Abstract
Meta-analysis is one of the most useful research approaches, the relevance of which relies on its credibility. Reproducibility of scientific results could be considered as the minimal threshold of this credibility. We assessed the reproducibility of a sample of meta-analyses published between 2000 and 2020. From a random sample of 100 articles reporting results of meta-analyses of interventions in clinical psychology, 217 meta-analyses were selected. We first tried to retrieve the original data by recovering a data file, recoding the data from document files, or requesting it from original authors. Second, through a multistage workflow, we tried to reproduce the main results of each meta-analysis. The original data were retrieved for 67% (146/217) of meta-analyses. Although this rate showed an improvement over the years, in only 5% of these cases was it possible to retrieve a data file ready for reuse. Of these 146, 52 showed a discrepancy larger than 5% in the main results in the first stage. For 10 meta-analyses, this discrepancy was solved after fixing a coding error of our data-retrieval process, and for 15 of them, it was considered approximately reproduced in a qualitative assessment. In the remaining meta-analyses (18%, 27/146), different issues were identified in an in-depth review, such as reporting inconsistencies, lack of data, or transcription errors. Nevertheless, the numerical discrepancies were mostly minor and had little or no impact on the conclusions. Overall, one of the biggest threats to the reproducibility of meta-analysis is related to data availability and current data-sharing practices in meta-analysis.
Meta-analysis is widely considered an important approach to evaluate a body of work. Given the ongoing growth in the number of scientific publications (Bornmann et al., 2021), evidence-synthesis approaches—such as meta-analysis—are becoming increasingly relevant for a cumulative science. This relevance rests on the credibility of meta-analytic results, which can be threatened by a lack of rigorous methodology or poor-quality reporting (Gurevitch et al., 2018). Given the importance of meta-analyses for evidence-based practice, these threats to their credibility need to be closely monitored.
In recent years, different concerns on the credibility of empirical claims have emerged. Several projects have systematically attempted to assess the replicability and reproducibility of published scientific results (e.g., Artner et al., 2021; Errington et al., 2021; Open Science Collaboration, 2015). Those initiatives showed many failures to replicate or reproduce the published results. In this context, the empirical assessment of the credibility of published results has become a major task for the scientific community.
There are different approaches to the empirical assessment of scientific credibility. “Reproducibility” refers to the attempt to obtain the same results as in the original publication using the same data and the same procedure. “Robustness” refers to the assessment of the sensitivity of the originally published results and conclusions to variations in the original procedure using the same data. “Replicability” is a core principle of the scientific method and refers to the fact that the same scientific evidence should be observed when independent researchers try to answer the same research question from the same approach at different moments using different data; in other words, obtaining the same results, using different data, and answering the same question (National Academies of Sciences, Engineering, and Medicine, 2019; Nosek et al., 2022). In this project, we focus on the reproducibility of meta-analyses.
The reproducibility of published scientific results could be considered the minimal threshold of scientific credibility (Hardwicke et al., 2021). Different approaches can be adopted for the empirical assessment of reproducibility. For example Nosek et al. (2022) made the distinction between process reproducibility and outcome reproducibility. Following this framework, a process-reproducibility assessment could be carried out by reviewing the availability of the materials, data, or precise details of the analytical strategy in the report that are required to proceed with the reproduction attempt. An outcome-reproducibility assessment can be carried out when the required elements are retrievable by actually reproducing the analyses. Note that the difficulty of performing an outcome-reproducibility assessment depends on which analytical information is available. The availability of the original analysis code (i.e., the original computational instructions in a programming language) facilitates reproducibility analysis by enabling simply rerunning the code on the data. Regrettably, the analysis code is currently seldom available (Hardwicke et al., 2020, 2022; López-Nicolás et al., 2022). When only a verbal summary of the performed analyses is available in the research report (which is the most common scenario in practice), the original analysis needs to be reconstructed. The challenges and implications of failed reproductions in both cases may be of a different nature.
Several reproducibility analyses of meta-analyses have been performed in recent years. For example, some process-reproducibility assessments have shown an important lack of data availability in machine-readable formats and an almost complete absence of analysis-script-code availability (López-Nicolás et al., 2022; Polanin et al., 2020). Furthermore, some outcome-reproducibility assessments have shown a considerable number of failures when trying to reproduce the primary effect sizes of some published meta-analyses by recollecting primary data from primary studies (Gøtzsche et al., 2007; Maassen et al., 2020; Tendal et al., 2009), possibly because of lack of details on how primary effect sizes were selected and computed. In these outcome-reproducibility studies, the main task entails reconstructing the original data by retrieving them from the source, that is, the included primary studies. Thus, their assessment focus is on this stage of the analysis pipeline of a meta-analysis, which usually involves decisions on how to select the primary outcomes and how to deal with possible dependency, and the computation of (standardized) effect sizes. Figure 1 displays a summary of the basic meta-analysis pipeline in a flowchart that includes the different stages and previous work that has explored different facets of reproducibility and a summary of the required elements to be able to reproduce each stage. In this project, we focus on the last stage, related to the statistical analysis and quantitative results of the synthesis.
