Abstract
Deficiency of methyl donor nutrients folate, choline, and methionine (methyl deficiency) during gestation can impair fetal development and perturb DNA methylation. Here, we assessed genetic susceptibility to methyl deficiency by comparing effects in wildtype C57BL/6J (B6) mice to mutant mice carrying a 1.3 kb deletion at the H19/Igf2 Imprinting Control Region (ICR) (H19ICRΔ2,3). The H19ICRΔ2,3 mutation mimics microdeletions observed in Beckwith-Wiedemann syndrome (BWS) patients, who exhibit epimutations in cis that cause loss of imprinting and fetal overgrowth. Dams were treated during pregnancy with 1 of 4 methyl sufficient (MS) or methyl deficient (MD) diets, with or without the antibiotic commonly used to deplete folate producing gut microbes. As expected, after ~9 weeks of treatment, dams in MD and MD + antibiotic groups exhibited substantially reduced plasma folate concentrations. H19ICRΔ2,3 mutant lines were more susceptible to adverse pregnancy outcomes caused by methyl deficiency (reduced birth rate and increased pup lethality) and antibiotic (decreased litter size and litter survival). Surprisingly, pup growth/development was only minimally affected by methyl deficiency, while antibiotic treatment caused inverse effects on B6 and H19ICRΔ2,3 lines. B6 pups treated with antibiotic exhibited increased neonatal and weanling bodyweight, while both wildtype and mutant pups of heterozygous H19ICRΔ2,3/+ dams exhibited decreased neonatal bodyweight that persisted into adulthood. Interestingly, only antibiotic-treated pups carrying the H19ICRΔ2,3 mutation exhibited altered DNA methylation at the H19/Igf2 ICR, suggesting ICR epimutation was not sufficient to explain the altered phenotypes. These findings demonstrate that genetic mutation of the H19/Igf2 ICR increases offspring susceptibility to developmental perturbation in the methyl deficiency model, maternal and pup genotype play an essential role, and antibiotic treatment in the model also plays a key independent role.
Keywords
Introduction
Optimal dietary intake of methyl donor nutrients such as folate, choline, and betaine during pregnancy is essential to healthy fetal growth and development, as well as long-term health outcomes.1-7 Although there is widespread fortification of folate in the United States, in many other countries worldwide there is a high prevalence of suboptimal methyl donor nutrient intake among women of reproductive age, particularly in economically challenged communities.8-11 Methyl donor nutrients are essential substrates in the 1 carbon metabolism pathway that generates S-adenosylmethionine (SAM), 12 the primary methyl group donor for methylation of DNA, and histone residues. 13 Therefore, availability of methyl donor nutrients has the potential to modulate the DNA methylome and related health outcomes. 12
DNA methylation occurs primarily at CG dinucleotides and is generated by DNA methyltransferase (DNMT) activity through covalent attachment of methyl groups from SAM to the fifth carbon position of the DNA base cytosine to generate 5-methylcytocine (5mC). One of the primary functions of DNA methylation is to act in combination with other epigenetic modifications to regulate gene expression via transcriptional activation or repression depending on the proximity of the methylation mark to specific regulatory elements such as promoters, insulators, enhancers, transcription factor binding sites, etc. 14 During the susceptible window of gestation, germ, and somatic cells of the developing mammalian embryo are epigenetically reprogrammed such that DNA methylation patterns are erased and reestablished in a locus and cell-specific manner.15-17 Once established, many methylation marks must be maintained and transmitted from parent to daughter cells to maintain normal cellular function. 18 Therefore, perturbation of DNA methylation by suboptimal nutrient availability during development can substantially perturb development and cause disease. 13
Deficiency of methyl donor nutrients folate, methionine, and choline during pregnancy have been shown to perturb DNA methylation. For example, offspring born to C57BL/6J mouse dams fed low folate diet during mating, gestation, and lactation, exhibited significant loss of global DNA methylation of the small intestine compared to dams belonging to the control/high folate group. 19 Importantly, supplementation postweaning did not rescue these effects demonstrating persistent epimutations can occur as a result of in utero deficiency. 19 Other studies that have assessed methylation in a locus specific manner show that methyl deficiency in utero can lead to loss of methylation at some loci and gain of DNA methylation at other loci.20,21 One study in particular showed gain of methylation at Igf2 in response to choline deficiency during gestation. 22
The primary objective of this study was to use the in utero methyl deficiency model of environmental perturbation to determine whether genetic mutations at the H19/IGF2 imprinting control region (ICR) found in Beckwith Wiedemann Syndrome (BWS) patients, increases sensitivity to epigenetic perturbation during development. Human and animal model studies suggest imprinted domains may be particularly sensitive to environmental influences during gestation.23-26 The ICR is differentially methylated and acts as the key regulatory element of “imprinted” parent of origin dependent monoallelic gene expression. 27 The differing methylation status at the maternal versus paternal ICR alleles is established during germ cell development (maternal allele in oocytes and paternal allele in sperm) and maintained after fertilization in the soma of the developing embryo and throughout the lifespan. 27 Thus, ICRs represent a rigorous and reproducible model locus to investigate the role of DNA sequence context in regulating offspring epigenetic susceptibility to environmental perturbation. The H19/IGF2 ICR regulates imprinted expression.28-31 of insulin-like growth factor 2, IGF2, a protein coding gene with key roles in growth and metabolism, 32 and H19, a non-coding RNA. 33 Both genes have essential roles in embryonic and placental growth and development. 27 The H19/IGF2 ICR is the most well-characterized ICR, located in the intergenic region between H19 and IGF2, with conserved synteny and function between mouse and human. 27 The maternal ICR allele is normally hypomethylated and bound by the insulator protein CCCTC-binding factor (CTCF), while the paternal allele is hypermethylated and CTCF-free. 27 A combination of epigenetic markings and CTCF binding at the H19/IGF2 ICR regulates promoter-enhancer interactions required to maintain allele-specific expression of H19 and IGF2. 27 Aberrant gain of methylation at the maternal H19/IGF2 ICR during development causes biallelic overexpression of IGF2 and subsequent fetal and postnatal overgrowth in BWS patients.34,35 This epimutation is found in approximately 5% to 10% of BWS patients and is currently used to diagnose BWS.35-37
The cause of gain of methylation epimutations at the H19/IGF2 ICR that lead to BWS are unknown. However, it has been proposed that environmental insults and/or changes in spacing or number of CTCF sites caused by ICR microdeletions may epigenetically destabilize the locus.28,30,35,38 H19/IGF2 ICR microdeletions have been identified that coincide with these epimutations,28,30,38-41 and targeted mutant mouse models made to mimic or directly insert patient deletions (humanized models) have been shown to partially recapitulate the overgrowth phenotype observed in BWS.31,42
Here, we tested whether maternal H19/Igf2 ICR microdeletion makes the locus more sensitized to environmental epigenetic perturbation either during somatic cell development when the ICR methylation status must be maintained, or during male germcell (sperm) development when the embryo’s maternally hypomethylated allele must become hypermethylated.
