Abstract
Team-science projects have become the “gold standard” for assessing the replicability and variability of key findings in psychological science. However, we believe the typical meta-analytic approach in these projects fails to match the wealth of collected data. Instead, we advocate the use of Bayesian hierarchical modeling for team-science projects, potentially extended in a multiverse analysis. We illustrate this full-scale analysis by applying it to the recently published Many Labs 4 project. This project aimed to replicate the mortality-salience effect—that being reminded of one’s own death strengthens the own cultural identity. In a multiverse analysis, we assess the robustness of the results with varying data-inclusion criteria and prior settings. Bayesian model comparison results largely converge to a common conclusion: The data provide evidence against a mortality-salience effect across the majority of our analyses. We issue general recommendations to facilitate full-scale analyses in team-science projects.
A salient recent reform in psychological science is the trend toward “team science.” In crowd-sourced collaborative projects, many different sites across the globe jointly collect data to answer questions about replicability and variability of effects (Chartier et al., 2018; Forscher et al., 2023; Uhlmann et al., 2019). These team-science efforts have become the “gold standard” for assessing the robustness of key findings in the psychological literature. Noteworthy examples of such large-scale endeavours are The Reproducibility Project: Psychology (Open Science Collaboration, 2015), Many Labs (Ebersole et al., 2016; Klein et al., 2014, 2018, 2022), ManyBabies (Frank et al., 2017; The ManyBabies Consortium, 2020), the Pipeline Project (Schweinsberg et al., 2016), and the Psychological Science Accelerator (Chen et al., 2018; Jones et al., 2021; Moshontz et al., 2018). These crowd-sourcing data-collection efforts allow researchers to obtain larger samples, and hence increase statistical power, and reach traditionally less studied populations (i.e., non-Western participants; Henrich et al., 2010).
Given the wealth of data that are obtained in these collaborative projects, we believe it is important to fully make use of the available information in the statistical analysis. Unfortunately, the analytic strategies that are often taken in team-science projects may not do justice to the collected data. Although some projects, such as ManyBabies, have conducted sophisticated hierarchical analyses, most of the Many Labs projects and other large-scale team-science projects have used standard meta-analytic approaches. In these standard analyses, the data are summarized per lab or site, and a frequentist meta-analysis is conducted in which either a fixed- or random-effects structure is applied. We refer to this type of analysis with compressed data as a “minimal analysis.” We believe a minimal analysis constitutes a missed opportunity because it both limits analytic possibilities and compromises the informativeness of the data. For instance, in a meta-analysis, one cannot investigate participant-level predictors, and the data are reduced to mean effect size and its standard error per lab, thereby losing information about the primary data. A huge advantage of large-scale team-science projects is that participant-level and sometimes even trial-level data within a person are available, so we believe one should use their full potential.
In the following, we argue for what we call a “full-scale analysis” instead of the minimal analysis in team-science efforts. Specifically, we demonstrate the usefulness of Bayesian hierarchical modeling (also known as multilevel modeling; see also Rouder et al., 2019). We first highlight general advantages of the Bayesian modeling approach and then illustrate our method by applying it to the recently published Many Labs 4 project (Klein et al., 2022).
Many Labs 4 is a large-scale attempt to replicate the mortality-salience effect from terror-management theory (TMT; Greenberg et al., 1994): Reminders of one’s own death strengthen one’s cultural identity. In the classical demonstration of this effect, participants from the United States who were prompted to imagine their own death expressed more pro-American (i.e., in line with their worldview) beliefs than participants who were prompted to imagine watching TV. In addition to the question of replicability, Klein et al. (2022) wanted to assess the impact of involving the original authors in the study design. Therefore, some studies followed a standard protocol that was agreed on by experts in the field (“author-advised”), whereas other studies were designed by the labs conducting them (“in-house”). After data collection from more than 2,000 participants in 21 labs with and without involvement of the original authors, the project could not replicate the original finding of Study 1 of Greenberg et al. (1994) and reported an overall meta-analytic effect size of
Bayesian Hierarchical Modeling
So what should such a full-scale analysis look like for Many Labs 4 or other team-science projects? In the following, we describe four features that we believe a full-scale analysis for team-science projects should include.
First, we believe a Bayesian analysis is preferred over a frequentist analysis because the former allows one to obtain evidence for the null hypothesis and to quantify (posterior) uncertainty (Wagenmakers et al., 2018). Especially in replication studies, the chances of obtaining null results are considerable. We opt for a Bayesian analysis using Bayes-factor model comparison (Jeffreys, 1939; Kass & Raftery, 1995). In short, Bayes factors quantify the relative evidence for a model (e.g., the alternative) over another model (e.g., the null). For an introduction to Bayes-factor model comparison, we refer the reader to Wagenmakers et al. (2018) and Rouder et al. (2018).
The main advantage of Bayesian statistics in light of large-scale replication efforts is that it allows a distinction to be made between evidence for the absence of the effect of interest and the absence of evidence for or against the effect (Keysers et al., 2020). In other words, failure to successfully replicate a key effect could mean that the data are undiagnostic for determining whether the effect is present, or it could mean that the data provide substantial evidence against the presence of the effect. Obviously, this difference is highly consequential for interpreting the results of a study.
The second feature of a full-scale analysis relates to the hierarchical nature of data in team-science projects. That is, instead of a meta-analysis, we advocate the use of a hierarchical model including all primary data with participants nested within labs (Hoogeveen, Haaf, et al., 2022; Rouder et al., 2019). In a hierarchical model, the lowest-level data are nested within their higher-level groups, such as trials nested within participants or participants nested within labs or countries. This structure makes it possible to assess general or overall effects and individual or lab-specific deviations from those overall effects. For a tutorial on Bayesian hierarchical modeling, we refer the reader to Veenman et al. (2022). Additional demonstrations of the Bayesian hierarchical modeling approach for team-science efforts can be found in Hoogeveen, Haaf, et al. (2022), Gervais et al. (2017), Tierney et al. (2021), and Tierney et al. (2022). The hierarchical approach for team-science efforts brings several benefits. First, by capitalizing on the full resolution of the data, no information is lost in the interim aggregation process. For instance, in a meta-analysis, a relatively large standard error for a given lab or site might reflect either a heterogeneous sample or simply a small sample. In a hierarchical model, the source of the (im)precision of the estimate is retained and thus can be interpreted. Second and relatedly, hierarchical shrinkage reduces the influence of outlying labs with small samples, hence automatically weighing the contribution of the different labs toward the global estimate (Efron & Morris, 1977). Third, although study-level predictors may be included in a meta-analysis, the hierarchical model additionally allows for the inclusion of participant-level predictors and/or the assessment of interaction effects. Finally, in the hierarchical approach, one can easily evaluate whether effects meaningfully differ per site/lab (e.g., in terms of WEIRDness [Western, educated, industrialized, rich, and democratic] or cross-cultural robustness; e.g., Hoogeveen, Haaf, et al., 2022).
