Abstract
In eukaryotic cells, lipid transfer can occur at membrane contact sites (MCS) to facilitate the exchange of various lipids between two adjacent cellular organelle membranes. Lipid transfer proteins (LTPs), including shuttle LTP or bridge-like LTP (BLTP), transport lipids at MCS and are critical for diverse cellular processes, including lipid metabolism, membrane trafficking, and cell signaling. BLTPs (BLTP1-5, including the ATG2 and VPS13 family proteins) contain lipid-accommodating hydrophobic repeating β-groove (RBG) domains that allow the bulk transfer of lipids through MCS. Compared with vesicular lipid transfer and shuttle LTP, BLTPs have been only recently identified. Their functions and regulatory mechanisms are currently being unraveled in various model organisms and by diverse approaches. In this review, we summarize the genetics, structural features, and biological functions of BLTP in the genetically tractable model organism C. elegans. We discuss our recent studies and findings on C. elegans LPD-3, a prototypical megaprotein ortholog of BLTP1, with identified lipid transfer functions that are evolutionarily conserved in multicellular organisms and in human cells. We also highlight areas for future research of BLTP using C. elegans and complementary model systems and approaches. Given the emerging links of BLTP to several human diseases, including Parkinson's disease and Alkuraya-Kučinskas syndrome, discovering evolutionarily conserved roles of BLTPs and their mechanisms of regulation and action should contribute to new advances in basic cell biology and potential therapeutic development for related human disorders.
Keywords
Introduction
Lipids are essential components of cell membranes and play fundamentally critical roles in a variety of cellular processes. The change in cell membrane properties and lipid composition allows cells to adapt to developmental, physiological, and environmental conditions, including ambient temperature fluctuations through an evolutionarily conserved mechanism termed homeoviscous adaptation (HVA) (Ernst et al., 2016). In both bacteria and metazoans such as the invertebrate model organism C. elegans, cell membrane fluidity or viscosity is maintained through the activation of temperature-triggered transcriptional regulation of genes encoding lipid desaturases (Ma et al., 2015; Ernst et al., 2016). Unsaturated lipids are biosynthesized by lipid desaturases and are transported to plasma membranes (PM) to maintain HVA. In eukaryotic cells, the endoplasmic reticulum (ER) is the major site of lipid biosynthesis and serves as a major hub for lipid metabolism including lipid desaturation by fatty acid desaturase enzymes within the cell. While canonical mechanisms for lipid transport include those mediated by lipid-containing vesicles and shuttle-type non-vesicular lipid transport proteins, recent studies have identified emerging roles of a novel family of bridge-like lipid transfer proteins (BLTPs) at MCS. These BLTPs appear to mediate fast, non-vesicular, bulk lipid transfer across organelle membranes, including ER and PM (Kumar et al., 2018; Osawa et al., 2019; Braschi et al., 2022; Levine, 2022; Neuman et al., 2022b; Wang et al., 2022). Despite these advances, mechanistic details (transfer rate, substrate specificity, and directionality) of the lipid transport at MCS in vivo still remain largely undefined.
BLTPs are structurally characterized by hydrophobic tunnels consisting of multiple repeating β-groove (RBG) domains (Braschi et al., 2022; Neuman et al., 2022). Based on structural homology, new nomenclature has recently been introduced by the HUGO Gene Nomenclature Committee for BLTP, including BLTP1 (also known as KIAA1109 in humans, Csf1 in yeast, lpd-3 in C. elegans, and tweek in Drosophila), and the well-characterized VPS13 and ATG2 protein families (Braschi et al., 2022). Current evidence supports the notion that BLTPs mediate the non-vesicular lipid trafficking among various intracellular organelles, including ER, lysosomes, mitochondria, and PM. Compared with vesicular or shuttle-type non-vesicular lipid transport mechanisms, BLTP may carry larger numbers of lipid molecules per protein and enable magnitudes of faster transport kinetics (Zhang et al., 2022). These unique features of BLTP may be particularly important for cells under conditions of high demands for lipid trafficking, such as during rapid cellular growth in development, organelle and subcellular compartment membrane expansion, autophagosome formation, and acute cellular responses to ambient thermal stress in HVA. However, our understanding of the regulatory pathways, physiological implications, and cellular functions of BLTPs is still far from complete.
In this article, we provide an overview of the current state of knowledge on BLTP in C. elegans, focusing particularly on genetic, structural, and functional aspects currently known from experimental studies and computational evidence (Figure 1A). We highlight how C. elegans studies may inform our broad understanding of BLTP in other organisms, and vice versa. We emphasize the need for further biochemical mechanistic studies to complement genetic and functional analyses in C. elegans, and present major unanswered outstanding questions in this burgeoning field.