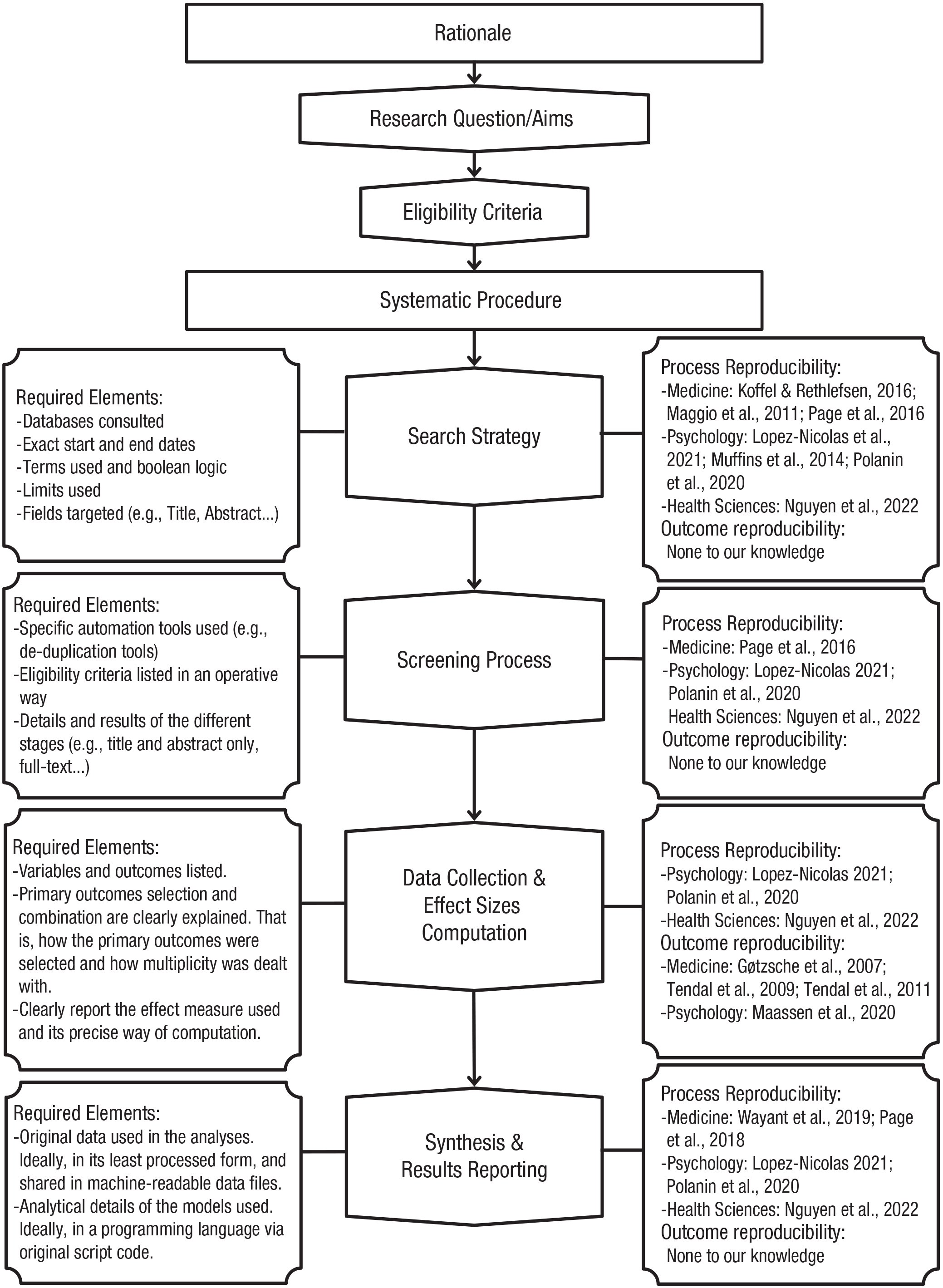
Flowchart displaying the basic pipeline of a meta-analysis. Each of the stages may be subject to reproducibility evaluation. (Left) The various elements that must be available to reproduce each stage are enumerated. (Right) Known studies that have evaluated some facet of the reproducibility of each stage are listed.
Reproducibility analysis of reported quantitative results typically uses the original data available from the original authors (e.g., Artner et al., 2021; Hardwicke et al., 2018, 2021). This puts the focus of the assessment on factors such as the reusability of the available data, challenges for the reconstruction of the original analysis scheme, and reporting errors. Although data availability seems to have improved in the last years (Hardwicke et al., 2018; Tedersoo et al., 2021; Wallach et al., 2018), systematic reviews and meta-analyses appear to be a special case. Typically, the data collected for a meta-analysis is study-level summary data extracted from published primary studies and commonly reported in the article through tables or forest plots. This may lead to the idea that common data-sharing practices do not apply to meta-analysis. For example, Page et al. (2022) analyzed the content of data-availability statements from a set of meta-analyses published in 2020. Only 31% included a data-availability statement, and only 13% of these included a link to access the data openly; 23% stated that all relevant data were available in the article itself, 10% stated that data sharing was not applicable because no data sets were generated, 8% stated that data sharing was not applicable because the data were drawn from already published literature, and 42% stated that data were available on request. It is surprising that even just considering meta-analyses that included a data-availability statement, the authors of these meta-analysis assume that such practices do not apply to meta-analyses or that the data in the article itself are sufficient.
Purpose
Previous research has revealed that there is room for improvement at different stages of the meta-analytic-process pipeline. In this study, our purpose is twofold. First, we broadened previous process-reproducibility assessments by considering data availability on request and contacting original authors to request required information to reproduce the meta-analysis. Second, we verified the outcome reproducibility of the meta-analyses that were process reproducible using the available data. For cases in which previous work focused on the reproducibility of primary effect sizes by recoding data from primary studies, we explored meta-analysis outcome reproducibility using the primary data already coded by the original authors. Therefore, we attempted to retrieve the data shared by the authors of the meta-analysis.
Disclosures
Preregistration
The pre-data-analysis protocol (https://doi.org/10.17605/OSF.IO/79J2T) was preregistered on October 19, 2021. Any deviation from this protocol is explicitly acknowledged.
Data, materials, and online resources
Data and analysis script code are openly available at https://doi.org/10.17605/OSF.IO/6CMZH.