The third feature of a full-scale analysis concerns the inclusion of theoretical constraint in the statistical analysis (Haaf et al., 2018; Haaf & Rouder, 2017; Rouder et al., 2019). Psychological theories typically constrain behavioral data in the sense that theories dictate ordinal predictions; observed effects are described in the form of “manipulation X causes higher scores on Y or slower responses” or “higher scores on X are associated with lower scores on Y or faster responses.” Given the ordinal nature of the hypotheses, we believe statistical tests should reflect the theoretical predictions about the direction of effects. For instance, we expect participants who imagined their own death to identify more with American culture than participants who imagined watching TV rather than just a difference between conditions.
The hierarchical nature of the data in team-science projects allows for more informative testing of ordinal predictions beyond directional constraint at the aggregate level. Specifically, rather than testing whether on average, participants who imagined their own death identify more with American culture than participants who imagined watching TV, one can also test whether this pattern holds across every lab that is included in the analysis. This latter constitutes a much riskier prediction because the effect now needs to be present in every single lab (e.g., 21 times instead of once). This risky prediction is potentially rewarded in terms of evidence when the data reflect the predicted pattern, boosting the effect’s credibility. Rouder et al. (2019) referred to this “Does every study?” question as a test of qualitative differences because it provides information on whether the effect of interest is qualitatively equal across studies (i.e., in the same direction).
Bayesian modeling methods are particularly well suited to test ordinal constraints at different levels, such as “Is the overall mortality-salience effect positive?” or “Does every lab show a positive mortality-salience effect?” We would therefore advocate to include both versions of these theoretically motivated ordinal constraints in the statistical analysis for team-science projects (for application of these ordinal constraints in meta-analysis and individual cognitive performance, see also Rouder et al., 2019, and Haaf & Rouder, 2017, respectively).
Finally, the fourth feature of a full-scale analysis relates to assessing the robustness of the findings. That is, beyond using Bayesian hierarchical modeling, we believe team-science projects can at least sometimes benefit from conducting a multiverse analysis (Steegen et al., 2016). In a multiverse analysis, the researcher can evaluate different potential constellations of the data (e.g., exclusions, theoretically relevant subgroups), priors, and predictors without committing to one—perhaps arbitrarily—chosen analysis path. As is demonstrated by the Many Labs 4 example below, there are often multiple defensible analytic choices that can be considered. A complete assessment of the robustness of a given effect might thus require many labs and many analyses (Wagenmakers et al., 2022). The multiverse approach not only presents a broader and more complete picture of the results, but it also allows one to explore the consequences of analytic choices. For instance, does including only an ideal subgroup of participants indeed increase the evidence for the presence of the effect? Do exclusions based on manipulation checks affect the evidence? Does the particular operationalization of a construct make a difference? Furthermore, the Bayes-factor model comparison approach allows for a straightforward interpretation of the multiverse results. Although we do not integrate the evidence from the different multiverse paths directly, the nature of Bayes factors as ratios or odds makes them easily comparable across paths. For instance, given equal prior odds and Bayes factors in favor of the effect of interest ranging between, say, 50 to 1 and 200 to 1 across all paths, one can be fairly confident in the presence of the effect. In other words, with those posterior odds (and unit prior odds), one would probably be comfortable betting on the effect irrespective of the chosen analysis path. Moreover, Bayes factors automatically take into account the sample size and reflect the informativeness of the data.
Many Labs 4 Reanalysis
Given the outlined advantages of a full-scale analysis, in the following, we present a Bayesian multiverse reanalysis of the Many Labs 4 data using hierarchical models. Note that we also conducted a Bayesian model-averaged meta-analysis (Gronau et al., 2021), which is reported in the Appendix. The results of the model-averaged meta-analysis are qualitatively comparable with those of the hierarchical modeling reported below.
A brief history
In December 2019, the Many Labs 4 authors posted a preprint of the project on PsyArXiv (Klein et al., 2019). Soon after, a critique of the analysis emerged in which Chatard et al. (2020) pointed out that Klein and colleagues (2019) had not followed their own preregistered analysis. Chatard et al. argued that the preregistration specified a minimum of 40 participants per experimental cell as the threshold for sufficient power of any individual study and therefore determined a total of 80 participants as target sample size for each lab. When reanalyzing the data from the Many Labs 4 project including only studies with 40 participants per condition, Chatard et al. found a significant effect in line with the original results. Intrigued by these divergent reports, we then decided to conduct a Bayesian multiverse analysis. A preprint of this analysis was published in 2020 on PsyArXiv (Chatard et al., 2020). Then in 2022, Klein and colleagues published their final results in Collabra, after which we also revisited the data, resulting in the current article.
Include or exclude?
Which of the different proposed analyses—Klein et al. (2022) or Chatard et al. (2020)—is the correct one? Given the theoretical arguments and (interpretations of) the preregistered plan, there may be several valid answers to this question and several levels of exclusion criteria that ought to be considered to subset the full sample of 2,281 participants across 21 labs.
The full set of exclusion criteria employed by either Klein et al. (2022), Chatard et al. (2020), and ourselves consists of five layers of exclusion settings with each three or two specific choices, resulting in
Note that some of the criteria are completely overlapping (e.g., only author-advised labs recorded American identity, hence all in-house labs are excluded for the third participant-level exclusion set). As a result, there are 45 instead of 72 unique constellations. Table 1 shows all 45 unique constellations, the resulting number of studies, and total number of included participants (see the Appendix for a table with all 72 constellations).
Exclusion Constellations and Resulting Sample Sizes

Note: Orange rows refer to Klein et al.’s key analyses; green rows refer to Chatard et al.’s key analyses; purple rows refer to our currently chosen analyses; AA = author-advised; IH = in-house. “Apply P-based” indicates whether the participant-level exclusion criteria are applied to the author-advised labs only (retaining all in-house participants) or to both author-advised and in-house labs (missing data excluded).
