Abstract
Falcon adenovirus is a newly recognized member of the family Aviadenoviridae and includes 2 closely related strains that are pathogenic to several species of falcons. Peregrine falcons appear to be one of the primary reservoirs, but recent outbreaks suggest that other carrier species probably exist. To allow screening of captive birds for virus shedding and investigations of disease outbreaks, conventional and realtime, quantitative polymerase chain reaction (PCR) assays and an in situ hybridization technique were developed. The diagnostic protocols were used on tissue and fecal samples from 7 species or subspecies of falcons infected with adenovirus as well as adenoviruses from other birds and mammals. The assays were specific for falcon adenovirus and detected both strains of virus in fecal samples from living animals or frozen and formalin-fixed, paraffinized tissues. Together with established serologic tests for falcon adenovirus, these molecular assays are valuable tools for management and conservation of falcons in captivity and the wild.
Introduction
A new species of adenovirus that causes inclusion body hepatitis, enteritis, and death was recently described in falcons. The adenovirus was first discovered from an outbreak of disease in 1996 that killed Northern aplomado falcons (Falco femoralis septentrionalis) and Peregrine falcons (Falco peregrinus). Subsequently, disease and death were documented in Orange-breasted falcons (Falco deiroleucus), Teita falcons (Falco fasciinucha), a Merlin (Falco columbarius), a Vanuatu peregrine falcon (Falco peregrinus nesiotes), and Gyrfalcon − Peregrine falcons (Falco rusticolus/peregrinus). The most recent outbreak occurred in March 2005, and the virus has been detected in birds in 4 states (Wyoming, California, Idaho, and Minnesota). 3,10
Most data indicate that Peregrine falcons are the natural host and primary reservoir of falcon adenovirus. 7,10 However, it is possible that other species of falcons may also be carriers, and cohousing of any mixed falcon species should be avoided unless the status of adenovirus exposure and infection in all birds is known. 7,10 Adenovirus infection has been recognized in various falcon species for years. 9,10 Because of the ongoing disease threat to falcons, specific and sensitive molecular assays for detecting falcon adenovirus have been developed. These assays include conventional polymerase chain reaction (PCR), real-time PCR, and in situ hybridization. Together, these methods can be used on samples from live or deceased animals to help obviate future outbreaks and to allow accurate investigation of disease when deaths do occur.
Materials and methods
Animals and samples. Forty animals had tissues analyzed by PCR. Thirty-two animals were from the original study that described the novel falcon adenovirus. 10 These included 16 Northern aplomado falcons and 6 Peregrine falcons that died in the June 1996 outbreak at a raptor center in Boise, Idaho; 2 Orange-breasted falcons (Falco deiroleucus) from a private facility in Wyoming; a Vanuatu peregrine falcon (Falco peregrinus nesiotes) from a private home in San Diego, California; 2 Teita falcons (Falco fasciinucha) and 2 Gyrfalcon − Peregrine falcons (Falco rusticolus/peregrinus) that were cohoused in a facility in Minnesota; a Merlin (Falco columbarius) from Minnesota; and 2 adult Northern aplomado falcons from the San Diego Zoo's Wild Animal Park in Escondido, California. Additional animals tested were 5 Teita falcons that died with lesions consistent with adenovirus infection, 3 a red-tailed hawk (Buteo jamaicensis), a red panda (Ailurus fulgens), and a racoon (Procyon lotor), all of which had adenovirus infections as determined by PCR and DNA sequencing. Samples of frozen tissue, formalin-fixed paraffinized tissue, or feces were analyzed from 7 species or subspecies of falcons: Northern aplomado falcons, Peregrine falcons, Orange-breasted falcons (Falco deiroleucus), a Vanuatu peregrine falcon (Falco peregrinus nesiotes), Teita falcons (Falco fasciinucha), Gyrfalcon − Peregrine falcons (Falco rusticolus/peregrinus), and a Merlin (Falco columbarius). Tissues included portions of all major organs as previously described. 10 Frozen tissues and feces were stored at −80°C after collection. Other tissues collected at postmortem examination were immersion fixed in 10% neutral buffered formalin for 2 to 7 days, dehydrated, cleared, infiltrated with paraffin, and embedded.