Reporting
Below, we report how we determined our sample size, all data exclusions, all manipulations, and all measures in the study.
Method
Identification and selection of articles and meta-analyses
In previous research, we identified a pool of 664 meta-analytic reports on clinical-psychological interventions published between 2000 and 2020 through a systematic electronic search (López-Nicolás et al., 2022). Of this pool, 100 were randomly selected using a random-number generator between 1 and the total number of meta-analyses identified. The full search strategies and a summary of the screening process are available at https://osf.io/z5vrn/, and the workflow of the random-selection process is available at https://osf.io/cp293/. This sample size was based on our judgment of an acceptable trade-off between informativeness and feasibility. From these 100 articles, each independent pairwise meta-analytic model of aggregate data fitted on at least 10 primary studies was selected. In case no meta-analysis reported in an article had at least 10 studies, the meta-analysis with the highest number of primary studies was selected, which was the case for 29 of the articles included in this report. This criterion was established to focus on the main meta-analyses of each article based on the assumption that the search strategies would be designed to maximize the number of primary studies included that were related to the main aims of the article.
Our unit of analysis was each independent meta-analysis selected under these criteria. A total of 217 independent meta-analyses were selected.
Retrieval of primary data
To be able to reproduce meta-analyses of aggregate data, primary-level 1 effect sizes and their associated standard errors are required. These are generally computed from statistics retrieved from the primary studies, such as means, standard deviations, or sample sizes. We attempted to retrieve the least processed data shared by the authors of the meta-analysis. First, we sought for the statistics used to compute primary effect sizes (e.g., means, standard deviations). Second, we sought for the primary effect sizes already computed and their standard errors (or, alternatively, the sampling variances). Finally, we sought for the primary effect sizes and their confidence limits, from which the standard errors were approximated as follows:
with
On the other hand, efforts were also made to retrieve the most reusable data possible. First, we searched for machine-readable data files through links leading to third-party repositories or in supplementary material hosted by the journal. Second, we looked for available data through tables or forest plots in the meta-analytic report itself or in supplementary material. In these cases, the primary data had to be manually recoded to reuse it. Finally, if the primary data of a meta-analysis were not directly available after the previous steps, we attempted to obtain the data through a request to the corresponding author identified in the associated article. We sent an initial request in June 2021 and a subsequent reminder in October 2021 if there was no reply. This reminder was sent to a more recent alternative email address if we were able to find one. If we were unable to obtain the data through the email request, the associated meta-analysis was labeled as not process reproducible.
Reconstructing the original analytical scheme
To proceed with reproducibility attempts of the meta-analyses that were labeled as process reproducible, we first looked for the availability of the original analysis script. When it was available, reproducibility was checked by rerunning the original script on the associated primary data. When it was not available, we tried to reconstruct the original analytical scheme using the technical details reported in the article. Specifically, we collected information on (a) the meta-analytic model originally assumed, (b) the weighting scheme, (c) the between-studies variance estimator, (d) the method used to compute the confidence interval, and (e) the software used to perform the meta-analysis. If any of these details about the analytical methods were not reported but the software used was mentioned, we inferred the first four pieces of information from the default settings of the software used. If the software used was not reported, we inferred this information from the default settings of the most used software in the sample, which was Comprehensive Meta-Analysis. We designed this procedure to reconstruct the original analytical scheme when the original analysis script was not available instead of trying to request it from the original authors because of the following: (a) Not necessarily all authors of included meta-analyses will actually have an analysis script to share because many might have used point-and-click software, and (b) we expected analysis-script availability to be very low, and requesting it would have meant sending requests for virtually every article included in our reanalysis.
Additional information about the meta-analysis was collected that is not reported in this article. The full list of variables collected is available in the Protocol (https://osf.io/42r3p), and a Codebook describing these variables is available at https://osf.io/vrty7.
Data-collection procedure
Data-collection procedure was carried out by five of the authors. At a first pilot stage, a random sample of five articles of the total pool was independently coded by the five members, and subsequently, in a series of meetings, disagreements between the coders were resolved by consensus. Next, the initial pool of 100 included articles was split among four coders, 25 articles each. A random sample of 25 articles of the total pool was assigned to the fifth member to carry out independent double-coding to examine the reliability of the data-collection process. Disagreements were resolved by consensus and by double-checking the original materials. Details about intercoder agreement are reported in the supplementary file (https://osf.io/fjhpw).
Reproducibility outcomes
Each meta-analysis was labeled using the following two-level
2
reproducibility-success scheme. First, each meta-analysis was labeled as (a) process reproducible and (b) not process reproducible. In our study, “not process reproducible” refers to situations in which we were unable to access the primary data through neither direct extraction nor on request.
3
Second, those labeled as process reproducible were labeled as (a) reproducible, (b) numerical error, and (c) decision error. Similar to previous studies (Artner et al., 2021; Hardwicke et al., 2018, 2021), an index of numerical error was computed (see Protocol, https://osf.io/42r3p). This index expressed the difference between reproduced and original values as a percentage. To avoid labeling minor numerical discrepancies related to numerical rounding as reproducibility problems, a 5% discrepancy threshold was set. Thus, a meta-analysis was labeled as numerical error if it showed a discrepancy larger than 5%.
4
Finally, the label decision error refers to situations in which the
We focus on reproducibility of summary effects, their confidence bounds, and the result of the null hypothesis significance test. Secondarily, we also assessed reproducibility of other synthesis methods such as heterogeneity statistics.
Reproducibility-checks workflow
Reproducibility checks were carried out at different stages. First was through reported analytic details or script code. When the analysis script code was available, computational reproducibility was checked by rerunning the script with the available primary data. In most cases, the analysis script code was not available. Thus, in these cases, we coded the analytic details as explained above to fit equivalent meta-analytic models as a function of these details using the available primary data. This analysis scheme was programmed in the R environment (R Core Team, 2022) using the metafor package (Viechtbauer, 2010).