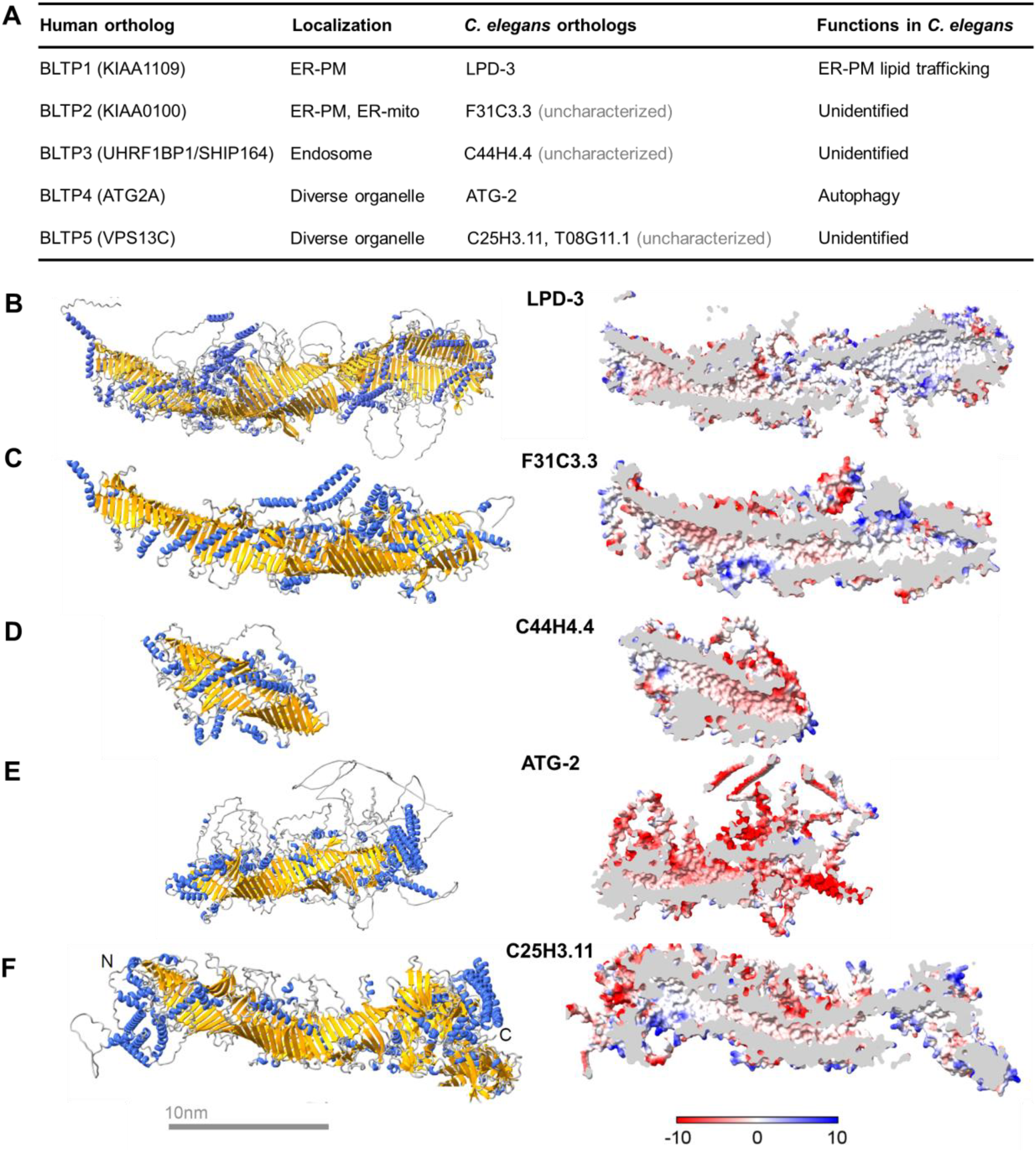
C. elegans BLTP. (A) Table listing human orthologs, currently known subcellular localizations, C. elegans orthologs, and currently known biological functions. (B–F) Predicted structures of BLTP in C. elegans, with ribbons (left) and cross-sectional (right) views. Predicted alpha helix (blue) and beta-sheet (yellow) segments in each ribbon structure, as well as an amphiphilic patch (hydrophobic, white; hydrophilic, blue and red) in cross-sectional views are also shown. The LPD-3 structure was predicted as in our previous studies (Wang et al., 2022). For C25H3.11, the full-length protein sequence was split into two segments: 1–2,000 and 1,000–3,302. Two segments were individually predicted by the AlphaFold v2.0 (https://cryonet.ai/af2/) program (Jumper et al., 2021). According to their overlapping sequence, predicted structures were aligned via Chimera (Pettersen et al., 2004), then combined in Coot (Emsley and Cowtan, 2004) for the full-length structure. The full-length structures for BLTP including ATG-2, F31C3.3, and C44H4.4 were directly obtained via prediction. The structural images were generated in ChimeraX (Pettersen et al., 2021).