Virus stocks. Poultry adenovirus serotypes: frozen or lyophilized tissue culture lysate for each of the 12 chicken serotypes (1−12) of group I adenovirus were obtained from Dr. S. Dhillon a and the SPAFAS Charles River Diagnostics Laboratories b for comparative molecular analyses. A vector-cloned portion of bovine adenovirus type 1 hexon was obtained from Dr. L. A. Hobbs. c Falcon adenovirus cell culture supernatant was obtained from Dr. L. Oaks. d
DNA isolation. DNA was extracted from 20 mg of frozen tissues and poultry adenovirus lysates using the Qiagen Tissue Kit, e according to the manufacturer's tissue-sample protocol, except that the recommended amounts of sample were first placed with the lysis buffer in 1.5-ml screw-cap FastPrep vials containing ceramic beads and lysed by agitation in a FastPrep shaker f at a setting of 4−5.5 m/sec for 40−60 sec, after which the lysate was transferred to a clean Eppendorf tube for continuation of the Qiagen protocol. DNA was extracted from formalin-fixed, paraffin-embedded samples of liver, lung, spleen, and intestine from those animals for which frozen tissues were not available by the same protocol as above, except that samples were first deparaffinized with successive washes in octane, xylene, and 100% ethyl alcohol and then air dried. DNA was extracted from feces using the Qiagen Stool Kit e according to the manufacturer's protocol.
PCR. Five PCR protocols were performed on DNA from frozen or paraffin-embedded tissues or feces. The PCR protocols and expected sizes of the amplicons are listed below. For all reactions, 50–500 ng of DNA were used. For conventional PCR; DNA was added to a 25-μl reaction mixture containing 10 mM Tris (pH 8.0), 50 mM KCl, 5 mM MgCl2, 200 μM each of dATP, dCTP, dGTP, and dTTP, 50 pmol/L of each primer, and AmpliTaq Gold DNA polymerase g at a final concentration of 0.05 U/μl. For real-time TaqMan PCR, the TaqMan Universal PCR Master Mix g was used at 1− with 0.9 uM each of unlabeled falcon-specific primer, 0.25 uM falcon-specific FAM-labeled probe, g and 1− TaqMan exogenous internal positive control (EIC) reagents with VIC-labeled probe, g including blocking reagent for negative control wells. Conventional PCRs were as follows: 1) Escherichia coli: ECA forward primer GGAAGAAGCTTGCTTCTTTGC TGAC and ECA reverse primer AGCCCGGGGATTTCA CATCTGACTTA (GenBank accession target U00096) were used at 95°C for 5 min then 95°C for 45 sec, 58°C for 45 sec, 72°C for 30 sec for 35 cycles, then 72°C for 8 min to generate a 544-base pair (bp) product. This PCR was used on fecal DNA to assess quality of DNA and presence of inhibitors; 2) beta-actin: beta-actin antisense GGGTGCTCCTCAGGGGCTACTCTCAG and beta actin sense CCCCCGTGCTGTGTTCCCATCTATCG primers (GenBank accession target X00182) were used at 95°C 5 min, 95°C for 1 min, 62°C for 45 sec, 72°C for 30 sec for 30 cycles, then 72°C for 8 min, to generate a 450-bp product. This PCR was used on DNA from frozen and paraffin-embedded tissues to assess quality of DNA and presence of inhibitors; 3) falcon specific adenovirus: the sense primer (AAGAAACACCACAACAGGG) and the antisense primer (GTAAGTAACCAGATCGAAGGTG) were used at 95°C for 6 min followed by 35 cycles of 95°C for 45 sec, 54°C for 1 min, 72°C for 1 min, and then 72°C for 8 min to generate a 385-bp product. This PCR was used to test for the presence of falcon adenovirus; 4) falcon-specific pVI/hexon region: forward primer CGCAGCC GACTACTATGGACTTAC and pVI/hexon region reverse primer CGTGGGTTGTGTCATCTGTTTG at 95°C for 6 min then 95°C for 45 sec, 57°C for 45 sec, 72°C for 30 sec for 35 cycles, and then 72°C for 8 min to generate a 595-bp product. This PCR was used to characterize the target area of the real-time assay in adenoviruses from different species of falcons.
The TaqMan assay was real-time falcon-specific pVI region with EIC: falcon adenovirus forward TaqMan primer CGCTACCCGTCATGCTACTTAA; falcon adenovirus reverse TaqMan primer CCCAAGCGCCG ACGTA; falcon adenovirus TaqMan probe FAMAAGGCCGGGTAACTGGAGAGGTCG, and TaqMan EIC reagents g were used in a multiplex reaction at 50°C for 2 min then 95°C for 10 min followed by 95°C for 15 sec and 60°C for 1 min for 40 cycles to generate a 64-bp product. This PCR was used to detect and quantitate falcon adenovirus (falcon-specific pVI region) and assess the presence of inhibitors (EIC).
DNA manipulation and sequencing. PCR products of the expected size from representative tissues and cases were purified and either direct-sequenced or cloned using the TOPO TA Cloning Kit. h Sequencing reactions were performed using the CEQ DTCS (dye terminator cycle sequencing) Quick-Start kit. i Sequences were acquired using a CEQ 2000XL capillary sequencer. i Novel DNA sequences were depositied into GenBank.