The secondary objective of this study was to determine whether the antibiotic routinely used in methyl donor deficiency mouse models contributes to any of the molecular or developmental phenotypes observed. Standard methyl deficiency mouse models include 1% succinylsulfathiozole (SS), an antibiotic used to kill folate-producing gut microbes, and thus further exacerbate folate deficiency. 43 Although Burgoon and colleagues previously reported that diets containing 1%SS alone did not lead to major developmental defects, 43 minor developmental disturbances and molecular effects such as epigenetic responses have not been extensively investigated. In light of what we now know about the seemingly pervasive role of the microbiome in health, and findings that antibiotic disruption of the intestinal microflora causes a wide range of developmental and metabolic disorders,44-47 it is important to reassess the impact of 1%SS in this model. Therefore, in this study, we have assessed the effect of methyl deficiency and the independent effect of 1%SS antibiotic treatment in perturbing development and methylation status of offspring.
Results
Maternal dietary depletion of methyl donors reduced maternal plasma folate levels but not SAM availability
To determine whether mice carrying a BWS-like microdeletion at the H19/Igf2 ICR are more susceptible to developmental and epigenetic perturbation caused by methyl donor deficiency, we compared the response of wild-type C57BL/6 (B6) mice to mice carrying a targeted 1.3 kb deletion at the H19/Igf2 Imprinting Control Region (ICR) (Figure 1). 31 To induce methyl donor deficiency but also allow for distinguishing potentially unrelated effects of antibiotic, we compared 4 diet groups (1) methyl sufficient (MS), (2) MS1%SS, (3) methyl deficient (MD), and (4) MD1%SS (Figure 1A). Dams from 3 genotypes (Figure 1B) were treated for 5 weeks before pregnancy to ensure pup deficiency at conception, throughout gestation, and until postnatal day 5 (PND 5). Thereafter, dams and pups were placed on standard rodent chow (Harlan 8604) (Figure 1A).

Dietary treatment scheme and experimental crosses: (A) dietary treatments: (1) methyl sufficient (MS), (2) MS with 1%SS (MS1%SS), (3) methyl deficient (MD), and (4) MD with 1%SS (MD1%SS). Solid arrow above indicates dietary treatment windows. Arrows below indicate the timepoint of experimental steps shown below, and (B) crosses used to generate G1 pups shown as dam x sire. Arrows indicate progeny genotype outcomes shown below. Wildtype C57BL/6 (B6, +/+), heterozygous (∆2,3/+), or homozygous (∆2,3/∆2,3) mutant dams were crossed with wildtype C7 sires to produce wildtype B6 (+/+) offspring, wildtype offspring from the ∆2,3 lineage (∆2,3(+/+)), and heterozygous offspring from the ∆2,3 lineage (∆2,3/+).
To evaluate the effect of the diets on maternal methyl donor metabolite availability to the offspring at birth and weaning, we measured maternal plasma folate levels after dietary depletion (~9 weeks, equivalent to status at birth) and after dietary repletion (~16 days restored to folate sufficient diet, equivalent to status at weaning). Since there was no significant effect of dam genotype on plasma folate levels expected or detected for the status after dietary depletion (P = .53) or repletion (P = .56), we combined the data for all genotypes. As expected, dietary depletion caused a significant reduction in maternal plasma folate concentrations such that concentrations in the MD and MD1%SS diet groups were significantly lower compared to the MS diet group (Figure 2A). The antibiotic treatment alone (MS1%SS) was not sufficient to significantly reduce plasma folate levels, however, the combined methyl donor deficiency and antibiotic treatment group (MD1%SS) exhibited the lowest concentration of folate (83% lower than MS) although not significantly lower than the MD group (Figure 2A). Plasma folate concentrations were completely restored in dams treated with 16 days of dietary repletion post-parturition using standard chow (Figure 2A).

Maternal folate, SAM, and SAH response to methyl donor dietary depletion and dietary repletion: (A) box and whiskers plot of maternal plasma folate levels for dams after dietary depletion. Sample sizes in order from left to right, (n = 6, 4, 7, 8) or dietary repletion (n = 4, 4, 5, 2), and (B) box and whiskers plot of maternal liver SAM, SAH, and SAM/SAH ratio (n = 5 for all groups). Each dot represents an individual sample, Black (MS base diet groups), Red (MD base diet groups).
Methyl donor nutrient availability is a key determinant of the concentrations of the primary methyl donor liver metabolite S-adenosylmethionine (SAM) and its metabolic byproduct S-adenosylhomocysteine (SAH). 48 The ratio of SAM/SAH has been proposed to be an indicator of the cellular methylation potential. 49 Folate-deficient diets have been reported to decrease SAM and SAM/SAH ratio and increase SAH concentrations.50-53 Therefore, we determined whether maternal folate depletion altered maternal availability of SAM, SAH, and the ratio of SAM/SAH. Surprisingly, the substantial decrease in maternal plasma folate observed in our model did not cause any significant changes in liver concentrations of SAM, SAH, or the SAM/SAH ratio. Although there was a slight decrease in average SAM levels in the MD1%SS group (23.0 ± 4.2 nmol/g, ±error represents standard error of the mean) compared to the MS group (30.8 ± 3.4 nmol/g), it was not statistically significant (Figure 2B).
