Abstract
The Purdue Make It system is a unique automated platform capable of small-scale in situ synthesis, screening small-molecule reactions, and performing direct label-free bioassays. The platform is based on desorption electrospray ionization (DESI), an ambient ionization method that allows for minimal sample workup and is capable of accelerating reactions in secondary droplets, thus conferring unique advantages compared with other high-throughput screening technologies. By combining DESI with liquid handling robotics, the system achieves throughputs of more than 1 sample/s, handling up to 6144 samples in a single run. As little as 100 fmol/spot of analyte is required to perform both initial analysis by mass spectrometry (MS) and further MSn structural characterization. The data obtained are processed using custom software so that results are easily visualized as interactive heatmaps of reaction plates based on the peak intensities of
Keywords
Introduction
High-throughput screening (HTS) technologies have become increasingly relevant both for industrial pharmaceutical lead discovery and for basic and applied organic synthesis and biomedical research in academic settings.1–3 Although some have argued that HTS is partially responsible for a decline in productivity in the pharmaceutical industry due to supposed inherent shortcomings of the technique, 1 the consistent stream of improvements to high-throughput analytical technologies has enabled researchers to acquire unprecedented amounts of high-quality data while enabling creative new experimentation. 4
High-throughput experiments originally entailed the screening of large chemical libraries against biological targets of interest to determine any modification in the target response through highly automated sample handling processes and instrumentation, miniaturized assays in terms of sample consumption and analysis time, and rapid data analysis. 5 HTS technologies have also been of increasing importance for reaction screening, wherein a broad and diverse array of chemical reaction variables (choice of reagents, catalyst, pH, solvent, stoichiometry, concentration, temperature, etc.) are rapidly screened to locate optimal conditions.6,7 The successful hits from reaction screening can then be used in bioactivity screening and further investigation.8–13
Traditional HTS methods for bioassay screening are based on optical detection techniques, such as colorimetry,14–17 fluorescence,18–25 radiometry,26–28 and luminescence.29–31 Most of these techniques accommodate high-density plates of up to 1536 wells and allow for sensitive and continuous detection with minimal analysis time (analysis is performed throughout incubation in most cases) via the use of commercially available plate readers. However, the fact that most biological systems do not involve species with distinctive optical properties is an intrinsic challenge for these techniques. As a result, the direct application of these methods is limited. In most cases, new assays must be specifically developed for each biological system. Common strategies to develop these optically based assays involve the use of radiolabeled compounds,26–28 nonnative substrates,25,32 and/or coupled reactions,24,33 all of which can in turn introduce interferences into the assay as optically based approaches suffer from low chemical specificity. Furthermore, these modifications to the system under study might change its behavior compared with the native state,33,34 as well as introduce safety concerns (e.g., radioactive material handling) and higher costs per assay.
In terms of chemical reaction screening, UV-visible spectrophotometry has long been incorporated into both flow-based and plate-based reactors for reaction monitoring and kinetic determinations,35–38 yet its role is limited, as other analytical techniques are still necessary for accurate product identification and quantitation. Fluorescence-based methods have also been developed for reaction monitoring, and they are particularly powerful in the case of direct asymmetric synthesis, where the use of chiral species to form fluorescent diastereomeric complexes with analytes allows for high-throughput determination of enantiomeric purity. 39
Alternatively, mass spectrometry (MS)-based systems are naturally label-free and have significantly higher molecular specificity in comparison with optical methodologies. This provides them with remarkable versatility and allows their use in the screening of a variety of chemical and biological reaction systems. Consequently, MS has become increasingly popular as the analytical platform of choice for HTS.40–42 Several MS-based platforms have been developed for this purpose, starting with well-known liquid chromatography (LC)–MS systems. The obvious trade-off of these platforms is speed, as LC-MS is significantly slower in comparison with optical detection techniques, given that the chromatographic separation is a bottleneck.40,41 Even with increased duty cycles, achieved by the use of multiple LC columns, clever scheduling of injection, flow diversion, and detection times (e.g., Thermo Scientific Transcend), LC-MS systems are inherently slow, requiring at least 10 s/sample.41,43,44 Novel LC sampling approaches that allow for mixing and monitoring of a target species, such as MISER and mixed-MISER, 45 have been incorporated in high-throughput systems, decreasing the analysis time to ~7 s/sample when analyzing 1536-density arrays. 46 Nonetheless, there is still a large gap in both the timescale and cost with respect to optically based techniques.
Higher throughputs can be readily achieved with LC-free MS-based platforms. Matrix effects and ionization suppression are inherent in these methods as a trade-off for eliminating the sample separation step, but novel ionization methods as well as advances in MS instrumentation have improved the performance of these LC-free methodologies so that they can be easily implemented for HTS without sacrifice in data quality.41,42 These LC-free MS-based platforms can be classified into four major categories: 41
Laser-based techniques, such as matrix-assisted laser desorption ionization (MALDI), which are well-established methods that provide high throughputs (>1 sample/s) but require time-consuming multistep plate preparation and for ionization to be performed under vacuum.47–49 A variant of MALDI, self-assembled monolayers for MALDI, uses monolayers to allow for precise control of surface chemistry to increase the sensitivity of MALDI to analytes of interest.50–52
Solid-phase extraction (SPE)-based platforms (e.g., Agilent RapidFire), which provide online enrichment and sample cleanup, reducing ionization suppression and matrix effects but increasing analysis time (~8 s/sample).13,41,42,53–55
Droplet-based systems, which are typically based on acoustic or microfluidic droplet generation and maneuvering. The latest iteration of acoustic-based systems (e.g., Sciex Echo MS) uses acoustic desorption ejection coupled to an open port interface (ADE-OPI), allowing for analysis times faster than 0.3 s/sample with special considerations regarding the choice of solvent.56–60 On the other hand, microfluidic-based approaches include mass-activated droplet sorting, a method based on the accurate sorting of split droplets according to results of parallel electrospray ionization (ESI)–MS screening with a throughput of ~1.5 s/sample. 61 Note that both of these techniques still suffer the drawbacks from direct ESI regarding ionization suppression and matrix effects, which, in the case of the ADE-OPI systems, are simply reduced through a 1000-fold online dilution.41,60
Desorption-based techniques, mainly represented by desorption ESI (DESI), the core of Purdue’s high-throughput Make It system. DESI provides contactless, ambient, fast (current throughput: >1 samples/s), and direct sample analysis, with great matrix tolerance and no requirement for sample preparation.62–64 Additionally, by relying on microdroplets, DESI allows reaction acceleration65–67 and in situ nanoscale synthesis, a feature unique to this platform compared with other HTS technologies.
