Abstract
Drift-diffusion models (DDMs) are pivotal in understanding evidence-accumulation processes during decision-making across psychology, behavioral economics, neuroscience, and psychiatry. Hierarchical DDMs (HDDMs), a Python library for hierarchical Bayesian estimation of DDMs, has been widely used among researchers, including researchers with limited coding proficiency, in fitting DDMs and other sequential sampling models. However, issues of compatibility in installation and lack of support for more recent Bayesian-modeling functionalities pose serious challenges for new users, limiting broader adaptation and reproducibility of HDDMs. To address these issues, we created dockerHDDM, a user-friendly computational environment for HDDMs with new features. dockerHDDM brings three improvements: (a) easy to install once docker is installed, ensuring reproducibility and saving time for researchers; (b) compatible with machines with Apple chips; (c) seamless integration with ArviZ, a state-of-the-art Bayesian-modeling library. This tutorial serves as a practical, hands-on guide for researchers to leverage dockerHDDM’s capabilities in conducting efficient Bayesian hierarchical analysis of DDMs. The notebook presented here and in the docker image will enable researchers with various programming levels to model their data with HDDMs.
Keywords
The drift-diffusion model (DDM) is one of the most widely used computational models (for an overview, see Ratcliff et al., 2016) to quantify the evidence-accumulation processes during decision-making in neuroscience (Cavanagh et al., 2011; Herz et al., 2017; Shadlen & Shohamy, 2016), psychology (Hu et al., 2020; D. J. Johnson et al., 2017; Kutlikova et al., 2023), behavioral economics (Desai & Krajbich, 2022; Sheng et al., 2020), and psychiatry (Ging-Jehli et al., 2021; Pedersen et al., 2022). According to the DDM, experimentally observed pairs of response times and choices arise from a process of stochastic evidence accumulation to a decision boundary (e.g., Voss et al., 2013; see Figure 1 and the related DDM glossary in Table 1). This theoretical framework has been shown not only to correlate robustly with established neural substrates (Chandrasekaran et al., 2017; Forstmann et al., 2016) but also to serve as a powerful measurement tool for examining individual differences across cognitive tasks, experimental manipulations, and participant populations (Boag et al., 2024; Donkin & Brown, 2018; Evans & Wagenmakers, 2020; but see Liu et al., 2023). Despite its theoretical contributions, the DDM is difficult to apply to experimental data in practice because the derivation of inference-relevant quantities (e.g., the likelihood function) requires a mathematical understanding of the complex stochastic process of evidence accumulation.