In the following, we report a reanalysis for the three exclusion constellations of the key analyses from Klein et al. (2022, orange rows in Table 1), the three exclusion constellations from Chatard et al. (2020, green rows), and our own choice of exclusion criteria (purple rows). Subsequently, lacking compelling argumentation for or against any of the criteria, we decided to conduct an analysis using the entire set of 45 unique constellations as a multiverse analysis (Steegen et al., 2016).
Disclosures
Preregistration
Our analyses, including prior settings, were preregistered on OSF (osf.io/ae4wx; see also Appendix D). However, we decided to deviate from the preregistration by including more constellations of exclusion criteria. Specifically, we originally planned to use only participant-level Exclusion Criterion 1 and later decided to include all of them. Moreover, two additional exclusion layers became apparent only after the final version of the Many Labs 4 report was published, specifically, those related to the timing-based exclusion criteria and the application of the participant-level criteria to the author-advised only or author-advised and in-house labs. We believe that including these additional paths in the multiverse analysis helps to provide a more complete analysis. We also note that the preregistration includes both the hierarchical analysis and the model-averaged meta-analysis. The latter is reported in Appendix B.
Data and materials
Readers can access the data and the R code to conduct all analyses (including all figures) at github.com/SuzanneHoogeveen/ml4-reanalysis.
Reporting
This study involved an analysis of existing data rather than new data collection.
Ethical approval
No ethical approval was required for this work because we did not collect any data.
Method
For Bayesian hierarchical modeling, we take advantage of the open availability of all collected data from the Many Labs 4 project. The dependent variable is the same across all studies (i.e., identification with American culture, operationalized through relative preference for American vs. non-American authors), and participants are nested in studies, resulting in a hierarchical data structure. We employed a modeling approach similar to the one developed for the embodied-cognition reanalysis by Rouder et al. (2019). That is, we used Bayes-factor model comparison with hierarchical models reflecting different structures of the data, varying in the extent to which they constrain their predictions. We believe this approach satisfies the analytic desiderata for team-science projects outlined before, that is, appropriately accounting for the nested structure of the data without compromising on informativeness, directly testing both the presence of an overall mortality-salience effect and the presence of between-studies heterogeneity, and reflecting theoretical constraints on the direction of the effect.
Concretely, there are four models under consideration. The null model (Model 1) corresponds to the notion that none of the studies show an effect; this model assumes no overall experimental effect or heterogeneity between studies. The common-effect model (Model 2) corresponds to the notion that all studies show the same effect in the expected direction; this model assumes no heterogeneity between studies. The positive-effects model (Model 3) corresponds to the notion that all studies show an effect in the expected direction yet to varying degrees. The unconstrained model (Model 4) refers to the notion that the overall effect and study effects may vary freely (in direction and size). We compute