Abstract
Maintenance of the cardiolipin (CL) level largely depends on Ups1-Mdm35 complex-mediated intramitochondrial phosphatidic acid transfer. In addition, the presence of an alternative CL accumulation pathway has been suggested in the yeast Saccharomyces cerevisiae. This pathway is independent of the Ups1-Mdm35 complex and stimulated by loss of Ups2, which forms a complex with Mdm35 and mediates intramitochondrial transfer of phosphatidylserine for phosphatidylethanolamine synthesis. Recently, we found that the alternative CL accumulation pathway is enhanced by a lowered phosphatidylethanolamine level, not by loss of Ups2 per se, and depends on three mitochondrial inner membrane proteins, Fmp30, Mdm31, and Mdm32.
Keywords
Mitochondria play an important role in phospholipid biosynthesis. Among phospholipids, cardiolipin (CL) and the majority of phosphatidylethanolamine (PE) are synthesized in the mitochondrial inner membrane (MIM). The MIM of the yeast Saccharomyces cerevisiae accommodates the enzymes required for CL and PE biosynthesis. The CL synthetic pathway from phosphatidic acid (PA) consists of four reactions as shown in Figure 1. On the other hand, PE is synthesized in the mitochondria through the decarboxylation of phosphatidylserine (PS), which is catalyzed by a PS decarboxylase, Psd1. Because mitochondria are incapable of synthesizing PA and PS, these phospholipids are transported to the MIM from other organelles such as the endoplasmic reticulum (ER) for the synthesis of CL and PE, respectively.
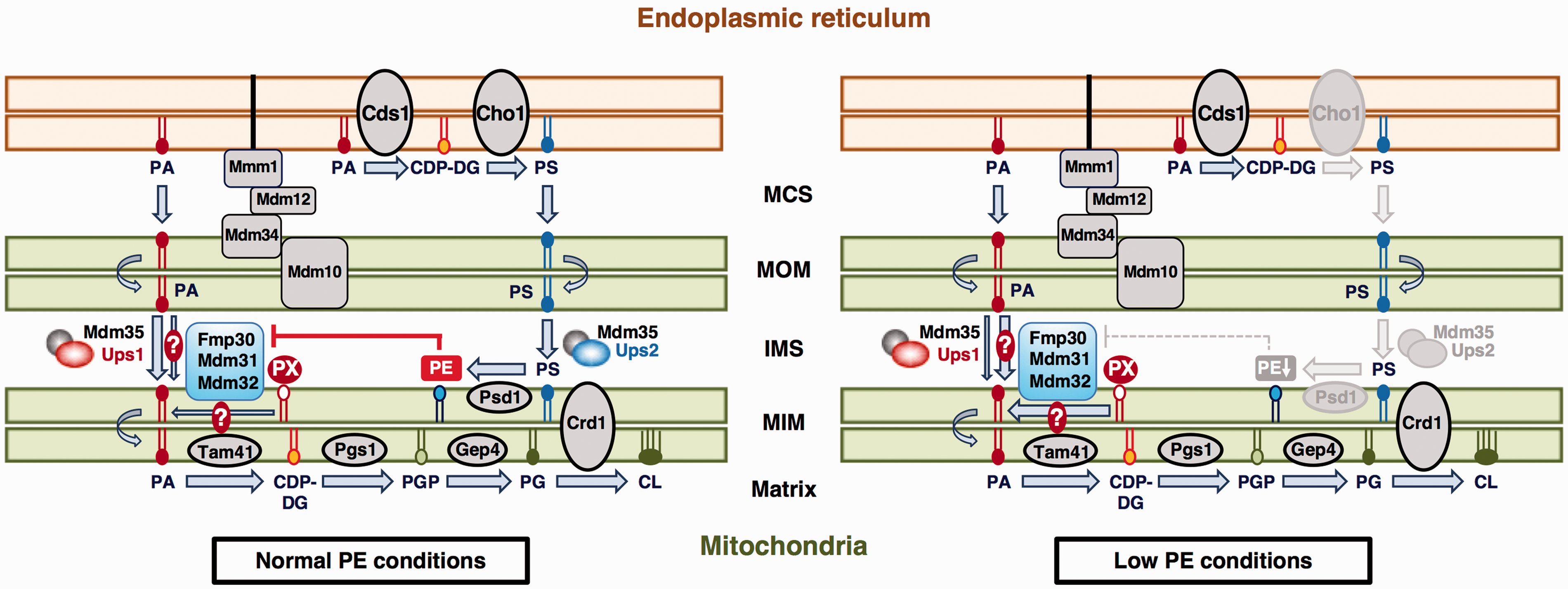
Working hypothesis for the Ups1-independent and low-level PE-enhanced CL accumulation pathway (FMM-dependent pathway). Under normal PE conditions, transfer of PA between the MOM and MIM is mainly mediated by the Ups1-Mdm35 complex. PE somehow inhibits the Fmp30-, Mdm31-, and Mdm32-dependent CL accumulation pathway (FMM-dependent pathway). Under the low PE conditions caused by loss of Cho1, Psd1, or Ups2, CL accumulation via the FMM-dependent pathway is enhanced, together with a decreased contribution of Ups1-Mdm35 