Illustration of the evidence-accumulation process assumed by the drift-diffusion model (DDM). DDM has four basic parameters: drift rate (
Drift-Diffusion Model Glossary
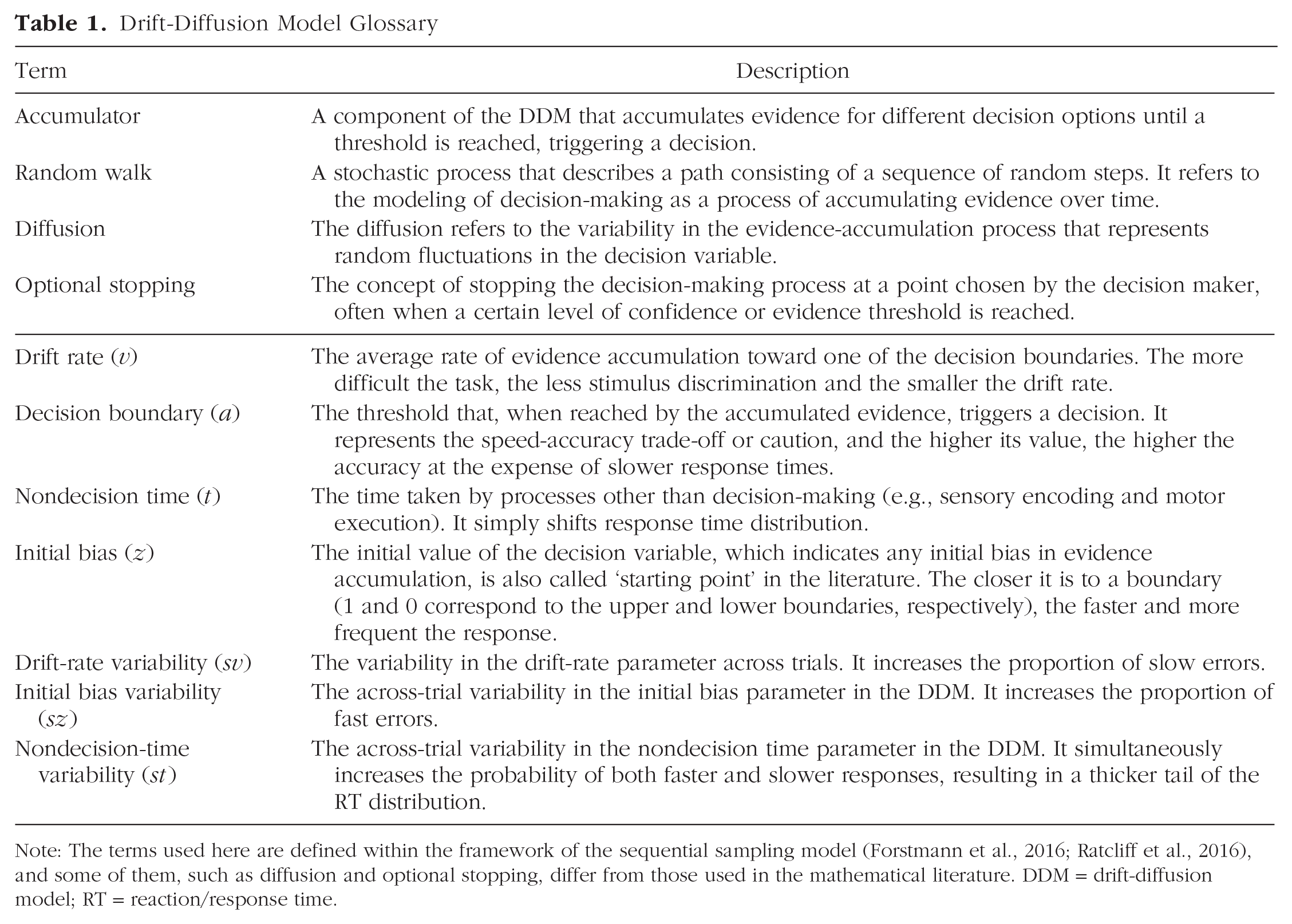
Note: The terms used here are defined within the framework of the sequential sampling model (Forstmann et al., 2016; Ratcliff et al., 2016), and some of them, such as diffusion and optional stopping, differ from those used in the mathematical literature. DDM = drift-diffusion model; RT = reaction/response time.
Several software packages have been developed to facilitate the application of DDM (see “Why Use dockerHDDM Among Tools” section), proving particularly beneficial for researchers with limited computational expertise. Among them, HDDM, a Python library for hierarchical DDM, is by far the most cited toolbox in the community (Wiecki et al., 2013; with 996 citations in Google Scholar as of August 26, 2024). Despite the success and popularity of HDDM, it suffers from several practical issues. First, the installation process of HDDM is cumbersome, exacerbated by its reliance on PyMC 2.3.8 for Markov chain Monte Carlo (MCMC) sampling, a package that is no longer supported and may clash with the latest computer modules. Second, and for the same reason, out-of-the-box HDDM is not compatible with Apple chips, which creates a significant barrier for Mac users. Third, although HDDM natively centers around Bayesian methods, it does not conveniently support all aspects of the evolved standards in Bayesian-modeling workflows (Ahn et al., 2017; Gelman et al., 2020; Kruschke, 2021). Significant progress has recently been made in supporting the principled Bayesian-modeling workflow in easy-to-use tool kits, such as the Python package ArviZ (Kumar et al., 2019). Bridging these new capabilities with HDDM facilitates a one-stop Bayesian-modeling pipeline for experimentalists and computational modelers interested in applying the DDM to their experimental data.
To address the above issues, we leveraged the Docker container technology to create dockerHDDM, a stable and complete virtualized Python computing environment that enables out-of-the-box implementations of Bayesian hierarchical DDMs. dockerHDDM has three major advantages (Table 2). First, it benefits from the easy-to-deploy nature of the Docker environment to avoid compatibility issues. Second, it is compatible with both Intel and Apple chips. Third, it augments HDDM with ArviZ, a Python module that enables a wide range of advanced Bayesian-modeling analyses. We expect dockerHDDM to provide an easy-to-use environment to help researchers across various backgrounds efficiently use DDM in their research.
Comparisons Between dockerHDDM and the Original HDDM Package

Note: HDI = high-density interval; ESS = effective sample size; LOO-CV = leave-one-out cross-validation; WAIC, widely applicable information criterion; PPC, posterior predictive checks.
Plotting, diagnosis, and model comparison are functions of ArviZ, including HDI, ESS, LOO, WAIC, and PPC.
How to Follow This Tutorial
The primary goal of this article is to present a practical guide to dockerHDDM for beginners with little modeling experience. In the tutorial, we start with step-by-step instructions on how to configure the dockerHDDM environment and how to use it in practical data analysis (Fig. 2). To assist reproducibility and easy application, a corresponding step-by-step video walk-through is available on YouTube at https://www.youtube.com/watch?v=ZU1fbXEuP8s or on OSF at https://osf.io/xz9m2.

dockerHDDM usage flowchart. The code in the figure is for demonstration purposes only. Specific instructions and copyable code can be found in the following corresponding sections. The top panel describes how to install Docker, corresponding to “Install Docker”; the middle panel describes how to pull and run dockerHDDM, corresponding to “Pull dockerHDDM Image” and “Run dockerHDDM Container”; and the bottom panel shows the workflow in dockerHDDM, corresponding to “Example of Workflow.” A video tutorial is available at https://www.youtube.com/watch?v=ZU1fbXEuP8s and https://osf.io/xz9m2.
Glossary of Terms Used in Bayesian Modeling
A
In the setup section (top panel in Fig. 2, corresponding to “Install Docker” section in this article), we provide instructions on how to install Docker. After that, we demonstrate how to obtain the dockerHDDM image and how to use this image to access the Jupyter notebook interface (middle panel in Fig. 2, corresponding to “Pull dockerHDDM Image” and “Run dockerHDDM Container” sections). Finally, within a working Jupyter notebook, we show how to analyze an example data set with dockerHDDM in a principled Bayesian workflow (bottom panel in Fig. 2, corresponding to “Example of Workflow” section).
Install and use dockerHDDM
Install Docker
Docker serves to create an all-in-one, fast, cross-platform computing environment. The Docker website provides easy-to-follow installation instructions (https://docs.docker.com/get-docker/) and supports Windows, MacOS, and Linux (see Box 2). Windows users should ensure their system version is 21H2 (build 19044) or higher and have either WSL or Hyper-V configured before installation (see https://docs.docker.com/desktop/install/windows-install/).
Basic Introduction to Docker
Docker is an open-source platform that automates the deployment, scaling, and management of applications. It achieves this through containerization, a process that packages an application and its dependencies into a single, portable, and consistent unit, known as a “container image.” Containers ensure that applications run reliably regardless of the environment (Peikert & Brandmaier, 2021; Wiebels & Moreau, 2021).
Docker uses a client-server architecture in which the Docker client communicates with the Docker daemon, responsible for building, running, and distributing containers. The core components of Docker are the Docker Engine, Docker Hub, and Docker Compose. The Docker Engine is the runtime that enables containerization, and Docker Hub is a cloud-based registry for sharing and managing container images. Docker Compose, on the other hand, is a tool for defining and running multicontainer Docker applications.
➢
After installing Docker Desktop (or Docker Engine for Linux users), one can verify the installation by running the following command in a terminal (Fig. 3). If the container starts and runs successfully, it will display a confirmation message and then exit (Fig. 3):

