Abstract
Testing the sensitivity of patient-derived tumor cells ex vivo can potentially help determining the appropriate treatment for each patient and spot the development of resistance to a given therapy. The number of cells obtainable from a biopsy is, however, often insufficient for performing ex vivo tests in conventional microtiter plates. Here, we introduce a novel Droplet-Microarray platform based on a hydrophilic-superhydrophobic patterned surface that enables screenings using only 100 cells and 30 picomoles of a drug per individual nanoliter-sized droplet. We demonstrate that the dose–response of as few as 100 primary patient-derived chronic lymphocytic leukemia (CLL) cells to anticancer compounds on the Droplet-Microarray platform resembles the dose–response obtained in 384-well plates requiring 20,000 tumor cells per experiment. The extremely miniaturized Droplet-Microarray platform thus carries great potential for ex vivo drug sensitivity and resistance tests on patient-derived tumor cells and potentially for implementing such tests in medical practice of precision medicine.
Keywords
Introduction
Cancer is the second leading cause of mortality worldwide and resulted in 8.8 million deaths worldwide in 2015. 1 Today’s common practice is for cancer patients to undergo therapy without having been examined for individual sensitivities or resistance of tumor cells to certain anticancer drugs. Drug response assays were shown to be predictive of patient response in vivo and even though such assay needs clinical evaluation before it could be used in practice, it carries a great potential for defining a given patient’s most suitable therapy. However, the amount of primary cells isolated from biopsies available for this test is not always enough. Therefore, there is a clear need for reducing cell numbers and increasing throughput per test. Here we introduce a Droplet-Microarray (DMA) platform based on a hydrophilic-superhydrophobic patterned surface that enables drug screenings using only 100 cells in individual 100 nL droplets. In this study, we report on our comparison of the drug response of primary patient-derived chronic lymphocytic leukemia (CLL) cells to anticancer compounds performed on the DMA platform and in microtiter plates.
Common anticancer drug treatment includes chemotherapeutic and targeted drugs that are often applied together in so-called combinatorial drug therapy. Chemotherapeutic compounds are cytotoxic agents that are not necessarily specific to cancer cells and may also be toxic to the body’s healthy cells. Targeted therapy, developed for some types of cancer, is more specific and blocks the growth of tumor cells by targeting the specific molecules involved in carcinogenesis and tumor progression 2 by delivering toxic agents specifically to tumor cells 3 or by stimulating the immune system to attack cancer cells. 4 Predicting which therapy a patient will most likely respond to remains a major challenge. Malignant tumors are very heterogeneous and possess high levels of intratumor (occurring in one patient) and intertumor (occurring between patients) heterogeneity. That is a reason why even the same type of tumor may respond differently to the same therapy. Methods that could help to define a given patient’s individual sensitivities to a therapy before treatment include the individual molecular profiling of a tumor and the drug sensitivity and resistance test (DSRT). Molecular profiling includes collecting genomic, transcriptomic, and proteomic data from a tumor with the aim of identifying targets and biomarkers that are specific to an individual patient and predictive of that patient’s response to a therapy.5 –9 However, due to the limited number of biomarkers and incomplete understanding of the mechanism of tumor progression, it is not always possible to predict the patient response.10 –12 Therefore, data from molecular profiling can be combined with results obtained in ex vivo DSRT, in which tumor cells obtained from a biopsy are tested against a panel of anticancer compounds for their sensitivity to different types of therapy.8,11,13 –17 DSRT could enable the identification of drug dependencies that are not predictable via molecular profiling alone. 11 In addition, DSRT could allow us to repurpose existing compounds when a drug not usually prescribed for a particular cancer type is discovered to be effective against that cancer.8,15,16 Repeated in vitro testing of samples obtained from patients during and after the therapy could help us identify clonal selection in a tumor during treatment and identify the individual sensitivity and resistance of cancer clones. This can facilitate the therapy’s adjustment to ensure the complete removal of cancer cells from the body. 13 About 80% of trials and clinical reports on DSRT have demonstrated that it is predictive of short-term clinical responses in individual patients,9,13,17 –24 making DSRT an important potential approach for identifying the most suitable therapy for individual patients.
DSRT is usually carried out in 96-, 384-, or 1536-well plates utilizing on average from 500 to 60,000 cells/well.8,13,24 –26 There are about 250 approved small-molecule anticancer compounds, and enormous amount of cells would be needed to test all the available compounds and especially their combinations. As fine-needle aspiration biopsies yield on average fewer than 500,000 cells after cells have been collected for histopathological analysis, 27 there are no cells left for additional ex vivo sensitivity testing. In addition, as compounds and reagents are so expensive, it is critical we lower the costs of such tests to make them affordable for each individual patient—a key factor for precision medicine. Therefore, we need highly miniaturized systems that enable the ex vivo testing of minute numbers of primary cells in small culturing volumes with large compound libraries.
We recently developed the DMA platform for cell-based screenings.28 –31 DMA consists of an array of hydrophilic spots on superhydrophobic background. Due to the extreme difference in wettability between hydrophilic spots (water contact angle [WCA] under 10°) and superhydrophobic background (both advancing and receding WCAs exceeding 150° and sliding WCA under 5°), 32 arrays of stable separated homogeneous droplets are easily formed on the planar DMA surface spontaneously either by applying aqueous solutions or by printing via nanoliter dispensers.29,30,33 These individual droplets can serve as miniaturized reservoirs for culturing cells. We have demonstrated the use of the DMA platform for the screening of cells of adherent30,33 and suspension 29 nature in 2D, as well as in 3D environments.34,35 In this paper, we have extended application of our miniaturized platform for the miniaturized DSRT on suspension primary patient-derived CLL cells. By using the DMA platform, we were able to reduce the consumption of cells by a factor of 200 compared with 384-well plates. We have validated our system with cells obtained from five different donors and nine anticancer compounds in a manual and automated setup.