Purdue’s high-throughput DESI-MS system has been developed over the last 5 years by an interdisciplinary team funded by the Defense Advanced Research Project (DARPA) as part of the agency’s Make It program, with the objective of automating small-molecule discovery and synthesis by efficiently exploring large domains of possible reaction types and conditions.68,69 The development and application of this high-throughput platform has resulted in several dozen peer-reviewed publications. The system has been successfully used to study multiple organic reactions through HTS campaigns. The reactions include N-alkylation,
70
Here we seek to illustrate the utility and broad applicability of Purdue’s Make It platform by describing two new experiments in the context of earlier work. The first is a combinatorial synthesis and reaction screening campaign, where 188 unique reactions are rapidly explored without a prior reaction period. The second is a label-free study of the nicotinamide
Experimental Methods
High-Throughput DESI-MS System Overview
Purdue’s high-throughput DESI-MS system is an automated platform developed initially for small-molecule discovery and synthesis. Its main goal was to extend knowledge of small-molecule organic synthesis beyond that available from conventional batch-based, intuition-driven experiments. Throughout its development (since 2015), multiple advances and iterations have taken place, yielding the synthesis, screening, and (bio)analysis platform that we describe.
The core of the high-throughput system is DESI, an ambient ionization technique. As with all other methods within this family, DESI creates ions directly in the ambient environment, with minimal or no requirement for sample workup.74,75 Additionally, DESI provides noncontact analysis (no capillaries carrying sample that can clog and also no carryover) 41 as well as a high tolerance to complex matrices, including those typically not MS compatible.62,76 DESI uses pneumatically driven charged microdroplets of solvent to impact the sample and transfer dissolved compounds in secondary splashed microdroplets to a mass spectrometer, where they produce ions representative of the sample.74,77 In cases where the sample contains reactive molecules and the conditions are appropriate, a reaction can occur before mass analysis, allowing observation of reaction products generated during analysis. Moreover, in the secondary microdroplets generated by the DESI spray, reactions can be accelerated by orders of magnitude,65–67 allowing rapid screening of reaction mixtures without a prior reaction period. The DESI-MS platform can thus be used both as an analytical tool, for screening reactions already carried out, and as a synthesis-analysis system in which the input mixtures of reactants undergo accelerated reactions in the microdroplets used for DESI analysis. In both cases, reactants, products, intermediates, and by-products can be rapidly analyzed by MS, or characterized further by MSn experiments. The main output of these screening experiments is typically a yes/no answer to the question of whether or not a specific reaction generates an expected product under certain conditions, allowing for optimization of synthetic conditions and providing a starting point for scale-up.6,7,63,70,71 Typically, no quantitative data are sought through organic reaction screening, although MS information can be readily utilized to assess compound purity.
The same platform as that used as a reaction screening tool can be adapted to perform bioassays, an application that requires robust methods with good quantitative performance. In this case, samples are quenched assays that are analyzed directly without workup, while typical analytes include the substrate and product of the enzyme-catalyzed reaction. DESI-MS-based bioassays are consequently discontinuous (as are most MS-based assays, opposed to traditional colorimetric/fluorometric methods),41,62 which implies that reactions must be quenched at precise endpoints for kinetic studies. Note, however, that quenching is a simple single step (e.g., addition of an organic solvent or a chaotropic agent) that requires no significant increase in the overall analysis time. Furthermore, the use of quenched samples allows for storage of the aliquots and reexamination in a confirmatory analysis, if necessary.
Regardless of the type of application, automated HTS using the DESI-MS platform involves three main steps: sample preparation and spotting, DESI-MS analysis, and data processing ( Step 1: Samples are added to a 384-well plate (master plate) aided by a robotic liquid handling system (Biomek i7; Beckman Coulter, Brea, CA). Note that these samples can be reaction mixtures prepared directly on the plate by addition of individual reactant stock solutions with no bulk incubation, or they can be previously incubated reaction mixtures. Samples can also be bioassay mixtures, after a reaction that is performed directly on the plate or previously prepared and quenched. With the master plate ready, 50 nL samples are spotted onto a DESI plate, using a slotted 384-pin tool (V&P Scientific, San Diego, CA). DESI plates are prepared in-house using customized soda-lime glass slides (Abrisa Technologies, Santa Paula, CA) with the same footprint of a standard well plate. A porous polytetrafluoroethylene (PTFE) membrane (Zytex G-115; Saint-Gobain, Wayne, NJ) is attached to each slide using a light film of spray adhesive (Scotch Spray Mount; 3M, St. Paul, MN). The Biomek robot allows pinning with 50 µm positional accuracy at densities up to 6144 reaction spots per plate. The high density of spots is achieved through shifting the pinning position of the 384-pin tool on the slide; up to 16 programmed shifts are available to users of the Biomek robot, which results in nominal center-to-center distances of 1.125 mm between spots at the highest density. The pinning is versatile and user-defined, meaning that it is not necessary to pin or analyze all positions. Note that as many as 16 independent master plates can be pinned on a single DESI plate (i.e., 6144 unique samples). DESI plates can also be pinned with the same sample in multiple positions to allow for running of instrumental replicates. For all experiments, three dye marks are also pinned at predefined positions in the corners of the DESI plate. These dye spots are used to calibrate the positions of the spotted samples. Step 2: The DESI plate is automatically transferred, using a Selective Compliance Articulated Robot Arm (SCARA) (PF3400; Precise Automation, San Diego, CA), to the DESI stage. Prior to analysis, through use of the dye points on three corners of the plate, a calibration is performed to allow positions on the plate to be recognized with sufficient precision that one can return to any particular spot if confirmation requires. The calibration process is simple, involving only the aligning of the DESI spray with the dye marks. After this, the user can select the pinning position(s) to be analyzed (from the 16 possible) and the instrumental method to be used for the analysis. This latter option provides great versatility, considering that the user can define any MS experiment type available with the chosen mass spectrometer. Next, the analysis starts. During analysis, the DESI stage is moved underneath the DESI sprayer in a spot-to-spot fashion (i.e., jumping from one spot to the next), saving scanning time, especially if low densities are pinned. At least 0.3 s is spent on each spot (effective analysis time), with a variable dead time (used for moving of the stage) of less than 0.7 s/sample. This dead time per spot is reduced as higher densities are analyzed. The DESI solvent used in the analysis can be selected and even changed on the fly by means of a 16-channel high-precision piezoelectric solvent delivery system.