Dams carrying the Δ2,3 mutation are more susceptible to adverse pregnancy outcomes caused by methyl donor depletion and antibiotic treatment
Methyl donor deficiency during the periconceptual period has been associated with poor reproductive and birth outcomes. 54 To determine whether mice carrying the BWS-like Δ2,3 microdeletion at the H19/Igf2 ICR are more susceptible to adverse pregnancy outcomes of methyl donor deficiency and whether the antibiotic treatment plays a role, for each dam genotype (+/+; Δ2,3/+, and Δ2,3/Δ2,3) we measured the (i) Methyl donor depletion effect (Diet) by comparing MS + MS1%SS versus MD + MD1%SS samples; and (ii) Antibiotic treatment effect determined by comparing MS + MD versus MS1%SS + MD1%SS samples.
Methyl donor depletion did not significantly alter litter size at birth for any of the genotypes (Figure 3A). However, for Δ2,3/+ depleted dams, there were significantly more females with that did not produce a litter, 32% (8/25) compared to 6% (1/15) of methyl sufficient Δ2,3/+ dams (Figure 3B). Methyl donor depletion also did not have a significant effect on litter survival to weaning for any of the genotypes (Figure 3C), but did result in a significantly higher rate of pup lethality for Δ2,3/Δ2,3 dams only, with an average of 24% of pups dying before weaning compared to 0% of methyl sufficient Δ2,3/Δ2,3 dams (Figure 3D). A similar trend of pup lethality was observed for methyl donor depleted Δ2,3/+ dams but was not statistically significant (Figure 3D). These data demonstrate that Δ2,3 pregnancies are more susceptible to adverse effects of methyl donor depletion compared to wild-type B6 pregnancies.

H19ICRΔ2,3 genetic susceptibility to pregnancy outcomes of methyl donor depletion and antibiotic treatment: (A) box and whiskers plot of litter sizes at birth for each treatment group and cross [Sample sizes in order from left to right, B6(+/+) n = 4, 7, 5, 8; ∆2,3/+ n = 10, 5, 12, 5; ∆2,3/∆2,3 n = 6, 4, 6, 4]. Each dot represents a litter, (B) proportion of matings resulting in live birth [B6(+/+) n = 4, 9, 6, 12; ∆2,3/+ n = 11, 5, 16, 9; ∆2,3/∆2,3 n = 8, 4, 6, 4], (C) pup fitness as determined by proportion of litter that survived to weaning [B6(+/+) n = 4, 7, 5, 9; ∆2,3/+ n = 10, 5, 12, 5; ∆2,3/∆2,3 n = 6, 4, 6, 4], and (D) for litters that survived to weaning [B6(+/+) n = 4, 5, 5, 6; ∆2,3/+ n = 10, 3, 11, 3; ∆2,3/∆2,3 n =3, 3, 3, 1], the percent of pups for each litter that died before weaning. Error bars indicate standard error of the mean (SEM).
Unlike methyl donor depletion, antibiotic treatment did not alter number of females that gave birth to a litter or partial litter survival to weaning (Figure 3B and D
Due to the high rate of pup lethality that led to very few pups surviving past birth, we did not further analyze developmental or epigenetic outcomes for Δ2,3/Δ2,3 pregnancies.
Offspring of Δ2,3 dams are more susceptible to developmental delay/growth restriction response to antibiotic treatment
Suboptimal methyl donor intake during pregnancy has been linked to reduced neonatal weight. 48 On the other hand, we have shown that mutant offspring of Δ2,3/+ dams exhibit classic BWS-like increased neonatal weights compared to their wildtype littermates, likely due to loss of imprinting at the H19/Igf2 locus and increased expression of the growth factor Igf2. To determine whether offspring of Δ2,3/+ dams exhibit increased susceptibility to impaired growth/development due to methyl donor deficiency or antibiotic treatment, we measured offspring bodyweight at PND5 (neonate), PND21 (weaning, juvenile), and PND56 (young adult) and compared to that of wildtype B6 offspring. For each mouse lineage (B6 or Δ2,3, G1 lineage) and pup genotype (+/+ or −/+) we measured the (i) Methyl donor depletion effect (Diet) by comparing MS+MS1%SS versus MD + MD1%SS samples; and (ii) Antibiotic treatment effect determined by comparing MS + MD versus MS1%SS + MD1%SS samples.
Methyl donor depletion in utero did not alter neonatal bodyweight of B6 offspring (Figure 4A). On the other hand, Δ2,3 neonates (combined genotypes as not genotyped at this stage) were significantly affected by methyl donor depletion and weighed on average ~6% (~180 mg) less than methyl sufficient pups (2.64 ± 0.03 g vs. 2.82 ± 0.04 g, respectively), with male and female pups similarly affected (Figure 4A). This is unlikely biased by combining genotypes since genotyping at PND21 revealed similar rates of transmission for each treatment group (MS, n=28:39; MS1%SS, n = 7:8; MD, n = 24:27, MD1%SS, n = 7:7). At weaning, methyl depleted B6 offspring weighed on average ~7% (~0.64 g) more than sufficient pups (9.29 ± 0.16 g vs. 8.65 ± 0.16 g, respectively), with male and female juvenile pups similarly affected (Figure 4B). MD1%SS B6 weanlings weighed the most on average at 9.45 ± 0.25 g compared to 8.27 ± 0.21g for MS pups, (~1.2 g more on average) (Figure 4B). For Δ2,3 juvenile pups, there was no significant effect of methyl depletion alone for either (+/+) or (−/+) offspring genotypes (Figure 4B). Developmental effects of methyl donor depletion did not last into adulthood for any of the genotypes (Figure 4C).