Materials and Methods
Preparation of Polymer DMA Slides
Preparation of hydrophilic-superhydrophobic patterned surfaces of DMA slides has been described in detail elsewhere.32,33 Briefly, patterns were prepared as follows. Glass slides (Schott Nexterion, Jena, Germany) were first activated with 1 M NaOH (Carl Roth GmbH + Co. KG, Karsruhe, Germany) for 1 h, followed by neutralization with 1 M HCl (Merck KGaA, Darmstadt, Germany) for 30 min. Afterward, activated glass slides were modified with 20% v/v solution of 3-(trimethoxysilyl)propyl methacrylate (Sigma-Aldrich Chemie, Munich, Germany) in ethanol for 30 min at room temperature. A polymer layer was formed by applying 25 μL of a polymerization mixture (24 wt% 2-hydroxyethyl methacrylate [HEMA], 16 wt% ethylene dimethacrylate [EDMA], 12 wt% 1-decanol, 48 wt% cyclohexanol, and 0.4 wt% 2,2-dimethoxy-2-phenylacetophenone; all from Sigma-Aldrich Chemie) onto an inert glass slide, covering it with a modified glass slide and cross-linking the polymer by UV irradiation with a 12 mW/cm2 intensity and 260 nm wavelength for 15 min. Inert glass slides were prepared by fluorination in trichloro(1H,1H,2H,2H-perfluorooctyl)silane (Sigma-Aldrich Chemie) vapor in desiccators under a 50 mbar vacuum overnight. A polymer surface was modified with alkyne groups by incubating the slides in solution containing 45 mL of dichloromethane (Merck KGaA), 56 mg of 4-(dimethylamino)pyridine (Novabiochem, Merck KGaA), 111.6 mg of pentynoic acid (Sigma-Aldrich Chemie), and 180 µL of N,N′-diisopropylcarbodiimine (Alfa Aesar GmbH & Co KG, Karlsruhe, Germany) for 4 h under stirring at room temperature. A superhydrophobic background was created by applying a 5% v/v solution of 1H,1H,2H,2H-perfluorodecanethiol (Sigma-Aldrich Chemie) in acetone onto the polymer surface and irradiating the slide through a photomask (Rose fotomasken, Bergisch Gladbach, Germany) with 260 nm of UV light at 12 mW/cm2 for 1 min. Afterward, hydrophilic spots were created by applying 10% v/v β-mercaptoethanol (Alfa Aesar) solution in 1:1 water–ethanol onto the patterned surface and irradiating the slide with 260 nm of UV light at 12 mW/cm2 for 1 min.
Printing of Compounds
Anticancer compounds in different concentrations and four repeats per each concentration were printed onto hydrophobic glass slides or directly to hydrophilic spots of polymer or nonpolymer DMA. Hydrophobic glass slides were prepared as follows. Microscope glass slides (Schott Nexterion, Jena, Germany) were fluorinated by trichloro(1H,1H,2H,2H-perfluorooctyl)silane (Sigma-Aldrich, Chemie) under a 50 mbar vacuum in a sealed desiccator containing an open vial of the silane overnight. Frozen aliquots of antineoplastic compounds in DMSO were provided by Molecular Therapy in Hematology and Oncology, and the Department of Translational Oncology, National Center for Tumor Diseases and German Cancer Research Centre, Heidelberg, Germany and are listed in
Primary CLL Cells
Peripheral blood samples from five leukemia and lymphoma patients were obtained from Molecular Therapy in Hematology and Oncology, and the Department of Translational Oncology, National Center for Tumor Diseases and German Cancer Research Centre, Heidelberg, Germany. Blood was separated by a Ficoll gradient (GE Healthcare, Munich, Germany), and mononuclear cells were cryopreserved. 8 All procedures for obtaining and preserving patient-derived cells were approved by the Ethics Committee Heidelberg (University of Heidelberg, Germany; S-356/2013).
Culturing Cells on DMA Platform
Jurkat human T-cell lymphocyte cells were cultured in RPMI-1640 medium (Gibco, Life Technologies) supplemented with 10% heat-inactivated fetal bovine serum (FBS; Sigma-Aldrich Chemie) and 1% penicillin/streptomycin (Gibco, Life Technologies). Cells were cultured in T25 flasks and diluted every 2–3 days until a cell density of 2 × 105 cells/mL was achieved.
Primary patient-derived CLL cells were obtained from National Center for Tumor Diseases, Heidelberg, Germany. CLL cells were isolated and frozen as described in the “Isolation of Primary CLL Cells” section. CLL cells were thawed by placing the vial containing cells in a 37 °C water bath for a few seconds before most of the liquid was thawed and only small ice clumps were still left. Cells were resuspended in 10 mL of RPMI-1640 medium (Gibco, Life Technologies) supplemented with 10% heat-inactivated FBS (Sigma-Aldrich Chemie) and 1% penicillin/streptomycin (Gibco, Life Technologies). Afterward, cells were centrifuged with 1000 rpm for 5 min. Supernatant was removed and cell pellet was resuspended in 10 mL of RPMI-1640 medium (Gibco, Life Technologies) supplemented with 15% heat-inactivated AB-type human serum (RPMI-HS, MP Biomedicals), 1% glutamine (Invitrogen), and 1% penicillin/streptomycin (Gibco, Life Technologies). Cells were counted with the automated cell counter Countess II FL (Thermo Fisher Scientific Inc., Darmstadt, Germany) and diluted to a desired cell concentration.
Cells were seeded on polymer or nonpolymer DMA slides. Polymer DMA slides were prepared as described in the “Preparation of Polymer DMA Slides” section and preconditioned before cell culture as follows. First, DMA slides were sterilized by immersion in 100% ethanol, followed by drying under a sterile bench for at least 20 min before use. Afterward, DMA slides were coated with 2.2% bovine gelatin (Sigma-Aldrich Chemie) in RPMI medium containing 1% heat-inactivated FBS (Sigma-Aldrich Chemie) and 1% penicillin/streptomycin (Gibco, Life Technologies). Coating was done manually by rolling the droplet of gelatin across the array, resulting in spontaneous droplet formation. Afterward, slides were incubated in a cell culture incubator for 1 h and dried for 20 min under a sterile bench before use. Nonpolymer DMA slides were obtained from Aquarray GmbH (Eggenstein-Leopoldshafen, Germany). Slides were sterilized by immersion in 100% ethanol, followed by drying under a sterile bench for at least 20 min before use. Afterward, nonpolymer DMA slides were either directly used for cell seeding or coated with gelatin as described above for polymer DMA.