78
When data acquisition is complete, the DESI plate can be (1) discarded, (2) stored for later reanalysis, or (3) immediately reanalyzed (e.g., to obtain MSn information from spots of interest). This latter capability is remarkable considering that only 50 nL of each sample is transferred to each spot on the DESI plate. Under typical conditions, with concentrations in the order of micromolar to millimolar,
79
this represents only a few nanograms of material. The low sample consumption also allows for storage of the master plates used for preparing the samples, so that further or confirmatory analyses can also be performed by spotting new slides. Step 3: The spectra obtained are processed using CHRIS, which extracts peak intensities associated with
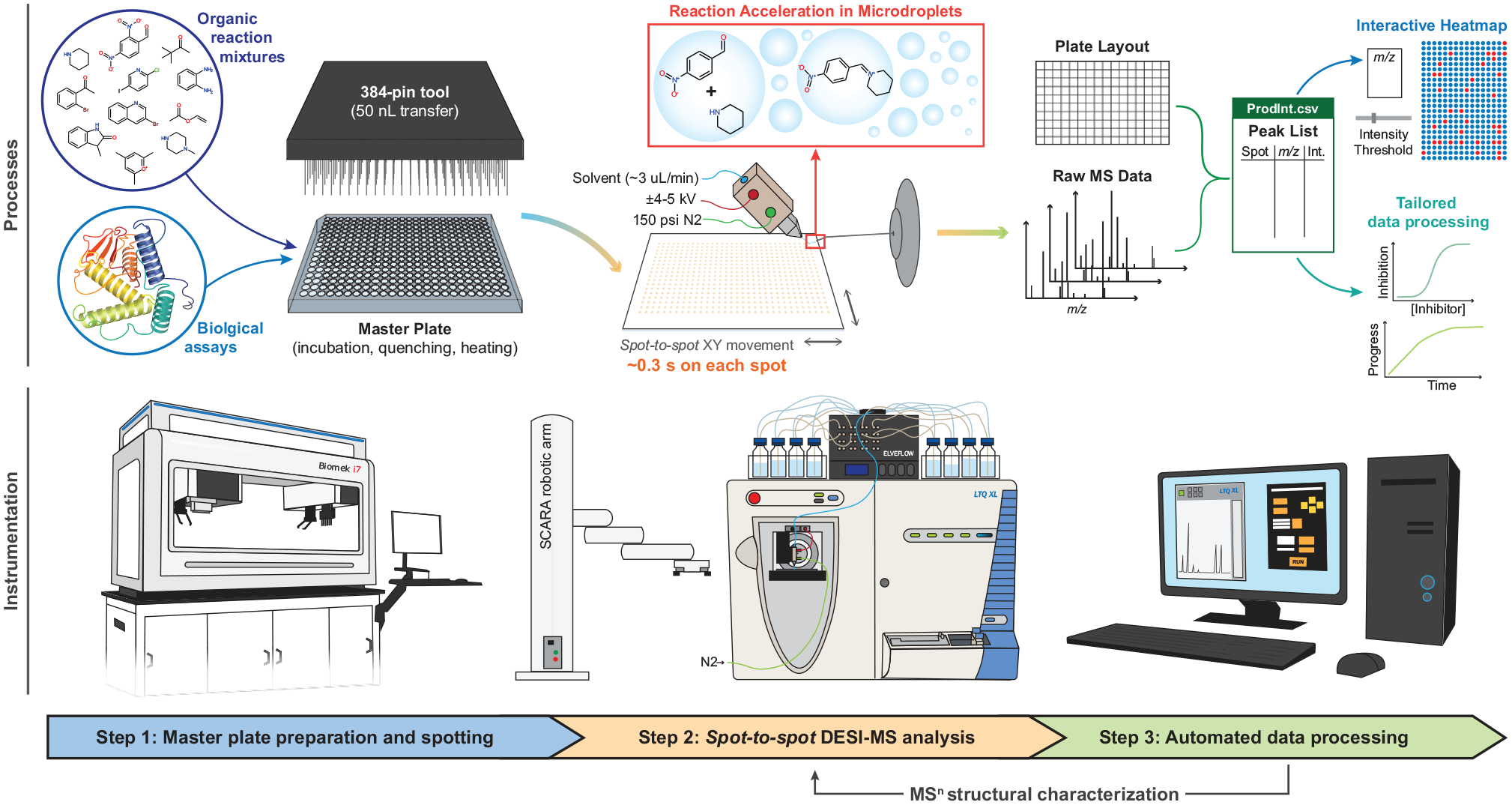
General workflow using the high-throughput DESI-MS platform. Three main steps are followed regardless of the particular characteristics of an experiment: (1) prepare one or more 384-well master plates and spot samples of interest onto a DESI analysis plate, (2) perform DESI-MS analysis directly from the spotted DESI plate, and (3) process the raw data obtained using custom software. Both the instrumentation and the detailed processes involved in each step are presented in the figure.
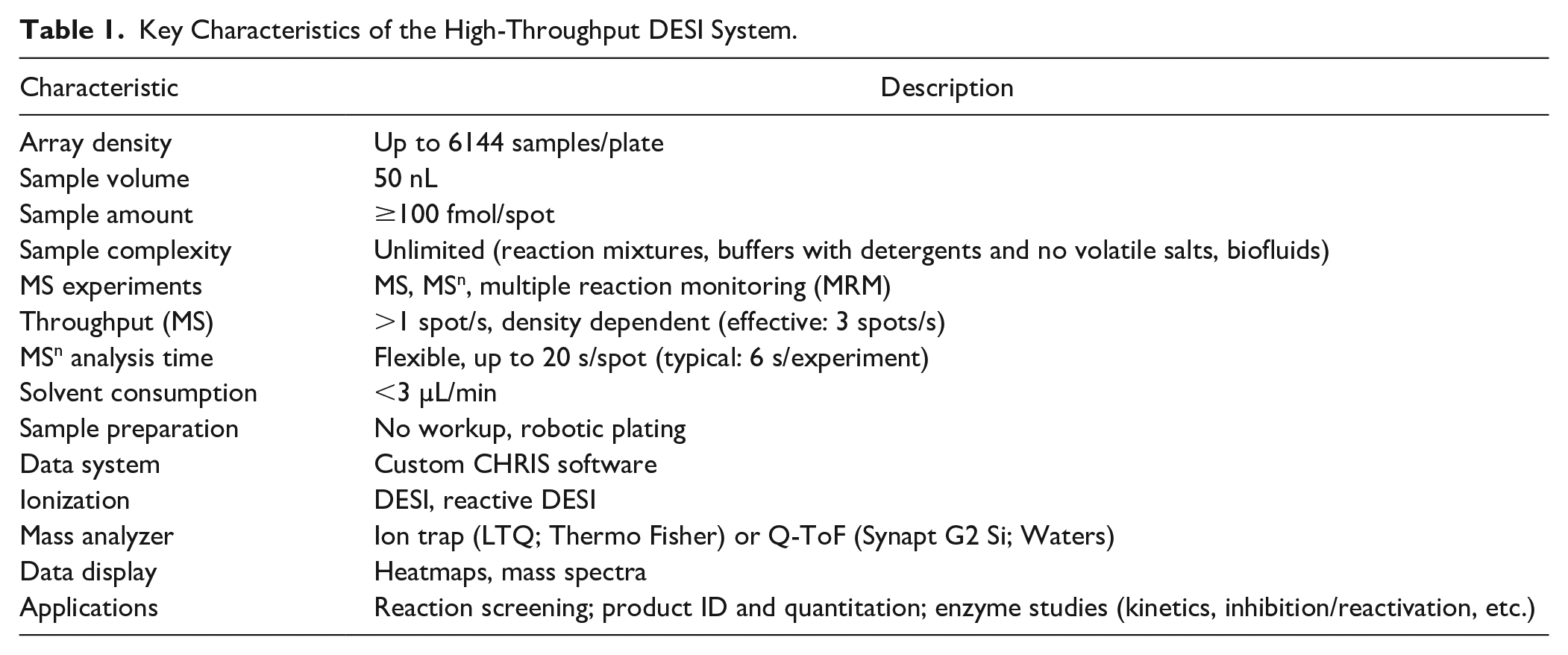
The main characteristics of the system are summarized in Table 1 .