LPD-3, the Prototypical BLTP, in C. elegans
C. elegans is a free-living invertebrate model organism that has been widely used for the discovery of genes and pathways underlying fundamental and evolutionarily conserved biological processes (Corsi et al., 2015). For many areas of biological and biomedical research, C. elegans can be advantageous as it is highly genetically tractable, has a short life cycle, is very easy to handle, and shares similarity and orthologous relationships with humans for a large majority of genes encoded in both C. elegans and human genomes. In our recent studies in C. elegans, we identified the gene lpd-3 in an unbiased mutagenesis screen as a novel regulator of HVA and discovered its highly conserved cellular functions in ER-to-PM phospholipid trafficking (Ma et al., 2015; Wang et al., 2022). Mutations in BLTP1, the human ortholog of lpd-3, can cause a rare autosomal recessive genetic disorder, Alkuraya-Kucinskas syndrome (AKS) (Bouzid et al., 2014; Gueneau et al., 2018; Kane et al., 2019; Kumar et al., 2020). We found that the C. elegans lpd-3 mutant, as an invertebrate animal model for AKS, can be pharmacologically rescued by Lecithin, a mixture of unsaturated phospholipids, suggesting a possible route for treating AKS in humans.
Encoding the ortholog of mammalian BLTP1, lpd-3 (lipid depletion 3) was originally named from a genome-wide RNAi screen for genes with a phenotype in lipid depletion, although the functional role of the gene was not identified (Ashrafi et al., 2003). In a forward genetic screen to discover novel regulators of the fatty acid desaturase FAT-7 and HVA (Ma et al., 2015; Wang et al. 2022), we isolated two mutants that failed to complement each other, based on the phenotypic suppression of strong FAT-7::GFP expression in acdh-11 mutants. Positional cloning and whole-genome sequencing led to the molecular identification of both mutations in C. elegans lpd-3 (Wang et al. 2022). The encoded LPD-3 protein is predicted as a megaprotein of 4,520 a.a. in length that structurally predicted to fold into a bridge-like tunnel configuration consisting of numerous RBG domains (Figure 1B). CRISPR-mediated GFP tagging of endogenous LPD-3 shows striking co-localization of LPD-3::GFP with ER-PM junction markers (Wang et al., 2022), consistent with an ER-targeting α-helix at its N-terminus and a membrane-anchoring amphipathic patch at its C-terminus. It is worth noting that endogenously tagged Csf1, the yeast ortholog of LPD-3, appears also localized to ER-PM junctions, although precise molecular functions of Csf1 remain unidentified (Tokai et al., 2000; John Peter et al., 2022; Toulmay et al., 2022). In assays based on genetically-encoded fluorescent protein sensors and phospholipid metabolic labeling, lpd-3 mutants exhibit profound defects in proper subcellular localization of phospholipids, including phosphatidylinositol species and phosphatidylcholine (Wang et al., 2022). Such assays further revealed similar defects of Zebrafish and human cells deficient in BLTP1 homologs, supporting highly evolutionarily conserved roles of LPD-3/BLTP1 in mediating ER-to-PM phospholipid trafficking. As lpd-3 mutants also show defects in glycosylphosphatidylinositol (GPI) anchors on the surface of embryonic cells (Toulmay et al., 2022), further studies may elucidate whether this reflects a role of LPD-3 in ER-to-PM lipid transfer or its distinct functions in a different biological context.
In addition to a molecular role in lipid transfer, thorough phenotypic analyses revealed that LPD-3 is essential for normal larval developmental progression, fertility, lipogenesis, cold stress resilience, and regulation of insulin-mTOR signaling during aging (Wang et al., 2022; Pandey et al., 2023). While several phenotypes including developmental delay and cold sensitivity are likely direct consequences of defective phospholipid trafficking, reduced phospholipids in PM can lead to a wide range of secondary consequences including sphingolipid dysregulation, insulin and mTOR hyperactivation, and shortened lifespan in C. elegans. Nonetheless, the highly penetrant lpd-3 mutant phenotypes may serve as a useful C. elegans model to help understand how BLTP1 mutations contribute to AKS etiology and to develop potential interventions for treating AKS. Indeed, we found that most lpd-3 mutant phenotypes can be rescued by the dietary supplementation of Lecithin, a naturally occurring mixture with highly enriched unsaturated phospholipids (Fiume, 2001; Scott, 2005). Lecithin is present in milk and represents a commonly used dietary supplement known for its excellent bioavailability and safety profiles in humans (Fiume, 2001; Scott, 2005). Despite this, its potential therapeutic effects in alleviating symptoms of AKS patients remain unknown, but pose a fascinating future research avenue. Conducting large animal and clinical trials could shed light on the efficacy of lecithin or similar compound supplementation with better-defined lipid compositions in managing AKS symptoms.
