Abstract
Chelonid herpesvirus (ChHV) infection in tortoises associated with stomatitis-rhinitis complex is a severe, mostly epizootic disease characterized by proliferative and diphtheroid-necrotizing glossitis, pharyngitis, rhinitis, and tracheitis, often occurring with pneumonia and encephalitis. The UL5 gene from a German ChHV isolate was used to generate a digoxigenin-labeled 307-base-pair DNA probe by polymerase chain reaction (PCR). ChHV DNA was detected in paraffin-embedded tissues of five naturally infected tortoises (two Afghan tortoises [Testudo horsfieldii], USA; two Hermann's tortoises [Testudo hermanni], Switzerland; one T. hermanni, Germany) by means of in situ hybridization (ISH) and PCR. Distribution of ChHV DNA exhibits many characteristics of alphaherpesvirus but also some characteristics of betaherpesvirus infections. The amino acid sequence of a portion of the ChHV UL5 homolog exhibited more than 50% similarity to alphaherpesvirus UL5 proteins. Nuclear hybridization signals were detected in epithelial cells of the lingual mucosa and glands. Furthermore, ChHV DNA was observed in tracheal epithelium, pneumocytes, hepatocytes, the renal tubular epithelium, cerebral glia cells and neurons, and intramural intestinal ganglia. ChHV DNA in endothelial cells of many organs underlines the systemic character of the disease. Importantly, ChHV DNA was detected by ISH in multiple tissues of tortoises originating from different geographic provenances. This indicates a high degree of conservation of the UL5 gene fragment among viruses prevalent in tortoises on different continents. With the described ISH, a molecular biological tool is available for rapid and specific diagnosis of ChHV infections and, more importantly, comparative pathogenetic studies of ChHV isolates from geographically unrelated regions.
Keywords
Herpesvirus infections are widespread and occur in most classes of vertebrates including fish, amphibians, and reptiles. In chelonians, herpesviruses have been associated with several disease complexes, which are characterized by diphtheroid-necrotizing stomatitis, hepatitis, rhinitis, tracheitis, and pneumonia in tortoises. Herpesviruses have also been associated with skin diseases in sea turtles, such as gray patch disease and fibropapillomatosis. 12 15 29 30 Herpesviruses identified in various species of turtles and tortoises have been preliminarily named chelonid herpesviruses (ChHVs), but represent an up-to-now unassigned species in the herpesvirus family. 1 32 Classification of ChHVs is mainly based on putative differences in the host spectrum. ChHV-1 was first described in association with gray patch disease in captive green sea turtles (Chelonia mydas) in the West Indies. This disease is characterized by patchy gray areas of hyperkeratotic and necrotic papules that occur over the head, neck, and flippers. 30 ChHV-2 was seen in two Pacific pond turtles (Clemmys marmorata) with fatal hepatic necrosis. 8 A similar disease has been seen in painted turtles (Chrysemys picta) and in map turtles (Graptemys pseudogeographica) in association with herpesvirus-like particles. These viruses have been preliminarily named ChHV-3. 5 14 ChHV-4 was seen in tissues of Argentinian tortoises (Geochelone chilensis) with necrotizing stomatitis or mouth rot. Interestingly, red-footed tortoises (Geochelone carbonaria) kept together with the diseased Argentinian tortoises remained clinically healthy. 13 ChHV infections have been associated with glossitis, stomatitis, enteritis, and meningoencephalitis in Hermann's tortoises (Testudo hermanni); with stomatitis and enteritis in Afghan tortoises (Testudo horsfieldii); 2–4, 6 10 16 18 19 21 22 with stomatitis, tracheitis, and pneumonia in desert tortoises (Gopherus agasizii); 9 28 and with stomatitis and encephalitis in spur-thighed tortoises (Testudo graeca). 3 4 11 22 Epizootics of chronic seromucous rhinitis (running nose syndrome) were described in large populations of captive T. graeca. 23 This outbreak was part of a series of epidemic ChHV infections that have occurred in Europe during the last decade. In most cases, outbreaks followed shared housing of different tortoise species after addition of new animals. In all of these cases, presumed carrier species, especially T. graeca, remained healthy, whereas other, presumably less resistant species, became sick or died. Kabisch and Frost 16 initially reported the isolation of herpesviruses from T. hermanni and T. horsfieldii with stomatitis–rhinitis complex and developed serologic tests for diagnosis in cell culture. Today, several ChHV isolates from tortoises from different geographic regions have been reported, but molecular biological data on ChHVs have not been available to date. 20 21 The data presented here could serve as a basis for the needed classification of ChHVs. The developed in situ hybridization (ISH) assay could provide a tool for further investigations leading to a more profound understanding of the pathogenesis of this important disease in tortoises.
Materials and Methods
Animals
Formalin-fixed, paraffin-embedded tissues of five tortoises from different geographic regions were included in this study. All tortoises had clinical and pathologic findings consistent with the stomatitis–rhinitis complex. 3 Tortoises No. 1 and No. 2 were captive, female T. horsfieldii, which were held together and submitted (No. 2 4 weeks later than No. 1) for necropsy at Washington Animal Disease Diagnostic Laboratory, Washington State University, Pullman, Washington, USA. Tortoises No. 3 and No. 4, both T. hermanni, suffered from stomatitis and No. 3 was serologically positive for herpesvirus antibodies. Both animals died and the heads were submitted for histopathology to the Institut für Tierpathologie, Universität Bern, Bern, Switzerland. Tortoise No. 5 was one of 13 diseased T. hermanni that were part of a collection of tortoises in southern Germany in which an outbreak of stomatitis–rhinitis complex occurred in 1995. 20 Necropsy and tissue sampling for virus isolation were conducted at the Institut für Geflügelkrankheiten and histopathology was performed at the Institut für Veterinär-Pathologie, Justus-Liebig-Universität Giessen, Giessen, Germany.
Tissue processing
Samples collected for histopathologic investigation and electron microscopy (Table 1) were fixed in 4% neutral buffered formalin or 2.5% glutaraldehyde in 0.1 M sodium cacodylate (pH 7.4) for 24 hours. Formalin-fixed tissues were routinely embedded in paraffin, cut at 4 μm, and stained with hematoxylin and eosin (HE). Glutaraldehyde-fixed samples were embedded in Epon-Araldite and stained with lead citrate and uranyl acetate. For ISH, sections were mounted on organosilane-coated slides. For polymerase chain reaction (PCR) on paraffin material, 5-μm-thick sections were placed in individual 1.5-ml microfuge tubes and processed as described elsewhere. 34
Results of in situ hybridization in tissues of five tortoises naturally infected with chelonid herpesvirus (ChHV).∗
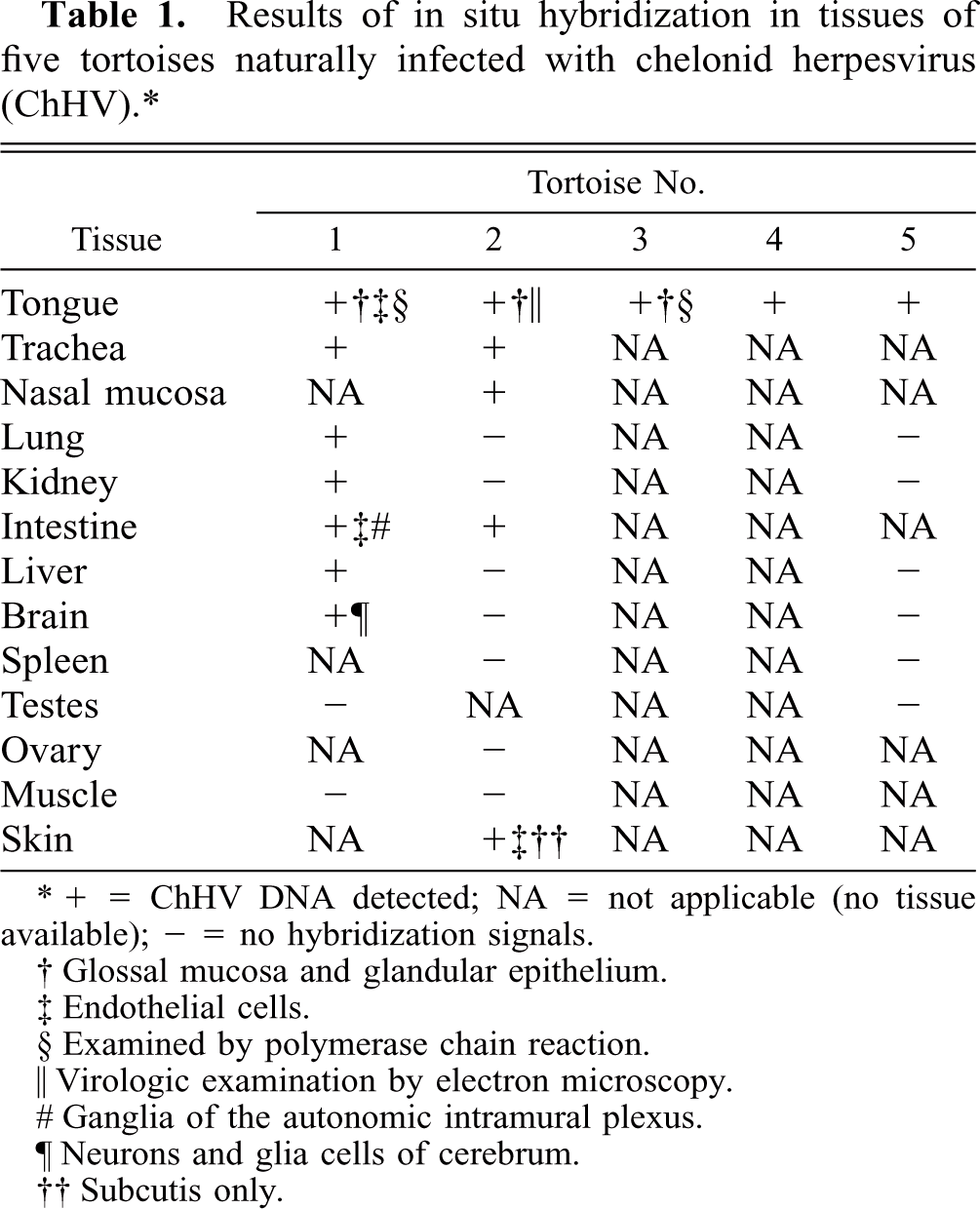
+ = ChHV DNA detected; NA = not applicable (no tissue available); 2 = no hybridization signals.