Key Characteristics of the High-Throughput DESI System.

Versatile Combinatorial Synthesis and Reaction Acceleration
To demonstrate the versatility of the high-throughput synthesis and screening capabilities of the system, a total of 24 types of reactions were screened: reductive amination, nucleophilic aromatic substitution (SNAr), Suzuki–Miyaura coupling, amine alkylation, amine acylation, amine arylation, Katritzky transamination, Claisen–Schmidt condensation, Friedel–Crafts alkylation, Friedel–Crafts acylation, exhaustive methylation followed by Hofmann elimination, Ugi reaction, Schiff base reaction, Biginelli reaction, Hantzsch pyridine synthesis, Malaprade oxidation, Baeyer–Villiger oxidation, Fischer esterification, Diels–Alder reaction, Sonagashira coupling, benzimidazole formation, Negishi coupling, alkyne cycloaddition, 80 and Buchwald–Hartwig amination. For each of these reaction types, between 2 and 16 combinations of several variables (reactants, catalysts, and solvents) were selected for analysis, yielding a total of 188 unique reactions (24 reaction types, 8 replicates per sample, 1500 samples in total) to be analyzed in a single campaign. Specific reactants, solvents, conditions, and catalysts for each reaction are included in the Supplemental Material.
The final concentration of each reactant in the reaction mixture was 50 mM, whereas the concentrations of catalysts, if used, were fixed at 10 mol%. All the reactant and catalyst stock solutions were prepared separately (in independent 384-well plates) and only mixed in a master plate immediately before pinning. This was done to minimize the reaction time in bulk. All the plating and mixing steps were carried out using the automated liquid handling robot. Manual work was limited to the preparation of stock solutions. Duplicate mixtures were prepared for each reaction, and four replicate spots of each mixture were transferred onto a DESI plate and immediately analyzed. The total time between initial mixing and full MS analysis was less than 2 min. MS analysis of ~1500 samples was completed in 25 min. Further MSn analysis was later carried out using the same spotted plates. Replicate spots were used when performing multiple MSn experiments. All analyses were carried out using a Prosolia DESI 2D stage (Zionsville, IN) and a Thermo Fisher LTQ XL ion trap (Waltham, MA) in the positive ion mode using as DESI solvent methanol (2.75 µL/min) and a spray voltage of 4 kV. The transfer capillary temperature was set at 300 °C, while the capillary and tube lens voltages were fixed at 14 V and 47 V, respectively. The DESI impact angle was 55°. The nitrogen pressure for the DESI spray was set at 150 psi. Collision-induced dissociation (CID), with helium as collision gas, was used for MSn experiments.
NNMT Activity Assay
NNMT activity was assessed using conditions previously utilized in the literature with regard to buffer (5 mM Tris pH 8.6, 1 mM DTT), incubation temperature (37 °C), and final concentrations of enzyme (human NNMT, 100 nM),
Reaction monitoring was carried out discontinuously for 3 h, whereas endpoint assays were performed after incubation for 1 h. Quenching was achieved by the addition of a reaction volume of acetonitrile containing 1% formic acid. The quenching solvent also included neostigmine (10 µM), chosen as the internal standard. No sample preparation was required prior to analysis. Assay mixtures were pinned directly after quenching from a 384-well plate onto a DESI slide. MS analysis was performed using a Prosolia DESI 2D stage and a Thermo Fisher LTQ XL ion trap operating in full-scan MS-positive mode. The DESI solvent used was methanol with 0.1% formic acid (2.75 µL/min), and the spray voltage was 4 kV. The transfer capillary temperature was set at 300 °C and its voltage at 38 V. The tube lens voltage was 68 V. The DESI impact angle was fixed at 55°, and the nitrogen pressure for the spray was kept at 150 psi.
Results and Discussion
Versatile Combinatorial Synthesis and Reaction Acceleration
A unique capability of the DESI-MS high-throughput system arises from the reaction acceleration phenomenon that occurs in small-volume microdroplets, such as those originating during nano-ESI, DESI, or electrosonic spray ionization.
67
This acceleration phenomenon has been linked to the unique conditions at the droplet–air interface (reaction rates increase strongly with decreasing droplet size), and not simply to the increased concentration of reagents in evaporating droplets.65,67,86–88 Furthermore, it has been demonstrated that simply collecting the droplets sprayed from a reaction mixture allows the collection of milligram amounts of reaction product.