Command to check Docker installation in terminal. After running the command
Pull dockerHDDM image
After ensuring that Docker has been successfully installed and the Docker engine is running (Fig. 3), you can pull the dockerHDDM image by simply running the command in the terminal (see the meaning of each argument in Fig. 4a):
or
This command will pull the latest default version of dockerHDDM, which corresponds to the image with the tag

Docker commands to download and run dockerHDDM. (a) Download/pull dockerHDDM from the Docker hub. The command by default downloads the latest version of
Run dockerHDDM container
After pulling the Docker image to a local machine, you can start a computing environment by running the dockerHDDM image with the command in the terminal (Fig. 4b):

This command creates a Docker container, which is a specialized environment encapsulated within the Docker platform. The
After running the
In the Jupyter interface, you will find two files and two folders (Fig. 2, middle). The notebook dockerHDDM_Workflow.ipynb offers a detailed reproduction of the analyses presented in this article, which we discuss further in “Example of Workflow.” In contrast, the notebook dockerHDDM_Quick_View.ipynb provides a brief overview of the dockerHDDM image’s new features and an introduction to basic modeling processes. One folder is “work,” which mounts the local path into the docker environment. The other folder, “OfficialTutorials,” contains notebooks that reproduce the official tutorials available at https://hddm.readthedocs.io/en/latest/tutorials.html. Beginners can follow HDDM_Basic_Tutorial.ipynb to get a basic understanding of HDDM, as discussed in Wiecki et al. (2013); HDDM_Regression_Stimcoding.ipynb covers more advanced models with regression, in which parameters can vary based on experimental conditions and other covariates; Posterior_Predictive_Checks.ipynb introduces posterior predictive checks (PPCs), showing how to generate predicted data from fitted parameter posteriors and how to analyze these predicted data; LAN_Tutorial.ipynb introduces advanced use of LAN functions that address the problematic likelihood of more complicated models based on neural-network methods (Fengler et al., 2021).
Novel Features of dockerHDDM
The

For all models defined by methods such as
In dockerHDDM, we included five extra arguments in
To preserve compatibility and consistent output with origin HDDM, the arguments are configured with the following defaults:
The
Finally, the
Example of Workflow
In this section (Fig. 2, bottom panel), we demonstrate how to use dockerHDDM (i.e., HDDM and ArviZ) to perform key steps of Bayesian modeling (Gelman et al., 2020; Martin et al., 2024): model specification and fitting, model diagnosis, model comparison, PPC, and statistical inference. The code reproduced in this section can be found in dockerHDDM_Workflow.ipynb in the dockerHDDM environment.
Example Data
For convenience, we use the data from Cavanagh et al. (2011), which is built within HDDM, as an example to demonstrate how to implement the modeling workflow. This data set contains response time and choice data from 14 Parkinson’s patients (see Table 3). In the experiment, participants were asked to choose between two options associated with either high or low reward values (i.e., reward probabilities in typical reinforcement-learning tasks). The relative value differences between the two options define two levels of conflict: high conflict for low-low and high-high trials (“HC” in variable “conf”) and low conflict for low-high trials (“LC” in variable “conf”).
Example Data Set From Cavanagh et al. (2011)

Note: The data structure required for HDDM is long-format data, where each row represents one trial. “subj_idx” is the subject index, “rt” is the response time (in seconds), and “response” in this case represents the accuracy, where 1 is correct and 0 is incorrect. These three columns of data are mandatory when using HDDM and must be kept consistent with the column names and the units (rt, seconds). “conf” is an optional variable, corresponding to the conflict level, where “HC” denotes high conflict and “LC” denotes low conflict. “conf” is not a mandatory variable or column, meaning that different factor names and levels can be used depending on the experimental design. In addition, multiple variables may be maintained in the data, which may be categorical or continuous.
Note that HDDM requires the inclusion of three columns of variables, “subj_idx,” “rt,” and “response,” to construct the hierarchical model. This means that when analyzing your own data, these three columns of variables must appear in the data set with identical column names. In addition, the unit of “rt” must be seconds, and “response” is coded as 1 for the upper boundary of the corresponding choice and 0 for the lower boundary (for more details, see https://hddm.readthedocs.io/en/latest/howto.html).
Model Specification
As a demonstration of model specification, we test an example question: Is there an effect of conflict levels on drift rate (Wiecki et al., 2013). To answer the question, we constructed three computational models (see Table 4).
Models Used in This Tutorial