Glossal mucosa and glandular epithelium.
Endothelial cells.
Examined by polymerase chain reaction.
Virologic examination by electron microscopy.
Ganglia of the autonomic intramural plexus.
Neurons and glia cells of cerebrum.
Subcutis only.
Virus isolation
The virus used in this study was isolated from the pharyngeal swab of a T. hermanni that had been housed together with tortoise No. 5 of this study and has been analyzed and described in detail previously. 21 Briefly, the swab was placed in basal medium Eagle with Earle's salt solution (Seromed, Biochrom KG, Berlin, Germany) supplemented with 10% (v/v) tryptose phosphate broth, 0.1% (v/v) HEPES buffer (Seromed), 0.005% (v/v) enrofloxacine, and 0.05% (v/v) nystatin. After sonication and centrifugation for 5 minutes at 1,000 × g, the supernatants were inoculated onto Terrapene Heart Cells (TH-1, American Type Culture Collection [ATCC], Rockville, MD, USA), and incubated at 28 C. A cytopathic effect consisting of focal rounding of cells and cell lysis was observed after 4 days. The isolated virus was identified as a herpesvirus by sensitivity to chloroform and to 5-iodo-2′-desoxyuridine and by electron microscopy. 21 The isolate was end-point titrated on TH-1 twice and assigned the number 4295/7R/95.
Molecular cloning and sequencing of ChHV DNA
ChHV strain 4295/7R/95 was propagated on heart muscle cells of Pseudemys scripta elegans (SKH-R, CLCV 483, Cell Line Collection in Veterinary Medicine, Insel Riems, Germany). Cells were grown at 28 C in Dulbecco's modified essential medium supplemented with 10% fetal bovine serum. After complete cytopathic effect had developed, infected cells were freeze-thawed twice to release intracellular virus. After removal of cellular debris by low-speed centrifugation (8,000 × g, 4 C, 15 minutes), supernatants were layered on a 40% sucrose cushion. Virus was pelleted by ultracentrifugation (Beckman SW 28 rotor, 25,000 rpm, 1.5 hours, 4 C). Virus was suspended in phosphate-buffered saline and virus DNA was prepared after proteinase K–sodium dodecyl sulfate treatment. 27 Ten micrograms of ChHV DNA was cleaved with restriction endonuclease BamHI and cloned shotgun in vector pTZ18R (Amersham-Pharmacia Biotech, Inc., Piscataway, NJ, USA). Thirty-nine BamHI clones were isolated and subjected to automated sequencing using fluorescent-labeled M13 forward and reverse primers using the Licor system (MWG-Biotech, Ebersberg, Germany). 26 Subsequently, clone pChHV20 was chosen for designing a ChHV-specific probe for ISH because it contained sequences encoding the UL5 homolog, a highly conserved protein of the helicase/primase complex. 35
Probe synthesis for ISH by PCR
As template for probe synthesis by PCR, 50 ng of pChHV20, the sense primer F2 (5′-TGGTTGGACAACAACATGGC-3′), and the antisense primer R2 (5′-TCATTCCGTTATACATGAGCG-3′) were used, resulting in the expected 307-base-pair (bp)-long amplification product (Fig. 1). PCR was carried out for 35 cycles using 2.5 U/100 μl Taq DNA polymerase (Promega Corp., Madison, WI, USA) and the PTC-200 PCR thermocycler (MJ Research, Watertown, MA, USA). One hundred microliters of the amplification mixture contained 10 mM Tris-HCl (pH 9.0); 50 mM KCl; 0.1% Triton®X-100; 200 μM each of deoxyadenosine (dATP), deoxyguanosine triphosphate (dGTP), deoxycytodine triphosphate (dCTP), and deoxythymidine triphosphate (dTTP), 0.25 μM of both primers, 3.5 mM MgCl2, and autoclaved water. After initial heating (3 minutes at 95 C), each further cycle consisted of a denaturation step (30 seconds at 95 C), followed by primer annealing (30 seconds at 55 C) and primer extension (30 seconds at 72 C). The final extension step after the last cycle was set at 10 minutes at 72 C. Specificity of PCR was shown by restriction endonuclease digestion with SnaBI (New England Biolabs, Schwalbach, Germany) yielding 126-bp- and 181-bp-long cleavage products. For probe preparation, dTTP was substituted with a ratio of digoxigenin-11-deoxyuridine triphosphate (dUTP) to dTTP of 1:2 (DIG DNA Labeling Mix, Roche Diagnostics, Mannheim, Germany). 7
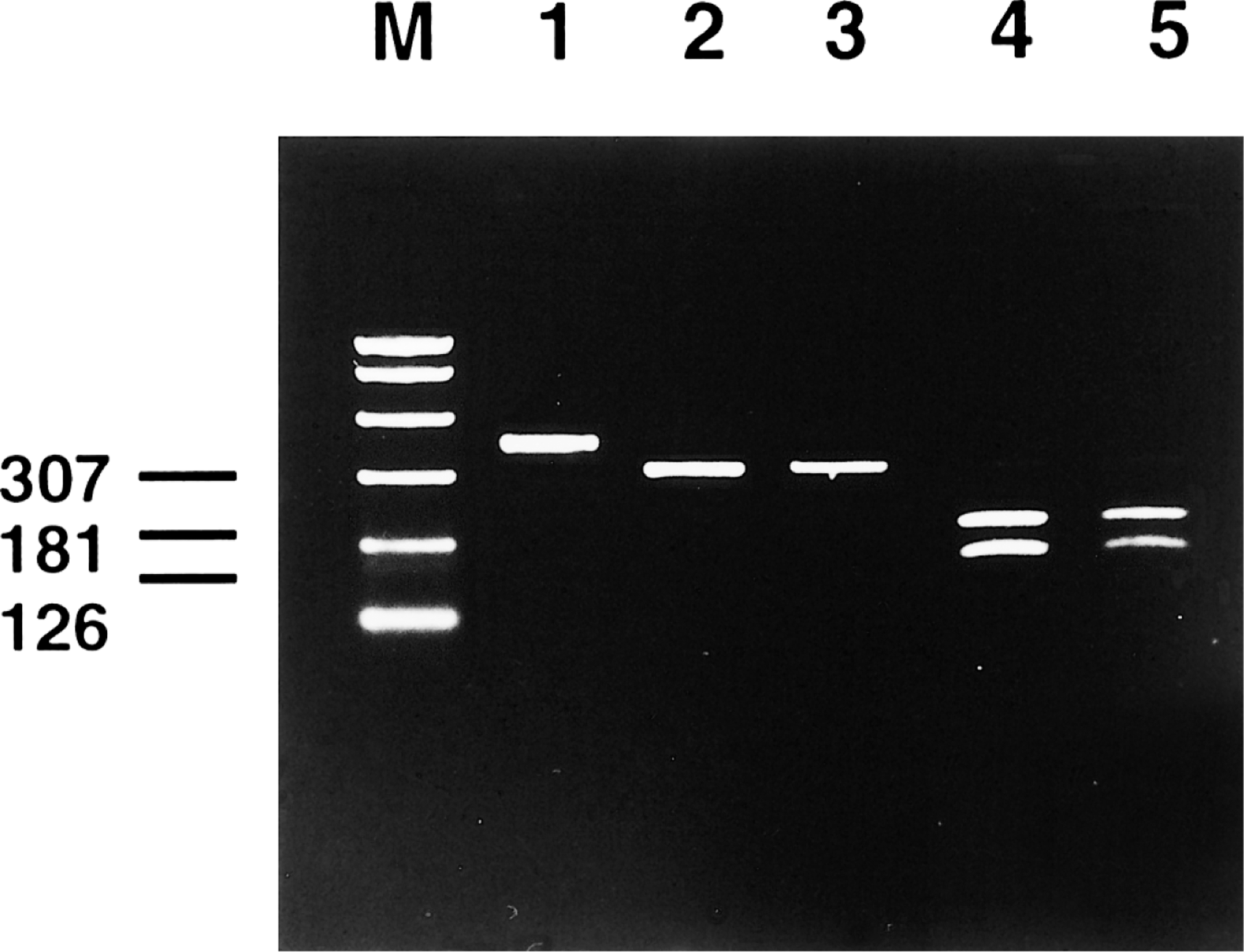
Agarose gel electrophoresis. Preparative PCR for 307-base-pair (bp) probe synthesis using cloned chelonid herpesvirus (ChHV) DNA as target. Slower migration in comparison to lanes 2 and 3 is due to digoxigenin labeling (lane 1); PCR of cloned ChHV DNA resulting in a 307-bp amplicon (lane 2); PCR of paraffin-embedded glossal tissue (lane 3); amplicon of lane 2 after restriction endonuclease cleavage with SnaBI, resulting in a 181-bp and a 126-bp fragment (lane 4); amplicon of lane 3 after restriction endonuclease cleavage with SnaBI, 181- and 126-bp fragment (lane 5).