89
This capability confers a clear advantage for reaction screening: considerable product yields (obtained through a microdroplet-accelerated reaction) can be obtained with little to no incubation time prior to analysis. This is exemplified in the results of the 188 combinatorial reactions on a single plate screened using microdroplet-accelerated reactions with no bulk incubation time (
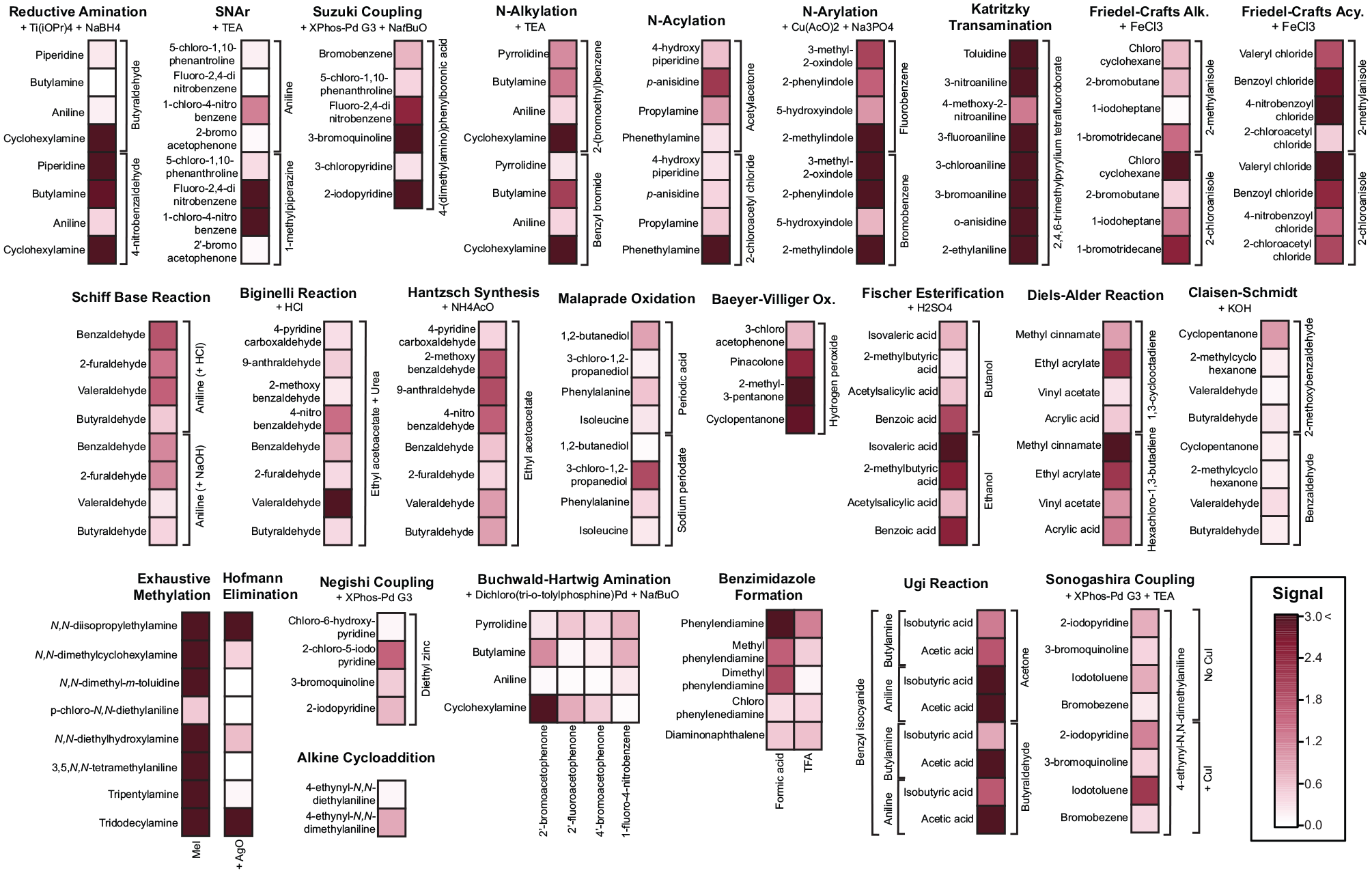
Segmented heatmap obtained through the screening of 24 reaction types, with 188 unique reactions. Eight replicates were run for all reactions. The results represent averages across all replicates. Response corresponds to signal in mass spectrum expressed relative to the average blank response.
Some representative spectra from diverse successful reactions are shown in
Figure 3
and
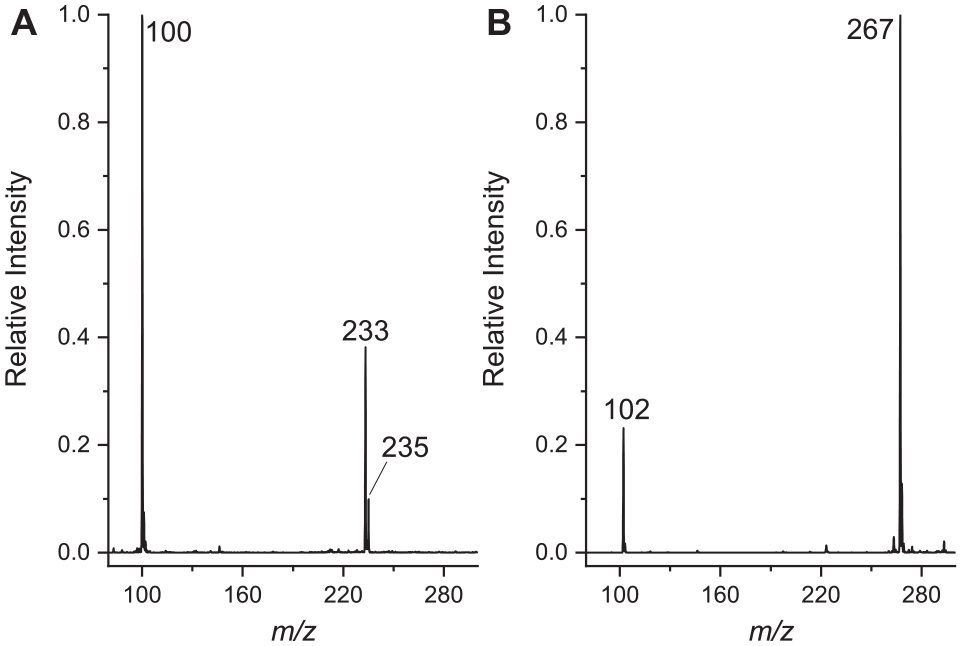
Representative MS spectra from successful reactions. (
Within each reaction type, the reactivity trend of the individual reagent combinations reflects the expected behavior of the chemical species, indicating that the system is reliable in capturing the chemical reactivity. For instance, in the case of reductive amination, 4-nitrobenzaldehyde shows a higher reactivity than butyraldehyde, owing to the nitro electron-withdrawing group, which favors nucleophilic attack.
6
Similarly, aliphatic amines, as expected, are more reactive compared with aromatic amines. Another clear example is the unsuccessful reaction of aniline with 4-nitrobenzaldehyde, and this effect also explains why the imine formation reactions, all using aniline, were unsuccessful. Note that the same trend is observed in the SNAr reactions, where 1-methylpiperazine was more reactive than aniline. This agrees well with the expected trend that product formation in SNAr is favored by amines with electron-donating groups and aryl halides with electron-withdrawing groups (compare the cases of the nitro substituted aryl halides with, for instance, 5-chloro-1,10-phenanthroline).
7
A further example of predictable structure–reactivity relationships is found in the
Further examination of the full MS spectra of individual reactions provides insight into the causes of unsuccessful reactions. For example, in the case of Hofmann elimination, several reactions yielded no elimination product, but upon inspection of the MS spectra, the presence of a strong signal for the reaction intermediate was noted. These intermediates are simply the result of exhaustive methylation with methyl iodide—the first step in the Hofmann elimination reaction pathway. Note that two of the substrates (both
MSn Characterization of Products, Intermediates, and Side Products
The high-throughput DESI-MS platform, when coupled to an ion trap mass spectrometer, allows ready product characterization through MSn experiments. The capability of performing full high-throughput MS screening followed by MS2 characterization has been reported previously, 6 but here we extend it by performing MS3 and MS4 experiments directly on the DESI samples. The only software modification required is a change in the method file to which CHRIS refers when performing the analysis. Single raw files with the MSn data are obtained for each spot analyzed. These files can be analyzed manually or automatically parsed using MSConvert 96 and custom MATLAB scripts. Due to the limited amount (low nanograms) of sample at each spot, a maximum of two stages of the MSn experiment can be performed while still obtaining high-quality spectra. This means that MS2 and MS3 experiments can be performed on the same spot, but the MS4 analysis must be performed in a replicate spot. Reasonably, this requirement depends on the concentration of the sample and the acquisition time of each experiment. In the data shown, acquisition times were limited to 7 s/experiment using a single normalized collision energy (25 arbitrary units). Note the order of magnitude greater time required for MS/MS versus single-stage MS.