Bayes factors for Models 2, 3, and 4 against Model 1, the null model. Evidence for Model 1 would indicate the absence of a mortality-salience effect across all labs; evidence for Model 2 would indicate that on average, people who contemplate their own death identify more strongly with their culture than people who contemplate watching TV to a similar degree across labs; evidence for Model 3 would indicate that in all of the labs, people who contemplate their own death identify more strongly with their culture than people who contemplate watching TV but to varying degrees across labs; and evidence for Model 4 would indicate that in some labs, people who contemplate their own death identify more strongly with their culture than people who contemplate watching TV, whereas in other labs, people who contemplate watching TV identify more strongly with their culture than people who contemplate their own death.
The Bayesian hierarchical modeling is conducted using the R package BayesFactor (Morey & Rouder, 2018). See Box 1 for a mathematical specification of the model.
Box 1
Hierarchical Model Specifications
The base model for the mortality-salience effect is a mixed linear model. Let
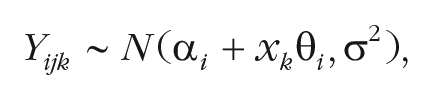
where
There are two critical prior settings to consider: the scale setting on the overall effect
Results
In the following, we first reanalyze the data from the key findings reported by Klein et al. (2022) using our proposed full-scale analysis and then those from Chatard et al. (2020). Finally, we report the analysis of the data based on our own choice of exclusion-criteria constellations.
Bayesian reanalysis of Klein et al.’s (2022) key findings
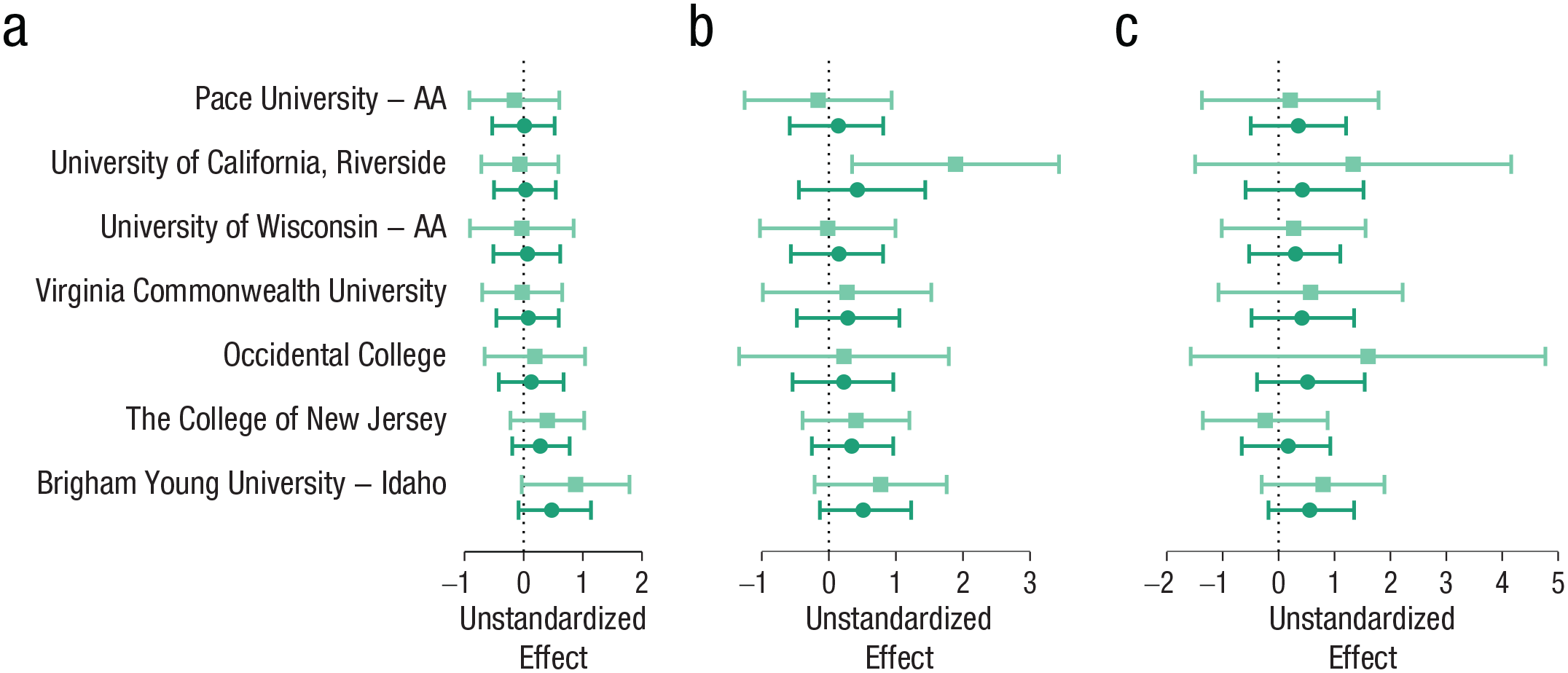
Figure 1a shows the observed, unstandardized effects and the estimates from the unconstrained hierarchical model for the first participant-level exclusion criterion. This is the main analysis that is the basis for the key claims of the Many Labs 4 project, as reported in the published article (Klein et al., 2022). Klein et al. (2022) included participants whose data were collected after the lead team posted its preregistration and only studies that featured more than 60 observations (before participant-level exclusions). The participant-level exclusion criteria were applied only to author-advised studies, whereas all participants from the in-house studies were retained.

Forest plot with Bayesian parameter estimates for the key analyses by Klein et al. (2022) for the three participant-level exclusion sets (applied to author-advised protocol participants only) with data collected after the lead team posted their preregistration and only from studies that featured more than 60 observations. (a) Participant-level Exclusion Set 1. The light orange squares represent unstandardized observed effects for each study with 95% confidence intervals. The dark orange points represent estimated unstandardized effects from the unconstrained model with 95% credible intervals. (b) Participant-level Exclusion Set 2. (c) Participant-level Exclusion Set 3. The estimates are sorted by the size of the observed effects for participant-level Exclusion Set 1 (Fig. 1a).
As is shown in Figure 1, there is considerable hierarchical shrinkage reducing the variability of estimated effects compared with observed effects. Effect-size estimates from the unconstrained model (similar to Cohen’s d) are 0.02, 95%
Bayes factors are shown in the first three rows of Table 2. “
Bayes Factors for Key Analyses
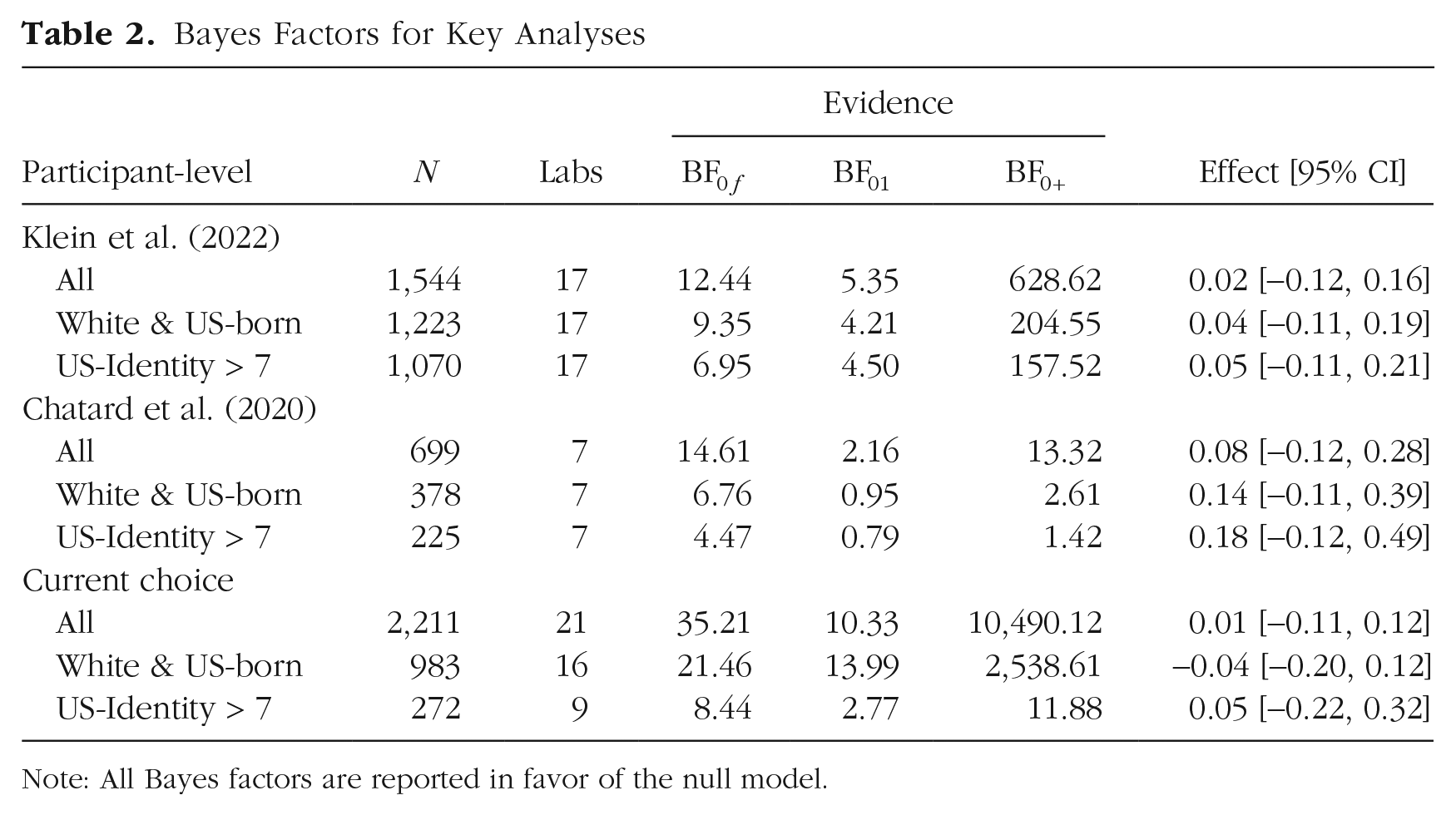
Note: All Bayes factors are reported in favor of the null model.