To test specificity of the generated PCR probe, ChHV DNA was digested with BamHI or EcoRI (Roche), separated by 0.8% agarose gel electrophoresis, and transferred to positively charged Nylon membranes (Amersham-Pharmacia). The membrane was hybridized with the digoxigenin-labeled PCR probe overnight at 45 C. Chemoluminescent detection of hybridized fragments was done according to the supplier's instructions (Roche). In case of BamHI-digested DNA, a 1.1-kbp fragment hybridized with the probe. This BamHI fragment was of the same size and was consistent with the fragment previously cloned into pTZ18R. With EcoRI-digested DNA, two fragments of 6.5 and 7.5 kbp reacted positive with the PCR-generated digoxigenin-labeled probe (data not shown).
In situ hybridization
Nonradioactive ISH was performed according to a standardized protocol. 33 Proteolytic digestion of tissue sections was carried out with 10 μg/ml proteinase K (Roche) in 50 mM Tris-HCl (pH 7.4) for 15 minutes at 37 C. A 1:50 dilution of the PCR product was directly applied as a probe. Tissue sections were briefly counterstained with hematoxylin. Tissue areas without histopathologic lesions or tissues of uninfected tortoises and a double-stranded DNA probe specific for the VP-4-gene of infectious bursal disease virus (kindly provided by E. Mundt, Insel Riems, Germany) served as negative controls. 25
Results
Gross findings
At necropsy, all tortoises showed multiple, raised, rough, tan to red plaques on and underneath the tongue and on the sides and roof of the oral cavity. Similar red mucosal plaques extended into the esophagus in tortoises No. 1, No. 2, and No. 5. The oral cavity and, where present, the esophagus and proximal trachea had thick layers of mucoid to caseous pale yellow material. Tortoise No. 2 had severe subcutaneous edema reaching from the submandibular area to the thoracic inlet. Stomach and intestines were empty in all necropsied animals (Nos. 1, 2 and 5). Tortoise No. 5 had rhinitis with foamy nasal discharge.
Histopathologic findings
The oral surface epithelium of all tortoises was largely ulcerated or moderately hyperplastic. In hyperplastic areas, epithelial cells had large, intranuclear, amphophilic inclusions in all but tortoise No. 5. In all cases, the subadjacent propria mucosae was severely edematous and multifocally thickened by a fibrin meshwork and moderate infiltrates of heterophils, with fewer lymphocytes and macrophages. Many of the vessels within the oral propria mucosae were thrombosed with necrosis of the vessel walls and were surrounded by hemorrhage. Intranuclear inclusions were present in respiratory epithelial cells of the nasal cavity, the tracheal mucosa, the bronchiolar epithelium and a few pneumocytes. Most alveolar septa of the lungs were mildly widened with heterophils, which multifocally expanded into and obliterated alveolar lumina. Multifocally, intestinal submucosal lymphoid follicles were necrotic, often in association with thrombosis of small, adjacent blood vessels. The overlying lamina mucosae was mildly necrotic and enterocytes contained intranuclear inclusions. A few renal tubular epithelial cells adjacent to areas of focal interstitial lymphocytic and plasmocytic infiltrates were degenerate and contained intranuclear inclusions. Nuclei of a few swollen submucosal and myenteric ganglia were dilated with intranuclear inclusions and had pronounced margination of chromatin (Fig. 8). Scattered cerebral (Nos. 1 and 2) and cerebellar (Nos. 3 and 4) neurons and glia cells had intranuclear inclusions. In cutaneous sections taken from the ventral neck and head of tortoise No. 2, the dermis was diffusely and severely edematous. Many subcutaneous arteries were thrombosed and surrounded by moderate numbers of lymphocytes and a few heterophils.
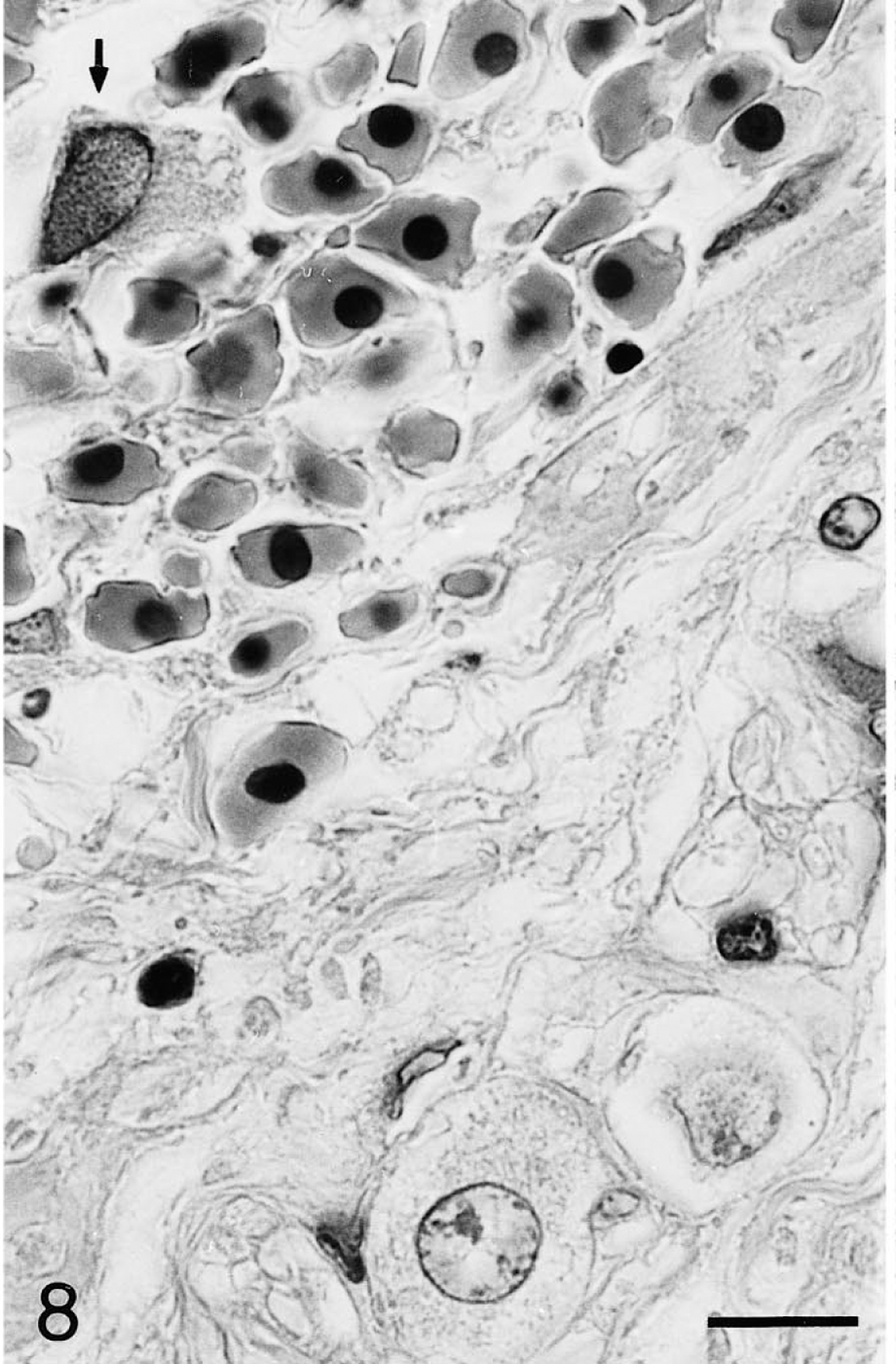
Intestine; Testudo horsfieldii (tortoise No. 1). A neuron in a myenteric autonomic ganglion is distorted by a large, amphophilic, intranuclear inclusion (arrow). Note unaffected neuron at the lower margin. Degenerate neurons are surrounded by hemorrhage from a larger intramural vessel. HE. Bar = 12 μm.