Second, given that the manual-recoding process is an error-prone task, some mistakes can appear. Thus, those meta-analyses labeled as numerical error and/or decision error in the previous stage were reassessed by a different member of the team. In cases in which an error was found in the originally coded results, analytic methods, and/or primary data, the meta-analyses were reproduced once again and relabeled according to the updated results. In addition, a qualitative assessment of the meta-analyses still labeled as numerical error and/or decision error was also carried out. The same reviewers who checked for errors produced individual reports on the possible source of the discrepancy, and its reproducibility was judged qualitatively by four of the other authors. This stage was a deviation from the preregistered protocol and made it possible to identify situations with obvious explanations, such as rounding issues or inverted signs.
In addition, for meta-analyses that remained labeled as nonreproducible, an email was sent to the corresponding author of the associated article explaining our aims, our approach, and our results regarding the author’s meta-analysis and requesting additional information that could explain the mismatch between the original reported results and the reproduced results. We tried to solve the reproducibility issues within a month after the request, and we updated the label accordingly.
Finally, the association between publication year and the possibility of retrieving the data in one of the ways conducted in this project were explored by fitting binary logistic regression models with publication year as predictor and process reproducibility as dependent variable. We quantified the strength of the association by calculating odds ratios and 95% confidence intervals based on the profile likelihood. These exploratory analyses were not preregistered. Details and results are reported in the supplementary file (https://osf.io/fjhpw).
Results
From the 100 included articles, 217 independent meta-analyses were selected following the criteria explained above. These meta-analyses included 18.35 primary studies on average (SD = 17.25; Mdn = 13; interquartile range = 10–19; range = 3–134) and were cited 108.39 times on average (SD = 151.00; Mdn = 57; interquartile range = 29–128; range = 3–1,036). 5 Figure 2 displays the distribution of number of primary studies among the meta-analyses included in our sample, the publication-year distribution among the articles included in our sample, and the citation-count distribution of those articles. Original results and characteristics of these meta-analyses are available at https://osf.io/8jzbk
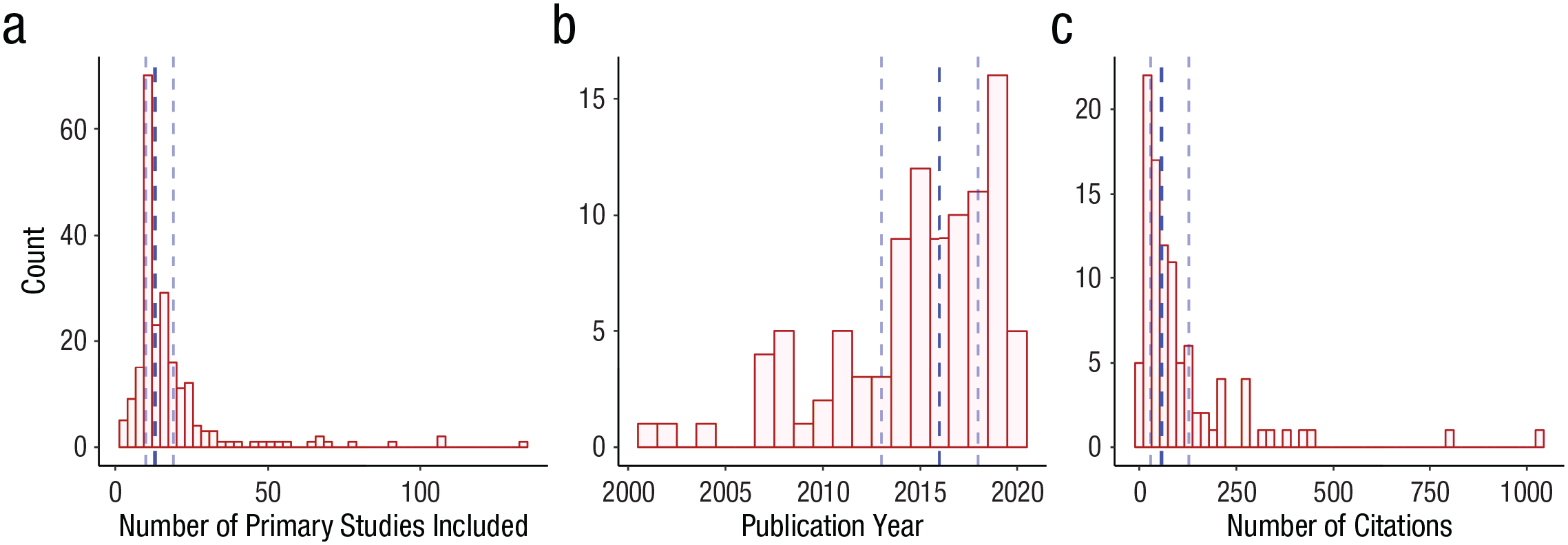
Distribution of (a) the number of primary studies included in each of the meta-analyses, (b) the publication year of the included articles, and (c) the citation count of the included articles. Vertical blue dotted lines represent the first quartile, median, and third quartile, respectively.
Process reproducibility
Figure 3 summarizes the primary data-retrieval results. On the basis of the availability of primary data, retrieved either directly from the article or on request, we labeled 146 meta-analyses (67%; see Fig. 3a) process reproducible. In addition, because the time span covered is fairly wide, the process-reproducible rate was also computed for different time periods. The meta-analyses were grouped into 5-year periods, except for the initial period, which was grouped into a 10-year period because of the limited number of meta-analyses available during the first 5-year period, which consisted of only five meta-analyses. The process-reproducibility rate was 41% (12/29), 59% (44/75), and 80% (90/113) for meta-analyses published between 2000 and 2010, 2011 and 2015, and 2016 and 2020, respectively (see Fig. 3a). This trend is further explored in the supplementary file available at https://osf.io/fjhpw.