In summary, the null results are consistent across participant-level exclusion criteria. Even though the evidence against an effect is more pronounced when all participants are included in the analysis, this pattern is easily explained by the resolution of the analysis with increasing numbers of observations: The smaller the number of observations, the less evidence there is in any direction and the wider the estimated posterior distribution of the overall effect.
Bayesian reanalysis of Chatard et al.’s (2020) key findings
We also reanalyzed Chatard et al.’s (2020) findings with a hierarchical-modeling approach. Figure 2 shows study estimates from the unconstrained model for the unstandardized effects. All confidence intervals and credible intervals cover zero.
Forest plot with Bayesian parameter estimates for the key analyses by Chatard et al. (2020) for the three participant-level exclusion sets, only studies that featured more than 80 observations, and author-advised labs only. (a) Participant-level Exclusion Set 1. The light green squares represent unstandardized observed effects for each study with 95% confidence intervals. The dark green points represent estimated unstandardized effects from the unconstrained model with 95% credible intervals. (b) Participant-level Exclusion Set 2. (c) Participant-level Exclusion Set 3. The estimates are sorted by the size of the observed effects for participant-level Exclusion Set 1 (i.e., Fig. 2a).
Effect-size estimates from the unconstrained model (similar to Cohen’s d) are 0.08, 95%
The pattern of Bayes factors is somewhat less consistent across exclusions than the estimation results. Bayes factors are shown in the middle three rows of Table 2. The pattern of Bayes factors depends on the participant-level exclusion criterion. Under participant-level Exclusion Criterion 1, the preferred model is the null model, and it is weakly preferred over the second-best model, the common-effect model, by a Bayes factor of
Bayesian analysis of our current choice
We also included an analysis of the Many Labs 4 data using our own choice of the exclusion criteria. Following Klein et al. (2022) and Chatard et al. (2020), we looked at all three participant-level exclusion criteria while settling on one particular choice for the other factors that seemed most sensible to us. The goal for this choice was to include the maximum number of participants but still adhere to the recommendations by the original authors ( Greenberg et al., 1994) to give the effect the best chance. Specifically, we included all complete data from all labs and protocols and applied the participant-level exclusions to both author-advised labs and in-house labs, discarding missing values. For Exclusion Criterion 1—completeness of the measures—we did retain participants for labs in which no explicit information on missingness was available as long as they were assigned to an experimental condition and answered both items of the dependent variable.
1
Note that our choice of analysis paths leads to quite variable numbers of participants (between
Figure 3 shows the study estimates from the unconstrained model for the unstandardized effects. Again, all confidence and credible intervals include zero. Effect size estimates from the unconstrained model are

Forest plot with Bayesian parameter estimates for the key analyses of our choice for the three participant-level exclusion sets (applied to both author-advised and in-house protocol participants), including all participants and all labs. (a) Participant-level Exclusion Set 1. The light purple squares represent unstandardized observed effects for each study with 95% confidence intervals. The dark purple points represent estimated unstandardized effects from the unconstrained model with 95% credible intervals. (b) Participant-level Exclusion Set 2. (c) Participant-level Exclusion Set 3. The estimates are sorted by the size of the observed effects for participant-level Exclusion Set 1 (i.e., Fig. 3a).
The Bayes factors paint a similar picture; the evidence against the presence of the mortality-salience effect is stronger with a larger sample size. For the most inclusive sample, with Exclusion Criterion 1
Bayesian multiverse analysis across all exclusion criteria
To assess the robustness of the previously reported results, we conducted a multiverse analysis using the 45 unique data sets from Table 1. We used the same hierarchical model construction as reported above and report here the Bayes factors for the presence of an effect against its absence. The Bayes factors are plotted in Figure 4 (y-axis). Bayes factors in favor of the mortality-salience effect are above the horizontal line, and Bayes factors against the mortality salience effect are below the horizontal line. The BFeffect0 is the weighted average of the evidence for the common-effect versus the null model and the unconstrained model (varying effect) versus the null model. The x-axis refers to the evidence for between-studies heterogeneity in the data. The BFheterogeneity0 is calculated by taking the evidence for the unconstrained model versus the common-effect model (i.e.,

Results from the Bayesian multiverse analysis. Bayes factors in favor of a mortality-salience effect are above the horizontal line; Bayes factors against the mortality-salience effect are below the horizontal line. All analyses provide evidence against between-studies heterogeneity as shown by all heterogeneity Bayes factors being smaller than 1 on the x-axis. The color of the points refers to the different key analyses sets, and the size of the points refers to the number of participants the analysis is based on. All but two of analyses provide evidence against the mortality-salience effect. BFeffect0 reflects the model-averaged evidence for common effect and the varying effect versus the null, and BFheterogeneity0 reflects the evidence for the varying effect versus the common effect (i.e.,
The majority of Bayes factors are in line with the absence of the mortality-salience effect. Because the Bayes factor depends on the sample size, more evidence against morality salience comes from analyses that are based on more data (i.e., larger number of included participants and studies). Only two constellations of exclusion criteria provide evidence for the mortality-salience effect. In addition, none of the analysis paths provide evidence for heterogeneity (all BFheterogeneity0 < 1).
In sum, the evidence against the mortality-salience effect appears relatively robust against choices of exclusion criteria. When conducting a large number of analyses on the same data, some of these analyses will almost inevitably lead to evidence in the opposite direction than the overall results. This is especially the case when the data provide relatively weak evidence (Bayes factors less than 5 to 1 against an effect). Bayes factors close to 1 signal a lack of resolution of the data and therefore the absence of evidence for or against an effect. When the number of participants is high and many studies are included, there is convincing evidence against the mortality-salience effect. The two Bayes factors that are weakly in favor of the mortality-salience effect are based on less than half of the original data and prefer the presence of the effect only by factors of 1.5 and 1.7.