Molecular cloning of ChHV DNA
From a total of 39 BamHI clones obtained by shotgun cloning of ChHV DNA, clone pChHV 20, containing a 1.1-kb fragment of ChHV, was chosen. The sequenced region did not exhibit any detectable homology with herpesviral or other helicases at the nucleotide level. Translation of the nucleotide sequence into the corresponding amino acid sequence resulted in 42% identity and 59% similarity to equine herpesvirus 1 (amino acids 410–691) and 4 (amino acids 419–692) UL5 proteins, 35 36 and 41% identity and 57% similarity to Marek's disease virus (amino acids 382–667) UL5 proteins, respectively.
PCR and ISH
PCR on formalin-fixed, paraffin-embedded tissues of two glossal samples yielded the expected amplification product (tortoises No. 1 and No. 3; Fig. 1). Using the ChHV DNA–specific probe, ISH revealed an intense nuclear signal in tissues of all five examined tortoises. No hybridization signal was obtained with adjacent, histologically normal tissues, tissues from uninfected animals, and the nonspecific control probe. The results of the in situ hybridization are summarized in Table 1. Positive cells were detectable in glossal epithelia of all tortoises. The strong hybridization signal localized to the nuclei of keratinocytes in the glossal hyperplastic squamous mucosa (Fig. 2) corresponded with areas in which nuclear inclusions were present in HE-stained sections and was consistent with the presence of viral particles in electron microscopic specimens (Fig. 10). A strong signal was also seen in desquamated cells and cellular debris covering the tongue, and in the glandular epithelium of the glossal propria mucosae (Fig. 3). In tortoise No. 2, nuclei of endothelial cells of larger glossal arteries also showed strong hybridization signals. Corresponding to intranuclear inclusions observed in HE-stained sections, clusters of epithelial cells in the tracheal mucosa of tortoise No. 1 (Fig. 4) were ISH positive. In the lung, a similar hybridization signal was seen in a large number of pneumocytes lining alveolar septa (Fig. 5), exceeding by far the number of pneumocytes containing intranuclear inclusions in HE-stained sections. In intestinal sections, hybridization signals were detected either in epithelial cells of the lamina mucosae or in endothelial cells of thrombosed vessels (Fig. 6). Nuclear inclusions were found in submucosal intramural ganglia of the intestine (Fig. 8), in which strong hybridization signals were also obtained. Small numbers of hepatocytes and renal tubular epithelial cells showed positive hybridization signals (Fig. 7). Hybridization signals were also detected in the nuclei of cerebral neurons and glia cells (Fig. 9).
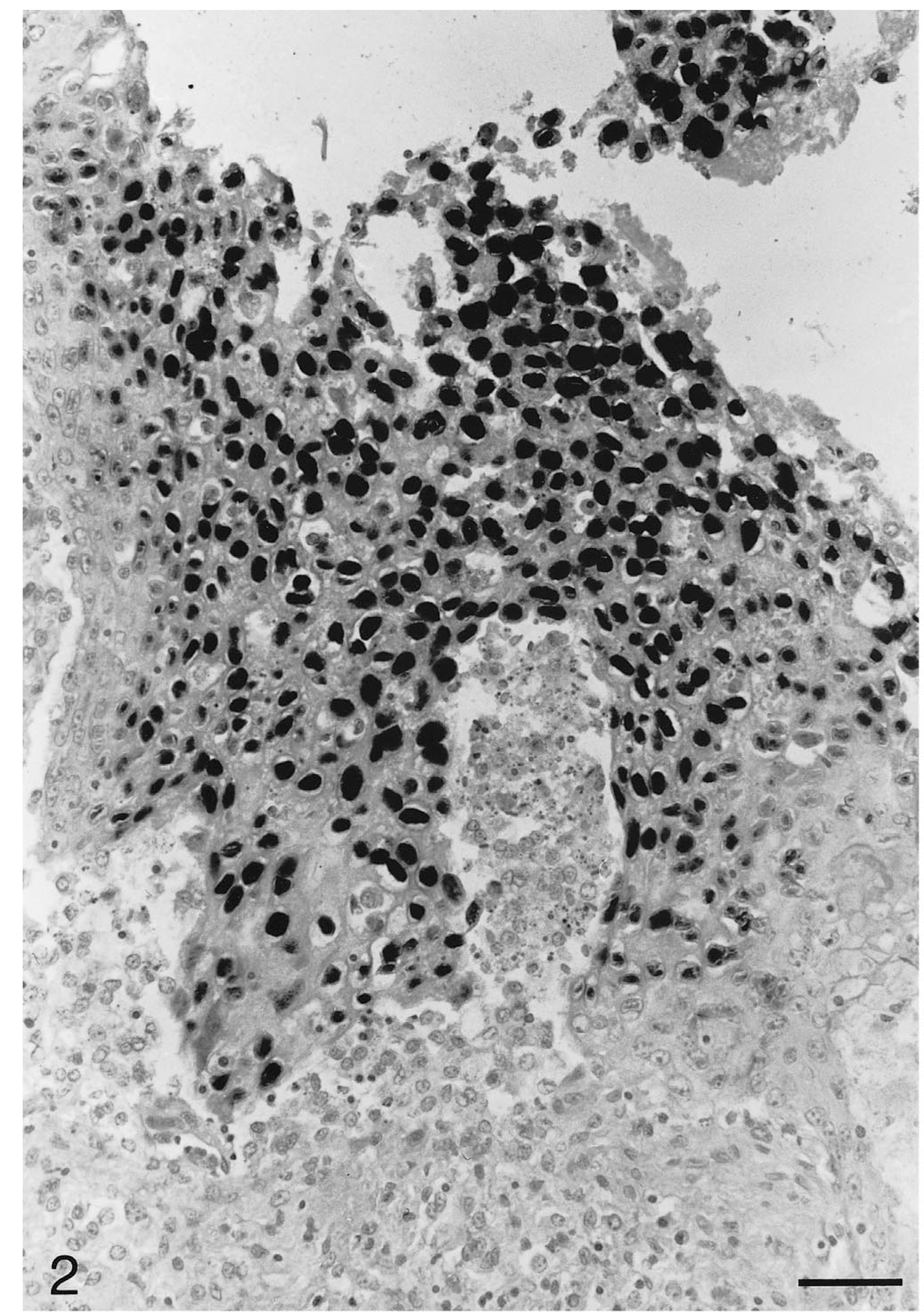
Tongue; Testudo horsfieldii (tortoise No. 1). Subacute diffuse severe proliferative and necrotizing glossitis. Focally, cells in the hyperplastic squamous mucosa and desquamated epithelial cells have strong black chelonid herpesvirus-DNA-specific intranuclear hybridization signals. In situ hybridization; hematoxylin counterstain. Bar = 50 μm.