Offspring growth response to in utero methyl donor depletion and antibiotic treatment: (A) top panel: Bar graph of offspring PND5 (neonatal) bodyweight separated by wildtype (B6, left panel) versus mutant (∆2,3, right panel) dam lineage and pup genotype (+/+ or +/− in parentheses). Mice not genotyped at PND5 therefore, data for both ∆2,3 pup genotypes are combined [sample sizes in order from left to right, B6(+/+) n = 19, 29, 20, 19; ∆2,3(+/+ and −/+) n = 80, 9, 64, 14]. Bottom panel: Individual pup weights (dots) separated by sex, male (M, black dots), female (F, grey dots); lines connect means between groups for males (solid line) or females (dashed line), (B) top panel: Bar graph of offspring PND21 (weaning) bodyweight separated by dam lineage and pup genotype [B6(+/+) n = 19, 25, 19, 19; ∆2,3(+/+) n = 23, 7, 23, 6; ∆2,3(−/+) n = 38, 7, 27, 6]. Bottom panel: Individual pup weights (dots) separated by sex, and (C) bar graph of offspring PND56 (adult) bodyweight (males only) separated by dam lineage and pup genotype. [B6(+/+) n = 9, 14, 7, 11; ∆2,3(+/+) n = 17, 2, 18, 4; ∆2,3(−/+) n = 19, 2, 16, 2].
Antibiotic treatment in utero, compared to untreated mice, led to a significant increase in B6 offspring bodyweight for neonatal and juvenile stages (neonatal, ~10%/~260 mg increase, 2.97 ± 0.0.05 g vs. 2.71 ± 0.05 g; juvenile, ~5%/0.46 g increase, 9.16 ± 0.16 g vs. 8.70 ± 0.16 g, respectively), with both males and females similarly affected (Figure 4A and B). However, this developmental effect did not last into adulthood (Figure 4C). In contrast, Δ2,3 lineage offspring exposed to antibiotic, compared to untreated mice, exhibited a significant decrease in bodyweight at neonatal, juvenile, and adult stages (Figure 4A to C). On average, antibiotic treated (+/+) and (−/+) neonates of Δ2,3/+ dams (combined as not genotyped at this stage) weighed ~12% (340 mg) less than untreated mice (2.45 ± 0.05 g vs. 2.79 ± 0.03 g, respectively) (Figure 4A). Antibiotic treated (+/+) juveniles of Δ2,3/+ dams weighed ~20% (1.9 g) less on average (7.31 ± 0.27 g vs. 9.17 ± 0.13 g, respectively), while adults weighed ~7% (1.6 g) less on average (22.27 ± 0.26 g vs. 23.90 ± 0.33 g) (Figure 4B and C). However, the effect of antibiotic treatment was seemingly more severe for males compared to female (+/+) juveniles of Δ2,3/+ dams (Figure 4B). Antibiotic treated (−/+) juveniles of Δ2,3/+ dams weighed ~17% (1.6 g) less on average (7.49 ± 0.27 g vs. 9.07 ± 0.13 g, respectively), while adults weighed only ~5% (1.4 g) less on average (24.82 ± 0.39 g vs. 26.18 ± 0.56 g), but the latter was not statistically significant (Figure 4B and C).
Similar effect of methyl donor depletion and antibiotic on both wildtype (+/+) and mutant (−/+) offspring of Δ2,3 dams implicates that these developmentally-induced changes in bodyweight are caused by a maternal effect.
Offspring from H19ICRΔ2,3 dams are more susceptible to subtle epigenetic perturbation in cis
Methyl donor nutrients play a key role in several transmethylation pathways essential for metabolic and epigenetic regulation, especially during gestational reprogramming events. 55 To determine whether increased susceptibility of the Δ2,3 lineage to methyl deficiency and antibiotic treatment is caused by epigenetic perturbation of the mutated H19/Igf2 locus, we assessed the DNA methylation state across key regulatory elements within the domain in offspring liver and sperm. We measured methylation in liver at : (i) CTCF binding sites at the beginning and end of the imprinting control region (ICR) that regulates H19 and Igf2 expression (H19CTS1 and H19CTS4); (ii) the region immediately upstream of the H19 promoter that regulates H19 expression (H19PromProx); and (iii) a differentially methylated region at Igf2 that regulates Igf2 expression (Figure 5A).

Male offspring liver DNA methylation status at the H19/Igf2 domain in response to in utero methyl donor depletion and antibiotic treatment: (A) schematic of H19/Igf2 domain showing expected allelic expression and methylation status for liver. Blue arrows indicate regions assessed for DNA methylation changes, (B) bar graph of methylation levels at the H19/Igf2 ICR at CTCF site 1 (H19CTS1), (C) bar graph of methylation levels at CTCF site 4, (H19CTS4) (D) bar graph of methylation levels at the H19 promoter proximal region, and (E) bar graph of methylation levels at the Igf2DMR1. Sample sizes in order from left to right, B6(+/+) n = 8, 10, 10, 9; ∆2,3(+/+) n = 3, 2, 5, 4; ∆2,3(−/+) n = 3, 2, 1, 2.
In adult offspring liver, which is normally partially methylated (~50%) at the regions tested, due to gain of methylation on the paternal allele during gestation (Figure 5A), we found that methyl donor depletion in utero did not significantly alter liver DNA methylation status at any of the key regulatory regions tested at the H19/Igf2 domain for any of the genotypes (Figure 5B to E). However, a trend of increased DNA methylation was detected for Δ2,3 (+/+) offspring at the ICR CTCF binding site 1 (Figure 5B) and for Δ2,3 (+/+) and (−/+) offspring at the Igf2DMR1 (Figure 5E). Antibiotic treatment in utero similarly had little effect on liver DNA methylation at the H19/Igf2 domain for any of the genotypes (Figure 5B to E). We did detect a subtle but significant decrease in DNA methylation at the ICR CTCF binding site 4 that was induced by antibiotic treatment but only in Δ2,3 (−/+) offspring (63.0 ± 1.6% vs. 60.4 ± 1.7%) (Figure 5C). B6 and wildtype Δ2,3 (+/+) offspring were not similarly affected (Figure 5C).