Prior sensitivity
In addition to assessing the effects of various data-exclusion decisions, one might also investigate the role of prior choices on inference. Specifically, we looked at the dependence of the Bayes factors on the prior settings for the overall effect and for the between-studies variability. Although some researchers have argued that the influence of the prior on the results should be minimized (e.g., by using uninformative default settings; Aitkin, 1991; Gelman et al., 2004; Kruschke, 2013), we believe the influence of the prior is a meaningful and inherently informative element of Bayesian inference (Rouder et al., 2018; Vanpaemel, 2010). Nevertheless, the extent to which reasonable prior choices affect the results clearly speaks to the robustness of the conclusions.
For the main analysis, we used a scale of 0.4 on the overall effect
Bayes Factors for Key Analyses (Participant-Level Exclusion Set 1) Under Different Prior Settings

Note: All Bayes factors are reported in favor of the null model.
Table 3 shows the Bayes factors resulting from crossing these combinations for the key analyses (for participant-level Exclusion Set 1), and Figure 5 shows the evidence across all 45 unique data-exclusion paths for each of the four different prior setting combinations. Most support for the effect is obtained under the expectation of a small effect, and most support for between-studies heterogeneity is obtained under the expectation of little between-studies variability. Nevertheless, across all settings, evidence somewhat in favor of a mortality-salience effect occurs only in 10 of 180 (5.6%) paths. Evidence in favor of heterogeneity across studies is obtained under three of 180 (1.7%) paths. Another observation from these plots is that although the prior setting for the overall effect changes the global strength of the evidence, it does not appear to affect the multiverse paths differentially, given that the dots seem to move upward or downward uniformly. In contrast, the prior setting for the between-studies variability not only affects the overall evidence for heterogeneity, but it also influences the range of the Bayes factors between multiverse paths. Specifically, the prior expectation of little variability reduces the evidence against heterogeneity and makes the Bayes factors more similar across paths, whereas the expectation of much variability not only leads to more evidence against heterogeneity but also enhances the differences between paths. In sum, choices of prior scales can slightly boost or reduce the evidence in favor of the effect. Yet in this case, the effects of reasonable prior choices are rather contained; the null model is still consistently preferred over models with a mortality-salience effect.

Results from the Bayesian multiverse analysis under different prior settings for the overall effect and the between-studies variance in the effect. The arrows show the overall trend relative to the main analysis with the primary prior settings.
Conclusion
We conducted a Bayesian reanalysis of the Many Labs 4 project with varying exclusion criteria and prior settings. In a Bayesian multiverse analysis using hierarchical models, we calculated a total of 45 sets of Bayes factors based on different combinations of five layers of data-exclusion criteria derived from the Many Labs 4 preregistration, the comment by Chatard et al. (2020), the published article by Klein et al. (2022), and our own judgments. Forty-three out of 45 Bayes factors provide evidence against an overall mortality-salience effect, ranging between 1.32 to 1 and 16.94 to 1 in favor of the absence of an effect. The remaining two Bayes factors provide only weak evidence for the presence of such an effect (i.e., Bayes factors of 1.45 to 1 and 1.68 to 1). In addition, we find some evidence against heterogeneity of effects across studies. Finally, the pattern of results remains qualitatively equal under different reasonable prior settings for the overall effect and the between-studies variability. In combination, we would argue we conducted a full-scale analysis of the data provided by the Many Labs 4 project, an inspection from various different angles. Even if we do not believe the evidence from this full-scale analysis and assume there is an effect, this effect is so small (between
Our analyses revealed that the evidence is relatively consistent across different exclusion criteria. For the current analysis, we assumed that all exclusion criteria were equally plausible. With this assumption, we implicitly assigned an equal weight to all analyses. However, we admit that this may not be the case. Chatard et al. (2020) argued that their chosen criteria are superior when considering theoretical arguments and study planning. With their analysis, they implicitly introduced a weighing in which all other exclusion options received a weight of zero. Readers can choose these weights themselves when they consider how to interpret the results reported here. 2
There are additional issues with selectively subsetting and reanalyzing data sets. A key danger is that for some subsets, one always finds results opposite of the conclusions from the analysis of the full data set. On the study level, researchers should therefore first ensure that there is evidence for variability of studies that warrants such subsetting (Harrer et al., 2021, Chapter 5). In the current analysis, we found evidence against study heterogeneity. When interpreting the results, we therefore recommend to rely mainly on the estimates from the full data set. In addition, subsetting the data inevitably reduces the resolution to detect an effect. The critics of the Many Labs 4 project (Chatard et al., 2020) based their main conclusions on analyses with smaller sample sizes. Ironically, although Chatard et al. (2020) argued that sample size should be considered when including studies, their exclusion criteria actually reduced the power of the meta-analysis overall. To tackle this issue—and if there was evidence for study heterogeneity—one could include some of the subsetting criteria as dummy-coded predictors in the hierarchical model instead of disregarding the data altogether (e.g., author-advised vs. in-house).
Furthermore, we believe the Many Labs 4 case and its development from preprint to published article highlights an important potential drawback of preregistration. In the final article, the Many Labs 4 lead team decided to discard all observations collected before the preregistration date, resulting in the removal of more than a quarter of the data. As mentioned, we consider this removal of data wasteful and unnecessary. In this case, we believe the fact that data collection was crowdsourced and the added value of retaining 556 perfectly valid observations justify “breaking the strict rules of preregistation” that data collection should start only after the analysis plan has been preregistered. As noted by DeHaven (2017), preregistration is “a plan, not a prison.” So rather than discarding a large portion of the data for the main analyses, we believe a transparent statement on the timing issue would have sufficed in this case. In general, preregistration by definition should not trump common sense and researchers’ judgment.
In summary, the multiverse analysis conducted here shows a certain convergence of results. Even though the degree of evidence varies, models with no effect of mortality salience are mostly preferred over models with an effect of mortality salience. This result highlights the robustness against choices of exclusion criteria. The Bayesian multiverse approach using hierarchical models provides rich results that go much beyond the original analyses by the Many Labs 4 team. In particular, we believe the current approach satisfies the desiderata of a full-scale analysis in team-science projects: (a) providing evidence on a continuous scale from evidence against the crucial effect through inconclusive evidence to evidence in favor of the crucial effect, (b) applying hierarchical modeling to appropriately account for the nested structure of the data, (c) evaluating both the evidence for the experimental effect and the evidence for between-labs heterogeneity, (d) reflecting theoretical constraints on the effect of interest (i.e., ordinal constraints), and (e) evaluating the robustness of the findings by exploring a multitude of relevant analysis paths.