Southern hybridization and detection. Southern blotting of all PCR products was performed with a PosiBlot pressure apparatus j to positively charged nylon membranes. k , l DNA was fixed to the membrane using a Stratalinker. j DNA probes were generated by labeling with digoxigenin using the PCR DIG Probe Synthesis Kit, m and hybridized products were exposed to Omat LS X-ray film. n
In situ hybridization. Five micron sections of selected formalin-fixed, paraffin-embedded tissues on Platinum slides o were deparaffmized, hydrated, treated with trypsin digest-all p at 37°C for 10 min, washed in Tris-buffered saline (TBS), heated at 98°C for 12 min in TBS, placed immediately in 4°C TBS, and prehybridized with Dig Easy Hyb Granule solution m at 42°C for 1 hr. A 385-bp cloned and sequenced segment of the falcon adenovirus hexon gene obtained from the falcon adenovirus-specific hexon region PCR was labeled with digoxigenin using an incorporation method m and used in Dig Easy Hyb Granule solution m for hybridization at 42°C for 16 hr at 50 pmol/L of labeled probe per milliliter of hybridization solution. Slides were washed in 2− SSC with 0.1% sodium dodecyl sulfate (SDS) 2 times for 5 min at room temperature (RT) followed by 2 washes in 0.5− SSC with 0.1% SDS at 68°C for 15 min. Slides were blocked in blocking solution m for 30 min, washed in TBS, treated with antidigoxigenin antibody p diluted 1:5000 (150 mU/ml) in TBS for 1 hr at RT, washed in TBS, treated with nitro blue tretrazolium chloride/5-bromo-4-chloro-3-indolyl phophate, toluidine salt (NBT/BCIP) j for 10 min, washed in TBS and distilled water, counter-stained with Gills hematoxylin q and mounted with crystal mount. r Duplicate control slides received identical treatment except no labeled probe was added to the hybridization solution. Additional negative control slides consisted of tissues infected with herpes simplex virus 1.
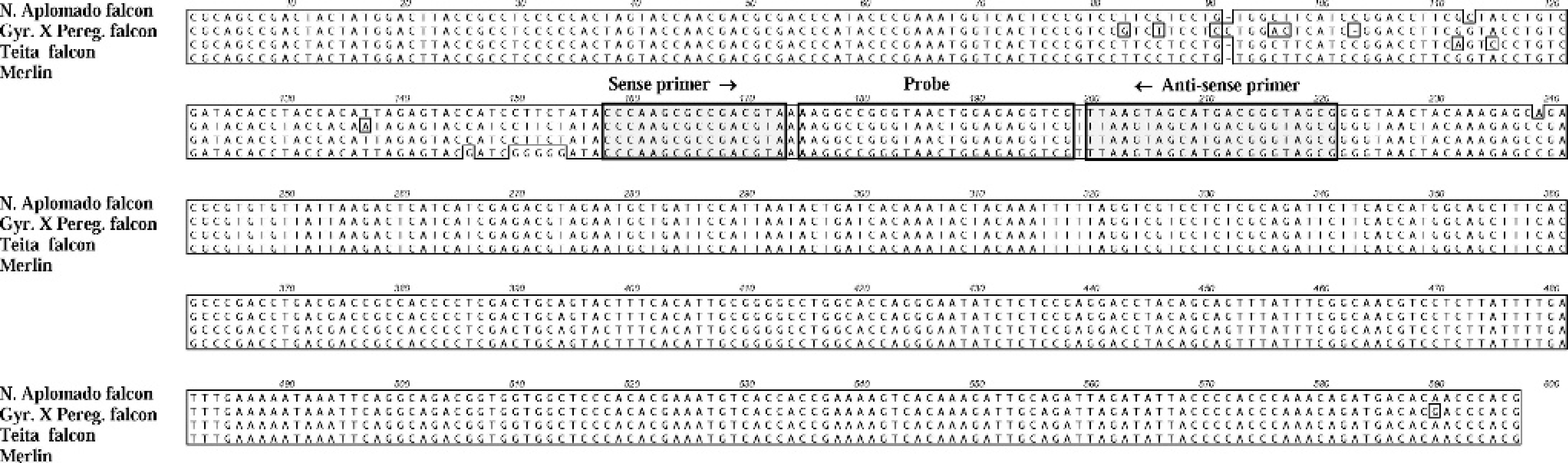
Alignment of 4 falcon adenoviral genotypes at the pVI/hexon region for development of the real-time falcon-specific pVI region with EIC. Shown are the primer and probe binding sites for the real-time PCR. The nucleotide sequences in this region for adenovirus from Northern aplomado falcon, Orange-breasted falcon, and Peregrine falcon were identical.