In adult offspring sperm, which is normally fully methylated (90% to 100%) at the regions tested, due to gain of methylation on both maternal and paternal alleles during gestation (Figure 6A), we found that methyl donor depletion in utero caused a subtle (<2%) but significant decrease in DNA methylation at the ICR CTCF binding site 1 in offspring from B6 (+/+) dams (Figure 6B). However, this effect was not observed for any other regions tested at the H19/Igf2 domain or for any other genotypes(Figure 6B to E). In contrast, a subtle but significant increase in DNA methylation was observed for methyl donor depleted Δ2,3 (+/+) offspring at the Igf2DMR1 (Figure 6E). Other than a subtle significant increase in methylation for Δ2,3 (−/+) offspring (Figure 6B), antibiotic treatment in utero had little effect on sperm DNA methylation at the H19/Igf2 domain for any of the genotypes (Figure 5B to E).
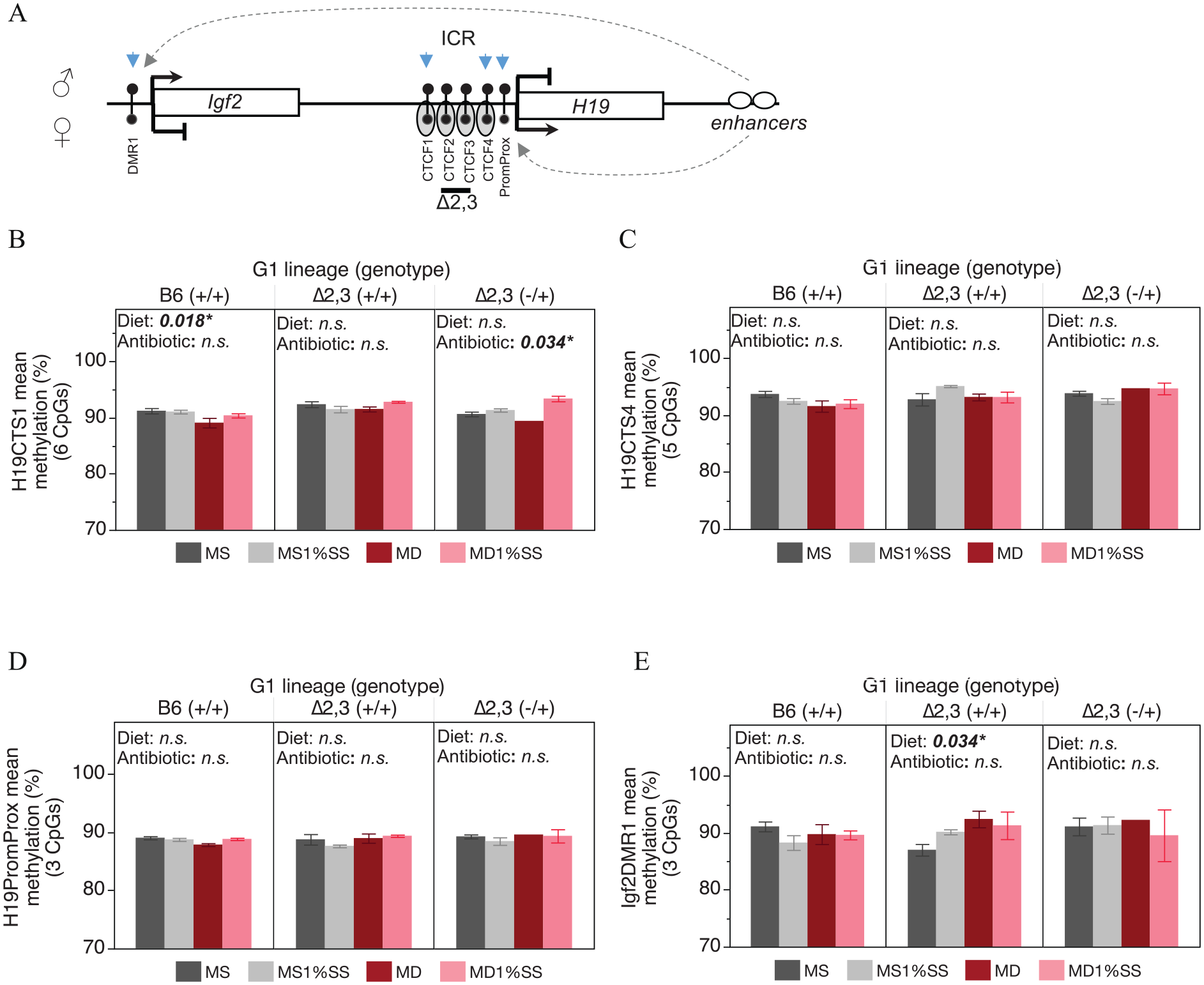
Male offspring sperm DNA methylation status at the H19/Igf2 domain in response to in utero methyl donor depletion and antibiotic treatment: (A) schematic of H19/Igf2 domain showing expected allelic expression and methylation status for sperm. Blue arrows indicate regions assessed for DNA methylation changes, (B) bar graph of methylation levels at the H19/Igf2 ICR at CTCF site 1 (H19CTS1), (C) bar graph of methylation levels at CTCF site 4 (H19CTS4), (D) bar graph of methylation levels at the H19 promoter proximal region, and (E) bar graph of methylation levels at the Igf2DMR1. Sample sizes in order from left to right, B6(+/+) n = 8, 10, 7, 9; ∆2,3(+/+) n = 3, 2, 5, 3; ∆2,3(−/+) n = 4, 3, 1, 2.
Interestingly, we found that methylation status at the H19/Igf2 domain also differed by mouse line and genotype independent of diet or antibiotic treatment. After adjustment for diet and antibiotic treatment effects in the regression model, Δ2,3 (−/+) mutant offspring liver exhibited significantly higher levels of DNA methylation than B6 (+/+) wildtype offspring at all regulatory regions tested across the H19/Igf2 domain, and significantly higher levels than Δ2,3 (+/+) wildtype offspring at all regions except the distal portion of the H19/Igf2 ICR (H19CTS4) (Figure 7A). On the other hand, Δ2,3 (+/+) wildtype offspring were only significantly higher than B6 (+/+) wildtype offspring at H19CTS4 and the region upstream of the H19 promoter (H19PromProx), while similar methylation levels were observed at the proximal portion of the ICR and Igf2 (H19CTS1 and Igf2DMR1) (Figure 7A). After adjustment for treatment effects, adult body weight was significantly lower in Δ2,3 (−/+) and Δ2,3 (+/+) offspring compared to B6 (+/+) (Figure 7B) and adult body weight was significantly correlated with H19CTS4 methylation levels (Figure 7C). In contrast to liver, genotype had only a subtle effect on the methylation status of sperm at the H19/Igf2 domain, with only a significant increase of <2% at the H19CTS1 locus (Figure 8A) that was not associated with bodyweight (Figure 8B).