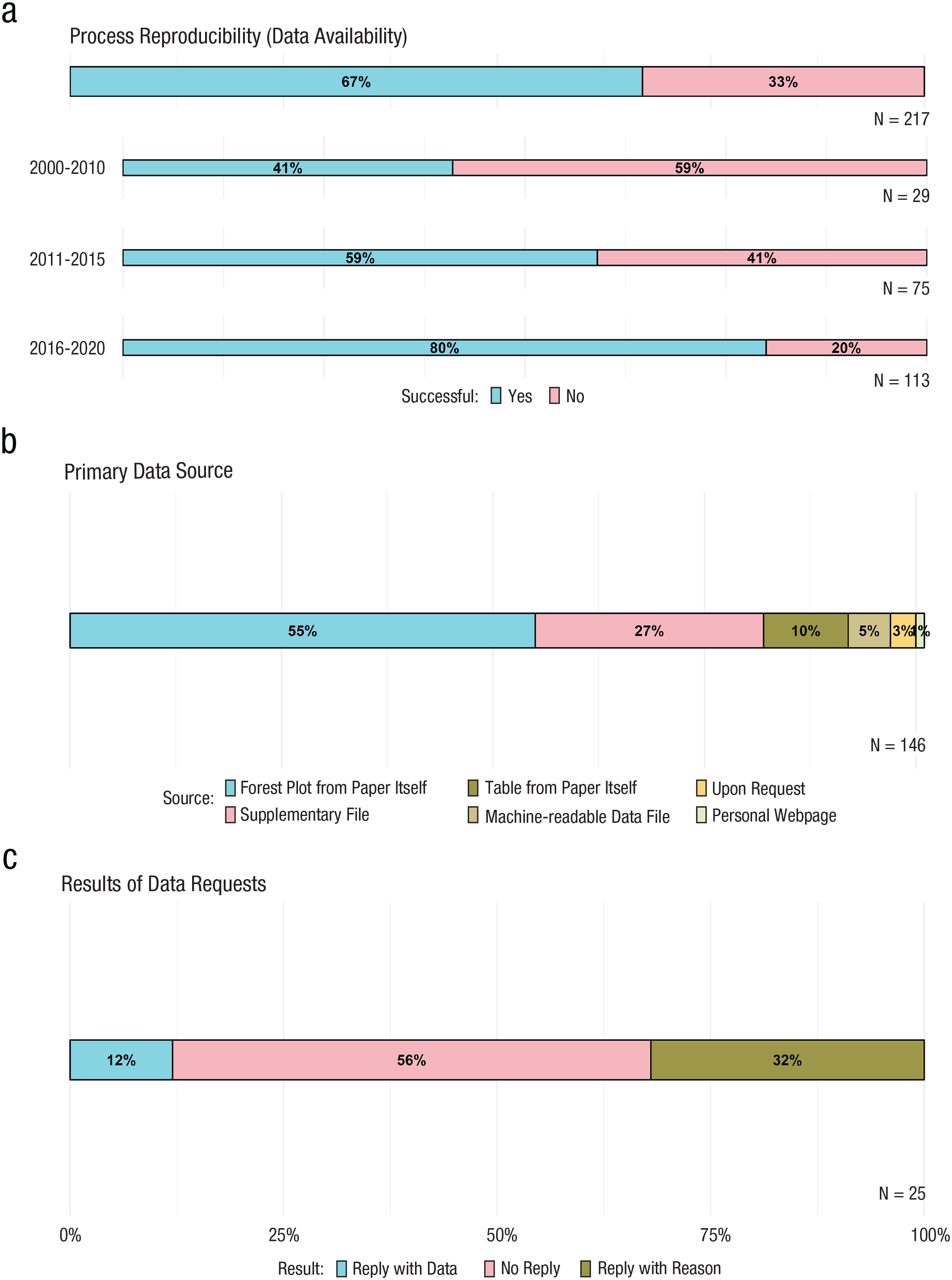
Percentage of (a) process-reproducible meta-analyses, (b) different types of sources of original data, and (c) data-request results.
Of these 146 meta-analyses, in about half of the cases, the primary data were retrieved from a forest plot in the article itself, and in about a third of the cases, the primary data were retrieved from supplementary files (for further details, see Fig. 3b). Although attempts were made to retrieve data for 78 meta-analyses from 25 different articles by emailing the corresponding authors, data were retrieved for only seven meta-analyses from three different articles (12%, 3/25; see Fig. 3c). For the remaining 71, for 22 different articles, a reply providing some reasons not to share was received in 32% (8/25, see Fig. 3c) of cases, whereas no reply was received for the remainder of the meta-analyses. Table 1 summarizes the different reasons corresponding authors gave when data were not provided on request.
Reasons Given When Data Were Not Received on Request

Challenges faced retrieving primary data
In most cases, when the meta-analytic data were available, it was shared in document formats. Data were retrieved from tables or forest plots in pdf or docx format—either in the document itself or in the supplementary materials—in 92% (134/146) of the cases. This required a manual recoding of the primary data to be able to reuse them. Furthermore, when data were reported through general tables (i.e., tables listing all the primary studies included with their characteristics), the meta-analysis associated with each data entry was not always obvious, leading to the time-consuming task of matching each data entry with each independent meta-analytic result reported in the article. There were only seven meta-analyses (from three different articles) of the 146 meta-analyses labeled as process reproducible (5%) in which the task of retrieving the data required simply downloading the data in a machine-readable data file format. On the other hand, as shown in Figure 3c, when the necessary data were not available, retrieving it on request to the original authors led to a low response rate.