Calculations. Virus quantities were determined by generating standard curves from known masses of vector-cloned target gene; absolute quantities of DNA in nanograms were converted to virus genome equivalents using the standard formula for double-stranded DNA equating 660 grams per mole per base pair of DNA. The TOPO-TA vector h with the target gene was 4020 bp.
DNA sequence analysis. Analyses for primer design for falcon-specific adenovirus hexon region was done with the software program MacVector. s Analysis for the falcon-specific TaqMan assay was done with the software program Primer Express. g GenBank searches and comparisons were done with the basic local alignment search tool (BLAST) at the National Center for Biotechnology Information Web site (www.ncbi.nlm.gov).
Results
The falcon-specific hexon region assay was designed using 2806 bp of DNA sequence of the adenovirus hexon gene (GenBank accession AY683541) obtained from tissues of infected Northern aplomado falcons from the original 1996 outbreak. 10 The primers targeted the L3 region of the hexon. BLAST analyses indicated that the primers were specific for falcon adenovirus. The real-time falcon-specific pVI region with EIC assay was designed using 6257 bp of the adenovirus sequence (GenBank accession AY683541). After identifying a suitable region for real-time PCR, additional DNA sequences were generated by the falcon-specific pVI/hexon region PCR for this region from different animals and demonstrated 4 unique viral genotypes in 7 lineages of falcons (Northern aplomado, Peregrine, Orange-breasted, Vanuatu, Gyrfalcon − Peregrine, and Teita falcons, and a Merlin) (GenBank accessions EF675478–EF675482). Adenovirus from Northern aplomado falcon, Orange-breasted falcon, and Peregrine falcon had identical sequence in the pVI/hexon region. Nucleotide variations were present