Both CLL and Jurkat cells were seeded on DMA slides either manually, using the effect of spontaneous droplet formation, or with an I-DOT nanoliter dispenser (Dispendix GmbH, Stuttgart, Germany). For manual seeding, cells were diluted to a final concentration of 75 × 104 cells/mL. A preconditioned DMA slide was placed into a petri dish. Cells were seeded by applying 1.7 mL of cell suspension onto one array field containing 14 × 14 spots (total 196) and a petri dish was closed with a humidifying lid, which contained a cloth pad wetted with buffer. Cell suspension was incubated on the DMA slide for 90 s, followed by slight (~45°) tilting of the petri dish with the DMA slide to allow for the spontaneous formation of an array of droplets containing cells. Afterward, a petri dish with the DMA slide was placed in a cell culture incubator. For seeding with I-DOT nanoliter dispenser cells were diluted to a final concentration of 100 × 104 cells/mL and dispensed directly to hydrophilic spots in a 100 nL volume obtaining on average 100 cells/spot. Afterward, the DMA slide containing cells was placed into a petri dish with a humidified lid and placed in a cell culture incubator.
All experimental protocols performed with patient-derived cells were approved by the Ethics Committee Heidelberg (University of Heidelberg, Germany; S-356/2013).
Manual DSRT on DMA Platform
The range of compound concentrations (four repeats per each concentration) were printed as described in the “Printing of Compounds” section. Cells were thawed and seeded manually as described in the “Culturing Cells on DMA Platform” section. After seeding, cells were incubated in a cell culture incubator for 2 h prior to introduction to compounds. Compounds were introduced to cells using the sandwiching method and aligning the frame, as described previously (
Imaging and Image Analysis
Imaging was performed using an automated screening Olympus IX81 microscope. The grid of the DMA pattern was defined in the microscope software. The autofocus function “interpolate AF” was used to minimize defects that can arise from uneven surface thickness and unparalleled slide positioning. Z stacks of 10 slices were made to obtain cells in focus as suspension cells are on slightly different focal planes. The images of one central field of view per each SL spot were taken at 10× magnification in three different channels: bright field, GFP (for calcein AM), and RFP (for PI). For analysis, images obtained from 10 Z stacks were merged using the “Extended Depth” algorithm in ImageJ software (NIH, Bethesda, MD). Numbers of calcein AM and PI-positive cells were calculated using a specially developed algorithm for the automated counting of cells described previously. 29
Estimation of Cell Viability in Flask
The viability of cells cultured in flasks was estimated by two different methods. In the first method, an aliquot of cell suspension from the flask was stained with Trypan blue and cells were counted in a Countess II FL cell counting machine. In the second method, the cells cultured in flask were seeded on an empty DMA slide and cells were immediately stained with calcein AM/PI using the sandwiching method, followed by imaging. Both methods to estimate viability in the control flask yielded similar results.
Automated DSRT on DMA Platform
Nonpolymer DMA slides were purchased from Aquarray. The range of compound concentrations was printed directly into hydrophilic spots as described in the “Printing of Compounds” section. Cells were thawed and seeded using an I-DOT dispenser (Dispendix) as described in the “Culturing Cells on DMA Platform” section. Cells were incubated for 48 h with compounds before doing the readout assay. For the readout assay, cells were stained with calcein AM (Thermo Scientific) and PI (Invitrogene) by dispensing 50 nL of staining solution (1.5 µg/mL calcein AM and 3 µg/mL PI in PBS) directly into each droplet and incubating for 15 min in the cell culture incubator. Cell imaging and image analysis took place as described in the “Manual DSRT on DMA Platform” section.
Obtaining Dose-Dependent Curves in Different Volumes
Compound doxorubicin in water was printed with an I-DOT dispenser (Dispendix) directly in hydrophilic spots of nonpolymer, noncoated DMA slides or in wells of microtiter plates in different amounts to ensure the following range of final concentrations in each volume tested: 0.041, 0.123, 0.37, 1, 3, 10, 30, 50, 100, and 200 µM. Afterward, the Jurkat cells were seeded in 100 cells/spot or cells/well in volumes ranging from 0.1 to 100 µL using an I-DOT dispenser (Dispendix) either on nonpolymer, noncoated DMA slides or in microtiter plates. To obtain 100 cells in each volume, several cell dilutions were done to ensure an average of 100 cells per each volume tested. Volumes 0.1 and 0.2 µL were tested on DMA slides with spot sizes 1 × 1 mm; volumes 0.5 and 1 µL on DMA slides with spot sizes 1.4 × 1.4 mm; volumes 5 and 10 µL on DMA slides with spot sizes 3 × 3 mm; volumes 10, 20, and 30 µL in a 384-well plate; and volume 100 µL in a 96-well plate (
CellTiter Glo Assay in 384-Well Plate
Cells were seeded in amounts of 100 or 20,000 cells/well in 30 µL of medium in wells preprinted with compounds. After 48 h of incubation with compounds, 30 µL of CellTiter Glo reagent (Promega, Madison, WI) was added to each well, and the plate was shaken for 2 min and incubated for 30 min. Luminescence was then measured using a 1420 Luminescence Counter Victor Light (PerkinElmer, Waltham, MA) and integration time of 2 s/well.
Statistical Analysis
Error bars on all published graphs represent standard deviations.
Dose–response curves in
The Z′ factor was calculated using DMSO as a negative control and doxorubicin treatment as a positive control. The following formula was used: Z′ factor = 1 – (3*(σ0 + σDox)/(µ0 – µDox)), where σ0 is the standard deviation of viabilities of control, σDox is the standard deviation of viabilities of cells after treatment with doxorubicin at a concentration of 10 µM for nonpolymer and 200 µM for polymer DMAs, µ0 is the average viabilities of the control, and µDox10 is the average viabilities of cells treated with doxorubicin at a concentration of 10 µM for nonpolymer and 200 µM for polymer DMAs.