We assessed the MSn analysis capability by evaluating in some detail two interesting cases found in the reaction plate screened. The first is the reductive amination reaction between 4-nitrobenzaldehyde and piperidine to form 1-(4-nitrobenzyl)piperidine (
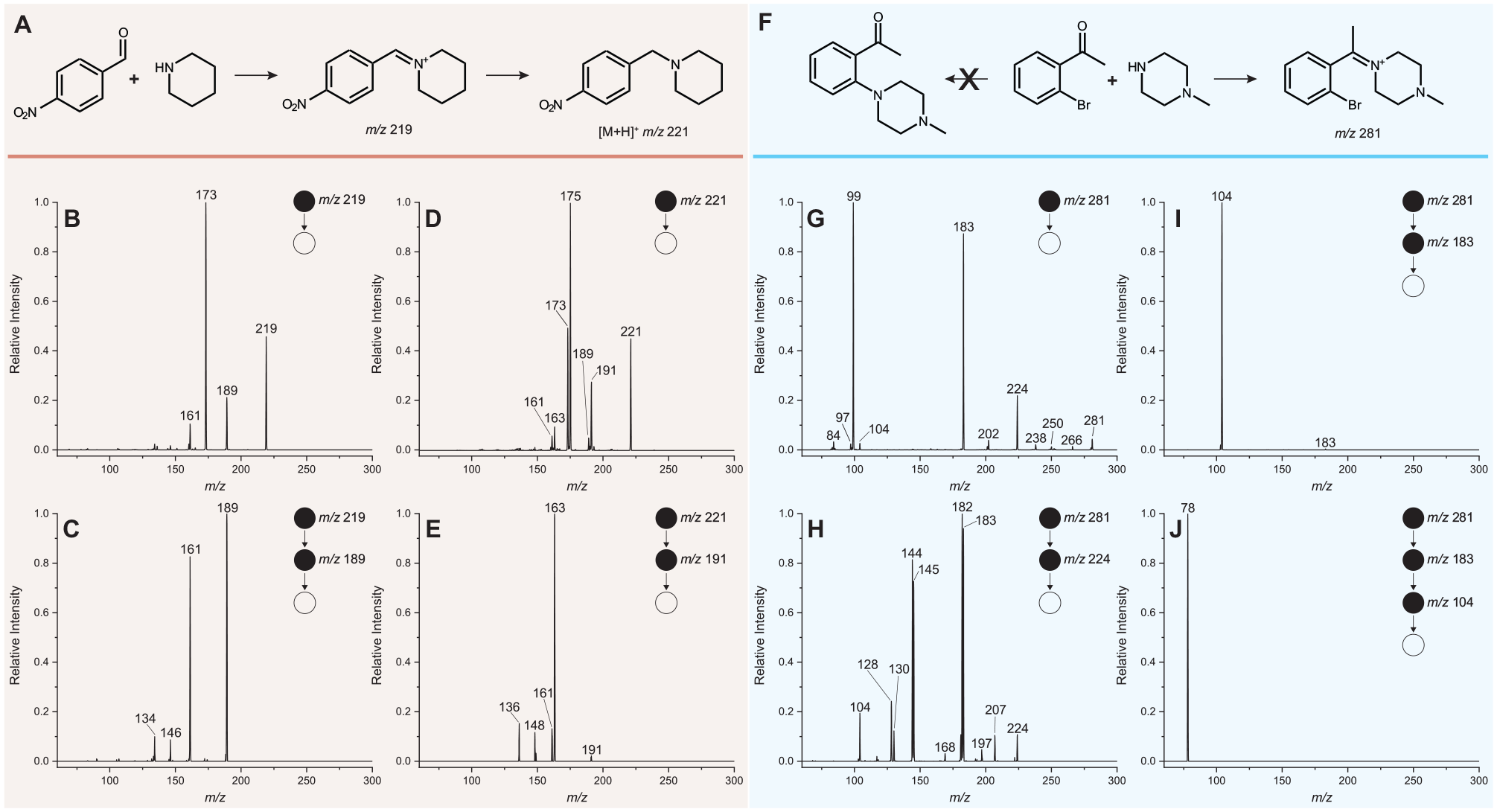
MSn experiments covering two interesting cases found in screening the XY plate. (
A second case of interest is the reaction of 2′-bromoacetophenone and 1-methylpiperazine. This reaction mixture was explored under the category of SNAr; however, it did not yield the expected product, 2′-(4-methylpiperazin-1-yl)acetophenone (
The utility of MSn experiments is evident, even in these simple cases where some ideas exist about the nature of the potential intermediates or side products. However, for more complicated situations, ongoing efforts are oriented toward developing and incorporating machine learning algorithms into the high-throughput data system. These algorithms are oriented toward assisting in the structure elucidation of unknowns based on MS/MS information. The training of these methods clearly relies on the high-throughput platform itself and specifically on the ability to create large spectral libraries rapidly and efficiently. 104
Limitations of DESI-MS for Reaction Screening
As previously described, DESI doubles as an analytical and a synthetic tool, allowing the screening of already carried out reactions, as well as accelerating transformations without bulk incubation time (as here demonstrated). However, this highlights the question of where the reactions are occurring and raises a subsequent concern regarding the translation of DESI results to conventional synthetic methods of scale-up. The first question is challenging, considering that there are three locations in which the reaction can occur: (1) in the microtiter array (i.e., in bulk), with no acceleration; (2) in the thin film generated when nanoliter aliquots of the reaction mixture are spotted on the DESI plate, with some acceleration; or (3) in the DESI droplets, with the most acceleration. 67 In some cases, the observed output might be a combination of effects, whereas in others one of the reaction locales might be dominant. For instance, some of the reactions explored in this work showed little acceleration (whether in droplets or thin films); thus, when screened in a conventional DESI fashion, their results are ascribed to the bulk reaction. On the other hand, for reactions strongly accelerated in droplets, such as the Katritzky transamination, this will be the overwhelming cause of reaction acceleration. Nonetheless, it is important to highlight that despite this uncertainty in reaction location, DESI-based screening of organic reactions is valid and useful, as the conditions found optimal using DESI-MS can be readily translated to more conventional scaled-up synthesis methods such as flow chemistry (besides already existing droplet-based scale-up systems). 89 We have demonstrated this compatibility extensively for the synthesis of products of pharmaceutical interest, such as lomustine, a first-line agent for the treatment of brain tumors and second-line agent for Hodgkin’s lymphoma, 9 and HSN608, a potent pan-active FLT3 inhibitor lead compound for the treatment of acute myeloid leukemia. 12 Furthermore, we have shown in multiple cases that the results obtained by high-throughput DESI-MS are directly comparable to traditional LC-MS screening, although the DESI-based analysis is orders of magnitude faster.6,70
Another limitation arises from the ability to determine quantitative product yields. The direct use of ion counts (peak intensity) from full MS analysis, in the absence of internal or external standardization, does not necessarily provide quantitative information regarding the yield of a particular reaction across different reaction conditions or when comparing diverse reaction types. This is mainly due to the differences in ionization efficiencies across the various species screened and the dependence of this parameter on the composition of the solution. Consequently, the reaction screening presented here is intended to be qualitative, even a yes/no reaction output, and not a precise quantitation of reaction products (which would demand specific standards and calibration for each potential reaction in each reaction environment). However, note that in our particular combinatorial experiment the observation of trends across a single reaction type (i.e., same reaction conditions and at least one identical reactant), although not quantitative, is still highly informative. This claim is supported by considering (1) the excellent agreement of the experimental results with the expected chemical reactivity trends, as described above, and (2) the similarity in ionization efficiencies across the expected product species in each selected subset. For instance, within the Suzuki coupling reactions, all expected products are aromatic analogs with an