Other BLTP Family Members in C. elegans
Besides LPD-3 (ortholog of BLTP1), the C. elegans genome encodes multiple other BLTP members (Figure 1) that together with LPD-3 are respectively orthologous to BLTP1–5 in humans. While most BLTP-encoding genes remain still functionally uncharacterized, except lpd-3 and atg-2, large-scale RNAi and expression studies have implicated C. elegans BLTP genes in diverse cellular and developmental roles, including embryonic development, neuronal function, and fat storage regulation (Kamath and Ahringer, 2003). Recent single-nucleus RNA sequencing studies also show that all BLTP genes are widely expressed in different tissues and exhibit age-dependent changes (Gao et al., 2023).
F31C3.3 encodes the C. elegans ortholog of BLTP2. Like LPD-3, the predicted structure of F31C3.3 contains many RBG domains following an N-terminal transmembrane helix that may anchor it in the ER (Figure 1C). Inside of the protein is a hydrophobic tunnel, characteristic of BLTP, extending along the entire length of the protein. Sites of cell type and tissue expression for F31C3.3 include mainly the germline, sperms, body wall muscles, neurons, and coelomocytes (Gao et al., 2023). Interestingly, F31C3.3 is located on chromosome I within an operon together with another two genes, zipt-1 (zinc ion transporter) and cyn-5 (peptidyl-prolyl cis-trans isomerase). Although little is known about the biological function of F31C3.3 in C. elegans, studies of the Drosophila ortholog HOBBIT/BLTP-2 identified its roles in intracellular membrane trafficking to drive insulin-dependent growth during development (Neuman and Bashirullah, 2018). Hobbit can localize to the ER-PM junction in both yeast cells and Drosophila larval salivary glands, and its ER-PM localization is required for function in Drosophila (Neuman et al., 2022). It remains to be investigated whether F31C3.3 may have similar subcellular localization in C. elegans and how its localization may underlie any important roles in non-vesicular lipid transport, membrane trafficking, and organismal functions.
C44H4.4 encodes the C. elegans ortholog of BLTP3. Compared with LPD-3 and F31C3.3, C44H4.4 is predicted to form a much shorter tunnel, retaining the RBG domains and hydrophobic cavity extending from its N to C terminus (Figure 1D). C44H4.4 is mostly expressed in embryonic cells, coelomocytes, distal tip cells, excretory glands, and intestinal cells (Gao et al., 2023). C44H4.4 inhibition through RNAi or mutations has been reported to cause embryonic and L1 lethality, implicating its importance in embryonic and larval development (Meneely and Herman, 1979; Maeda et al., 2001; Tischler et al., 2006). Neither the subcellular localization nor the molecular function of C44H4.4 is known in C. elegans, while its mammalian counterpart BLTP3 (aka SHIP164 or UHRF1BP1L) exhibits an endocytic association that supports a role in endosome-Golgi membrane trafficking or sorting (Otto et al., 2010; Hanna et al., 2022).
ATG-2 is the C. elegans ortholog of BLTP4 and has been well-studied for its important roles in autophagy. Like its orthologs in yeast and mammalian cells, C. elegans ATG-2 is essential for the maturation of the autophagosome, the organelle responsible for the degradation and recycling of cellular materials in cellular autophagy. Specifically, mutations in atg-2 cause defects in autophagic degradation of PGL granules during embryonic development (Tian et al., 2010). Though not shown for ATG-2 in C. elegans, yeast Atg2 can use its hydrophobic cavity to accommodate phospholipid acyl chains and direct lipid transfer from ER membranes for autophagosome formation (Osawa et al., 2019), suggesting similar molecular functions of ATG-2 in C. elegans autophagy.