in samples from Teita falcons, the Merlin, Gyrfalcon − Peregrine falcons, and the Vanuatu falcon. However, no nucleotide differences were present among any samples from the 7 falcon lineages within the binding sites for the real-time falcon-specific pVI region with EIC assay (Fig. 1).
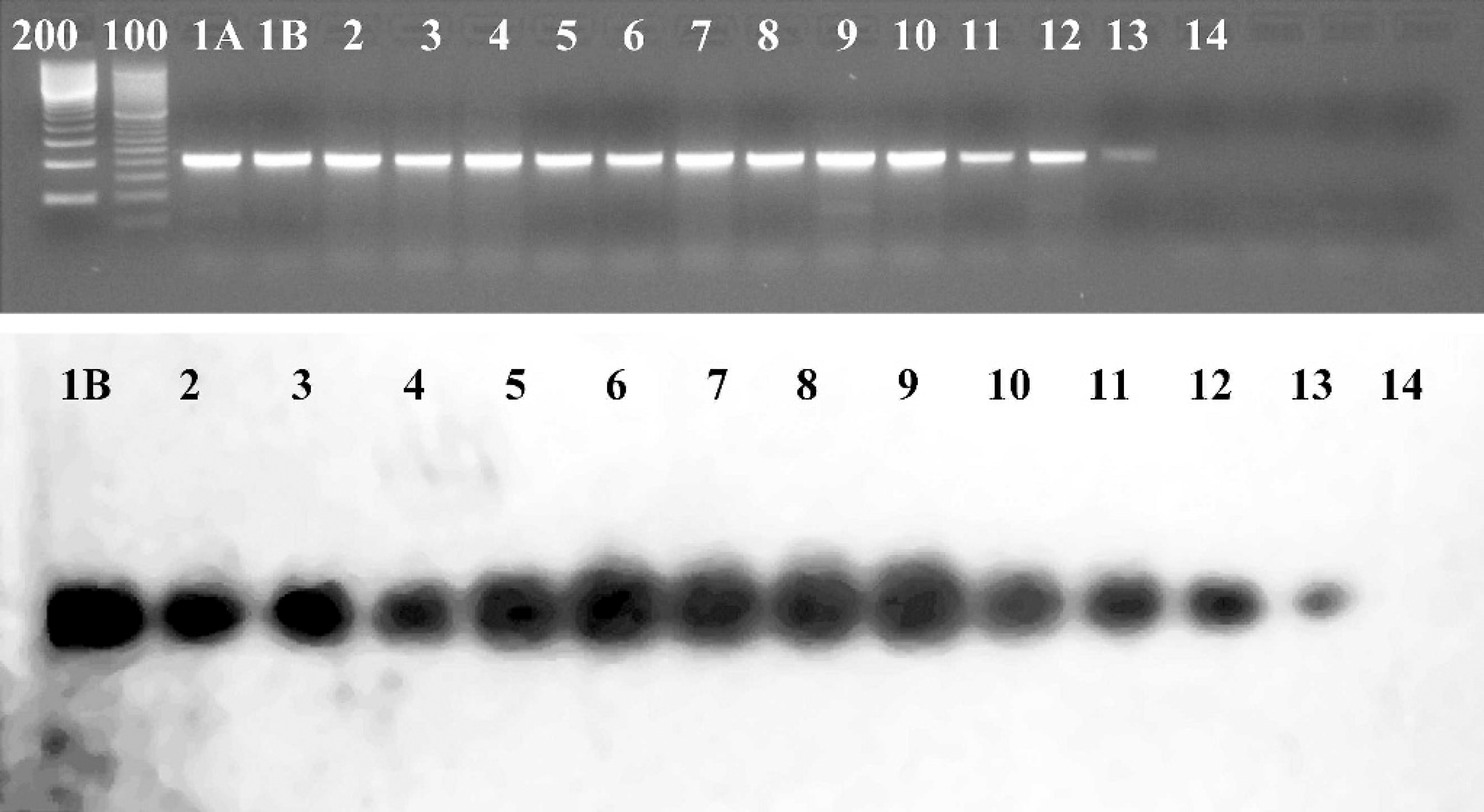
The falcon-specific hexon region PCR was performed on DNA extracted from 137 frozen tissues from 40 animals. One hundred and four samples representing 35 animals, including all 7 lineages of falcons, were positive by PCR. All 104 were confirmed with Southern blot hybridization or cloning and sequencing of amplicons (Fig. 2). Positive results were obtained from DNA samples from many organs (brain, kidney, spleen, liver, intestine, lung, trachea, bursa of Fabricius, pancreas, and ventriculus). Bone marrow, skin, eye, and adrenal gland were negative. Both hexon genotypes of falcon adenovirus (GenBank accessions AY683554 and AY683555) were detected. The 2 falcon adenovirus genotypyes were confirmed by cloning and sequencing. DNA from paraffin-embedded tissues from 14 animals that had positive PCR on frozen tissues were also tested, and samples from 11 of these animals were positive. PCR on DNA from frozen fecal samples from 11 healthy Northern aplomado falcons were negative. Six frozen fecal samples were spiked with 1 μL of cell culture supernatant containing whole virus, and 5 were positive by PCR. PCR on DNA from the 12 poultry adenovirus serotypes, hemorrhagic enteritis virus (HEV) of turkeys, egg drop syndrome (EDS) virus, the bovine adenovirus 1 hexon gene, and tissues from the red-tailed hawk, red panda, and raccoon were negative. PCR for actin was positive on all frozen and paraffin-embedded tissue samples. PCR for E. coli was positive on all fecal samples.
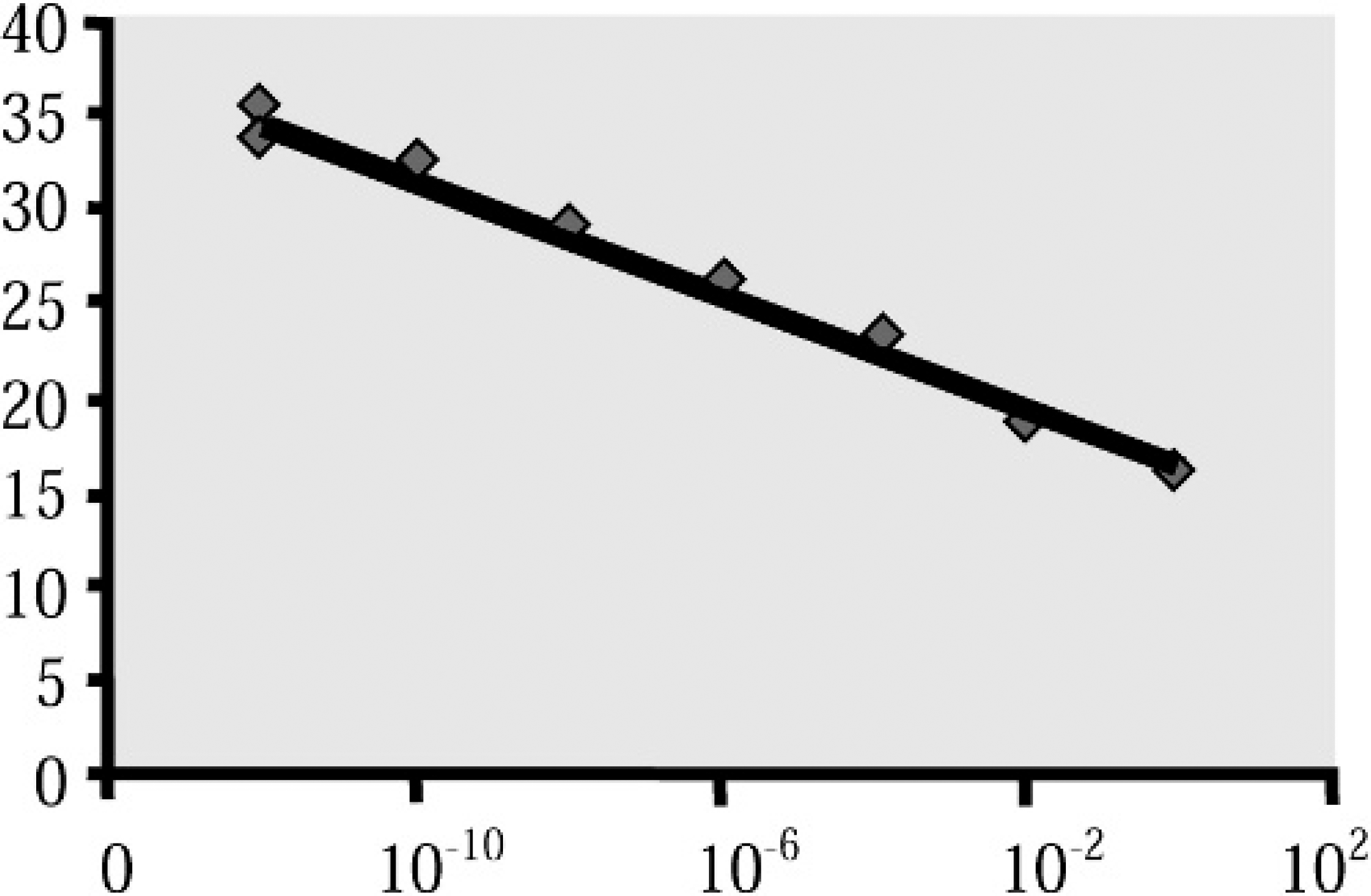