Note:
Model 0 served as the baseline without considering the effect of conflict level on the model parameters. The model contains the seven parameters, referred to as the full DDM, including the decision boundary (
By default, HDDM considers the hierarchical-modeling approach that includes parameters at both the individual and the group levels (see Box 3). Model 0 has 11 population-level parameters, including the means and the standard deviations for the four basic parameters (
Parameters in Hierarchical Drift-Diffusion Models
HDDM employs hierarchical Bayesian modeling by default, where each participant’s free parameters are sampled from population-level distributions (Wiecki et al., 2013). Taking full drift-diffusion model (DDM; Model 0) as an example, nondecision time
Note: The hierarchical structure of the full DDM in HDDM. The parameters inside and outside the rectangle are subject and population level parameters, respectively.
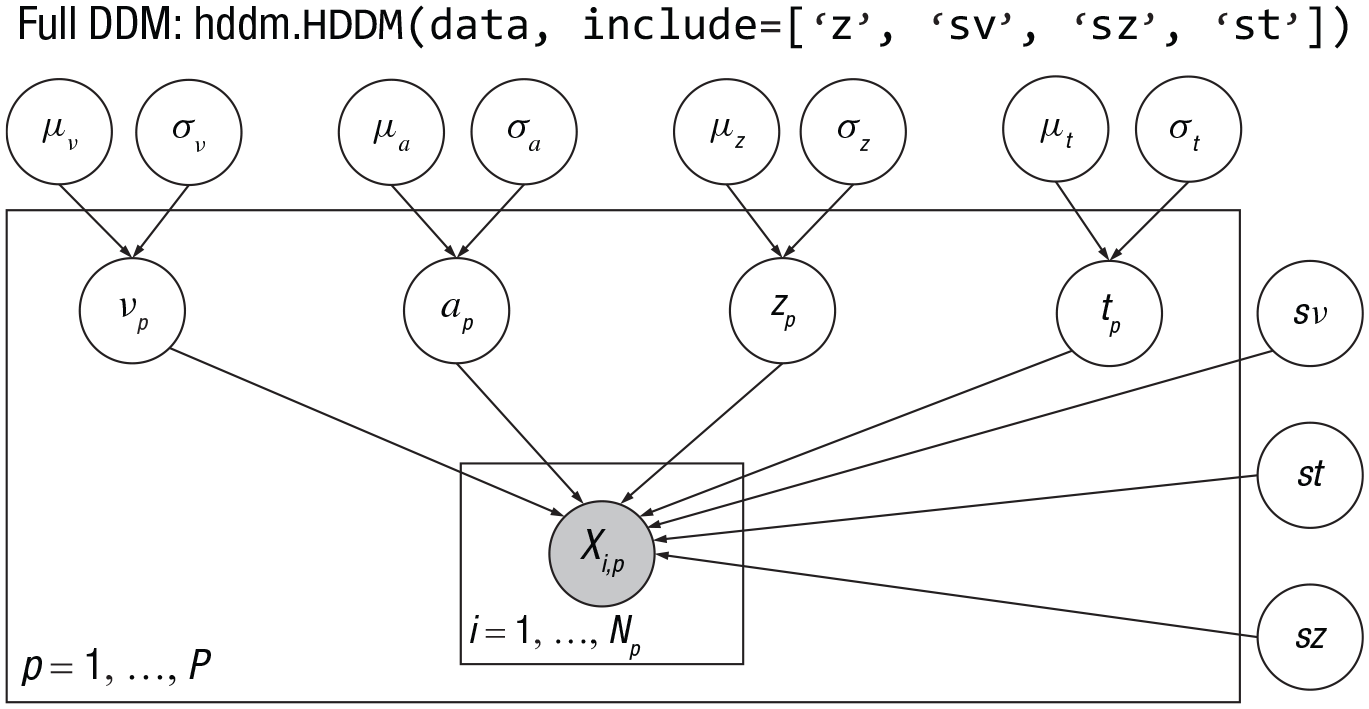
Consequently, there are a total of 11 population-level parameters. At the subject level, subjects have their own estimate of the parameter of a, v, t, z, leading to a total of
HDDM provides two types of priors: weakly informative priors and noninformative priors. By default, dockerHDDM uses weakly informative priors as summarized in the table below (Wiecki et al., 2013). The default informative priors are suitable for most perceptual tasks. However, for tasks with longer response times, it is recommended to use noninformative priors. In this case, one has to set the parameter

Note: Table extracted and refined from Wiecki et al. (2013).
HDDM also allows parameters to vary with variables by integrating hierarchical linear regression models (also called “linear mixed models” or “multilevel models”). Specifically, the
Although both the
Model 1 allows the drift rate to vary as a function of the conflict levels (i.e.,
Note that Model 1 assumes complete independence between high and low levels of conflict within subjects. This assumption may be inappropriate because it is likely that a person who responded relatively quickly in the “LC” condition will also respond relatively quickly in the “HC” condition and vice versa. For more detailed differences between Model 1 and Model 2, see Box 3.
Model 2 was constructed to include correlations between drift rates across conflicting levels. In Model 2, we use a hierarchical regression model with
Model fitting
The defined HDDM model allows the MCMC algorithm to be run using the

