Abstract
Quickly and easily producing uniform populations of microsphere-based 3D cell cultures using droplet-based templating methods has the potential to enable widespread use of such platforms in drug discovery or cancer research. Here, we advance the design of centrifuge-based droplet generation devices, describe the use of this platform for droplet generation with controlled cell occupancy, and demonstrate weeklong culture duration. Using simple-to-construct devices and easily implemented protocols, the initial concentration of encapsulated cells is adjustable up to hundreds of cells per microsphere. This work demonstrates the first instance of using centrifugal droplet-generating devices to produce large numbers of cell-encapsulating microspheres. Applications of this versatile methodology include the rapid formation of templated 3D cell culture populations suitable for suspension culture or large batch bioreactor studies that require uniform populations.
Introduction
Appropriately designed 3D culture systems can bridge the gap between 2D cell culture and animal models in drug discovery and cancer research. 1 In such applications, particle-based 3D culture systems provide important advantages, including the ability to maintain 3D cultures in suspension, the use of particle analysis methods such as imaging or cytometry-based approaches, and control of the presence or absence of nutrient or metabolite concentration gradients through adjustment of particle size or cell density. The spectrum of particle-based 3D constructs spans from templated 3D cell aggregates to fully mature multicellular tumor spheroids (MTSs) formed by aggregation and subsequent long-time cell proliferation. 2 MTSs are used as in vitro culture systems that mimic in vivo tissues for studies of growth kinetics, cellular heterogeneity, nutrient and waste gradients, hypoxia, acidosis, drug penetration, drug response/resistance, and metabolism.3,4 However, all particle-based 3D culture systems provide advantages relative to overly simplistic 2D systems or complex and potentially unrealistic xenograft models.1,2 Currently, though, the widespread use of such 3D cell culture methods as standard approaches to replace 2D cell culture or xenograft systems is limited by technical challenges.
In conventional aggregation-based 3D cell culture particle formation (scaffold-free methods), cells are placed into culture conditions in stirred suspension, as a liquid overlay on top of agarose, or in well-based hanging-drop formats.5,6 For many cancer cell lines, aggregation and proliferation occur under these conditions. If desired, proliferation for long times is possible, eventually leading to fully mature MTSs with particle size and cell density that result in intraparticle concentration gradients of nutrients and wastes. In such systems, necrosis often occurs in the nutrient-starved particle interior. 7 Although such spatial heterogeneity is useful in some contexts of tissue mimicry, there are many other situations where avoidance of gradients and cell heterogeneity is desirable, requiring control of particle size distributions. Suspension-based and liquid overlay approaches generally lead to polydisperse size distributions. Size selection is possible, but reduces the overall yield of MTSs or cell-based particles and still produces populations with >10% variation in diameter. 8 Hanging-drop methods provide better size control but require cumbersome pipetting, and the surface area of the dish or wells limits the number of 3D culture particles that can be produced. Finally, scaffold-free formation of 3D cell culture particles is limited to adherent cell types that naturally form cell–cell contacts and produce extracellular matrix, thus excluding many other nonadherent cell types potentially of interest in 3D co-cultures. Taken together, conventional scaffold-free 3D culture methods (1) require long culture times to generate, (2) lead to polydisperse populations, and (3) generate relatively small numbers of cell-type restricted constructs. These considerations hinder potential applications such as high-throughput MTS-based screening or incorporation of 3D cultures into macroscale tissue mimics.