Both Bayes-factor model comparison and Bayesian hierarchical modeling are gaining popularity in psychological science. Recent tutorial articles make these approaches more accessible; for instance, see Wagenmakers et al. (2018) and Rouder et al. (2018) for an introduction to Bayes-factor model comparison and Veenman et al. (2022) and Rouder and Province (2019) for tutorials on Bayesian hierarchical modeling. Finally, the ease and informativeness of Bayesian multiverse analyses show that this approach should be more generally used to analyze team-science projects. The current analyses were conducted in R, and the code is provided at github.com/SuzanneHoogeveen/ml4-reanalysis.
General Recommendations
In sum, we believe the amount of time and effort spent on team-science projects and the resulting wealth of data deserve a full-scale analysis. We believe a Bayesian hierarchical-modeling approach is ideally suited for such an analysis because it allows evidence to be quantified both for and against an effect of interest, and it facilitates the consideration of theoretical constraint in the data. In the following, we highlight four additional general recommendations for team science that facilitate a full-scale analysis.
Our first recommendation is to use all data that are available. Most directly, this means using a hierarchical model with all primary data nested in studies rather than a meta-analysis based on compressed and aggregated data. Furthermore, although participant-level exclusions may be explored (see Point b above), we would advise never to apply study-level exclusions based on sample size. In particular, more data always means more statistical power and more resolution. In addition, hierarchical shrinkage will automatically reduce the influence of outlying labs with relatively few observations by more strongly pulling these observations toward the global estimate.
Our second recommendation is to conduct a multiverse analysis (Steegen et al., 2016) to investigate the evidence across different reasonable exclusion criteria, model choices, or prior settings. As illustrated by the Many Labs 4 project, team-science efforts often involve a range of reasonable options for data-exclusion criteria, prior settings, and perhaps other analytic choices. To get a full picture of the robustness and potential relevant dimensions of the data affecting the outcomes, analysts could explore multiple analytic paths (see also Tierney et al., 2021, 2022). In some cases, it might make sense to apply different weights to different paths of the multiverse, for instance, based on theoretical or methodological grounds.
Our third recommendation is to preregister but remain open to justifiable deviations. Especially in highly complex projects with crowd-sourced data collection and many involved parties, unexpected events and deviations are the norm rather than the exception. At least in our personal experience, none of the team-science projects went exactly as planned, and many required reconsideration of preregistered choices (e.g., Hoogeveen, Haaf, et al., 2022; Hoogeveen, Sarafoglou, et al., 2023; Tierney et al., 2021, 2022). Although full transparency is clearly key in these situations, we believe the quality of the eventual analysis and hence the validity of the conclusions should outweigh the strict adherence to the preregistration. Another option to ensure uncontaminated data analysis would be to use “blinded analysis” (MacCoun & Perlmutter, 2015, 2018), in which analysts perform their analysis on an altered version of the data (e.g., shuffling the dependent variable, adding noise to the data, or switching labels of categorical variables). Only after the analysts are fully satisfied with the analysis, the blind is lifted and the real data are revealed (for more information on analysis blinding, see Dutilh et al., 2021; Sarafoglou et al., 2023).
Our fourth recommendation is to consider collaborating with methodologists on the statistical analysis. Typically, team-science efforts involve relatively extensive and complex data (e.g., hierarchically structured). We believe the time and effort put into data collection and study design also justify spending some additional time, effort, and resources on data-analysis expertise. For the sake of illustration, imagine that each participating lab in the Many Labs 4 project invested 15 min per participant; this comes down to 21 labs spending about 1,589 min on data collection for a total of 556 hr. 3 Given this huge investment of time and effort, the overall project quality might benefit from also matching the investment into the analysis, potentially by outsourcing the analysis to methodological and statistical experts. At least in our personal experience, experts are often eager to help out (and get their hands on “real data” for a change). For example, we have been involved in the data analysis for a couple of team-science projects (e.g., Camerer et al., 2018; Tierney et al., 2021, 2022). Having an independent analysis team may also make it easier to justify deviations from the preregistration and to apply differential weights to paths in the multiverse analysis given that either of these decisions can be made independently from the analysts.
The idea of team-science efforts such as the Many Labs projects is that the robustness of empirical phenomena becomes clear when data are collected across several labs. Likewise, the robustness of statistical conclusions becomes clear when data are analyzed using several thoughtfully selected models in a full-scale analysis (Wagenmakers et al., 2022). A complete assessment of robustness and uncertainty therefore requires many labs, many models, perhaps many analysis paths, and ideally, many collaborating experts.