identical 4-(dimethylamino)phenyl moiety, whose tertiary nitrogen (shared by all products) is the most likely protonation site. Comparisons between diverse reaction types, with significantly different products, are less direct due to the confounding effect of the dissimilarities in ionization efficiencies. However, even in these cases, the MS information is useful in assessing compound purity. If required, the addition of internal standards to obtain quantitative reaction information from quenched reaction mixtures, as it has been shown for MALDI-based screening, 105 is directly compatible with the workflow of the high-throughput DESI-MS platform. Note also that the limitation imposed by ionization efficiency is not exclusive to DESI-MS, but is shared by all methods based on MS.
The absence of quantitative concentration measurements does not always represent a limitation. For instance, in the case of most biological assays, the test compounds, for example, inhibitors (which change from sample to sample), are not the species being determined through the assay. Hence, when testing enzymatic inhibitors, the inhibitor itself is not quantified, but the substrate and/or product of the target enzyme is, and this is the same across all assays. The ratio between product and substrate (which is precisely determined by DESI) can then be utilized as a proxy for the bioactivity measurement, so that trends can be established even without the use of calibration procedures. Calibration in these cases allows for processing of the ion ratios in order to obtain accurate numeric responses, directly comparable to other methods. An example of successful standardization of DESI-MS data utilizing an internal standard to obtain quantitative results is presented in the following section for the study of the NNMT enzyme.
NNMT Activity Assay
NNMT is a small-molecule methyltransferase, primarily involved in the
In contrast, through use of the high-throughput DESI-MS platform, the enzymatic assay can be directly performed without labels at rates faster than 1 s/sample (~0.3 s/sample of effective analysis time). The NNMT assay was easily adapted using reported conditions for buffer, concentrations, incubation, and quenching (see the “Experimental Methods” section). The only modification necessary was the addition of an internal standard to the quench solvent. Neostigmine (
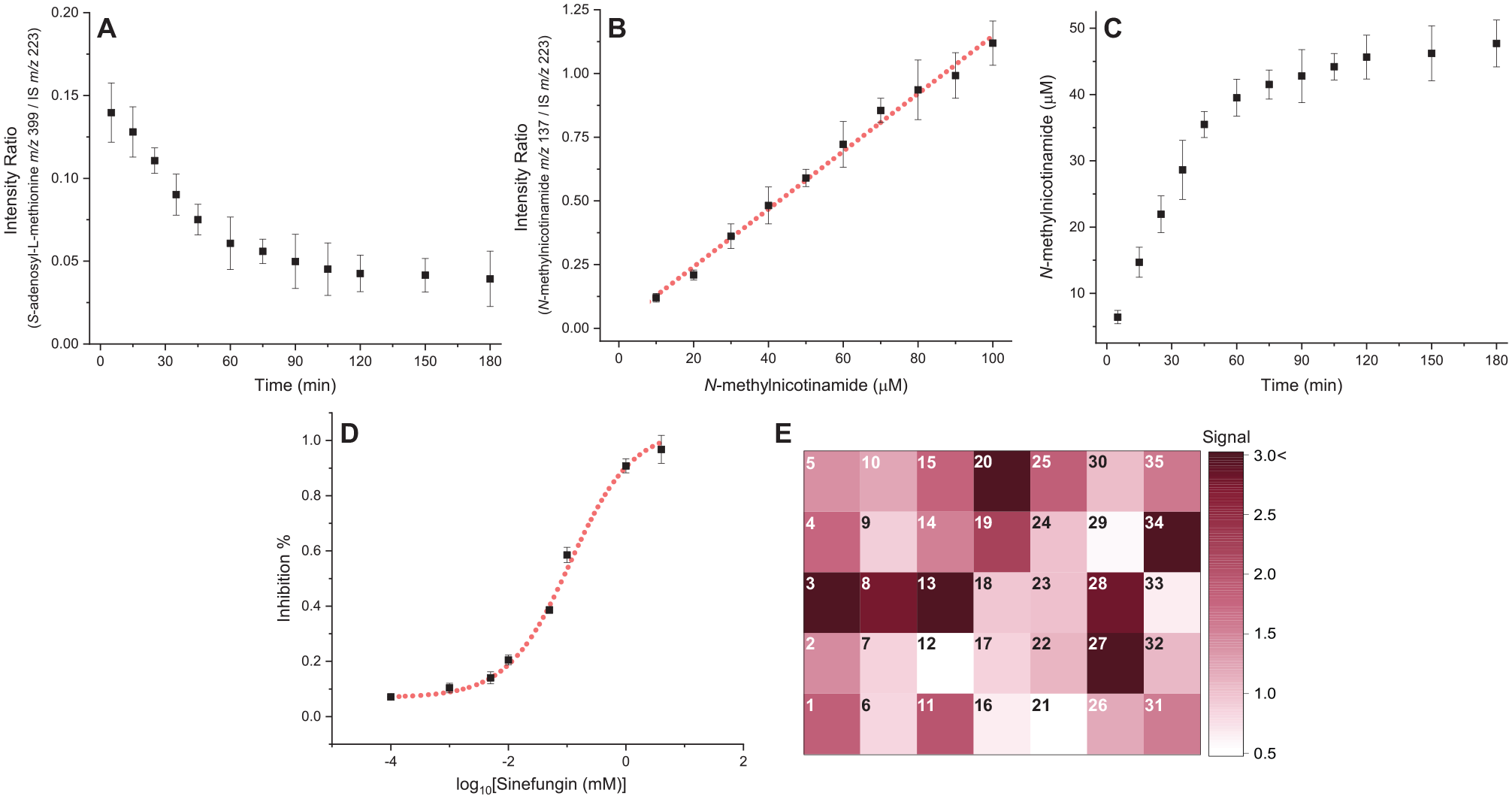
Results of the NMMT activity assays carried out using the high-throughput DESI-MS platform. The enzymatic reaction was monitored discontinuously following the response (ion intensity) of (
DESI-MS analysis in full MS mode was used to simultaneously monitor product generation and reactant consumption. Due to their ionic character,
In a similar fashion, inhibitors can be screened and characterized using the DESI-MS-based assay. Here we demonstrate this capability through a dose–response study of sinefungin (
Finally, thanks to the versatility of the high-throughput DESI-MS assay, multiple pyridine substrates can be screened rapidly without any changes to the method. This exploration of NNMT substrate tolerance is a relevant problem considering that the enzyme has a role in xenobiotic detoxification.84,85 Previous studies have explored several pyridines, quinolines, and cyclic amines,85,113 finding, for instance, that quinoline can be a substrate of NNMT,
82
albeit with lower kinetic efficiency than pyridine.85,109 However, the substrate scope tested has been modest (<30 compounds), yielding few trends in its description. Therefore, this is a challenging case in which label-free HTS would be greatly beneficial. Here, as a proof of concept, we screened the 35 pyridines/quinolines available in our laboratory (complete list available in the Supplemental Material) to test the promiscuity of the enzyme. The success of the methyl transfer was assessed in this case by evaluating the presence of the expected pyridinium/quinolinium product (substrate + CH3:
Three of the successful substrates had substituents in the third position: 3-chloropyridine (13), 3-phenylpyridine (20), and 3-aminoquinoline (27), whereas two were substituted in the fourth position: 4-(dimethylamino)pyridine (3) and 4-tert-butylpyridine (34). These results are in line with previous reports that suggest some favorability for substrates substituted in the third position (as in nicotinamide, the native substrate). 85 However, this favorability is highly dependent on the substituent, as in both our results and previous reports not all substrates substituted in the third position led to product (e.g., 3-ethylpyridine (29), 3-butylpyridine (19), 3-cyanopyridine (1)). Significantly few examples of substrates substituted in the fourth position have been evaluated previously, but our results seem to indicate certain tolerance for particular groups in that position. Large-scale screening efforts, which could be rapidly carried out using the high-throughput DESI-MS platform, might provide greater insight into the substrate scope of NNMT, adding valuable information to the existing knowledge on xenobiotic metabolism and detoxification.