Real-time falcon-specific pVI region with EIC assay standard curve plot of 10-fold serial dilutions of plasmid-cloned adenovirus DNA concentrations in ng/uL (x-axis) and threshold cycle value for PCR (y-axis).
The real-time falcon-specific pVI region with EIC assay was performed on plasmid DNA containing the pVI gene and sample genomic DNA. Sixteen serial dilutions of plasmid DNA ranging from 102 ng to 10−14 ng per reaction assayed in triplicate allowed construction of a standard curve with an R 2 value of .9953 and a slope of −3.14 (Fig. 3). The lower range of detection was 100 copies of target based on plasmid copy number. From this value, the lower range of virions detectable was determined to be 100, as there is 1 target gene present in each virion. The upper limit of detection was not determined. The realtime assay was performed on the same samples analyzed by the falcon-specific hexon assay. Among the 137 frozen tissue samples, real-time PCR detected virus in the same 104 samples that were positive by conventional PCR and an additional 4 samples that appeared negative (Table 1). The 4 samples positive only by real-time PCR had relatively low values for quantity of virus DNA (1.42 − 104, 1.25 − 105, 8.89 −103, and 8.54 − 104 virus genome equivalents). Real-time PCR results for DNA from paraffin-embedded tissue and fecal samples spiked with whole virus were the same as those for conventional PCR. DNA from the 12 poultry adenovirus serotypes, HEV of turkeys, EDS virus, the bovine adenovirus 1 hexon gene, and from tissues of the red-tailed hawk, red panda, and raccoon were negative. The EIC PCR was positive on all samples, except those to which blocker was added.
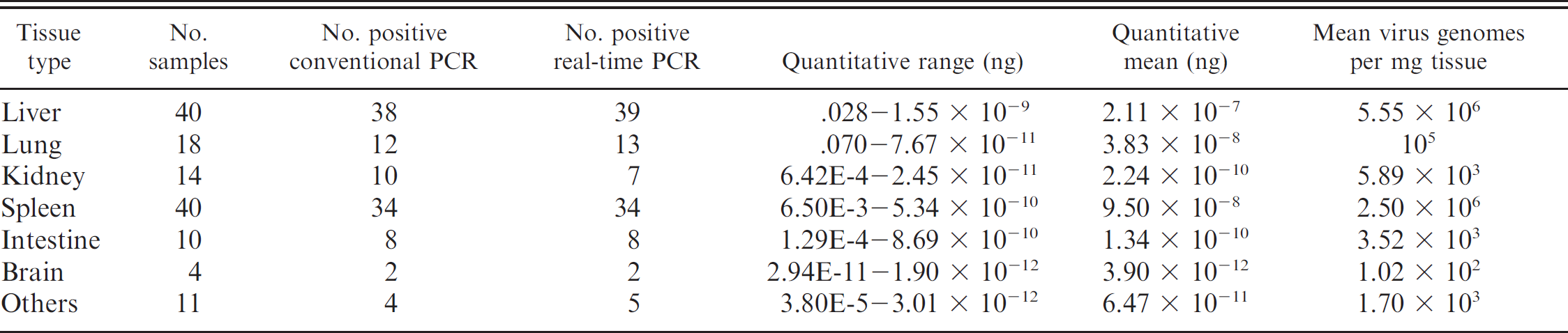
Comparison of conventional and real-time PCR assays for falcon adenovirus and summary of quantitative real-time PCR values.