Results and Discussion
Workflow of DSRT on the DMA Platform Using a Manual Setup
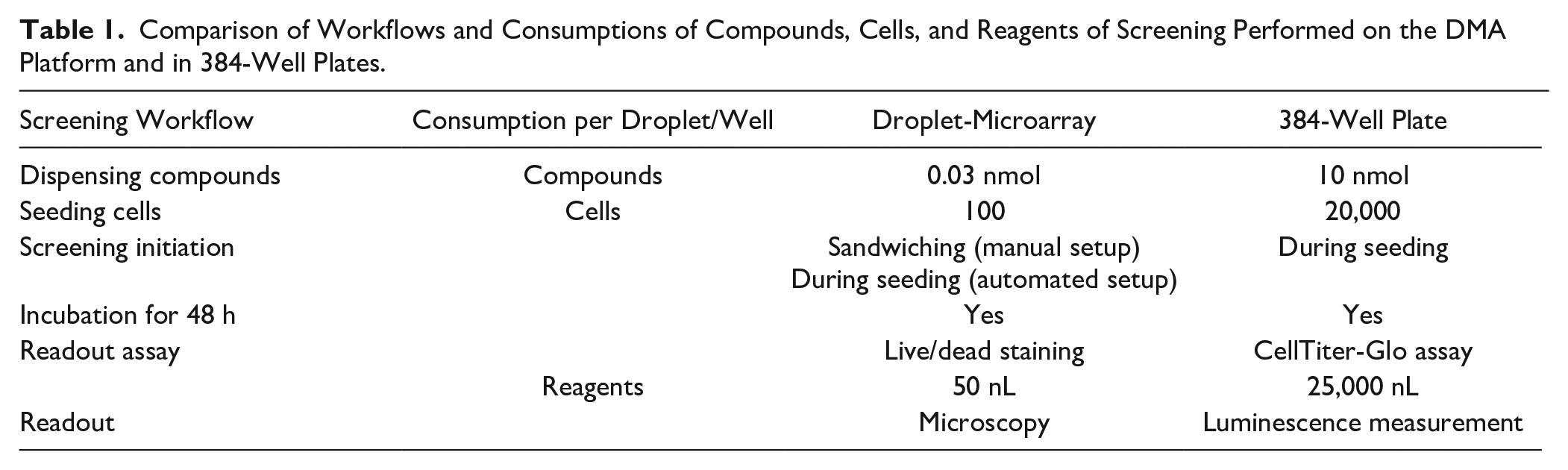
The workflow of a miniaturized DSRT and its step-by-step comparison with a workflow used in 384-well plates is presented in
Table 1
. For the miniaturized DSRT using the manual setup, we used DMA slides29
–31,33,34,36 with a polymer surface coating, manufactured by coating standard glass slides (7.5 × 2.5 cm) with a thin layer of rough and porous methacrylate-based polymer, followed by selectively modified hydrophilic and superhydrophobic areas with hydrophilic and hydrophobic chemical groups, respectively (see Materials and Methods). The array grid’s design was defined by a photomask and was precisely controllable. In the current study, we used DMAs containing 588 square hydrophilic spots with a side length of 1 mm capable of trapping about 80 nL of volume (
Comparison of Workflows and Consumptions of Compounds, Cells, and Reagents of Screening Performed on the DMA Platform and in 384-Well Plates.
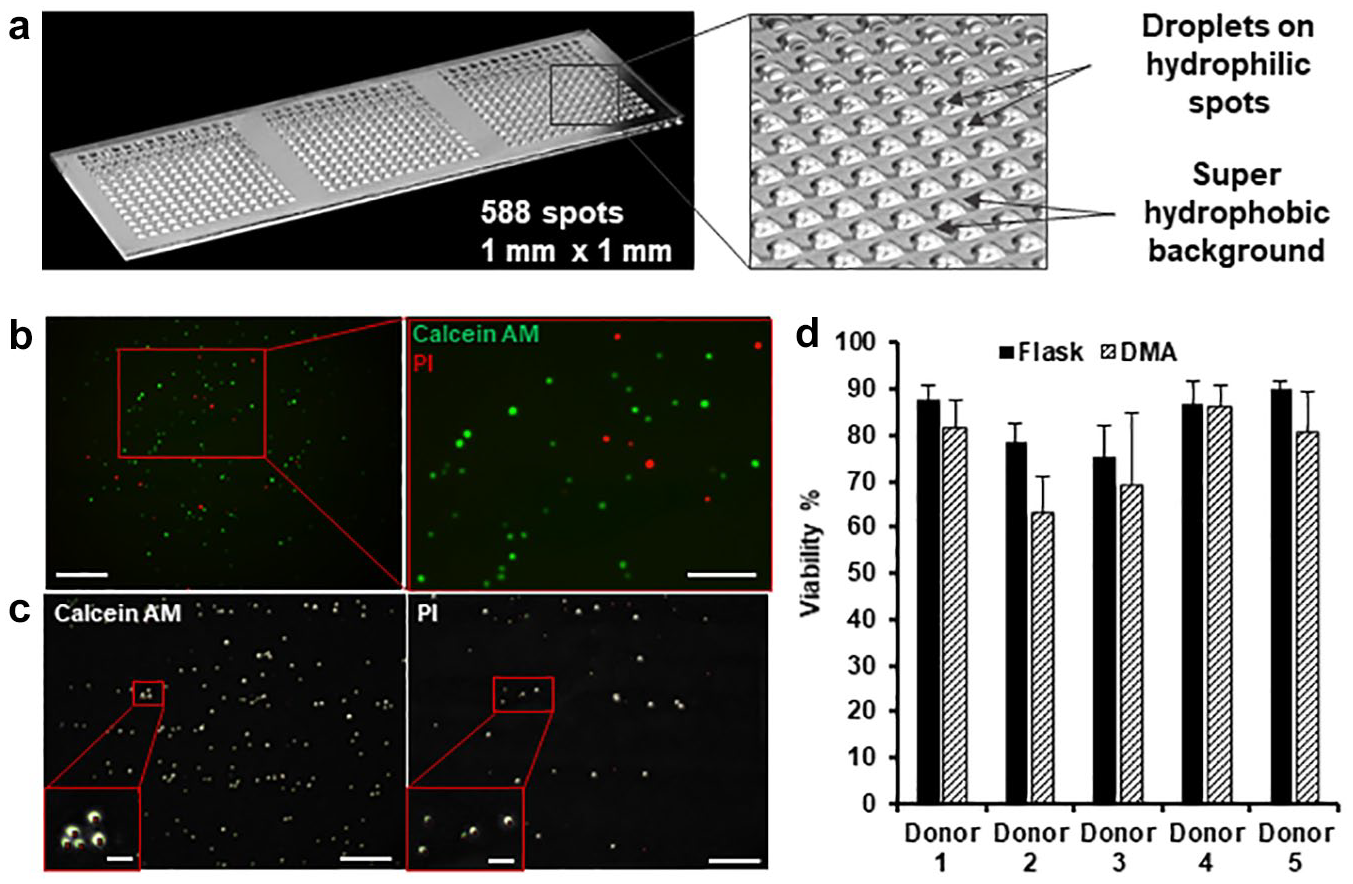
The DMA platform. (
Primary patient-derived CLL cells were seeded on DMAs by applying cell suspension onto the polymer surface to cover the whole array of spots followed by tilting the DMA slide, which resulted in spontaneous formation of an array of separated droplets containing cells (
The cell screening workflow based on our DMA platform has several advantages over the state-of-the-art microtiter plates. First, the same screen done in 384-well plates required 200 times more cells (20,000 vs 100), 300 times more compounds (10 nmol vs 0.03 nmol), and 500 times more reagents (25,000 nL vs 50 nL) per each well ( Table 1 ). Second, the DMA platform does not necessarily require expensive equipment and can be used manually, which requires neither multiple pipetting steps nor liquid handling automation. When provided with DMAs containing a compound library, laboratories not equipped with liquid handling robotics can perform cell screenings manually without a need for additional expensive machinery.