Effect of genotype on adult liver methylation status at the H19/Igf2 domain independent of diet: (A) box and whiskers plot of offspring adult liver methylation levels at regulatory regions across the H19/Igf2 domain, (B) adult (PND56) bodyweight stratified by genotype, and (C) relationship between adult (PND56) bodyweight and DNA methylation at H19CTS4. Each dot represents an individual sample.
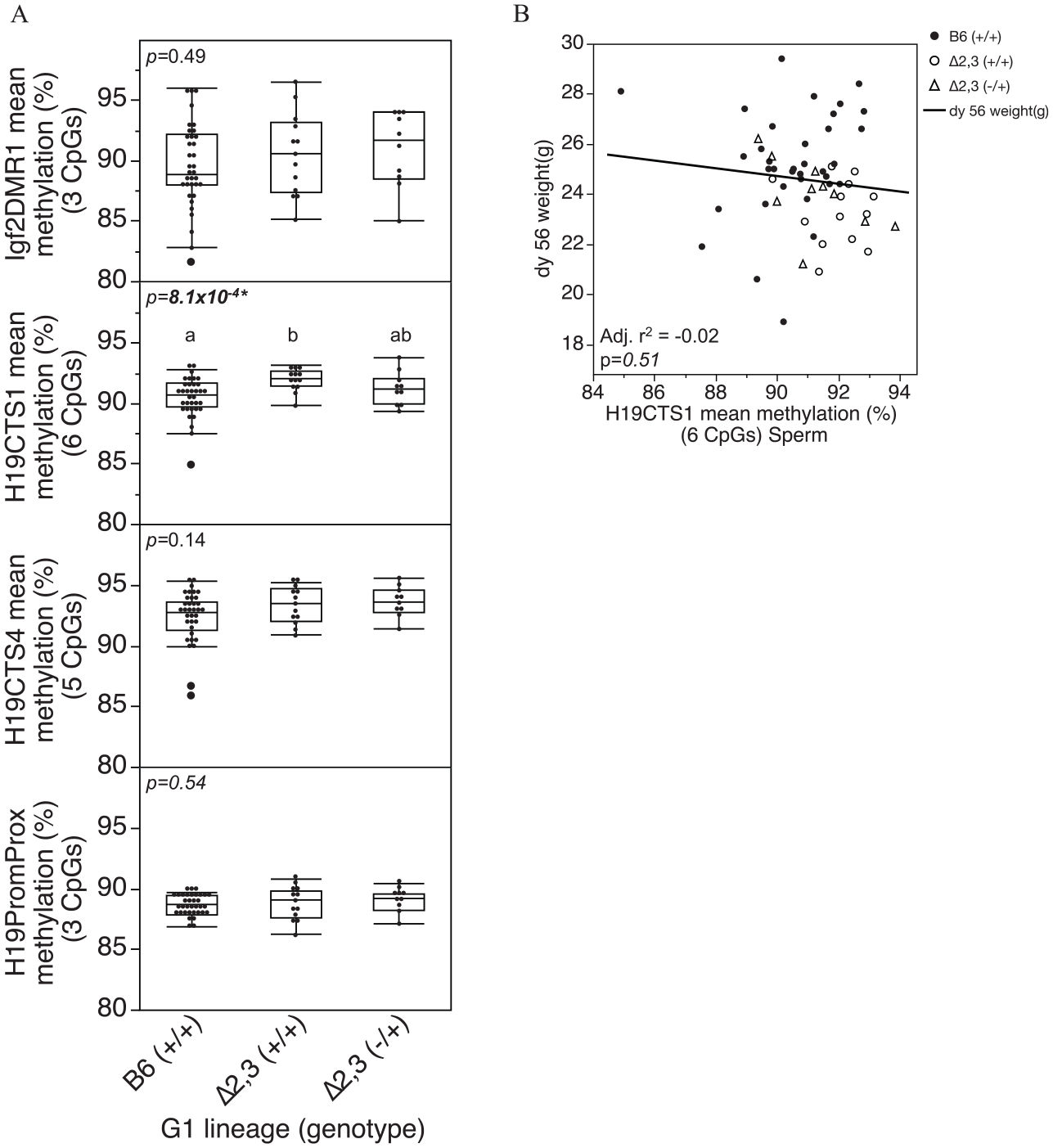
Effect of genotype on adult sperm methylation status at the H19/Igf2 domain: (A) box and whiskers plot of offspring adult sperm methylation levels at regulatory regions across the H19/Igf2 domain, and (B) relationship between adult (PND56) bodyweight and DNA methylation at H19CTS1. Each dot represents an individual sample.
Discussion
Here, we have demonstrated that the H19ICRΔ2,3 mutation makes mice more susceptible to environmentally-induced developmental effects including adverse pregnancy outcomes, restricted pup growth/development, and subtle loss of DNA methylation in cis. The H19ICRΔ2,3 mutation mimics microdeletions at the H19/IGF2 ICR observed in BWS patients 39 and was previously shown to mimic BWS overgrowth but only in fetal/neonatal stages and only when pups carried the mutation. 31 Therefore, it was surprising to find that in utero antibiotic treatment caused reduced bodyweight persisting into adulthood for offspring of H19ICRΔ2,3 dams and that both wildtype and mutant offspring were similarly affected (Figure 4). We also found that antibiotic treatment seemingly had a greater effect on pregnancy outcomes and postnatal development than methyl donor deficiency.