Conclusions and Future Directions
As demonstrated, the Purdue Make It DESI-MS system has distinctive capabilities (i.e., ambient ionization, direct analysis of complex samples, microdroplet-accelerated reactions, and rapid MSn characterization) that allow it to be used to perform high-throughput reaction screening, nanoscale synthesis, and (bio)analysis in a manner that is (presently) unmatched by other technologies. The DESI system serves as both a robust analytical tool and a synthetic methodology, while maintaining a competitive throughput (>1 sample/s), comparable to techniques that have more complex technical requirements. Additionally, it combines the versatility, specificity, and sensitivity of MS with the simplicity and convenience of the direct ambient analysis from complex mixtures, regardless of the presence of buffers, detergents, or nonvolatile salts. These qualities exceed those of many systems, especially those that utilize LC for sample workup or those that use optical methods of detection. These features, in addition to maintaining an automated sample handling, analysis, and data processing workflow, permit and encourage the user to employ creative and innovative experiments that are not available in any current technologies. Due to the modularity and accessibility of the Make It system, it is typically straightforward to implement small changes to the instrumentation or methodology to explore new systems and types of experiments.
One such new endeavor, currently under development, is based on the concept that nanogram amounts of reaction product can in principle be collected (by capturing a fraction of the secondary DESI droplets instead of directing all of them to the mass spectrometer) while performing HTS. Their collection will allow the high-throughput identification of optimal reaction conditions, as well as permitting small amounts of products to be available for subsequent high-throughput biological testing or orthogonal analytical characterization. This collection step will represent a bridge between all the current demonstrated capabilities of the platform—nanoscale synthesis, reaction screening, and label-free biological testing—yielding a whole integrated system suitable, for example, for a complete DESI-MS-based drug discovery campaign.
Other efforts underway are oriented toward implementation of machine learning for automated identification of reaction products from MS/MS spectra. This capability requires large data sets, which are readily obtained from the automated system, as already described. Furthermore, the Purdue Make It system capabilities for bioanalysis are not limited to traditional enzyme kinetic characterization and modulation by small molecules, as shown above. Possibilities exist to easily adapt the instrumentation and methodology to allow for adaptation of a variety of bioassays (e.g., permeability assays, receptor binding measurements) to be carried out in a high-throughput, label-free fashion, exploiting the versatility, selectivity, and sensitivity of MS analysis.
We anticipate that ongoing work, together with the demonstrated capabilities of the system for bioanalysis, synthesis, and reaction screening, as detailed in this paper, have the potential to dramatically assist in pharmaceutical, biotechnological, and biomedical diagnostic pipelines.
Supplemental Material
sj-pdf-2-jla-10.1177_24726303211047839 – Supplemental material for Automated High-Throughput System Combining Small-Scale Synthesis with Bioassays and Reaction Screening
Supplemental material, sj-pdf-2-jla-10.1177_24726303211047839 for Automated High-Throughput System Combining Small-Scale Synthesis with Bioassays and Reaction Screening by Nicolás M. Morato, MyPhuong T. Le, Dylan T. Holden and R. Graham Cooks in SLAS Technology
Research Data
sj-xlsx-1-jla-10.1177_24726303211047839 – for Automated High-Throughput System Combining Small-Scale Synthesis with Bioassays and Reaction Screening
sj-xlsx-1-jla-10.1177_24726303211047839 for Automated High-Throughput System Combining Small-Scale Synthesis with Bioassays and Reaction Screening by Nicolás M. Morato, MyPhuong T. Le, Dylan T. Holden and R. Graham Cooks in SLAS Technology
Footnotes
Acknowledgements
The authors acknowledge the skilled technical assistance of Larisa Avramova, Tiago J. P. Sobreira, Christina R. Ferreira, and Bharath Keshavemurthy.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the U.S. Department of Defense: Defense Advanced Research Project Agency (DARPA) under award W911NF-16-2-0020. The authors gratefully acknowledge the support of Waters Corporation through grant 40002775. Nicolás M. Morato acknowledges support from the Charles H. Viol Memorial Fellowship and the Eastman Summer Fellowship in Analytical Chemistry.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