Culturing Primary Patient-Derived CLL Cells Using the DMA Platform
In the present study, we used primary patient-derived CLL cells from frozen peripheral blood samples. Blood samples were obtained from five leukemia and lymphoma patients and provided by the University Hospital Heidelberg. CCL cells do not proliferate in vitro or in the blood. We have stained CLL cells cultured in 384-well plates for Ki-67, a proliferation marker, which showed that less than 1% of cells were proliferating (
As a first step, we ensured that CLL cells could be cultured in 80 nL droplets for 48 h, our planned incubation time. We checked the stability of droplets on DMA for 48 h in a cell culture incubator and showed that droplets of culture medium on DMA are stable for at least 48 h (
DSRT on the DMA Platform Using Manual Setup
As a next step, we carried out the compound treatment of primary CLL cells on polymer DMA slides using a manual setup according to the workflow described in
Table 1
. The cells from five different donors were previously tested with a library of anticancer compounds using a previously published workflow in 384-well plates (
Table 1
) at the University Hospital Heidelberg.
8
We chose nine anticancer compounds commonly used to treat this cancer (
The compound treatment results are presented in
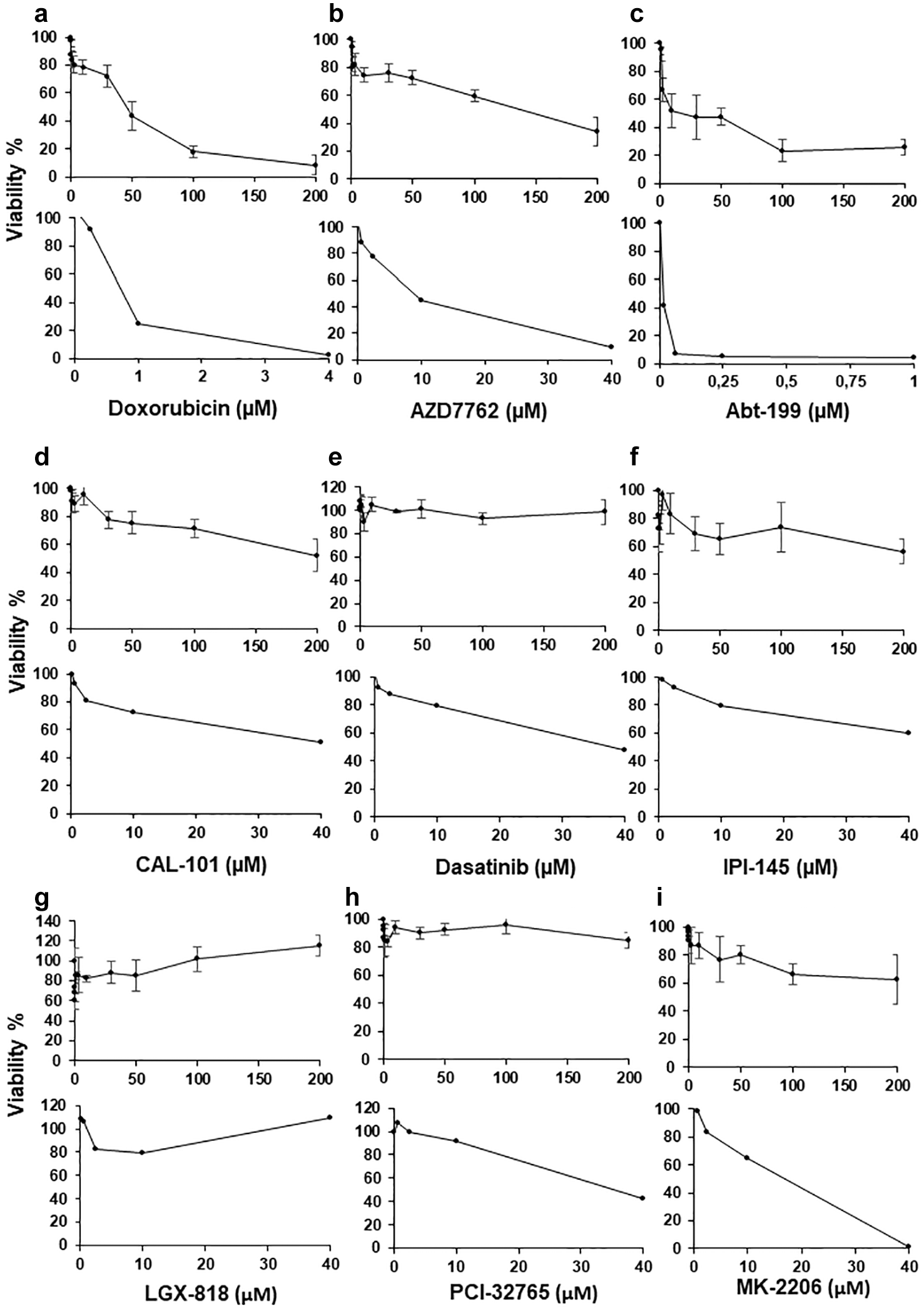