Outcome reproducibility
The outcome reproducibility was checked in 146 meta-analyses from 82 different articles. As mentioned above, in five of these meta-analyses (3%), all from the same published article, the original script code was available. Therefore, in these five cases, outcome reproducibility was checked running the original analysis script on the original primary data. In the remaining cases, the original analytical framework was reconstructed as explained in the method section. Figure 4 summarizes the results of the whole process of outcome-reproducibility assessment.

Results of the different stages carried out in the evaluation of the outcome reproducibility.
Following the first stage of reanalysis, 52 meta-analyses were reassessed because they were labeled as numerical error and/or decision error. Of these, 17 were reanalyzed again because some coding errors were found in the second stage. After this, 10 were relabeled as reproduced, and seven still had relevant discrepancies. Furthermore, 15 were labeled as approximately reproduced or reproduced with minor adjustment in a qualitative check because the discrepancy was probably explained by rounding issues, inverted signs for results (when effect sizes were reported in absolute values) and primary data, minor reporting errors, or minor adjustments in the analytical scheme. 6 In the remaining 20 and in the seven reanalyzed again without success, some issues or relevant discrepancies without apparent explanation were found. Figure 5 displays in a scatterplot the consistency between the original and reproduced summary effect size and their confidence bounds of these 52 meta-analyses. In addition, as a secondary analysis, the reproducibility of the I2 heterogeneity statistic was explored. Figure 6 displays in a scatterplot the consistency between the original and reproduced I2 statistics. As shown in Figures 5 and 6, the discrepancies found in the heterogeneity statistic I2 are larger than those found in the summary effects and their confidence intervals. The Pearson’s correlation between the summary effect and I2 discrepancies was .172. The lack of precision of the available data (rounded data) or incomplete information on aspects such as the tau-squared estimator applied seem to have a substantial impact on the reproducibility of this result.

Scatterplot of the reproduced values as a function of the original values classified by whether decision error was found. Only the results of the 52 meta-analyses with a discrepancy of more than 5% identified in the first stage are displayed, but with the corrections made in the second stage. (a) Summary effects. (b) Confidence intervals (colors represent lower or upper bound of the confidence interval).
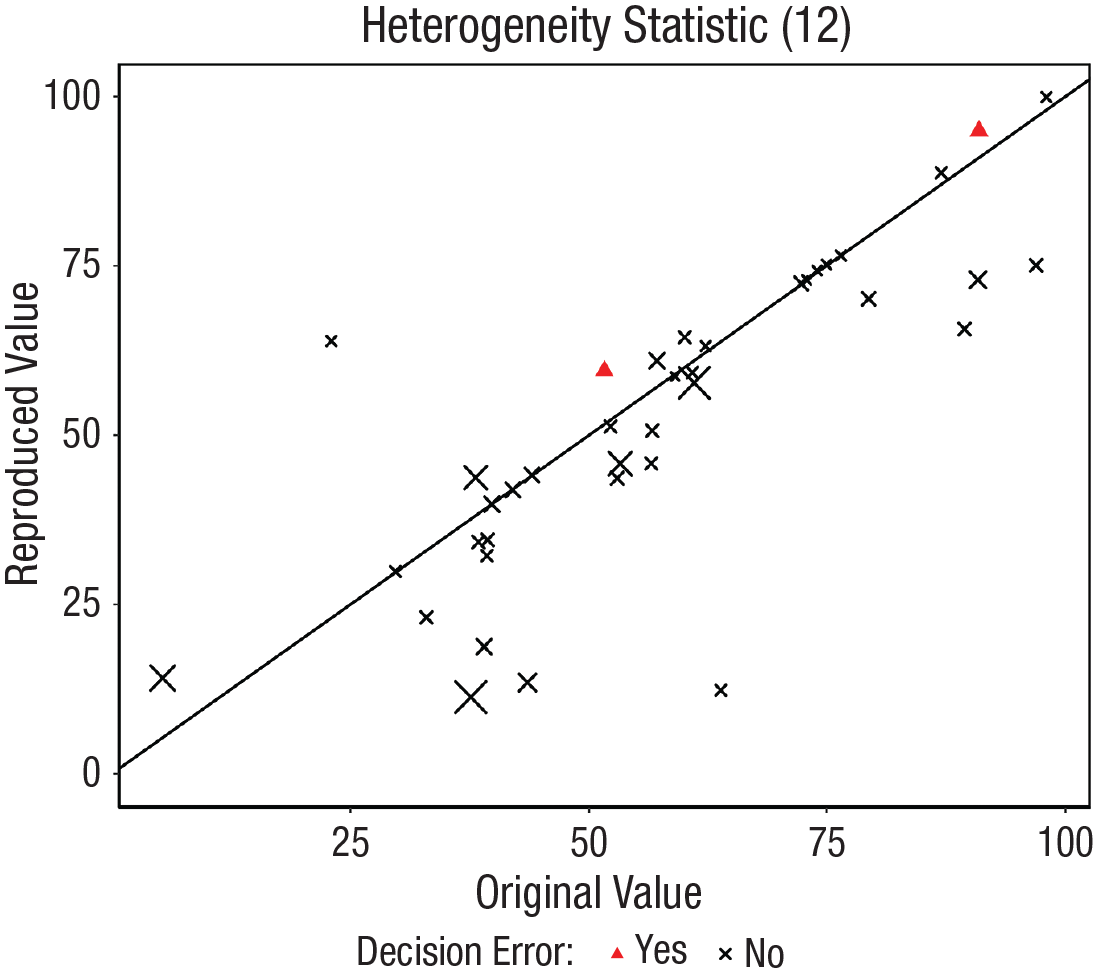
Scatterplot of the reproduced values as a function of the original values classified by whether decision error was found. Only the results of the 52 meta-analyses with a discrepancy of more than 5% identified in the first stage are displayed, but with the corrections made in the second stage. The values displayed are I2 heterogeneity statistics. The size of the crosses is a function of the discrepancy in the summary effect.