The C. elegans genome encodes two uncharacterized orthologs of VPS13 (aka BLTP5): T08G11 which is more closely related to VPS13A/C, and C25H3.11 which is more closely related to VPS13D (Velayos-Baeza et al., 2004; Levine, 2022). Humans have four homologs of the VPS13 gene: VPS13A, B, C, and D localized at different MCS within the cell, and specific mutations in these genes correspond to distinct neurological disorders: chorea-acanthocytosis (VPS13A) (Rampoldi et al., 2001), Cohen syndrome (VPS13B) (Kolehmainen et al., 2003), early onset Parkinson's disease (VPS13C) (Lesage et al., 2016) and ataxia (VPS13D) (Seong et al., 2018). Both T08G11.1 and C25H3.11 are large proteins of over 3,000 a.a. in length and differ in tissue expression patterns and chromosomal locations in the C. elegans genome (wormbase). While C25H3.11 is mainly expressed in neuron, germline, and head mesodermal cells, T08G11.1 is expressed mostly in the reproductive system, body musculature, and dorsal nerve cords. The C. elegans genome appears to lack a VPS13B orthologue likely due to the common evolutionary process of loss of genes (Krylov et al., 2003). Interestingly, as structurally predicted by AlphaFold2, the C-terminus of C25H3.11 is surrounded by eight beta barrels yet does not contain the UBA domain commonly found in human and fly VPS13D orthologs (Figure 1F) (Levine, 2022). RNAi against C25H3.11 can cause slow growth and sterile progeny, whereas its homozygous deletion causes strong induction of the mitochondrial unfolded protein response reporter hsp-6p::GFP in the C. elegans intestine (Maeda et al., 2001; Rolland and Conradt, 2022). By contrast, RNAi against T08G11.1 does not appear to cause any apparent phenotypes in C. elegans. As mammalian VPS13D and VPS13A can localize to mitochondria-endosomes and mitochondria-ER (Guillén-Samander et al., 2021; Tornero-Écija et al., 2023), it remains to be investigated whether C. elegans counterparts also localize to these organelles to mediate lipid transport.
Future Directions and Concluding Remarks
The newly discovered BLTPs are an exciting family of bulk lipid transporters. They have been studied with the powerful approaches of genetics, cell biology, biophysics, and biochemistry. While the ATG2/VPS13 subfamily protein is relatively better understood with experimentally determined structures and functions, the biochemical, biophysical, cell biological, and molecular genetic analyses of BLTP have lagged behind the structural prediction and bioinformatics-based exploration of other BLTP family members. Major unanswered questions include several categories:
What drives lipid transport along BLTP? It is currently unclear if this requires certain unidentified ATPases, differential membrane tension across organelle membranes, or other unconventional biophysical mechanisms. What are the specific types and movement directions of lipids transported by each family member of BLTP? Although AlphaFold2-based structural prediction from protein sequence is highly informative to infer the overall shape and topology of BLTP, in vitro protein-liposome reconstitution and determined BLTP structures from biological samples by cryo-EM/ET will be still highly valuable and may provide key clues or answers concerning this category of questions. What are (static or dynamic) subcellular localizations of BLTP in vivo? Very little is known about whether and how BLTP localization and activity are regulated under different developmental, physiological, and pathological conditions. How do mutations in BLTP cause diseases in animal models and humans? What therapeutic strategy may modify or alleviate BLTP-linked diseases?
As C. elegans is highly genetically tractable, the approach of genetic screens and its genotype-organismal phenotype analysis can be very powerful to infer the biological functions of BLTPs. For a molecular mechanistic understanding of BLTP, tools from biochemistry and molecular biophysics will be highly complementary and fruitful, particularly for the first two questions. With loss- and gain-of-function (LOF, GOF) as well as conditional alleles of BLTP becoming readily available, C. elegans can also help address the question (2) by enabling studies of LOF and GOF effects on intracellular lipid distribution and dynamics in live animals using genetically encoded lipid sensors (e.g., Lact-C2-GFP for phosphatidylserine and AKT-PH-GFP for PIP2/3). Given the translucent body and facile CRISPR-tagging of endogenous alleles with fluorescent proteins, C. elegans is also poised to yield insights into the question (3). In addition, many of the human disease-linked mutations in BLTP can be modeled in C. elegans to help understand disease etiology and develop genetic or pharmacological modifiers using tractable phenotypic readouts in C. elegans. A case in point is the rescue of lpd-3 mutant phenotypes in C. elegans by Lecithin (Wang et al., 2022), further studies of which may lead to defined lipid species and compositions that can alleviate certain symptoms of human AKS patients. Overall, the research reviewed here highlights the significance of BLTP in lipid metabolism and biology in C. elegans which should continue to provide a fruitful avenue for further exploration.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Institute of General Medical Sciences (grant number R35GM139618, D.K.M.), UCSF PBBR New Frontier Research (D.K.M) and BARI Investigator Award (D.K.M.).