Comparison of dose-dependent effects of compounds on primary patient-derived CLL cells isolated from donor 1 on the polymer DMA platform (upper panel) and 384-well plate (lower panel): (
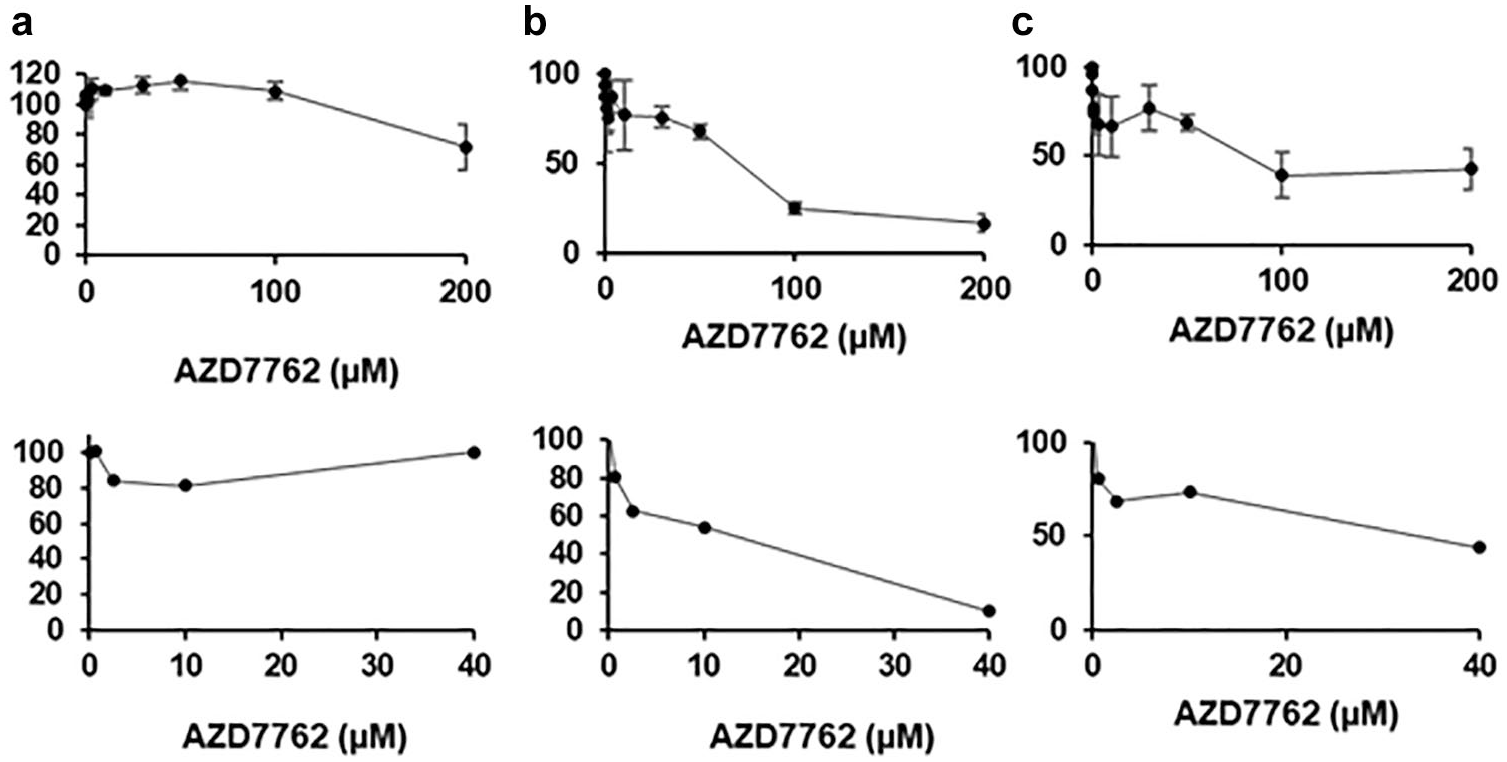
Comparison of dose-dependent effects of compounds on primary patient-derived CLL cells isolated from three different donors on the polymer DMA platform with manual setup (upper panel) and 384-well plate (lower panel): (
Comparison of Cell Responses Using Different Experimental Setups
We hypothesized that the higher effective concentrations of compounds on polymer DMA slides could be due to (1) the partial transfer of compounds into droplets, (2) nonspecific adsorption of compounds on the polymer surface, (3) a different cell response in extremely small culturing volumes of 80 nL, and (4) different readouts used to estimate viability on the DMA platform and 384-well plate ( Table 1 ). To test these hypotheses, we chose a simple experimental model where we compared the response of a human T-lymphocyte cell line (Jurkat cells) with a range of concentrations of doxorubicin under different experimental conditions. To precisely control the number of cells and culturing volumes per experiment, we used a noncontact liquid dispenser for dispensing compounds and cells in these experiments.