In situ hybridization with the falcon-specific hexon region probe demonstrated virus in tissues from 6 out of 6 Northern aplomado falcons that were positive by PCR. Hepatocytes and biliary epithelium were most consistently infected with virus. Positive staining was usually intranuclear, although some cells had significant cytoplasmic reactivity. Positive foci often comprised groups of 3–5 adjacent cells and were associated with variable (none to approximately 20) inflammatory cell infiltrates. Less commonly, positive cells were present in renal tubular epithelium, epithelium of the ventriculus and small intestine, parabronchiolar epithelium, splenic mononuclear leukocytes, pancreatic acinar cells, glial cells in the brain, and adipocytes (Fig. 4). No staining was seen in negative control sections or in control sections containing herpes simplex virus 1.
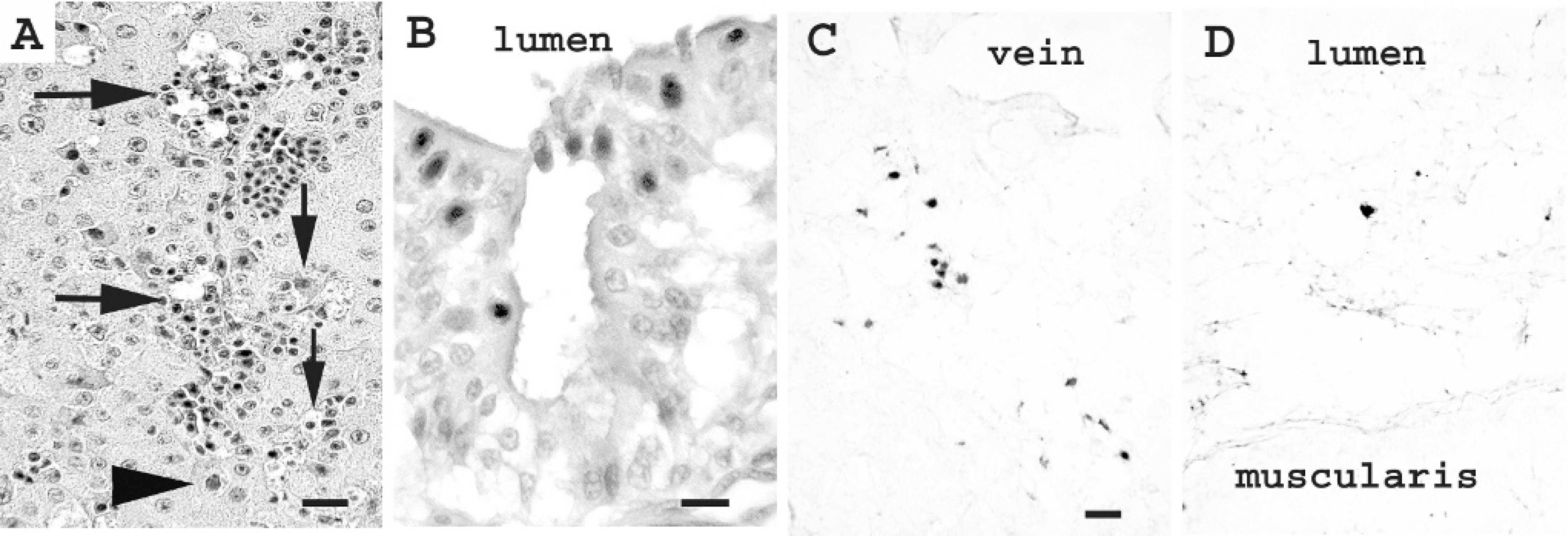
Photomicrographs of falcon organs infected with adenovirus:
Discussion
Since 1996, falcon adenovirus has emerged as a significant pathogen in the United Sates, killing at least 82 animals in 5 locales. 3,7,10 The epidemiology of the virus is not completely understood, but Peregrine falcons may be natural hosts and reservoirs. Because of the ongoing risk of adenoviral disease in mixed-species facilities, identification of carrier animals and accurate investigations of diseased animals is critical. The aim of this study was to develop specific and sensitive molecular assays for detecting falcon adenovirus.