Different approaches to address these 3D cell culture formation issues include cell encapsulation in droplet-based hydrogels (e.g., alginate) to produce templated 3D cell cultures (scaffold-based methods). Embedding cells into size-controlled 3D biomaterial scaffolds enables manipulation of cell seeding density, spatial organization, geometry, solute diffusion, and the effect of tissue-scale concentration gradients. 9 However, the complexity of implementation currently limits widespread adoption of these methods in traditional cell biology laboratories. For example, one method of high-throughput alginate droplet generation is to extrude an alginate solution from a nozzle under pressure and use acoustic modulation to break up the stream into droplets. Droplets are generated in air followed by submersion into a solidification solution containing calcium ions to form crosslinked hydrogels. In addition to producing larger-diameter droplets, 10 the resulting droplets are often of nonuniform shapes and sizes. Finally, this approach relies on complex electronic and acoustic instrumentation and a skilled user for operation. 11 Alternatively, droplets can be formed within microchannels in microfluidic systems by exploiting the surface tension between two immiscible fluids. In this case, a fluidic stream injected into a second immiscible fluid (possibly co-flowing in a sheath geometry) will break up into droplets. This approach can produce monodisperse template droplets, especially in cases of high surface tension such as immiscible aqueous/oil phases. Fully biocompatible all-aqueous two-phase systems in microfluidic devices have also been investigated to avoid the use of oils and inefficient postprocessing steps,12,13 though these systems generally require additional mechanical inputs such as acoustic energy because of their low surface tensions. 14 No matter the approach, however, microfluidic setups typically require methods and infrastructure that are not readily available in many cell biology laboratories (e.g., material fabrication techniques, fluidic valves and interconnects, high-precision pumps, and use of elastomeric polymers). Further, even if successfully implemented, the overall throughput of templated 3D cell culture production in such microfluidic systems is relatively low.15,16
The ideal scaffold-based 3D cell culture system would be simple to fabricate and operate in a standard cell biology lab, allow high-throughput droplet generation, produce monodisperse templated 3D cell culture populations, and be fully biocompatible (e.g., use only aqueous solutions). One such promising system for scaffold preparation is based on a centrifugal device to mass-produce droplets on a laboratory scale as described by Eral et al. 17 Their implementation used two nested tube compartments in a standard swinging arm centrifuge to provide centrifugal force for generation of sodium alginate droplets. The principle is similar to that of a dripping faucet, where droplet production occurs through the interplay of centrifugal force, droplet mass, and surface tension. Changing the centrifugal force, by adjusting the rotational speed of the centrifuge, changes the droplet size and the rate of production. The droplets are submerged in a CaCl2 catch solution for solidification. In prior work, commercially available needles, in some instances with silanization treatment, served as the nozzle component. However, the smallest inner diameter needles commercially available are 108 µm, which limits the conditions for droplet generation.
In this work we build enhanced centrifugal droplet-generating devices and provide a first demonstration of using these devices for encapsulating highly crowded cancer cells in alginate microspheres (diameter <300 µm), demonstrating laboratory-scale and quick production of uniform, scaffold-based 3D cell cultures. In order to achieve more tunable droplet size control, we modify the device design of Eral et al. 17 by using an easily built hybrid nozzle consisting of a needle with an attached pulled glass capillary. With this device, we are able to control nozzle as well as droplet diameter and obtain high-cell-occupancy 3D cell cultures. Using this approach to produce microspheres crowded with cancer cells offers many advantages for the scaffold-based production of templated 3D cell cultures. For example, nonadherent cell lines could be confined into size-controlled and monodisperse microspheres and included in co-cultures. This laboratory-scale approach, from device fabrication and operation to establishing 3D cell culture protocols, can be readily implemented using equipment and methods available in standard cell biology laboratories, thus enabling widespread use of standardized 3D cell culture. In summary, the assembly and operation of our devices is logistically simple because the methodology does not require skilled users and is relatively cost-effective as most supplies and equipment are readily available in standard cell biology laboratories. We anticipate that the realization of scaffold-supported 3D cell cultures will provide enhanced ability to study the chemical tissue microenvironment, implement large particle flow cytometry, enable tissue-based high-throughput drug discovery, and improve the fabrication of bioactive materials used for tissue engineering.
Materials and Methods
Hybrid Nozzles and Centrifuge Tube Assemblies
We assemble the hybrid nozzles using glass capillaries (1.12 mm ID and 1.5 mm OD; World Precision Instruments Inc., Sarasota, FL) that are pulled to a long taper using a Sutter Instrument Co. (Novato, CA) P-97 Micropipette Puller and cut to a specified nozzle inner diameter (50 µm) using a Narishige (Amityville, NY) MF-830 Microforge. The blunt end of the capillary is scored and cut to a total length of 12 mm. To complete the construction of the hybrid nozzle ( Suppl. Fig. S1 ), the capillary is placed over the end of a 20-gauge, ½-inch blunt-end needle (IntelliSpense; Integrated Dispensing Solutions Inc., Agoura Hills, CA) and epoxied in place (5 Minute Epoxy; Devcon Epoxy, Danvers, MA).