Main issues identified
Different issues in these 27 meta-analyses were identified in the second stage. For example, for one of the meta-analyses that showed a discrepancy in the confidence limits, inconsistencies were found in the original meta-analytic report itself. The confidence limits originally reported for that meta-analysis were different in the abstract, main text, and forest plot. Matching the reproduced results were those reported in the forest plot but not those reported in the text. Furthermore, inconsistencies in the original summary effect reported were found between the results reported in the abstract and the results reported in the main text and the forest plot. In addition, in an article in which primary data were available in both a table and a forest plot, minor inconsistencies were found between the primary data of the table and the forest plot. These examples of inconsistencies in original results or data were found in four cases (3%, 4/146). These appeared to be typos. Furthermore, some inconsistencies were found with respect to the number of primary studies included in each meta-analysis. For example, in one of the meta-analyses, the main text reported the inclusion of 10 comparisons in the meta-analysis, whereas in a table of results, 11 comparisons were reported for this meta-analysis. On the other hand, in 11 meta-analyses, the primary data retrieved from the supplementary materials were not sufficient to reach the number of primary studies stated as included in the meta-analysis in the original report.
Original authors’ clarifications
These 27 meta-analyses were from 10 different articles. Therefore, 10 clarification requests with information about the study aims, methods, and preliminary results were sent to the corresponding authors of the original articles. A reply was received in only two of the 10 cases. In one of them, the original authors sent back a link to an OSF repository, 7 where the original data and analysis script were stored. According to the authors, this link was not reported in the article by mistake. The script was run on these data, and the results were successfully reproduced. In this case, the data previously used were retrieved from a forest plot (means and standard deviations) and a table (sample sizes) reported in the article. The previous discrepancy was explained by two cases included in the original meta-analysis from the same primary study that were reported with the same ID in the forest plot and were not correctly matched with their corresponding sample size extracted from the table. This situation exemplifies the potential issues arising from having to reconstruct the original data from tables and figures and not having open access to the original data file.
In the other case, the original data were retrieved from a huge table in supplementary material with all effect sizes and their confidence limits. The original authors sent back this same table but increasing the number of decimal places of the effect sizes and after correcting some wrong values that they themselves detected in that process. This fixed the discrepancies for some of the meta-analyses in this article.
Discussion
The main aim of this study was to examine the reproducibility of a sample of published meta-analyses on the effectiveness of clinical-psychology interventions. We analyzed the availability and reusability of original data, assessed the reproducibility of the published results using these retrieved original data, and tried to reconstruct the original analysis plan. We encountered both difficulties in retrieving the original data and some problems with the reproducibility of the meta-analyses examined.
Even when we interpret data availability in the broad sense (i.e., retrieving data from tables and figures when no data file was available), for about a third of the included meta-analyses, no data were available. In these cases, attempts were made to obtain the data by request to the corresponding author, with little success. Authors shared data in only 12% of the requests that were made. This result is in line with what was found in a recent study in which data-availability statements from a set of primary studies were analyzed (Gabelica et al., 2022). Although 42% of primary studies in Gabelica et al. (2022) reported data were available on request (an identical percentage was found for meta-analyses in Page et al., 2022), only 6.8% of the authors shared the underlying data when requested. Although it is common to see authors state data are available on request, actually obtaining the data on request seems highly challenging. Although this problem of retrieving data on request is well known (Wicherts et al., 2006), the situation does not seem to have improved. Nowadays, there are straightforward, free, and open ways to share data, including meta-analytic data files. Several repositories (e.g., OSF, GitHub, Zenodo, Figshare) are available for researchers to openly share the data associated with published results. On-request availability has proven to be inadequate, and with the availability of data repositories, it is no longer necessary. Journals publishing meta-analyses should require that authors share the underlying data in a public data repository.
Nevertheless, a more positive sign comes from the positive association between publication year and the possibility of retrieving the data. The results tentatively suggest a trend of improving data availability over the years, with a notable rate of 80% observed in meta-analyses published between 2016 and 2020. This observation could be related to the existence of well-established meta-analysis reporting guidelines. For instance, the first PRISMA guideline (Moher et al., 2009) encouraged meta-analysts to report results of primary studies (e.g., primary effect sizes and their confidence interval through a forest plot, as was a common scenario among the cases included in this project), and the latest PRISMA guideline (Page et al., 2021) puts more emphasis on appropriate data sharing through data files ready for reuse. At the same time, in only 5% of the cases in which data were retrieved in our sample were we able to retrieve the data in a machine-readable data file that was ready for reuse (e.g., csv, xlsx). Most often, the data had to be retrieved from files in document format (e.g., docx, pdf). This forces people who want to reuse the data to manually recode the data, which is an inefficient and error-prone task. Even after partial double-coding was carried out, this procedure did not avoid some coding errors, which were detected only by double-checking meta-analyses with discrepancies. In our experience, the data-retrieval process can be difficult when results are presented in general tables because it involves matching subsets of these primary data with different meta-analytic results even though it is not always clear which studies were used in which meta-analysis reported in an article. Furthermore, because the tables in articles are often generated manually in document-file formats (e.g., Word), we observed examples in which this introduced another source of error. The foregoing discussion raises a key point about how time-consuming the appraisal of meta-analytic reproducibility currently is and how efficiency would be improved by having open access to the underlying meta-analytic data in data-file formats ready for reuse. The latest PRISMA guidelines and some initiatives that promote appropriate data sharing (e.g., Wilkinson et al., 2016) have the potential to generate significant improvements in the reuse of meta-analytic data in the years ahead. In this regard, our results provide a useful baseline for future assessments.