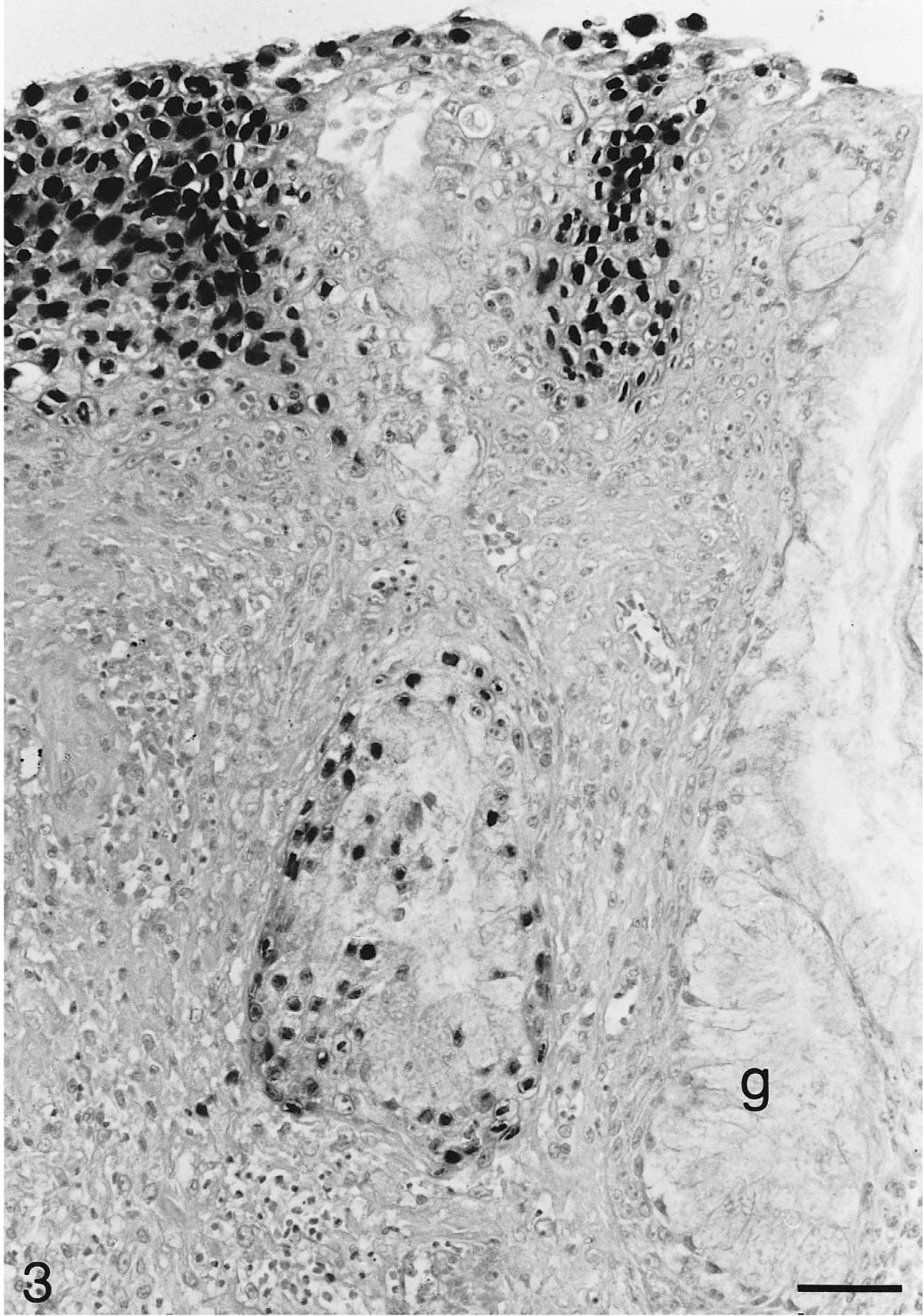
Tongue; Testudo hermanni (tortoise No. 3). Numerous nuclei of glossal epithelial cells and nuclei of epithelial cells in one glossal gland (g) show strong hybridization signals for chelonid herpesvirus DNA. In situ hybridization; hematoxylin counterstain. Bar = 50 μm.
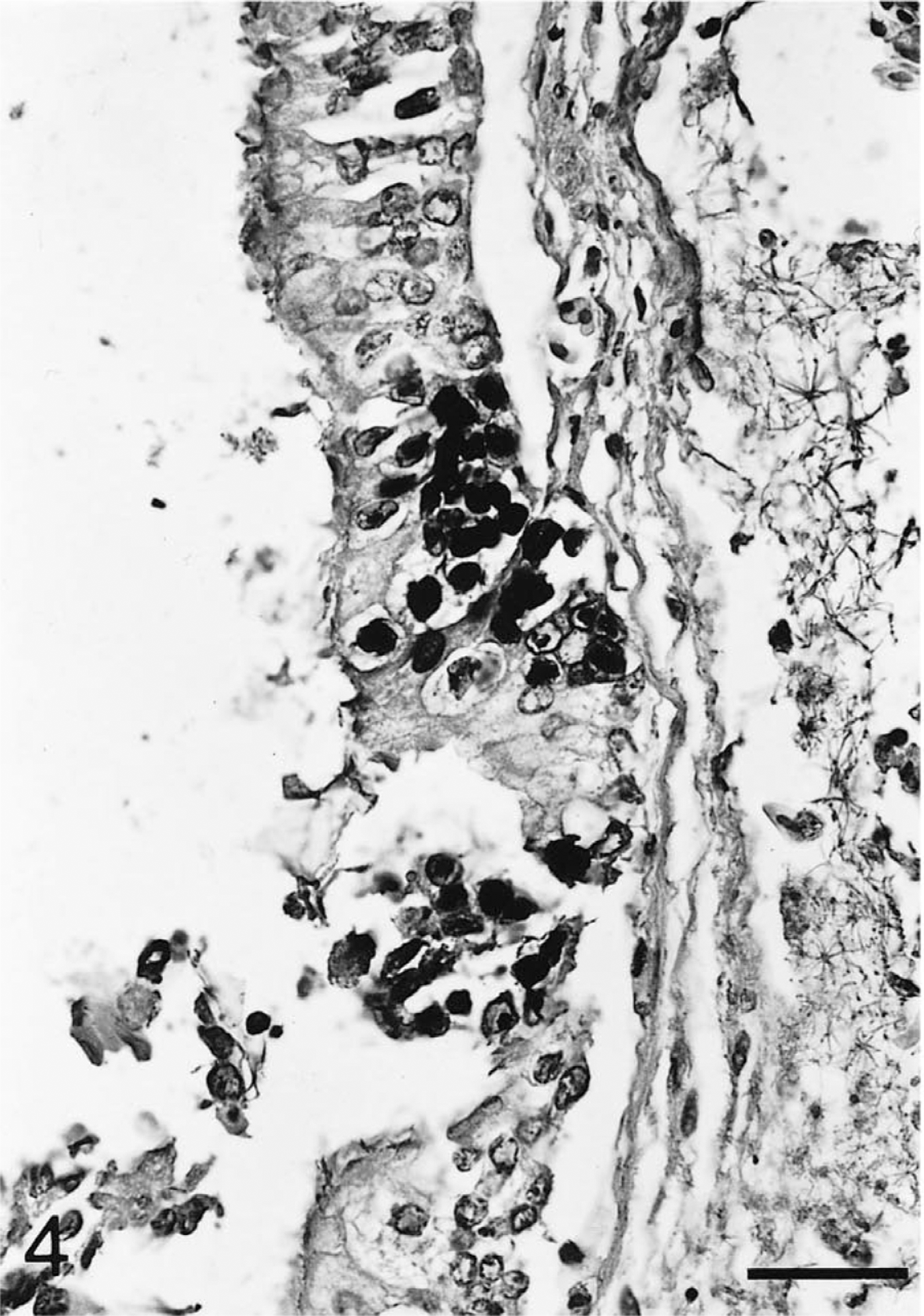
Trachea; Testudo horsfieldii (tortoise No. 2). Nuclei of focally detached tracheal mucosa cells are labeled with the hybridized chelonid herpesvirus DNA-specific probe. In the edematous submucosa, blood vessels are dilated with fibrin thrombi. In situ hybridization; hematoxylin counterstain. Bar = 40 μm.