To accurately estimate parameters and ensure convergence in hierarchical modeling, we set up four MCMC chains of 10,000 samples with 5,000 burn-ins (i.e., a total of 20,000 samples for each parameter). For the more detailed settings and arguments description, see “Novel Features of dockerHDDM.” With the new functionality introduced by dockerHDDM, we can calculate the log-likelihood of the model and generate posterior predictions after model fitting. Furthermore, the output of the model fitting can be converted into InferenceData,
We emphasize that model fitting is demanding in terms of computational resources and memory. For example, in our tests with the Apple M1 chip, Intel i7-10700 CPU, and AMD Ryzen 9-5900HX, model fitting took around 2 hr to 3 hr for 10,000 samples. Consequently, fitting three models took about 6 hr to 9 hr, and memory usage ranged between 8 GB and 12 GB. In addition, if pointwise likelihood calculations (i.e., with the argument
Model diagnosis
In Bayesian inference, it is crucial to ensure the convergence of MCMC chains. With ArivZ, dockerHDDM supports both visual inspection and quantitative convergence checks (Martin et al., 2024, Chapter 10).
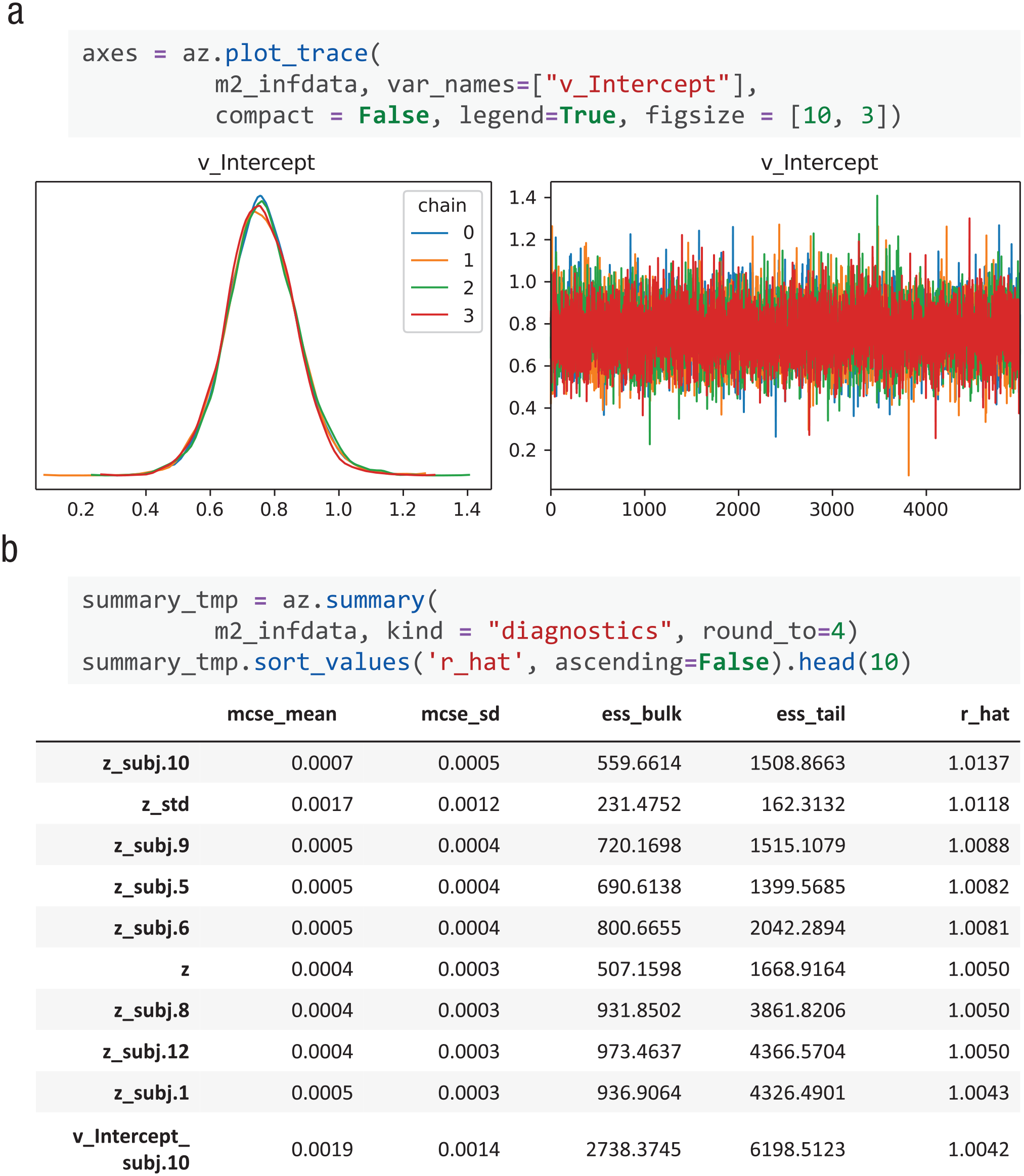
Model diagnosis. (a) Visualization of the traces of all chains using
The Gelman-Rubin statistics (
The latter two methods are covered by ArviZ’s
Model comparison
Upon verifying chain convergence, we proceed with model comparison to identify the best-fitting model. The evaluation metric provided in the original HDDM is deviance information criterion (Spiegelhalter et al., 2002). We include two more methods in dockerHDDM: widely applicable information criterion (WAIC; Watanabe, 2010) and Pareto-smoothed importance sampling leave-one-out cross-validation (PSIS-LOO-CV; Vehtari et al., 2017). These methods comprehensively integrate posterior samples for model comparison and evaluation (see Box 4).
Linking Deviance Information Criterion, Widely Applicable Information Criterion, and Pareto-Smoothed Importance Sampling Leave-One-Out Cross-Validation to Akaike Information Criterion
The deviance information criterion (DIC), widely applicable information criterion (WAIC), and Pareto-smoothed importance sampling leave-one-out cross-validation (PSIS-LOO-CV) are criteria founded on the concept of out-of-sample predictive accuracy, that is, the accuracy of using the fitted model to predict new data generated by the assumed data-generating process. Predictive accuracy is often encapsulated by the log predictive density (Box 1). However, the log predictive density approximated using the observed data and the posterior estimates of parameters is biased, and an adjustment is required to correct the bias. Thus, the key difference between DIC, WAIC, and PSIS-LOO-CV lies in the difference between the two terms of log predicted density and corrected bias (see the table below).