We build centrifuge tube assemblies as described previously
17
for high-cell-occupancy encapsulation using Falcon 50 mL conical polypropylene centrifuge tubes (Corning Inc., Corning, NY) as depicted in
The inner nozzle attachment is built by first using a drill press to make a 15 mm ID hole in the center of the twist cap from any standard 50 mL centrifuge tube. The plunger of a 10 mL BD Luer-Lok Tip syringe (BD, Franklin Lakes, NJ) is removed and the casing is smoothly cut at about 60 mm length, from needle attachment end to top, using a Dremel 4000 High Performance Variable Speed Rotary Tool (Dremel US, Mt. Prospect, IL). The inner nested tube assembly is achieved by inserting the modified 10 mL syringe into the hole in the centrifuge twist cap, Luer-Lok in the downward position, and arranging it to provide the desired height. The height, that is, the distance between the nozzle tip and catch solution, is such that the nozzle is not touching the crosslinking solution and is an important parameter that determines the sphericity of the solidified droplets. For example, if 2 mL of droplet precursor solution and 8 mL of crosslinking solution are used (as is the convention for the work described here), then the distance between the catch solution and nozzle tip should generally be between 3 and 6 mm for spherical droplets. This distance is measured during assembly by marking the expected catch solution surface in the 50 mL centrifuge tube and attaching a hybrid nozzle to the syringe Luer-Lok tip to estimate the distance between the catch solution surface and the nozzle tip. Once the appropriate height is achieved, a thin layer of hot melt adhesive is applied using an AdTech hot glue gun with mini size glue sticks (Adhesive Technologies Inc., Hampton, NH) at the junction of the syringe and twist cap to permanently set the syringe within the twist cap. In the left-hand side of
Loading of Centrifuge Tube Assemblies with Droplet Precursor and Catch Solution
A solution of ALG without cells is used to determine the parameters best suited for uniform droplet shape and size, which are then translated to cell encapsulation. All solutions and chemicals used for droplet generation are designed with buffering capacity, osmolarity, and sterility in mind to achieve biocompatible cell culture conditions (all reagents from Sigma-Aldrich, Co., St. Louis, MO, unless otherwise noted). In order to prepare the centrifuge tube assembly pairs for operation, 8 mL of the crosslinking catch solution comprising 1 M calcium chloride dihydrate (C2536), 25 mM HEPES BioPerformance Certified cell culture tested (H4034), 150 mM sodium chloride BioReagent suitable for cell culture (S5886), pH 7.4, and 0.2 µm sterile filtered using Nalgene APES bottle top vacuum filter apparatus (Thermo Fisher Scientific Nalgene Inc., Waltham, MA) is pipetted into the 50 mL centrifuge tubes. The nozzles are connected to the Luer-Lok attachment of the 10 mL syringe/screw top arrangements, carefully positioned into the 50 mL tubes, and locked into place by tightly securing the cap to the tube, making sure to keep the assemblies upright. Then 2 mL of ALG, prepared by dissolving 4% w/v alginic acid sodium salt, from brown algae (A1112) in physiological buffer (25 mM HEPES, 150 mM NaCl) with vigorous stirring overnight and pH adjustment to 7.4 using a specialized electrode for highly viscous solutions (Orion Ross Sure-Flow pH Electrode with Sure-Flow Junction; Thermo Fisher Scientific Inc.), is slowly pipetted into the syringe inner compartment. It is important to use a pipette tip that can be placed fully into the inner compartment while ejecting the solution so that the ALG fills the bottom of the syringe, avoiding the presence of bubbles that can influence droplet generation. Either two or higher even-numbered multiples of assemblies are made. Each pair of tubes is weighed and, as needed, adjusted for balance by removing fluid from one of the tubes, and then loaded into opposing positions of an Eppendorf 5810 Centrifuge with a swing arm rotor (Eppendorf North America, Hauppauge, NY).
Centrifuge operation is set using units of revolutions per minute (RPM), but the key performance parameter is acceleration. Acceleration (
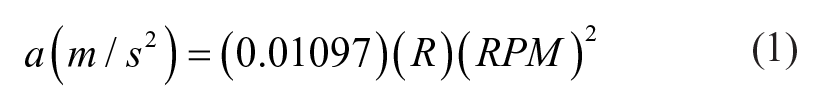
with
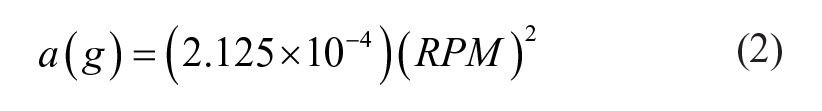
In the following, we report both RPM and acceleration values. For any given centrifuge and tube nozzle assembly, equivalent performance should be obtained by choosing operational RPM values to match the accelerations reported here.
Droplet Size Measurement
The size of microspheres generated is measured by imaging a population of microspheres under a brightfield microscope and measuring the major and minor axes of each microsphere using a calibrated reticle. The geometric mean diameter (GMD; square root of the product of the major and minor axes) for at least 25 microspheres is calculated.