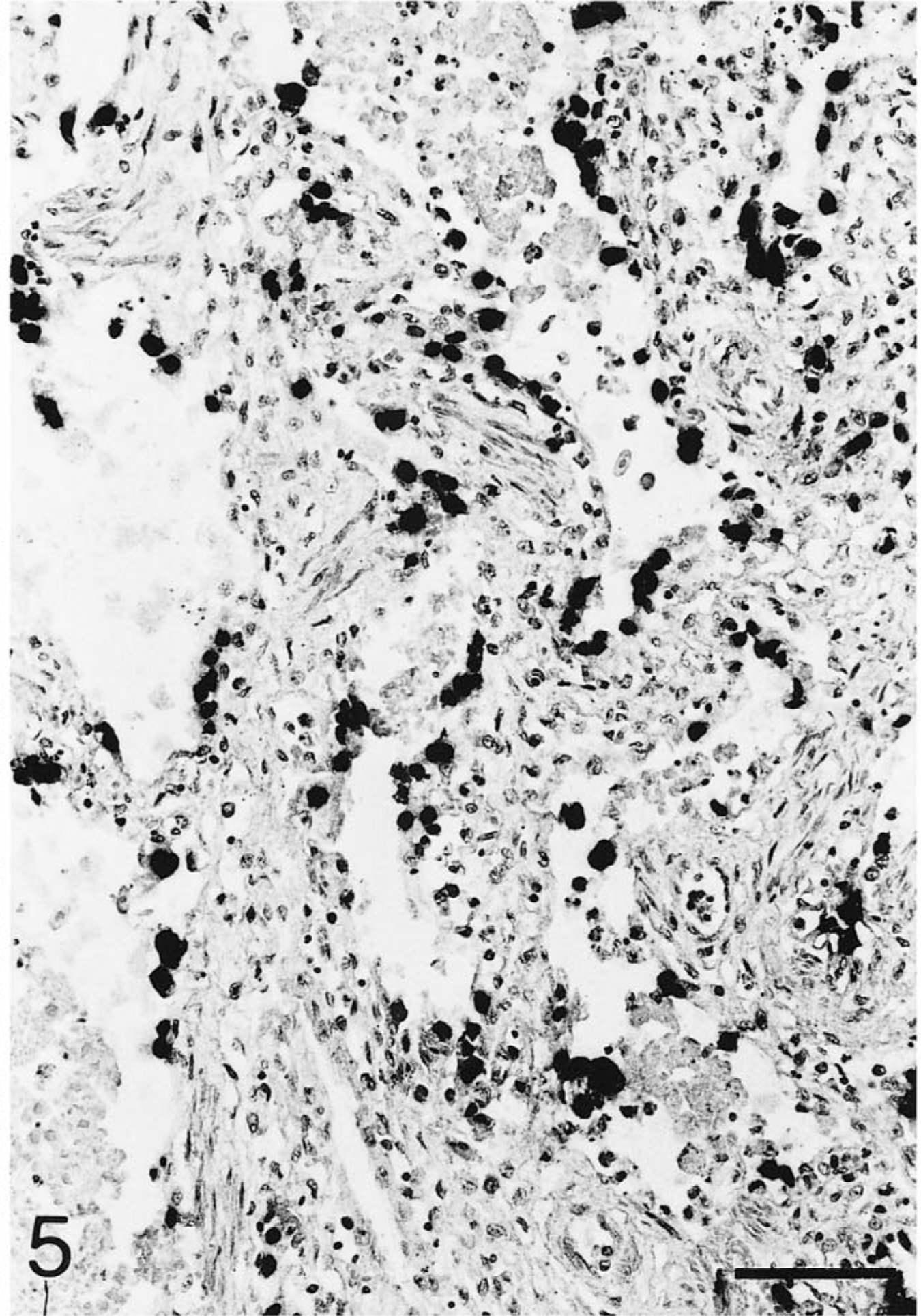
Lung; Testudo horsfieldii (tortoise No. 1). Numerous pneumocytes throughout the section display a strong hybridization signal. A few alveolar lumina are partially filled with exudate and cellular debris. In situ hybridization; hematoxylin counterstain. Bar = 80 μm.
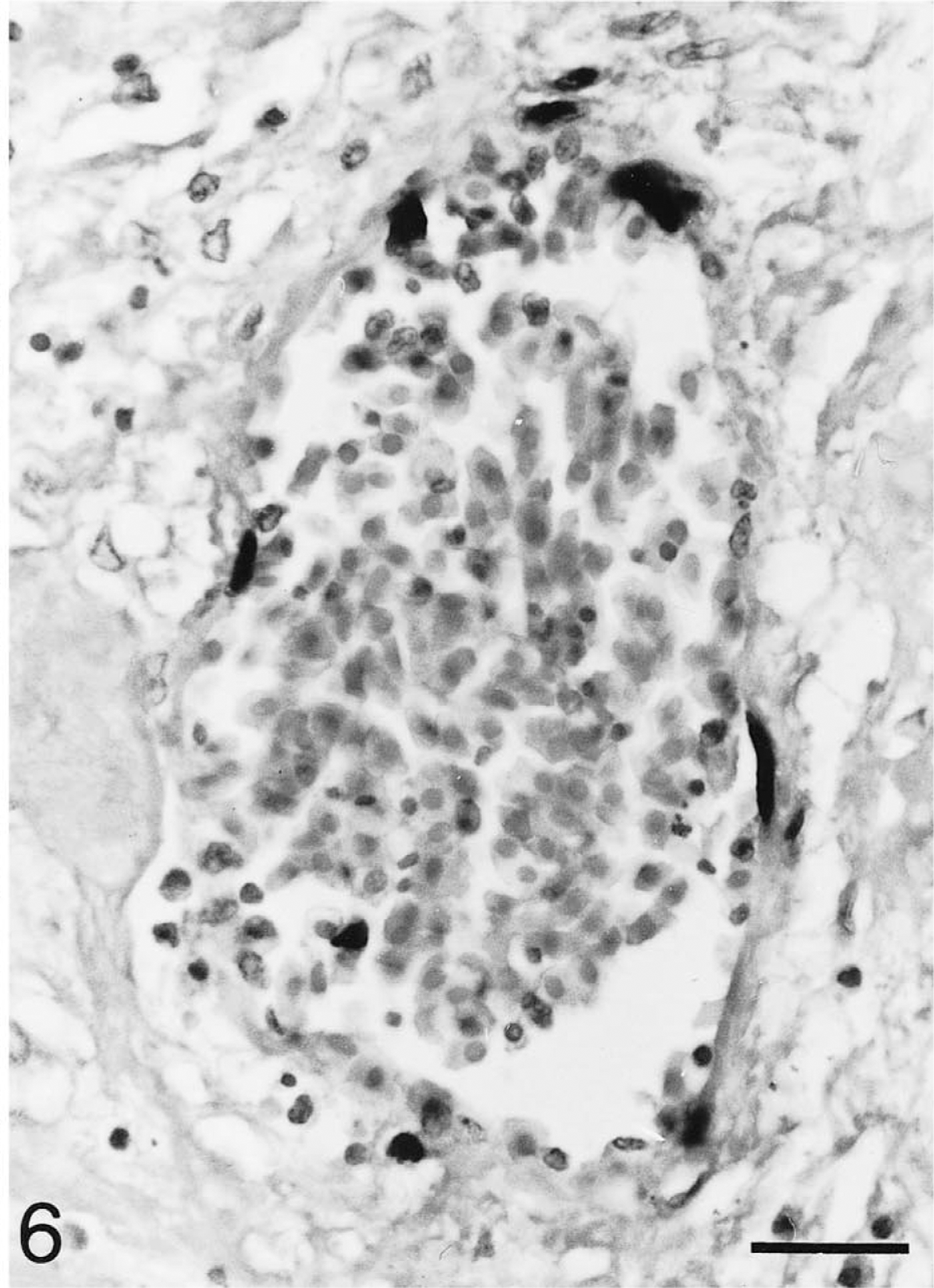
Intestine; Testudo horsfieldii (tortoise No. 1). Nuclei of nearly all endothelial cells of larger vessels have chelonid herpesvirus DNA-specific hybridization signals. In situ hybridization; hematoxylin counterstain. Bar = 25 μm.