Antibiotic treatment more severely and consistently impaired pregnancy and pup development in a manner that was seemingly independent of methyl deficiency. Both B6 and H19ICRΔ2,3 dams exhibited reduced littersize at birth and litter survival to weaning (Figure 3A and C). However, similar to methyl deficiency effects, B6 and H19ICRΔ2,3 offspring exhibited inverse response in pup development, with B6 pups exhibiting increased neonatal and weanling bodyweight and wildtype and mutant H19ICRΔ2,3 littermates exhibiting decreased bodyweight from neonatal stages into adulthood (Figure 4). Although it is unclear what causes this inverse effect in susceptibility, the similar response of wildtype and mutant H19ICRΔ2,3 littermates suggests a maternal/intrauterine effect. Furthermore, the finding that mutant H19ICRΔ2,3 pups exhibited significantly altered ICR methylation (Figure 5), demonstrates that this deletion in cis does sensitize the locus to epigenetic perturbation by in utero environment although the magnitude of effect is very small (<2%) and unlikely to alter gene expression. The fact that wildtype littermates were not similarly affected, suggests this epimutation alone is not sufficient to explain the pup growth/developmental phenotype.
Independent of treatment, adult mice carrying the H19ICRΔ2,3 mutation inherited from the dam exhibited increased DNA methylation in cis in the liver at regions both upstream and downstream of the deletion (Figure 7A). Interestingly, their wildtype littermates also exhibited higher methylation at the ICR than wildtype B6 mice but only at regions downstream of the deletion (Figure 7A). This finding of altered adult ICR methylation changes, not previously found at neonatal stages, strongly suggests the H19ICRΔ2,3 mutation has delayed onset effects on methylation and possibly even imprinting at the locus. Furthermore, ICR epigenetic perturbation of both wildtype and mutant pups of heterozygous H19ICRΔ2,3/+ dams suggests the potential for a maternal/intrauterine effect of this mutation on imprinting defects at this locus. This could explain why both wildtype and mutant offspring of heterozygous H19ICRΔ2,3/+ dams in this study exhibited similar susceptibility to phenotypic response to methyl deficiency and antibiotic treatment. Interestingly, adult H19ICRΔ2,3/+ offspring actually weighed less than B6 controls despite ~10% higher Igf2DMR1 methylation levels (Figure 7). Higher perinatal Igf2DMR1 methylation is usually associated with increased Igf2 expression and subsequent growth in fetal and neonatal stages. 31 The role of Igf2 in postnatal bodyweight remains poorly understood, however, previous studies show upregulation of postnatal Igf2 can result in reduced bodyweight. 56
We were surprised to find that despite more than 9 weeks of dietary depletion of methyl donor nutrients folate, choline, and methionine, and a more than 80% reduction in maternal plasma folate levels, methyl deficiency in this model seemingly had no effect on methylation potential (SAM availability) and only subtle effects on pregnancy and pup development, primarily observed for H19ICRΔ2,3 offspring. It is possible that a longer feeding time is necessary or further depletion of redundant methyl donors is necessary to alter methylation potential. Based on the minimal phenotypic response to methyl deficiency, it was not surprising that there was no significant effect of methyl deficiency on DNA methylation at the locus (Figure 5E). Although there was a nonsignificant trend of gain of methylation at the Igf2DMR1 for methyl deficient H19ICRΔ2,3 offspring, this epimutation is usually associated with increased Igf2 expression and increased fetal growth. 31 In contrast, methyl deficient H19ICRΔ2,3 offspring actually exhibited decreased neonatal bodyweight (Figure 4A) suggesting the 2 outcomes are likely not linked. On the other hand, methyl deficient wildtype B6 offspring that also showed no change in methylation at the H19/Igf2 domain, exhibited normal neonatal bodyweight but had increased bodyweight at weaning (PND21). This finding suggests the interactive gene x diet effects detected here are more complex than just the result of imprinting defects in the offspring implicating a potential role for maternal/intrauterine effects.
Taken together, this data supports the role of genetic alteration at the H19/Igf2 ICR in susceptibility to in utero environmental perturbation. Furthermore, adverse developmental effects detected here likely occur as a result of maternal and possibly intrauterine effects of these mutations. Although the exact mechanism of maternal influence remains unclear, previous studies show that reduced Igf2 expression in the placenta leads to reduced potential for placental nutrient transport to the fetus and subsequent growth restriction of the fetus. 57 Likewise, H19 expression in the placenta also plays an important role in placental growth and subsequent function, in part by restricting growth of the placenta. 58 Future studies will test whether these mechanisms are at play in this model. This data also show for the first time that treatment with 1% succinylsulfathiozole antibiotic during mouse pregnancy restricts pup growth/development beyond the perinatal period. Growth restriction has been linked to increased mortality, 59 neurodevelopmental disorders, 60 and metabolic disorders 61 later in life. This model can also be used to further understand the role of antibiotic in these outcomes.
Materials and Methods
Animals – sources, housing dietary treatments, and breeding scheme
Animal handling was performed in accordance with the Guide for the Care and Use of Laboratory Animals under the corresponding animal use protocol at the University of North Carolina at Chapel Hill and housed at the David H. Murdock Institute Vivarium in Kannapolis, NC. The mouse lines used were (i) wild-type fully inbred C57BL/6 (B6) mice, (ii) wild-type mice homozygous for CAST/EiJ across chromosome 7 and mixed CAST/EiJ-B6 genetic background (C7), and (iii) previously 31 described mutant H19ICRΔ2,3 mice carrying a targeted deletion of 1.3 kb at the H19/Igf2 imprinting control region on a mostly (>98%) B6 genetic background. As previously described, 62 C7 sires used here carry CAST genetic sequence at the ICR versus the B6 sequence of the dams and were used in the experimental design for the purpose of distinguishing parental origin of the ICR alleles, should substantial epigenetic changes suggest an imprinting error has occurred. B6 mice were purchased from The Jackson Laboratory (Bar Harbor, ME) and C7 and H19ICRΔ2,3 mice were derived in Dr. Marisa Bartolomei’s lab at the University of Pennsylvania. All were bred in-house at the David H. Murdoch Institute Vivarium in Kannapolis to generate experimental animals. Sterilized water and rodent chow were fed ad libitum. Vivarium temperature was maintained between 21 and 23 °C with a 12-h light cycle. Crosses and treatment schemes are illustrated in Figure 1. Dams were trio bred with stud males. Dams were treated 5 weeks before pregnancy, during pregnancy, and to postnatal day 5 (PND 5) with 1 of 4 purified pelleted diets purchased from Envigo (formerly Harlan/Teklad): (1)Methyl Sufficient (MS, TD.130225, ~0.6% methionine; ~2.5 g/kg choline; ~3 ppm folic acid), (2)MS + 1%SS (modified TD.130225 with addition of 1% succinylsulfathiozole), (3)Methyl Deficient (MD, TD.130224, ~0.4% methionine, no added choline, and ~0.2 ppm folic acid), and (4)MD + 1%SS (modified TD.130224 with addition of 1% succinylsulfathiozole).