An important finding is that the availability of the original analysis script was very limited. Only in five meta-analyses (3%, all from the same article) was the original script openly available. In most cases, the original analyses were reconstructed from the description provided in the article itself, which was not always rich in detail, so many of these computational details had to be inferred from the default settings of the software authors used. The availability of analysis scripts often shows similar rates in both meta-analyses (Page et al., 2022; Polanin et al., 2020) and primary research (Hardwicke et al., 2020, 2022). This makes it more difficult to easily check the computational reproducibility of the results from such studies. Reconstructing the analytical scheme adds to the workload, has the potential to introduce errors in both the original report and the reconstruction, and deals with the eventual lack of relevant analytical information. With the increasing availability of excellent open-source tools to perform meta-analysis (e.g., metafor in R; Viechtbauer, 2010) and useful templates (Moreau & Gamble, 2020), meta-analysts can use workflows that allow them to create and share analysis code for meta-analyses.
Despite these difficulties, we were able to recover the original data and reconstruct the original analysis approach for 146 meta-analyses, for which the reproducibility of the results was assessed. These attempts went through several stages as explained above of trying to minimize the impact of possible coding errors and requesting clarifications from the original authors. Nevertheless, even with these efforts, some discrepancies remained in the results. We identified different issues that hindered our reproducibility attempts. For example, in some cases, internal discrepancies were found in the article itself (e.g., text–figure discrepancies, text–abstract discrepancies, or text–table discrepancies). Furthermore, some problems were found with the lack of some primary data in which data available in the supplementary material included fewer cases than those finally reported in the results of the published article. These situations could be explained by typos in the manuscript or updates when performing the meta-analysis that produced different versions of the manuscript, data, or supplementary material. Although it is important to note that discrepancies in the summary effect results and their confidence intervals were mostly minor, with little or no impact on the conclusions, these situations are easily avoidable. Some of the problems identified could be explained by typos. Currently, there are tools that facilitate the production of so-called reproducible manuscripts, such as the R packages knitr (Xie, 2022), rmarkdown (Allaire et al., 2022), and papaja (Aust & Barth, 2022). A reproducible manuscript embeds analysis code, data, and results reporting in a single document, extracting and reporting the results from the output of the computational process itself and avoiding error-prone manual transcriptions.
Our results are complementary to those observed in previous research on the reproducibility of the primary effects of meta-analyses (Gøtzsche et al., 2007; Maassen et al., 2020) and related problems because of the multiplicity of primary effects (Tendal et al., 2009). These studies found problems in reproducing the primary effects of published meta-analyses or in reaching agreement between independent coders in computing them. Such problems, to a greater or lesser extent, had some impact on the meta-analytic results. Our results show that even when reusing the primary effects as originally coded, certain problems of reproducibility of the results may remain. Some of these problems are added error on the source of error found in previous research on reproducibility of primary effects, which, in turn, are added error on the sources of error types of primary estimates (e.g., measurement errors, sampling errors, or reporting errors). No scientific research is totally error free, but one of the main tasks of scientists is to minimize this error, and in some cases, such as those observed in this study, minimizing some potential sources of error can be straightforward.
Our study has some limitations. First, the time span covered is fairly wide. Thus, the findings may not capture the changes that have arisen in recent years. Therefore, future studies should examine more specific changes over years to evaluate whether better practices emerge that facilitate reproducibility. Second, most of the primary data were retrieved through manual recoding, which introduces some error. The reported data were rounded, which means we did not have access to precise values, and in many cases, the standard error had to be approximated from the confidence limits. These limitations of our study are caused by the suboptimal practices when sharing data, as we discussed above. Given the nonprecise nature of most of the data retrieved, we had to make a decision about which margin of discrepancy was acceptable. In this study, a margin of 5% was chosen. Because this cutoff is arbitrary, we have tried to focus more on possible issues in the results that fell above this margin than on establishing an exact ratio of nonreproduced meta-analyses based on this arbitrary cutoff. Finally, we examined only meta-analyses in clinical psychology because this is one of the areas that produces the most meta-analyses in psychology and these meta-analyses have a direct impact on applied practice, but it is unknown to which extent our conclusions generalize to meta-analyses in other subdisciplines in psychology.
In conclusion, we observed several difficulties when attempting to reproduce meta-analyses. Two aspects can be highlighted: (a) data availability and reusability of the data as they are shared and (b) apparent errors in the reporting of results. Because data collected for a meta-analysis can be especially useful for future research, direct and open access to such data sets allows for easy updates and reanalyses, which are valuable in a cumulative science. Meta-analytic data generally do not contain sensitive or personal information and can therefore almost always be shared openly because doing so does not involve ethical or legal conflicts. In addition, meta-analytic results often represent the state of the art of the evidence on a particular topic. These results guide applied practice, public policy, or future research directions. This prominent status entails a major responsibility for the credibility, reliability, and validity of published meta-analytic results.
Footnotes
Acknowledgements
We thank the research group of the Meta-Research Centre of Tilburg University for providing feedback on an earlier version of this article. We also thank all the original authors of the included meta-analyses who enabled access to the data used in this project and the authors who provided clarifications in response to requests for clarification in the third stage. A version of this article was posted as a preprint on PsyArxiv: ![]() .
.
Transparency
Action Editor: Jessica Kay Flake
Editor: David A. Sbarra
Author Contribution(s)