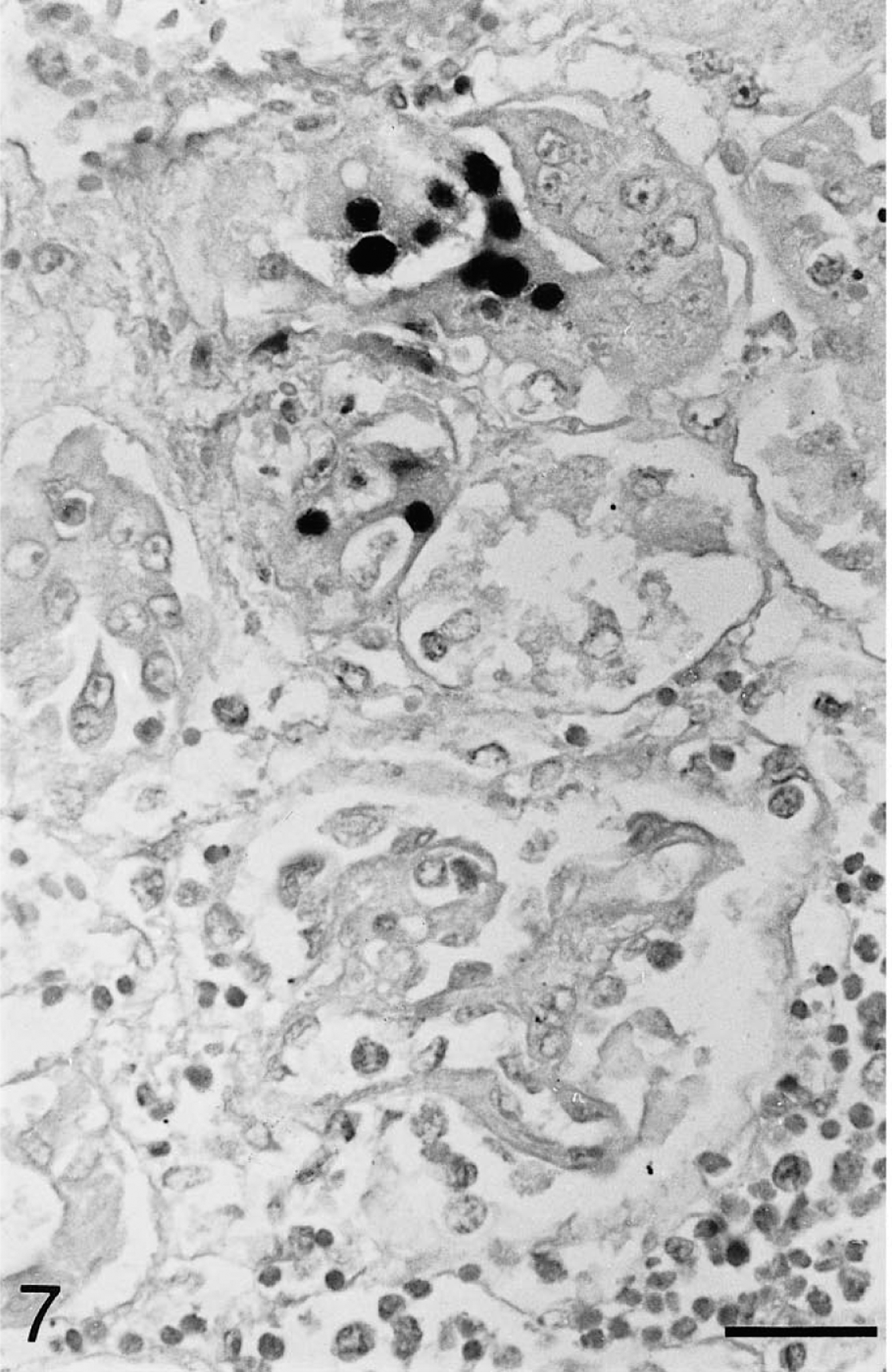
Kidney; Testudo horsfieldii (tortoise No. 1). Chelonid herpesvirus DNA is present in a small number of tubular epithelial cells in the renal cortex. Moderate periglomerular lymphocytic and histiocytic infiltration. Note lack of staining in the adjacent glomerulus. In situ hybridization; hematoxylin counterstain. Bar = 25 μm.
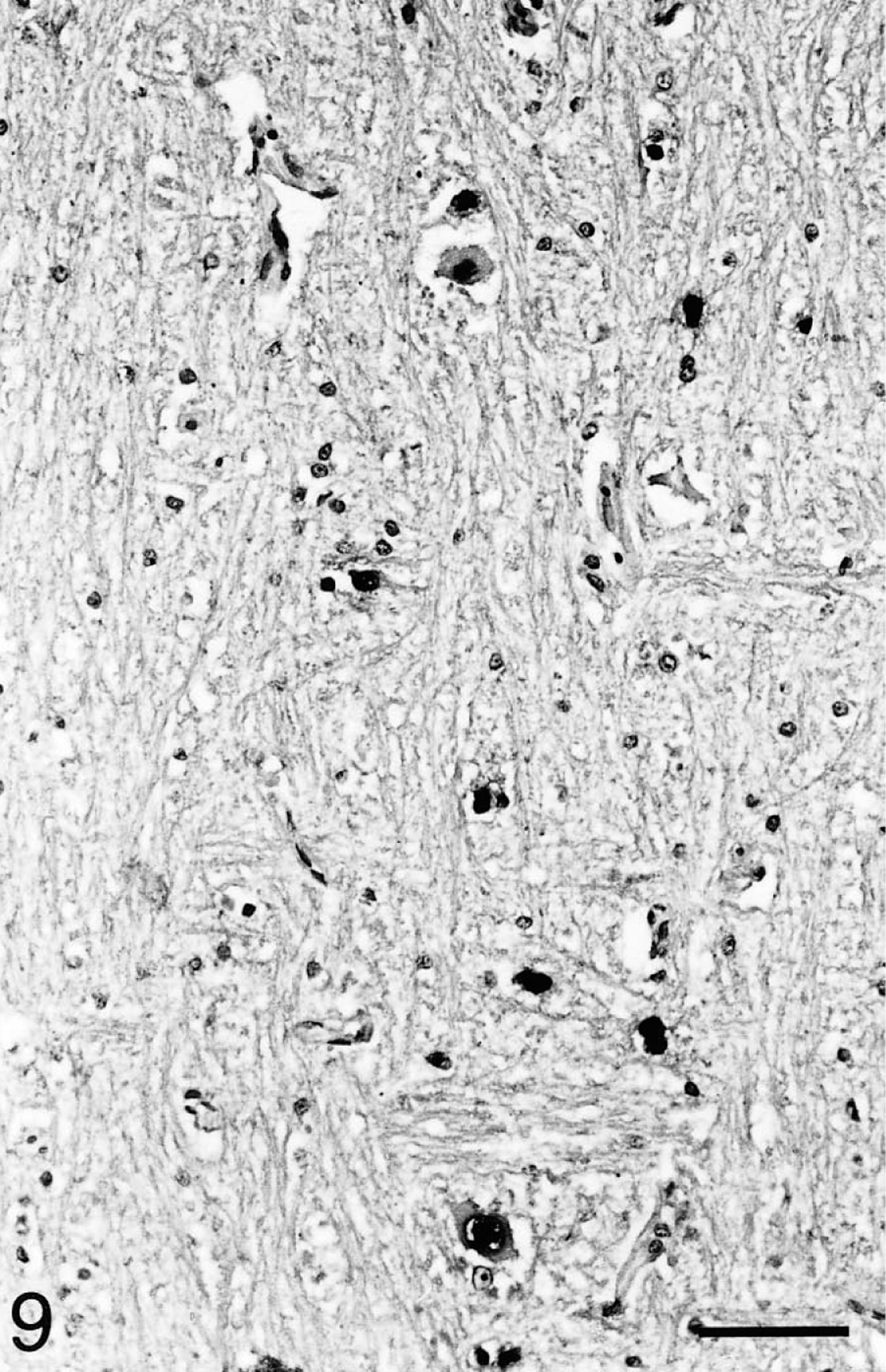
Brain; Testudo horsfieldii (tortoise No. 1). A few cerebral neurons and glia cells exhibit intranuclear hybridization signals without accompanying inflammatory reaction. In situ hybridization; hematoxylin counterstain. Bar = 80 μm.
