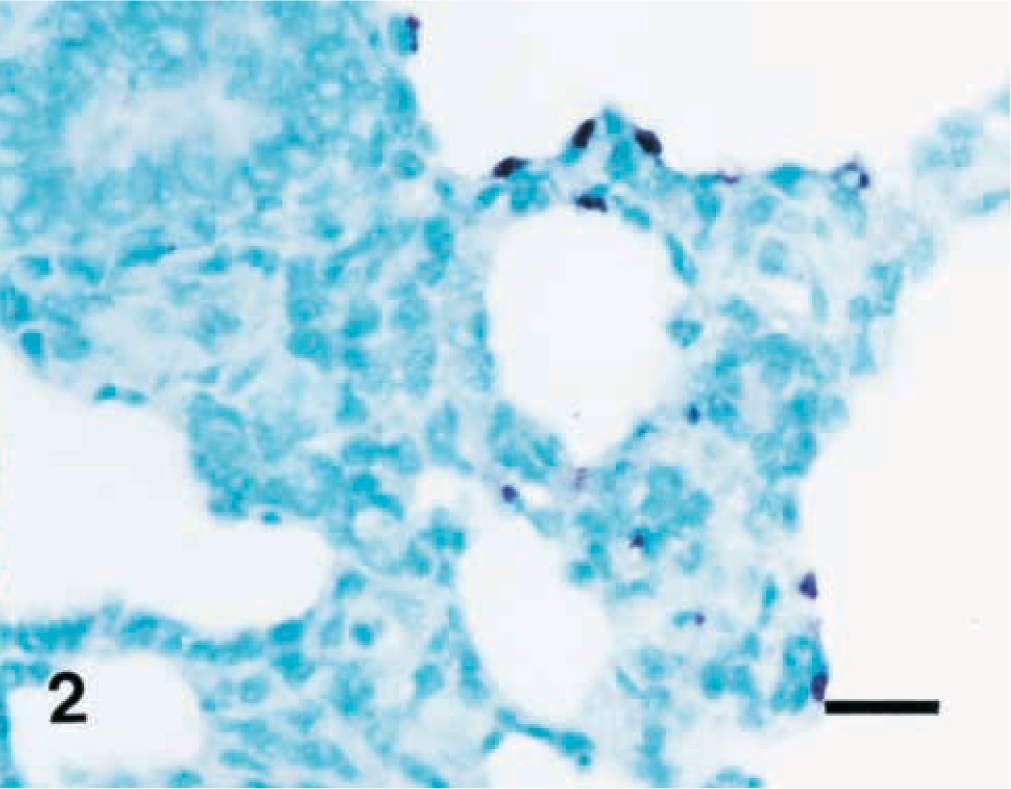
Lung; pig. Experimental infection with Mycoplasma hyopneumoniae, 21 days postinoculation. M. hyopneumoniae DNA was detected in the type I pneumocytes. In situ hybridization, nitroblue tetrazolium/5-bromocresyl-3-indolylphosphate, methyl green counterstain. Bar = 30 μm.
In situ hybridization results in pigs experimentally infected with Mycoplasma hyopneumoniae.
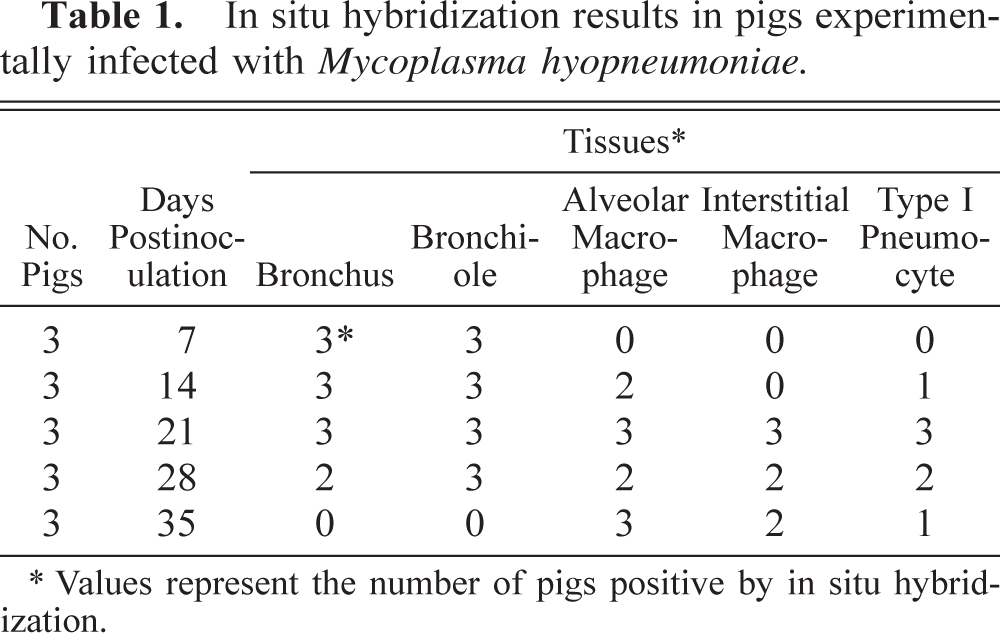

Values represent the number of pigs positive by in situ hybridization.