complex-mediated PA transfer to CL accumulation. The FMM-dependent pathway might provide PA to the MIM for CL synthesis by mediating intramitochondrial PA transfer or by enzymatically generating PA from an unidentified phospholipid (designated as PX). PA = phosphatidic acid; CDP-DG = CDP-diacylglycerol; PGP = phosphatidylglycerolphosphate; PG = phosphatidylglycerol; CL = cardiolipin; ER = endoplasmic reticulum; PS = phosphatidylserine; PE = phosphatidylethanolamine; MCS = membrane contact site; MOM = mitochondrial outer membrane; IMS = intermembrane space; MIM = mitochondrial inner membrane.
Phospholipid transfer between the ER and the mitochondrial outer membrane (MOM) is thought to occur at the membrane contact site (MCS) formed by membrane tethering protein complexes. The ER-mitochondria encounter structure (ERMES) is the most characterized MCS between the ER and mitochondria in yeast (Kornmann et al., 2009). The ERMES complex comprises Mmm1, Mdm12, Mdm34, and Mdm10. Mmm1, Mdm12, and Mdm34 include the synaptotagmin-like mitochondrial lipid-binding protein domain. Very recently, in vitro analysis revealed that the Mmm1-Mdm12 complex transfers phospholipids between liposomes, suggesting the direct involvement of the ERMES in phospholipid transport between the ER and mitochondria (Kawano et al., 2017).
Transfer of PA and PS between the MOM and MIM can be mediated by the Ups1-Mdm35 and Ups2-Mdm35 complexes, respectively (Aaltonen et al., 2016; Connerth et al., 2012; Miyata, Watanabe, Tamura, Endo, & Kuge, 2016). The intermembrane space (IMS) proteins Ups1 and Ups2 require the common cofactor Mdm35 for their stability and lipid transfer activity. Loss of Ups1 leads to a defect in CL synthesis. On the other hand, loss of Ups2 leads to a defect in mitochondrial PE synthesis. In ups1Δ yeast, CL level is decreased to ∼20% of that in wild-type yeast. However, concomitant deletion of UPS2 restores the CL level in ups1Δ cells, suggesting the presence of a Ups1-independent CL synthetic pathway that is enhanced by loss of Ups2 (Tamura, Endo, Iijima, & Sesaki, 2009).