Litter handling and sample collections
To reduce maternal stress and improve fecundity and pup survival, breeding cages, and litters were minimally disturbed until weaning. Therefore, we waited until PND5 to sex pups (using sexually dimorphic genitalia) and alter the diet. Since the goal of this study is to study developmentally programmed effects that persist after restoration of normal environmental conditions, at PND5 dams (and pups) were switched to standard chow diet (TD. 8604, ~0.4% methionine, ~2.5 g/kg choline, 3 ppm folate, Envigo, formerly Harlan/Teklad) (Figure 1). All offspring were weaned onto TD. 8604 and remained until the end of the study Litter size was measured at birth and weaning. Pups were genotyped at weaning using previously described assays, 63 and ear tagged for identification into adulthood. Adult G1 pups were euthanized at 8 weeks of age in accordance with current recommendations by the American Veterinary Medical Association (AVMA) guidelines. Fresh tissues and plasma were collected immediately after euthanasia and flash frozen in liquid nitrogen and stored at −80°C.
Body weight
Offspring bodyweight was measured by analytical balance at PND5, PND21, and PND56.
Plasma folate, SAM, and SAH measurements
Plasma were collected from wild-type and mutant dams by cardiac puncture immediately after CO2 exposure in heparin coated tubes. Plasma levels after dietary depletion, equivalent to maternal status at birth, was obtained from dams that did not produce a live litter and were on the diet for ~9 weeks (Figure 2A). Plasma levels after dietary repletion, equivalent to status after 16 days of methyl sufficient diet, was obtained from dams with litters that were euthanized at weaning (Figure 2A). Folate concentrations were measured by microbiological assay, which measures growth response of the bacteria Lactobacillus casei (ATCC 7469) to folic acid. 64 S-adenosylmethionine (SAM) and S-adenosylhomocysteine (SAH) were measured in 50 mg of liver first pulverized in liquid nitrogen with mortar and pestle and stored at −80°C. SAM and SAH concentrations were measured in liver homogenates using Ultra Performance Liquid Chromatography. All assays were performed at the UNC Nutrition Obesity Research Center Metabolic Molecular Phenotyping Core.
DNA extraction and methylation detection by pyrosequencing
Genomic DNA was isolated from frozen liver and sperm using an automated magnetic-bead based nucleic acid extraction system (Anaprep 12). DNA quality and concentration was assessed by NanoDrop 2000 spectrometer (Thermo Scientific, DE). Double stranded DNA was measured by Quant-it PicoGreen dsDNA assay (#P7589, Life technologies, NY). Genomic DNA was bisulfite-converted using the EZ DNA methylation Gold kit according to the manufacturer’s protocol (#D5006, Zymo research, CA). All pyrosequencing PCR and sequencing primers and sequenced regions were devoid of SNPs or deletions. Pyrosequencing was performed using the Pyromark Q96MD instrument according to the manufacturer’s instructions (Qiagen, MD). Assays for all loci have been previously described, including DMR1 at the insulin-like growth factor 2 gene (Igf2DMR1), 53 the H19/Igf2 ICR H19 CTCF site 1 (H19CTS1, previously H19ICR_Rp1), 65 and CTCF site 4 (H19CTS4, previously H19Cbs4) 66 and a DMR upstream of the H19 gene promoter (H19 promoter proximal H19PromProx). 53 The average methylation of all CpGs combined within the assayed region are presented as mean methylation (%). The number of CpGs assayed for each region was 3 (Igf2DMR1), 6 (H19CTS1), 5 (H19CTS4), and 3 (H19PromProx).
Statistical analysis
Statistical analyses were performed in JMP Pro software version 12.2.0 (SAS, NC). Where applicable, normality of data distribution was confirmed by Shapiro-Wilk goodness-of-fit test and variance tested by Bartlett’s test. Data that met assumptions of normality were analyzed by parametric tests (t-test, ANOVA, linear regression), whereas data that did not meet assumptions of normality were analyzed by nonparametric tests (Wilcoxon/Kruskal Wallis test) as indicated in each figure legend. For all comparisons, P-value <.05 are considered statistically significant. All error bars and “± values” represent standard error of the mean. Where figures indicate “Diet” and “Anti” we measured the (i) Methyl donor depletion effect (Diet) determined by comparing MS + MS1%SS versus MD + MD1%SS samples; and (ii) Antibiotic treatment effect determined by comparing MS + MD versus MS1%SS + MD1%SS samples.
Footnotes
Acknowledgements
We thank Dr. Marisa Bartolomei for the H19ICRΔ2,3 mouse lines and the C7 mice; Dr. Mihai Niculescu (UNC Nutrition Research Institute) for advice on the methyl donor deficiency mouse model; and Dr. Steve Orena at the Nutrition Obesity Research Center Metabolic Molecular Phenotyping Core for measuring folate, SAM, and SAH levels.
Funding:
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: NIEHS Transition to Independent Environmental Health Research Career Development Award [KES023849A] and NIEHS funded UNC Center for Environmental Health and Susceptibility Award [5P30ES010126] to F.Y.I.
Declaration of conflicting interests:
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
Study design was conceived and supervised by FI. Animal and molecular experiments and preliminary data analyses were performed by JO and AP (with sperm methylation data generated by ME). Final data analyses were performed by FI. Manuscript was drafted and revised by AP and FI.