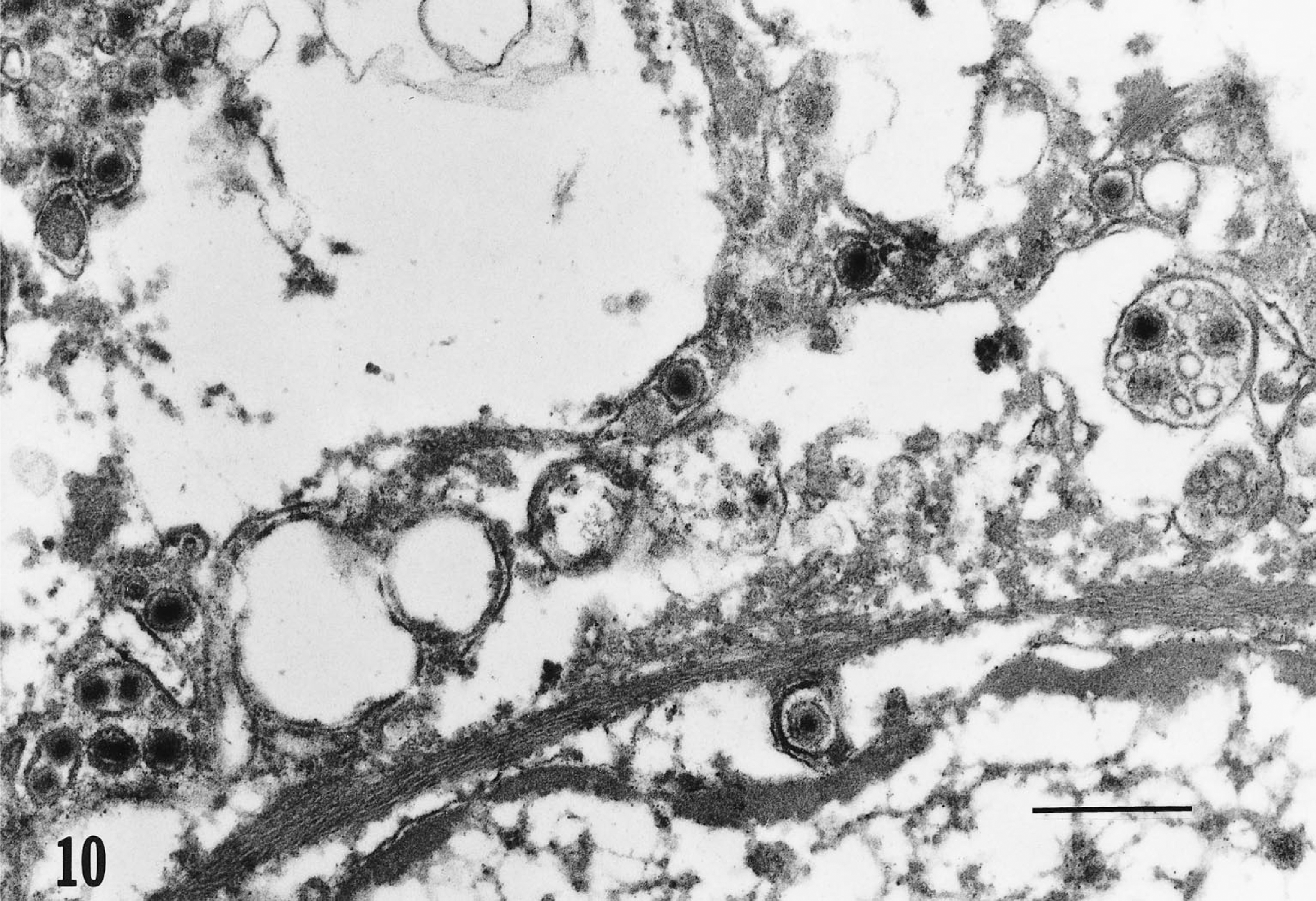
Transmission electron micrograph. Tongue; epithelial cell; Testudo horsfieldii (tortoise No. 1). The perinuclear cisterna and cytoplasm contain multiple enveloped viral particles consistent with herpesvirus morphology. Single virions bulge from the nuclear membrane. Lead citrate and uranyl acetate. Bar = 0.5 μm.
Discussion
The gross and histopathologic changes observed in this study corresponded largely with the literature and are thought to result from the viremic spread of a pantropic ChHV in the infected tortoises. 22 However, for the first time to our knowledge, ChHV DNA was demonstrated to be located in a broad range of tissues in tortoises suffering from stomatitis–rhinitis complex. The ISH signal colocalized to the same areas and cell types that contained intranuclear inclusions in HE-stained sections. In the lungs, the number of pneumocytes staining for ChHV DNA exceeded by far the number of pneumocytes containing inclusion bodies. In glossal sections of tortoise No. 5, ChHV DNA was present, even though inclusion bodies were not detectable in HE-stained slides. Autolytic changes may have obscured the presence of inclusion bodies in this animal. In areas with severe vascular necrosis, far more endothelial cells displayed hybridization signals than inclusion bodies. Pronounced degenerative changes can hamper detection of subcellular structures such as inclusion bodies. In all other tissues in which nuclear inclusions were not present in HE-stained slides, ISH also failed to detect herpesvirus DNA. The presented results show that infection with ChHV is not limited to the alimentary and respiratory tracts in tortoises suffering from stomatitis–rhinitis complex. In contrast to observations by others, infection of endothelial cells was generalized and was a main feature of the infection. 22 Therefore, ChHV described here appeared to exhibit endotheliotropism resulting in systemic necrotizing vasculitis with formation of thrombi and hemorrhages. Endotheliotropism has been reported with members of the alphaherpesvirus family including equine herpesvirus-1, bovine herpesvirus-1, and canine herpesvirus. 17 Interestingly, preferential infection of and replication in endothelial cells of ChHV also parallels the vascular tropism of the recently reported herpesvirus of Asian and African elephants. 31 Whereas infection with the elephant virus is fatal in Asian elephants, the same virus causes only mild infections with development of dermal fibropapillomas in African elephants. 31 Therefore, the African elephant may function as a virus carrier, as has been proposed for T. graeca and G. carbonaria. 13 16 21 This may explain, why, after mixing chelonids, fatal infections occur in T. hermanni and T. horsfieldii, whereas T. graeca remain clinically healthy. As reviewed in the introduction, ChHVs have been classified in the past based on isolation from different host species, which is in our opinion insufficient. Therefore, molecular biological data, as presented in this study, are urgently needed in order to establish a classification scheme for ChHVs.
Infection of respiratory epithelia, a finding in ChHV-infected tortoises, is a typical feature of alphaherpesviruses. The formation of multinucleated syncytial cells, as occasionally observed with the alphaherpesviruses bovine herpesvirus-1 and avian infectious laryngotracheitis virus, was not detected in this study. However, giant cells have been reported by others in herpesvirus infections in chelonids. 28 The occurrence of intranuclear inclusions in glandular epithelia of the tongue without inflammatory reaction or destruction of the lingual glands as well as the presence of inclusions in renal tubular epithelium resemble features of infections with betaherpesviruses such as porcine cytomegalovirus. Infection of lingual glands may explain virus excretion via saliva in acutely and latently infected animals resulting in the detection of herpesviruses using pharyngeal swabs even in the absence of stomatitis. 21 Localization of ChHV DNA to neurons and glia cells without concurrent encephalitis corresponds with observations by others and is also consistent with findings in alphaherpesvirus-infected animals. The lack of an inflammatory reaction could be associated with invasion of the central nervous system late in infection or may reflect persistent infection without clinical symptoms as in herpes simplex virus 1 infections. 22 Interestingly, inclusion bodies and herpesvirus DNA were also detected in ganglia of the autonomic myenteric plexus, which parallels the distribution pattern of pseudorabies virus in piglets. 24 Together with reduced food uptake due to stomatitis, malabsorption caused by intestinal dysfunction as consequence of neuronal damage may have led to weight loss and cachexia in these animals.
The pattern of histopathologic changes, especially number and distribution of inclusion bodies, and the ISH signals were slightly different among the examined animals. First, only limited samples were available from tortoises No. 3 and No. 4. Second, exact data on the course of disease were only available for two animals (Nos. 1 and 2). The more widespread infection in tortoise No. 1 could be attributable to lower innate resistance. Alternatively, tortoise No. 2 may have died at an earlier time point during infection, possibly due to secondary problems such as malnutrition.
Hybridization signals were obtained in tortoises from different geographic provenances. This indicates a high degree of conservation of the fragments of the helicase gene we used among the chelonid herpesviruses from Europe and North America. Further molecular biological studies are in progress to investigate the molecular evolution of this possibly archaic herpesvirus.
Footnotes
Acknowledgements
We would like to thank Dr. Th. C. Mettenleiter for helpful discussions and critical reading of the manuscript, Dr. J. R. Abbott at WADDL for the diagnostic work up of tortoise No. 1, Dr. R. Riebe for providing SKH-R cells, and Ruth Brown at WADDL-E.M. laboratory for electron microscopy. We gratefully acknowledge the skillful technical support of Gabriele Czerwinski, D. Schumacher, Kerstin Wink, H. Stephan, and Elke Zorn.