To test our first hypothesis, we compared the cell response to doxorubicin that had been either preprinted directly into hydrophilic spots prior to cell seeding (reverse method) or first printed onto a glass slide and then added to DMA containing cells by sandwiching (sandwiching method) (
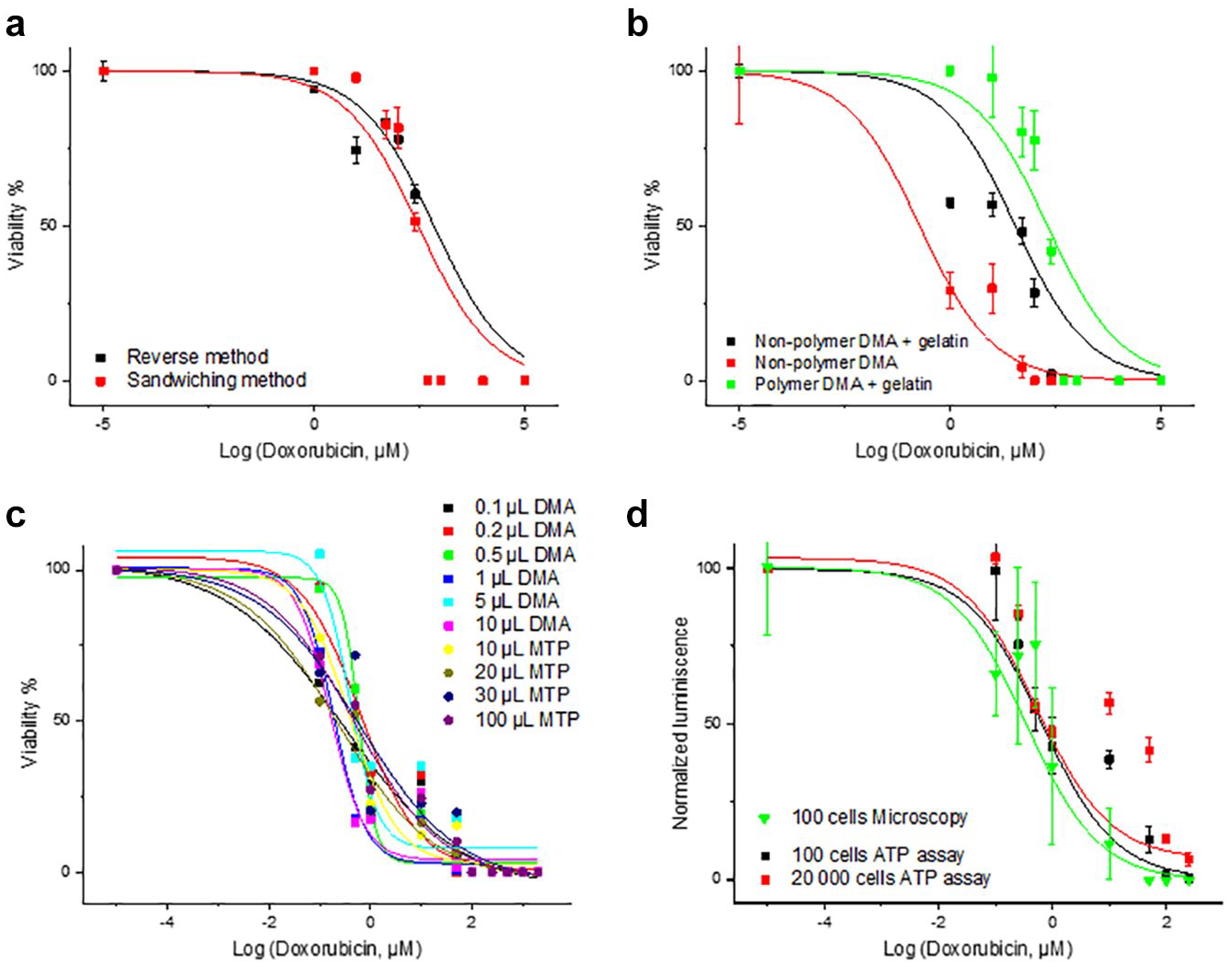
Comparison of dose-dependent effects of doxorubicin on Jurkat cells on nonpolymer DMA using different experimental setups: (
To test the second potential explanation, we compared the response of Jurkat cells to doxorubicin on a polymer DMA and on a nonpolymer DMA slide. Doxorubicin’s IC50 value on polymer DMA slides coated with gelatin was much higher than that obtained on nonpolymer DMA slides coated with gelatin (~202.51 µM vs 35.2 µM, respectively) (
To test our third hypothesis, we compared the response of 100 cells to different concentrations of doxorubicin in volumes ranging from 0.1 to 100 µL. To cover this range of volumes, we used both a nonpolymer DMA platform and polystyrene microtiter plates (
To investigate our fourth hypothesis, we compared the dose–response of Jurkat cells to doxorubicin in 384-well plates using microscopy and luminescence-based readout, which resulted in close dose–responses with both methods, with IC50 values of 0.35 and 0.66 µM for microscopy and ATP assay, respectively (
Taken together, the nonspecific adsorption of compounds onto functionalized polymer seems to be the main reason for the increase in the drugs’ working concentrations on the polymer DMA surface (
DSRT on DMA Platform Using Automated Setup
As a next step, we tested whether the response of primary CLL cells to nine previously tested anticancer drugs on nonpolymer, noncoated DMA slides is indeed comparable to the cell response in 384-well plates via microcopy and luminescence-based readouts. CLL cells were dispensed directly into hydrophilic 1 mm2 spots preprinted with various concentrations of anticancer drugs, followed by incubating the cells for 48 h. Staining solution containing calcein AM and PI was then dispensed into the droplets using a noncontact liquid dispenser. The dose–responses and IC50 values of eight drugs on the nonpolymer DMA slides concurred closely with the response observed in a 384-well plate (
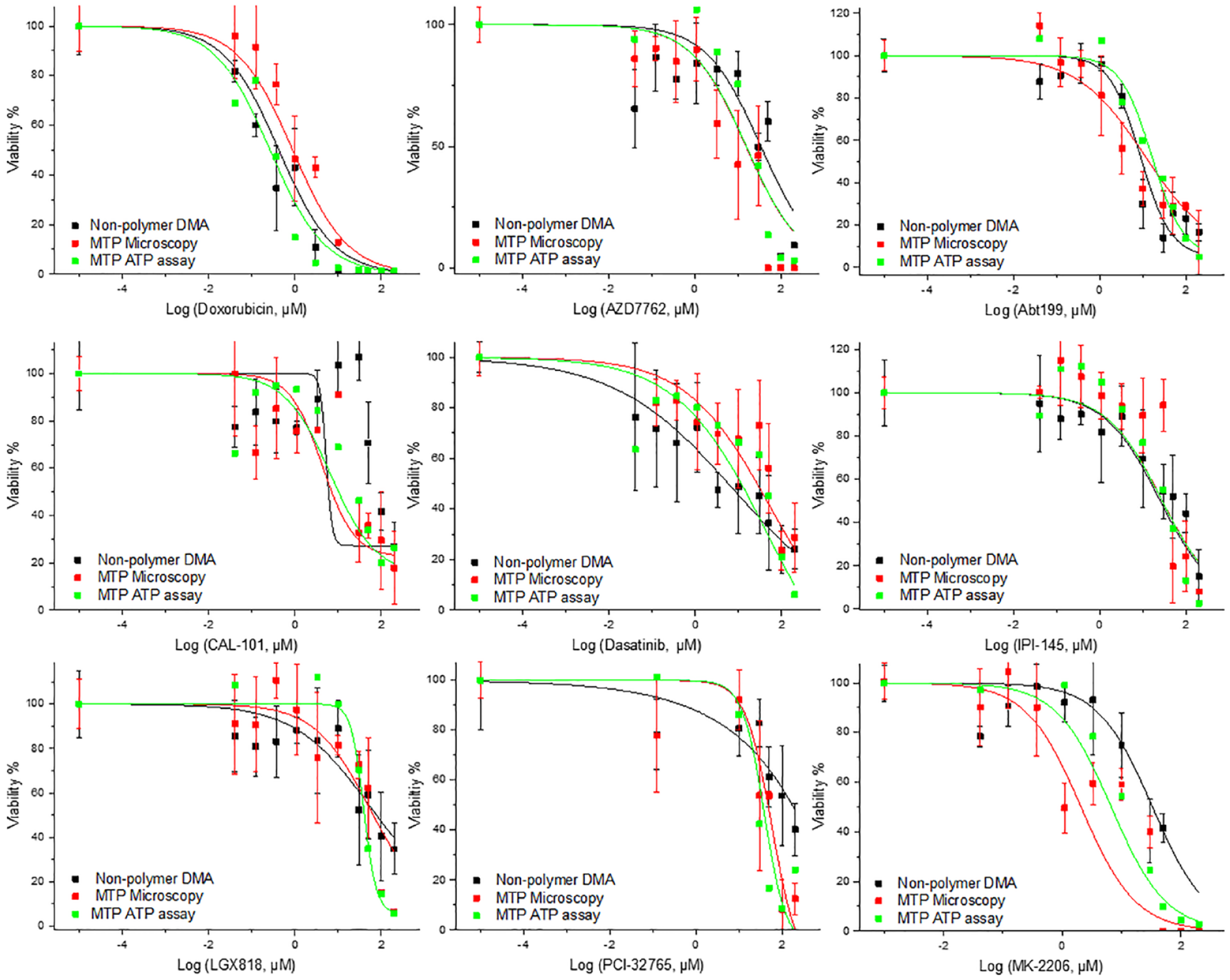
Comparison of dose-dependent effects of compounds on primary patient-derived CLL cells on nonpolymer DMA and in a 384-well plate using microscopy- and luminescence-based readouts. All experiments were performed on cells isolated from donor 4, except for the dose-dependent effect of Abt199, which was performed on cells from donor 2. In the case of the DMA platform, the average was taken from four repeats; error bars are standard deviations. In the case of a 384-well plate, the microscopy-based readout average was taken from three repeats; error bars are standard deviations. In the case of a 384-well plate luminescence-based readout, only one repeat per concentration was made.
Taking together, our results indicate that we can obtain comparable and reliable results on the DMA platform. In other words, by using primary patient-derived cells on the DMA platform with as few as 100 cells in 100 nL culturing volume, we obtained dose–responses comparable to those obtained in a 384-well plate using 20,000 cells in 30 µL of culturing volume. We have estimated the time and cost of performing the full protocol for each compound on DMA and 384-well plate (
Conclusions