Note:
DIC uses the Bayesian posterior means for estimating log predictive density and includes an adjustment based on the effective number of parameters (
WAIC further refines DIC, evaluating the log predictive density across the entire posterior and correcting bias via the variability of log predictive density (
PSIS-LOO-CV estimates the predictive density by simulating the leave-one-out cross-validation, which by definition is the out-of-sample predictive accuracy, so bias correction is no longer needed for PSIS-LOO-CV. For more details on these three indices, see Gelman et al. (2014) and Vehtari et al. (2017).
For the demonstration, we compared three models across all three evaluation metrics (lower value is better). 5 As shown in Table 5, Model 2 exhibits the lowest values on all three metrics, indicating it is the best model. The results of model comparison revealed that Models 1 and 2 are much better than the baseline Model 0, suggesting that experimental conflict conditions have a substantial effect on drift rates. Moreover, Model 2 is slightly better than Model 1, suggesting that regression model may suit the data better. Nevertheless, the similarities between Model 1 and Model 2 suggest that both models fit the data adequately in this case.
Model Comparison With Different Criteria

Note: DIC = deviance information criterion; PSIS-LOO-CV = Pareto-smoothed importance sampling leave-one-out cross-validation; WAIC widely applicable information criterion; m0 = Model 0; m1 = Model 1; m2 = Model 2.
Rank is ranging from the best model to the worst.
Note that WAIC and PSIS-LOO-CV require the pointwise log-likelihood of each data point given a posterior sample of parameters, which must be computed using the likelihood function and posterior trace (see Box 1). This variable is not directly provided in the HDDM object and must be customized to be calculated via the likelihood function and posterior trace.
In dockerHDDM, the pointwise log-likelihood can be computed at the sampling and fitting stage, via

Finally, we note that the model-comparison metrics allow only a relative ranking of alternatives. To assess the absolute goodness of fit of the model, we recommend performing the PPC, as discussed in the next section, alongside the diagnostic information provided by LOO and WAIC (see Martin et al., 2024, Chapter 5; Vehtari et al., 2017).
PPC
In addition to model comparison, which assesses relative performance, the PPC evaluates how well predictive data generated from posterior samples of parameters align with the actual data. PPC is crucial because model comparison evaluates only the “least worst” model, but this model may not necessarily account for the data very well (see Martin et al., 2024, Chapter 5).
ArviZ offers convenient visualization tools for inspecting PPC (Kumar et al., 2019). The function

Posterior predictive check plot
Statistical inference
A final step in Bayesian modeling is to draw statistical inferences from the posterior parameter distributions in the best-fitting model. In our example, we test the hypothesis of whether drift rates significantly differ between HC and LC conditions based on Model2 (“m2” in the Notebook). This hypothesis is tested using the posterior samples of the regression coefficient in “m2,” which has a variable name “v_C(conf, Treatment(‘LC’))[T.HC]”.
Note that there are several acceptable methods for Bayesian hypothesis testing, such as BFs (Boehm et al., 2023; Wagenmakers et al., 2010), maximum a posteriori based p value (Mills, 2018), directional probabilities (Makowski et al., 2019), and the full Bayesian significance test (Kelter, 2022). In cognitive science and psychology, although BFs are often advocated as a Bayesian alternative to frequentist p values (Kelter, 2021; van de Schoot et al., 2017; Wagenmakers et al., 2010), debate remains about which Bayesian measures should be used in which settings of scientific hypothesis testing (Kelter, 2023; Makowski et al., 2019). Therefore, it is useful to consider various Bayesian hypothesis-testing methods depending on the study objectives and design (Kelter, 2023; Kruschke, 2021; Makowski et al., 2019).
Here, we demonstrate Bayesian inference using an approach that combines the approach combining highest density interval (HDI) and the region of practical equivalence (ROPE; Kruschke, 2018; see Box 1). In addition, we provide methods for calculating BFs in the Appendix.
We define a ROPE of [–0.2, 0.2] to represent values practically equivalent to zero
6
and use the