The falcon-specific hexon region PCR was designed to amplify a segment within the L3 region of the hexon protein. The loops of adenovirus capsid proteins are variable segments supported by structurally conserved beta barrel platforms, and within the loops are hypervariable zones. 1 The L3 region primers covered areas encoding several variable residues and were, overall, highly specific for the falcon adenovirus. The specificity of the assay allows other viruses, such as the fowl adenoviruses or red-tailed hawk adenovirus, to be excluded as etiologies of outbreaks without sequencing amplicons. The PCR was also able to amplify both genotypes of falcon adenovirus. This capability provides a reliable means for detecting and characterizing strain variants and tracing origins and patterns of virus spread in disease outbreaks.
The real-time falcon-specific pVI region with EIC PCR was a rapid and efficient method for analyzing samples. Assay results were completed within 2.5 hours and included the increased fidelity of probe hybridization. Also, because the assay was a multiplex PCR, the presence of inhibitors was evaluated simultaneously with adenovirus testing. An additional advantage was the ability to quantitate virus, which could be important for assessing shedding loads in captive animals and understanding risk factors involved in the pathogenesis of disease. 3 The real-time falcon-specific pVI region with EIC PCR also provided a modest increase in sensitivity over conventional PCR. Unlike the conventional PCR, however, the real-time assay does not allow for strain typing of virus.
In situ hybridization provided the advantage of correlating the presence of virus with histology. Both diagnosis of infection and distribution of virus could be assessed, allowing insights into mechanisms of disease and possible modes of transmission. Although the specificity of the technique was not tested against other adenoviruses, it is likely that hybridization was specific for the falcon adenovirus at high stringency conditions because the probe targeted the hexon L3 region. In situ hybridization also allowed testing of particular pieces of tissue that had microscopic lesions suggestive of adenovirus infection and avoided the process of DNA extraction and has been found to be valuable for identifying infections in other avian species. 6
The ability to perform retrospective studies on paraffin-embedded tissues is important for understanding disease of wildlife. 11 In situ hybridization was a sensitive method for analyzing samples archived in paraffin. Conventional PCR for falcon adenovirus hexon and the real-time assay were also effective on DNA from paraffin tissues, although the efficiency was less than for DNA from frozen tissues or fecal samples. Because the real-time assay requires only 64 bp of intact target DNA, it is better suited for evaluating formalin-fixed samples than the conventional PCR, which amplifies 385 bp. Combining the 3 assays would be most useful in any retrospective analysis by providing the highest level of sensitivity, the ability to strain-type virus, and insights into virus distribution and latency during infections.
Prior to the development of molecular techniques, classical diagnostic tests for adenoviruses (i.e., virus morphology, serology, and virus isolation) were used for identifying viruses from a variety of animals. However, each has particular limitations. 2,4,5,8,10 Morphology and virus isolation are relatively insensitive techniques and do not allow characterization of viruses as species and strains. 5 Serology is a powerful method for detecting exposure and has been developed for falcon adenovirus. 7 It is highly sensitive and is often specific for adenovirus subtypes. 5,7 However, it does not provide information on active infection or virus shedding and has limited utility on postmortem studies. Molecular assays are both sensitive and specific for identification of adenoviruses and complement serologic techniques for diagnosing infection. 5 Because the falcon adenovirus is a widespread pathogen that poses significant risk to captive and wild falcons, conservation scientists should use these assays to minimize virus spread and disease outbreak.
Acknowledgements
This work was supported by the Zoological Society of San Diego. Technical assistance was provided by April Gorow and Yvonne Cates. We thank the Beckman Coulter Corporation for donation of the automated capillary DNA sequencer and high-speed centrifuge used in these studies, and Charles and Shirley Sykes of San Diego for financial support of our laboratory.
Footnotes
a.
Dr. S. Dhillon, Washington State University Research and Extension Center, Puyallup, WA.
b.
SPAFAS Charles River Diagnostics Laboratories, Storrs, CT.
c.
Dr. L. A. Hobbs, National Animal Disease Center, Ames, IA.
d.
Dr. Lindsay Oaks, Washington State University, Pullman, WA.
e.
Qiagen, Valencia, CA.
f.
Q-BIOgene, Carlsbad, CA.
g.
Applied Biosystems, Foster City, CA.
h.
Invitrogen, Carlsbad, CA.
i.
Beckman Coulter, Fullerton, CA.
j.
Stratagene, La Jolla, CA.
k.
Amersham Pharmacia Biotech, Piscataway, NT.
l.
Millipore, Bedford, MA.
m.
Roche Molecular Biochemicals, Indianapolis, IN.
n.
Eastman Kodak Co., Rochester, NY.
o.
Mercedes Medical, Sarasota, FL.
p.
Zymed Laboratories, South San Francisco, CA.
q.
Surgipath Instrumentation Inc., Richmond, IL.
r.
Biomeda Corp., Foster City, CA.
s.
Accelrys, San Diego, CA.