To elucidate the Ups1-independent pathway for CL synthesis, we recently looked for the factors that are involved in this pathway (Miyata, Goda, Matsuo, Hoketsu, & Kuge, 2017). Consequently, we found that loss of CHO1 and PSD1 encoding PS synthase and decarboxylase, respectively, as well as UPS2 restored the CL level in ups1Δ cells and that forced production of PE through overexpression of CHO1 and PSD1 decreased the CL level in ups1Δups2Δ cells. These findings suggest that low PE levels are factors responsible for the restoration of the CL level in the absence of Ups1. Furthermore, we identified three proteins located in the MIM, Fmp30, Mdm31, and Mdm32, as essential factors for the low-level PE-enhanced CL accumulation in the absence of Ups1. Depletion of Fmp30, Mdm31, or Mdm32 almost completely abolishes CL accumulation in ups1Δups2Δ cells. Importantly, even in UPS1 normal cells, accumulation of CL becomes more dependent on Fmp30, Mdm31, and Mdm32 than on Ups1 under lowered PE conditions caused by loss of Ups2, Psd1, or Cho1. In addition, Mdm31 and Mdm32 have been shown to interact physically with Fmp30, suggesting that these three proteins cooperatively function in maintenance of the CL level. Thus, the PE level is the critical determinant of whether Fmp30, Mdm31, and Mdm32 elicit their functions in maintenance of the CL level (Figure 1).
How a reduction of the PE level stimulates the activities of Fmp30, Mdm31, and Mdm32 remains to be determined. It has been reported that loss of UPS2 or PSD1 partially restores respiration growth and the mitochondrial ultrastructure in the cells defective in the mitochondrial contact site and cristae organizing system (MICOS) complex (Aaltonen et al., 2016). Similarly, loss of UPS2 also improves CL levels and growth in cells defective in the ERMES complex (Tamura et al., 2012). These observations imply that a reduction of PE affect the ultrastructure and functions of the MCSs within mitochondria, and possibly between the ER and mitochondria. Therefore, it is plausible that the activities of Fmp30, Mdm31, and Mdm32 are intimately linked with the mitochondrial ultrastructure such as the MCS influenced by the decrease in PE levels. In this regard, it is noteworthy that Fmp30, Mdm31, and Mdm32 exhibit strong genetic interactions with components of the ERMES and MICOS complexes (Hoppins et al., 2011; Kuroda et al., 2011).
Fmp30 exhibits homology with mammalian N-acylphosphatidylethanolamine (NAPE)-specific phospholipase D (NAPE-PLD), exposing its putative catalytic domain to the IMS. Importantly, Fmp30 harboring mutations of the amino acid residues corresponding to the catalytic center of NAPE-PLD are incapable of rescuing the growth of fmp30Δpsd1Δ cells, suggesting that the enzyme activity of Fmp30 is prerequisite for its function under low PE conditions (Kuroda et al., 2011). Mammalian NAPE-PLDs hydrolyze NAPE to produce N-acylethanolamine and PA. Accordingly, Fmp30 could potentially generate PA in the MIM, thereby bypassing Ups1-dependent intramitochondrial PA transfer. Alternatively, the phospholipid-hydrolyzing activity of Fmp30 might regulate the local phospholipid composition, which is relevant to the membrane dynamics, such as the formation of the MCS, facilitating the intramitochondrial PA transfer. Further studies including identification of the intrinsic substrate of Fmp30 and detailed determination of the mitochondrial lipidome in fmp30Δ cells are needed to address these possibilities.
Mdm31 and Mdm32 show homology to each other. Mdm31 and Mdm32 harbor two membrane-spanning regions at the both sides of mature proteins, exposing a middle region to the IMS. Mdm31 and Mdm32 are reported to be involved in maintenance of the mitochondrial morphology and mitochondrial DNA in addition to CL metabolism, suggesting that these proteins are multifunctional ones. It remains unknown how Mdm31 and Mdm32 function in CL metabolism cooperatively with Fmp30. Given that Mdm31 and Mdm32 physically interact with Fmp30, these proteins are possibly required for the proper function of Fmp30.
In conclusion, we identified a novel CL accumulation pathway, which is independent of the PA transfer mediated by Ups1-Mdm35 complex but dependent on Fmp30, Mdm31, and Mdm32. We termed this route the FMM (Fmp30, Mdm31, and Mdm32)-dependent pathway. The FMM-dependent pathway is crucial for maintenance of the CL level under low PE conditions in yeast. The Ups1-dependent and FMM-dependent pathways would reciprocally maintain the CL level in response to various cellular conditions.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported in part by MEXT/JSPS KAKENHI 16K07354 and 17K15120.