(a) Statistical inference of parameters. The high-density interval (HDI; black line and texts) is compared with the region of practical equivalence (ROPE; red line and text).
Therefore, considering the results from various aspects (model comparison, PPC, and posterior inference), we conclude that the model that takes into account the influence of conflict level on drift rate performs the best. Moreover, HC affects the cognitive process of decision-making by impeding the speed of evidence accumulation.
Discussion
In this tutorial, we focus on an easy-to-use computational environment for HDDM, including installation of the tool, its features, and case applications. Although some conceptual discussions have been addressed in other articles (Boag et al., 2024; Shinn et al., 2020; Voss et al., 2013), we nevertheless discuss some relevant issues below.
Why use dockerHDDM among tools?
Inference for the DDM can be implemented via multiple software/packages, such as fast-DM (Voss & Voss, 2007), flexDDM (LaFollette et al., 2024), rtdists (Singmann et al., 2022), EZ-DDM (Wagenmakers et al., 2007), and pyDDM (Shinn et al., 2020). For more details on tool and algorithm comparisons, see Shinn et al. (2020). Although all the above tools are estimated in a frequency framework and fit data at the individual-participant level, HDDM takes the Bayesian approach and estimates model parameters at both the individual and group levels (i.e., the hierarchical-model or multilevel-model approach; see Wiecki et al., 2013). Tools that also allow the Bayesian hierarchical modeling approach of DDM include brms based on RStan (Henrich et al., 2023), the Wiener module in JAGS (Wabersich & Vandekerckhove, 2014), EMC2 (Stevenson et al., 2024), and hBayesDM (Ahn et al., 2017). For comparison between these tools and HDDM, see Table 6.
Tools Comparison for Modeling Hierarchical DDM
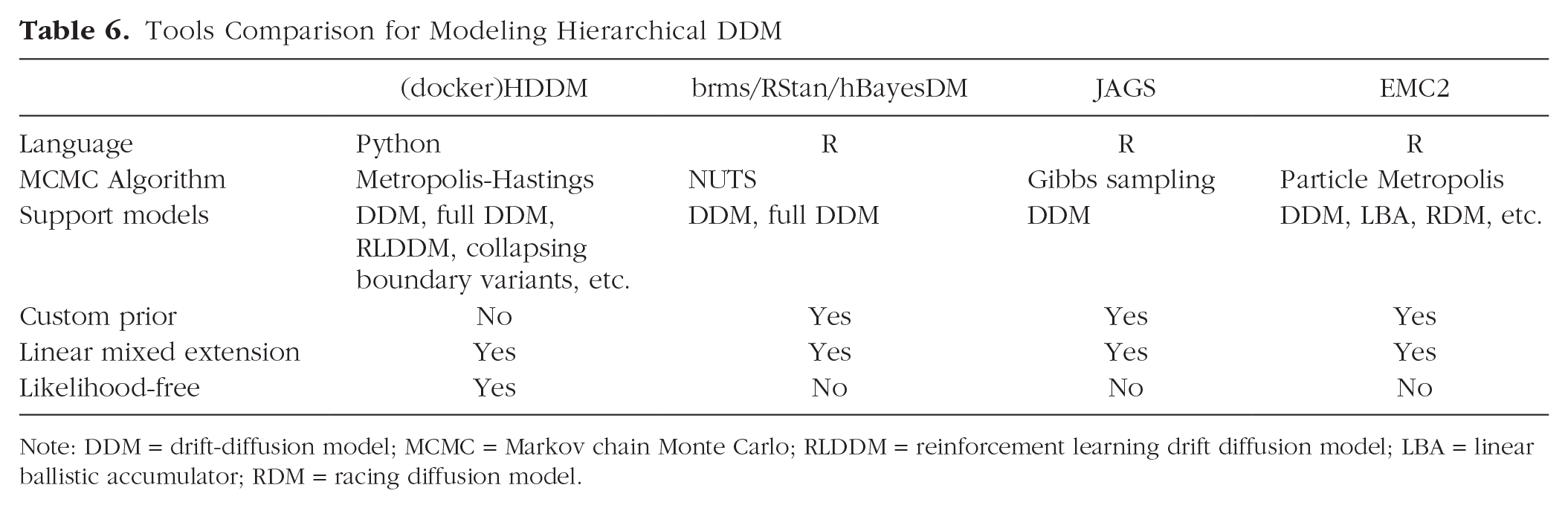
Note: DDM = drift-diffusion model; MCMC = Markov chain Monte Carlo; RLDDM = reinforcement learning drift diffusion model; LBA = linear ballistic accumulator; RDM = racing diffusion model.
HDDM stands out for its ease of use, enabling users to construct and fit basic models with just a few lines of code. It facilitates the definition of complex mixed-effects models without the need for prior specifications, making it more accessible for beginners. Although brms and EMC2 also define mixed-effects models well, they necessitate users to manually define prior distributions for random effects and covariance structures. In addition, RStan and JAGS require expertise in linear model reparameterization. The absence of this expertise may result in model-fitting failures or biased estimates. On the other hand, the simplicity of HDDM comes at the cost of flexibility because it restricts users to the default priors (see Box 3) and does not allow for customization. However, the weakly informative prior implemented in HDDM was based on previous meta-analyses of published results (Matzke & Wagenmakers, 2009) and applicable to typical cognitive experiments.
Another advantage of HDDM is its support for diverse accumulation models, including models with collapsing boundaries and those integrated with reinforcement learning, called “RLDDM” (Fengler et al., 2022; Pedersen & Frank, 2020; Pedersen et al., 2017). In addition, the latest version of HDDM provides many likelihood-free models, broadening its applications. For instance, its integration with neural networks, such as the LANs (likelihood approximation networks; Fengler et al., 2021), has greatly enhanced the efficiency of model design and development.
A notable limitation of dockerHDDM is its lack of integration with the most advanced parameter-estimation techniques. For instance, its successors, HSSM and EMC2, have begun incorporating advanced MCMC methods. Moreover, innovative neural-network approaches, such as LANs (Fengler et al., 2021), MNLE (Boelts et al., 2022), and Bayesflow (Radev et al., 2022), have the potential to significantly enhance these estimation procedures. However, the mastery of these cutting-edge techniques requires a higher level of expertise to prevent misuse.
Consequently, we propose that the mission of dockerHDDM should be to streamline operations and lower the barrier to entry, facilitating analogical learning and, ultimately, preparing users for the transition to the more sophisticated methods.
Whether to include parameters’ intertrial variability?
As a demonstration, we used the seven-parameter full DDM. If a user wishes to fit only the four-parameter model, the unnecessary parameters can be removed from the include argument, for example,
Consequently, the choice to include trial-by-trial variability requires a delicate balance between the prediction and complexity of the model and the specific requirements of the data. Given the extensive data requirements for inferring across-trial variability, our stance is to cautiously include across-trial variability in the model for a more robust fit and more precise inference of the basic parameters (see similar discussion in Boag et al., 2024). For instance, because the variability of the nondecision time tends to be easily recovered (e.g., the result of the parameter recovery in Appendix Figure S2), it may be prudent to include only this parameter but not the other variability parameters by default. Nevertheless, when the data set is substantial and the research objective prioritizes the analysis of specific response-time patterns, such as fast or slow errors, the selective integration (the parameter variability of drift and start point; also see Table 1) of these parameters may be warranted. We recommend reading the work by Boehm et al. (2018), which offers expert advice and recommendations on estimating across-trial variability parameters.
Data quantity and quality for fitting the DDM