No hybridization signal was consistently recorded for tissue sections treated with DNase I prior to in situ hybridization (data not shown). Sections from negative control pigs showed no hybridization signal for M. hyopneumoniae. Probes for PCV-2 gave consistently negative results in all tissues tested.
In the present study, M. hyopneumoniae was detected in all infected pigs until 35 DPI using in situ hybridization. These observations are in agreement with those of a previous study.26 In that study, M. hyopneumoniae was detected by PCR. But PCR cannot provide information regarding the specific location of the M. hyopneumoniae in affected tissues.26 In situ hybridization can identify host cell types infected with M. hyopneumoniae.18 The major site of infection was the bronchial and bronchiolar epithelium, although other cells were also infected. From a previous study, it was not possible to explain why variable localization of M. hyopneumoniae was seen in naturally infected pigs.18 Infected pigs at 7–28 DPI showed positive in situ hybridization for M. hyopneumoniae at the luminal surface of bronchial and bronchiolar epithelial cells. In contrast, positive type I pneumocytes and alveolar and interstitial macrophages were observed in pigs necropsied at 14–35 DPI. These observations suggest that changes in the localization of M. hyopneumoniae can occur during the course of infection. However, this difference in distribution of M. hyopneumoniae also could be due to infection with different strains of M. hyopneumoniae.
M. hyopneumoniae has been shown in vivo to be resident in the bronchial and bronchiolar epithelium.4,11,18,21 Although bronchiolar lesions are not a feature of mycoplasmal pneumonia, bronchiolar epithelial cells appear to be a significant route of exposure. M. hyopneumoniae colonizes the surface of ciliated epithelial cells in the bronchi and bronchioles of pigs and does not invade epithelial cells.1 Previous studies have shown that the adherence of M. hyopneumoniae to ciliated epithelial cells is mainly mediated by receptor–ligand interactions.17,28,29 In the present study, M. hyopneumoniae was detected by in situ hybridization at the luminal surface of epithelial cells in the bronchi and bronchioles but not in the cytoplasm of these cells. Intimate attachment of this organism to cilia during infection has been well documented.4,21 This association leads to a progressive loss of cilia, desquamation of the epithelium, and development of pneumonia.
Peribronchiolar lymphoid hyperplasia is a predominant histologic lesion induced in pigs by M. hyopneumoniae,12,22 and M. hyopneumoniae has a mitogenic effect on swine lymphocytes.22 In the present study, a hybridization signal was always detected at the luminal surface of bronchial and bronchiolar lining epithelial cells when a given bronchus or bronchiole had peribronchiolar lymphoid cuffs. However, hybridization signals were not seen in the peribronchiolar lymphoid cuffs. These findings suggest that M. hyopneumoniae binds to the luminal surface of bronchial and bronchiolar lining epithelial cells and then activates the immune system to induce peribronchiolar lymphoid hyperplasia of these bronchi and bronchioles.
Porcine respiratory disease complex (PRDC) is an economically significant respiratory disorder characterized by slow growth, decreased feeding efficiency, lethargy, anorexia, fever, cough, and dyspnea.15 Although pneumonia in pigs with PRDC is due to a combination of both viral and bacterial agents, M. hyopneumoniae is considered the most important causative agent of PRDC.27 Disruption of airway epithelium by M. hyopneumoniae is one of the factors that potentiate the combined infection commonly seen in PRDC.10 Under field conditions, disruption of the mucociliary apparatus and airway epithelium could potentially lead to increased secondary viral and bacterial infections. Combined respiratory infection in pigs is typically associated with porcine reproductive and respiratory syndrome virus, swine influenza virus, Pasteurella multocida, and Actinobacillus pleuropneumoniae and less frequently with Arcanobacterium pyogenes, streptococci, and staphylococci.2,15,19
Results of this experiment further clarify and confirm previous observations regarding the distribution of M. hyopneumoniae;11,18 the spatial distribution of M. hyopneumoniae changed during the course of the experimental infection. The in situ hybridization technique was useful for the detection of M. hyopneumoniae DNA in tissues and may also be a valuable technique for studying the pathogenesis of mycoplasmal infection.
Footnotes
Acknowledgements
The research reported here was supported by the Ministry of Agriculture, Forestry and Fisheries—Special Grants Research Program (MAFF-SGRP) and by the Brain Korea 21 Project, Republic of Korea.