Footnotes
Appendix A
Appendix B
Appendix C
Appendix D
Appendix E
Full Table of Data-Exclusion Constellations

| Participant level | N-based | Protocol | Timing-based | Apply P-based | Sample size | Labs |
|---|---|---|---|---|---|---|
| All | All | All | All | AA only | 2,225 | 21 |
| White & U.S.-born | All | All | All | AA only | 1,880 | 21 |
| U.S. identity > 7 | All | All | All | AA only | 1,699 | 21 |
| All | N > 60 | All | All | AA only | 2,067 | 17 |
| White & U.S.-born | N > 60 | All | All | AA only | 1,746 | 17 |
| U.S. identity > 7 | N > 60 | All | All | AA only | 1,593 | 17 |
| All | N > 80 | All | All | AA only | 1,866 | 14 |
| White & U.S.-born | N > 80 | All | All | AA only | 1,545 | 14 |
| U.S. identity > 7 | N > 80 | All | All | AA only | 1,392 | 14 |
| All | All | AA | All | AA only | 798 | 9 |
| White & U.S.-born | All | AA | All | AA only | 453 | 9 |
| U.S. identity > 7 | All | AA | All | AA only | 272 | 9 |
| All | N > 60 | AA | All | AA only | 699 | 7 |
| White & U.S.-born | N > 60 | AA | All | AA only | 378 | 7 |
| U.S. identity > 7 | N > 60 | AA | All | AA only | 225 | 7 |
| All | N > 80 | AA | All | AA only | 699 | 7 |
| White & U.S.-born | N > 80 | AA | All | AA only | 378 | 7 |
| U.S. identity > 7 | N > 80 | AA | All | AA only | 225 | 7 |
| All | All | All | After prereg | AA only | 1,659 | 20 |
| White & U.S.-born | All | All | After prereg | AA only | 1,314 | 20 |
| U.S. identity > 7 | All | All | After prereg | AA only | 1,133 | 20 |
| All | N > 60 | All | After prereg | AA only | 1,544 | 17 |
| White & U.S.-born | N > 60 | All | After prereg | AA only | 1,223 | 17 |
| U.S. identity > 7 | N > 60 | All | After prereg | AA only | 1,070 | 17 |
| All | N > 80 | All | After prereg | AA only | 1,343 | 14 |
| White & U.S.-born | N > 80 | All | After prereg | AA only | 1,022 | 14 |
| U.S. identity > 7 | N > 80 | All | After prereg | AA only | 869 | 14 |
| All | All | AA | After prereg | AA only | 797 | 9 |
| White & U.S.-born | All | AA | After prereg | AA only | 452 | 9 |
| U.S. identity > 7 | All | AA | After prereg | AA only | 271 | 9 |
| All | N > 60 | AA | After prereg | AA only | 698 | 7 |
| White & U.S.-born | N > 60 | AA | After prereg | AA only | 377 | 7 |
| U.S. identity > 7 | N > 60 | AA | After prereg | AA only | 224 | 7 |
| All | N > 80 | AA | After prereg | AA only | 698 | 7 |
| White & U.S.-born | N > 80 | AA | After prereg | AA only | 377 | 7 |
| U.S. identity > 7 | N > 80 | AA | After prereg | AA only | 224 | 7 |
| All | All | All | All | AA and IH | 2,211 | 21 |
| White & U.S.-born | All | All | All | AA and IH | 983 | 16 |
| U.S. identity > 7 | All | All | All | AA and IH | 272 | 9 |
| All | N > 60 | All | All | AA and IH | 2,053 | 17 |
| White & U.S.-born | N > 60 | All | All | AA and IH | 897 | 13 |
| U.S. identity > 7 | N > 60 | All | All | AA and IH | 225 | 7 |
| All | N > 80 | All | All | AA and IH | 1,852 | 14 |
| White & U.S.-born | N > 80 | All | All | AA and IH | 864 | 12 |
| U.S. identity > 7 | N > 80 | All | All | AA and IH | 225 | 7 |
| All | All | AA | All | AA and IH | 799 | 9 |
| White & U.S.-born | All | AA | All | AA and IH | 453 | 9 |
| U.S. identity > 7 | All | AA | All | AA and IH | 272 | 9 |
| All | N > 60 | AA | All | AA and IH | 700 | 7 |
| White & U.S.-born | N > 60 | AA | All | AA and IH | 378 | 7 |
| U.S. identity > 7 | N > 60 | AA | All | AA and IH | 225 | 7 |
| All | N > 80 | AA | All | AA and IH | 700 | 7 |
| White & U.S.-born | N > 80 | AA | All | AA and IH | 378 | 7 |
| U.S. identity > 7 | N > 80 | AA | All | AA and IH | 225 | 7 |
| All | All | All | After prereg | AA and IH | 1,650 | 20 |
| White & U.S.-born | All | All | After prereg | AA and IH | 777 | 15 |
| U.S. identity > 7 | All | All | After prereg | AA and IH | 271 | 9 |
| All | N > 60 | All | After prereg | AA and IH | 1,535 | 17 |
| White & U.S.-born | N > 60 | All | After prereg | AA and IH | 702 | 13 |
| U.S. identity > 7 | N > 60 | All | After prereg | AA and IH | 224 | 7 |
| All | N > 80 | All | After prereg | AA and IH | 1,334 | 14 |
| White & U.S.-born | N > 80 | All | After prereg | AA and IH | 669 | 12 |
| U.S. identity > 7 | N > 80 | All | After prereg | AA and IH | 224 | 7 |
| All | All | AA | After prereg | AA and IH | 798 | 9 |
| White & U.S.-born | All | AA | After prereg | AA and IH | 452 | 9 |
| U.S. identity > 7 | All | AA | After prereg | AA and IH | 271 | 9 |
| All | N > 60 | AA | After prereg | AA and IH | 699 | 7 |
| White & U.S.-born | N > 60 | AA | After prereg | AA and IH | 377 | 7 |
| U.S. identity > 7 | N > 60 | AA | After prereg | AA and IH | 224 | 7 |
| All | N > 80 | AA | After prereg | AA and IH | 699 | 7 |
| White & U.S.-born | N > 80 | AA | After prereg | AA and IH | 377 | 7 |
| U.S. identity > 7 | N > 80 | AA | After prereg | AA and IH | 224 | 7 |
Note: Orange rows refer to Klein et al.’s (2022) key analyses; green rows refer to Chatard et al.’s (2020) key analyses; purple rows refer to our chosen analyses; gray rows are repeated data sets and not included in the multiverse analysis. “Apply p-based” indicates whether the participant-level exclusion criteria are applied to the author-advised labs only (retaining all in-house participants) or to both author-advised and in-house labs (missing data excluded). AA = author-advised; IH = in-house; prereg = preregistration.
Acknowledgements
For all analyses, we used R (Version 4.1.2; R Core Team, 2021) and the R-packages BayesFactor (Morey & Rouder, 2021), coda (Plummer et al., 2006), cowplot (Wilke, 2020), dplyr (Wickham et al., 2022), ggplot2 (Wickham, 2016), kableExtra (Zhu, 2021), knitr (Xie, 2015), ks (Duong, 2022), LaplacesDemon (Statisticat LLC, 2021), lemon (Edwards, 2020), MASS (Venables & Ripley, 2002), Matrix (Bates et al., 2022), MCMCpack (Martin et al., 2011), metaBMA (Heck et al., 2019), metafor (Viechtbauer, 2010), papaja (Aust & Barth, 2022), Rcpp (Eddelbuettel & Balamuta, 2018; Eddelbuettel & François, 2011), and tinylabels (Barth, 2021).
Transparency
Action Editor: Pamela Davis-Kean
Editor: David A. Sbarra
Author Contribution(s)