Both the number of subjects and the number of trials should be considered. Because of the hierarchical nature of the model, hierarchical models typically require fewer trials than nonhierarchical models (Alexandrowicz & Gula, 2020; Wiecki et al., 2013). In general, 12 subjects are sufficient to obtain stable results (Wiecki et al., 2013), but we recommend collecting data from more than 20 subjects for a more robust fit. However, the number of sufficient trials varies depending on the parameters of interest. For the basic four-parameter model, the number of trials has a small effect on parameter estimates (Alexandrowicz & Gula, 2020). Twenty trials appears to be the minimum standard, and more than 50 trials tend to produce robust results (Wiecki et al., 2013). Estimates of
Note that parameter estimation can be affected by extreme values, such as very fast response times. HDDM addresses this issue by assuming a mixture model in which a proportion of the response times are from a uniform distribution (Ratcliff & Tuerlinckx, 2002; Wiecki et al., 2013). The proportion of response times is controlled by the parameter
Finally, it is essential to conduct PPCs to validate the model (see “PPC”). These checks help to ensure that the model is capable of accurately reproducing the observed data, thus providing confidence in the evaluation of the model and parameters.
Computational resources and tips
To achieve accurate estimates, more subjects, more trials, and often more samples are required, leading to increased demands for computational resources. This is not unique to dockerHDDM; other tools using MCMC algorithms, such as DMC and brms mentioned earlier, are also affected by these factors. In the examples provided in this article, fitting each model with 14 subjects and 3,988 trials takes 2 hr to 3 hr and requires 8 GB to 12 GB of memory. Running out of memory can cause the Jupyter kernel to suspend and restart, interrupting the process. Predictably, computational resources become a limiting factor with increasing data. To facilitate better model analysis, we offer the following tips and recommendations.
Initial testing
When initially building the model, use subset data from a small number of subjects and reduce the MCMC sample size to verify that the model definition and code are correct. Once validated, increase the data and sample sizes.
Adjust memory settings
If users experience a Jupyter kernel suspension or restart because of memory constraints, they can attempt to configure or increase virtual memory. For Windows users, it is necessary to check and remove the memory-usage limitations imposed by WSL (Windows Subsystem for Linux).
Separate execution
Model fitting, calculation of point-wise log likelihood, and generation of PPCs data can be executed separately. This approach helps prevent interrupting long-running processes because of errors and ensures that each step can be independently validated and debugged before proceeding to the next.
Notebook segmentation
Fit models into separate notebooks to reduce the resource load of loading multiple models.
Model saving
Save the fitted models and then load only the InferenceData files instead of the entire models to reduce resource usage.
Cloud deployment
Docker is easily deployed in cloud-computing environments (or use the docker image in Singularity). Use your institution’s computing services or rent cloud computing services to handle larger data sets.
Summary
In this article, we introduce dockerHDDM, a user-friendly, out-of-the-box, and one-stop Docker image for implementing HDDM analysis within a modern Bayesian hierarchical workflow. Our dockerHDDM has three major advantages: (a) It leverages Docker to solve compatibility issues and simplify the installation process, (b) it ensures broad support across different machines equipped with either Intel or Apple chips, and (c) it integrates state-of-the-art Bayesian modeling practices with ArviZ, facilitating a more principled Bayesian workflow. We also provide a step-by-step video tutorial on how to use dockerHDDM.
Although we have provided a step-by-step guide to using dockerHDDM, it is unfortunately not possible to provide a comprehensive introduction to computational modeling. Given the extensive knowledge required for principled computational modeling, we recommend readers refer to the materials in Box 5 for a deeper understanding of the DDM family, computational modeling, hierarchical models, and Bayesian modeling. We expect that dockerHDDM and this detailed tutorial will reduce the technical burden and help readers get started with computational modeling. Ultimately, we hope that this tool and the computational-modeling concepts presented in the tutorial will promote the computational reproducibility of drift-diffusion modeling for users of all levels of computational expertise.
Recommendation for Further Reading
A full understanding of how Bayesian hierarchical drift-diffusion modeling works requires not only basic knowledge of drift-diffusion modeling but also knowledge of Python programming, Bayesian statistics, and hierarchical regression models. This background knowledge is generally not part of the coursework in psychology or neuroscience education, although the situation has been changing in recent years. We recommend the following resources to quickly catch up and avoid misuse or abuse of hierarchical drift-diffusion modeling.

Footnotes
Appendix
Acknowledgements
Thanks to HDDM (Wiecki et al., 2013; Fengler et al., 2021; Fengler et al., 2022) and ArviZ (Kumar et al., 2019) for the open resource. We thank Dr. Mads Lund Pedersen for his open dockerfile, which insipre the current project. We also thank the netizens for their time in testing and valuable feedback, which allows us to continuously improve the tools and tutorials. We appreciate the help of Dr. Yuan Rui in the early stage of docker image development.
Transparency
Action Editor: Rogier Kievit
Editor: David A. Sbarra
Author Contributions