The present work illustrates how we established a workflow for a highly miniaturized DSRT using the DMA platform. We have demonstrated the promising potential of culturing primary CLL cells isolated from patients in 80 to 100 nL droplets for 48 h with high viability. We showed that by using the DMA platform and only 100 cells and 300 times fewer compounds and reagents, we obtained the same results as if we had employed conventional 384-well plates and 20,000 cells per each well. We have demonstrated our system in 2D, however, as we have previously shown that the DMA platform is compatible with culturing cells in 3D environments by means of scaffolds34,35 or by promoting the self-organization of cells. 35 The possibility of culturing cells in 3D gives rise to even more physiologically relevant applications using primary patient-derived cells.
We demonstrated the application of manual and automated operation of DMA slides for DSRT, both of which can facilitate various applications. The manual setup requires no liquid handling robotics and can be done in a laboratory lacking automation for low- to middle-throughput screenings. On the other hand, our automated setup is beneficial for experiments requiring higher throughputs or even smaller volumes and cell numbers. All the components of the described system for both manual and automated setup are available commercially, and established protocols can be adopted in different laboratories. We believe that highly miniaturized DSRT on our DMA platform has a potential to open new opportunities in the field of personalized and precision medicine by enabling tests with minute amount of primary patient-derived materials and reagents that were impossible in state-of-the-art microtiter plates.
Supplemental Material
Supporting_Info_TECH-19-0082_REVIEW_Popova – Supplemental material for Miniaturized Drug Sensitivity and Resistance Test on Patient-Derived Cells Using Droplet-Microarray
Supplemental material, Supporting_Info_TECH-19-0082_REVIEW_Popova for Miniaturized Drug Sensitivity and Resistance Test on Patient-Derived Cells Using Droplet-Microarray by Anna A. Popova, Sascha Dietrich, Wolfgang Huber, Markus Reischl, Ravindra Peravali and Pavel A. Levkin in SLAS Technology
Footnotes
Acknowledgements
The authors are grateful to Dr. Dietrich and Professor Zenz (Hospital University of Heidelberg) for providing compounds and CLL cells for the study. The authors are grateful to Dr. Christine Blattner (Institute of Toxicology and Genetics, KIT) for providing the Jurkat cell line. The authors are grateful to the Crossmedia service department at KIT and especially to Amadeus Bramsiepe and Willi Mueller for professional photos of DMA.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: A.A.P. and P.A.L., in addition to being employed by the Karlsruhe Institute of Technology, are (since March 2018) shareholders of Aquarray GmbH. S.D., T.Z., W.H., M.R., and R.P. declare that there is no conflict of interest regarding the publication of this article.
Ethics Statement
All the experiments were performed in accordance with relevant guidelines and regulations. The human material used in the current study was obtained in accordance with the Ethics Committee Heidelberg (University of Heidelberg, Germany; S-356/2013). Patients who donated tumor material provided written informed consent prior to study. In the current study, the patient information was not revealed and obtained results were used exclusively for comparison purposes between two experimental platforms (DMA and microtiter plates) and were not connected back to data of individual patients.
Funding
The authors received the following financial support for the research, authorship, and/or publication of this article: This research was supported by the ERC starting grant (no. 337077-DropCellArray), ERC-PoC grant (no. DLV-680913-CellScreenChip), and EXIST Forschungtransfer (Aquarray 03EFJBW155).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