Single-Cell Suspension, Cell Encapsulation, and 3D Culture Protocol
The centrifugal droplet assemblies are prepared as described above, but the ALG solution added to the inner compartment of the droplet-generating assembly now contains a cell suspension. In high-cell-occupancy experiments, the human malignant melanoma cell line MEL28 is used. The cells are handled using standard conditions: complete minimal essential media with 10% Cosmic calf serum and 1% penicillin and streptomycin (all from GE Healthcare Bio-Sciences, Marlborough, MA) and 37 °C, 5% CO2 in the incubator. Protocols for standard cell transfers or preparation into single-cell suspensions, using 0.25% trypsin with EDTA (GE Healthcare Bio-Sciences), referred to as “trypsin,” are used. Cells are plated at low density and cultured over 5 days on 150 mm dishes to generate cells for encapsulation. The dishes are trypsinized with 5 mL of trypsin for 20–30 min at 37 °C, 5% CO2 in the incubator and suspended by adding 5 mL of media to transfer the entire 10 mL suspension per dish into a 50 mL centrifuge tube. A small volume is taken to count the combined cell suspensions using a Z2 particle counter (Beckman Coulter Life Sciences, Indianapolis, IN) and calculate the concentration of cells. The cell suspension is pelleted by centrifugation at 2000 RPM for 10 min. The supernatant is aspirated and the cell pellet is washed by resuspending into 20–50 mL of a cell encapsulation buffer (1 mM EDTA, 25 mM HEPES, 150 mM NaCl, pH 7.4), depending on the cell concentration to be used. Use of EDTA in this buffer sequesters any low-concentration calcium ions and prevents premature gelation due to generation of crosslinks when the cells are added to the ALG. The cell suspension in encapsulation buffer is then centrifuged into a pellet for a second time at 2000 RPM for 10 min. The supernatant is aspirated once more, and the cell pellet is resuspended in 100 µL of encapsulation buffer with gentle pipetting up and down using a 1000 µL pipette tip. Four hundred microliters of 4% ALG is added to the 100 µL of cells and mixed. This approach is necessary as direct resuspension of the cell pellet into 4% ALG produces a suspension high in cell multiplicity (doublets, triplicates, and larger cell clumps) because of the viscosity. By resuspending cells into 100 µL of buffer (low viscosity) and adding dropwise into the ALG, clumps can be broken up and a single-cell suspension is achieved, important for uniform distribution of cells in the droplets and to prevent clumps from clogging the devices. The 500 µL of cells in ALG is then drawn up into an 18-gauge needle/10 mL syringe (BD) and added dropwise to 4.5 mL of 4% ALG while vortexing to produce a 5 mL total volume single-cell suspension. The 5 mL cell suspension is passed through a 40 µm Fisherbrand sterile cell strainer nylon filter mesh (Thermo Fisher Scientific Inc.) to remove any remaining aggregates. Depending on application-specific cell type and concentration, this straining step may not be necessary. Finally, 2 mL of the cell suspension in ALG is added to the inner compartments of centrifuge tube device assembly pairs to complete the final preparation for device operation. The throughput of the system can be enhanced by operating multiple paired devices in parallel, limited only by the number of positions available in the centrifuge.
A range of cell concentrations, using the same device construction and cell preparations described above, is investigated to determine the maximum number of cells in 4% ALG able to flow through a 50 µm hybrid nozzle–centrifuge tube assembly. For high cell occupancy, the cell distribution per droplet follows a near Gaussian distribution (obtained as the large number limit of a Poisson distribution), so the encapsulated concentration and thus the cells per droplet is a measure of the average number of cells expected to occupy a droplet. High cell occupancies are achieved for droplets generated with a 50 µm diameter nozzle in tube assemblies centrifuged 1250 RPM (332
After device operation is complete, for subsequent culture the cell-encapsulating microspheres are washed out of the catch solution by allowing the particles to settle, aspirating the solution, and resuspending in 15 mL of complete media. The example for extended cell culture shown here is generated using 1300 RPM (359
Results
The device schematic (
Fig. 1A
) and depiction of droplet generation in a swing-arm centrifuge (orthogonal position to the axis of rotation as depicted by the force vector as shown in
Fig. 1B
) is presented in
Figure 1
.
Figure 1B
also shows an example brightfield microscopy image of resultant gel microspheres produced by a 50 µm hybrid needle–glass capillary nozzle centrifuge device assembly with a 6 mm air gap at 1300 RPM (359
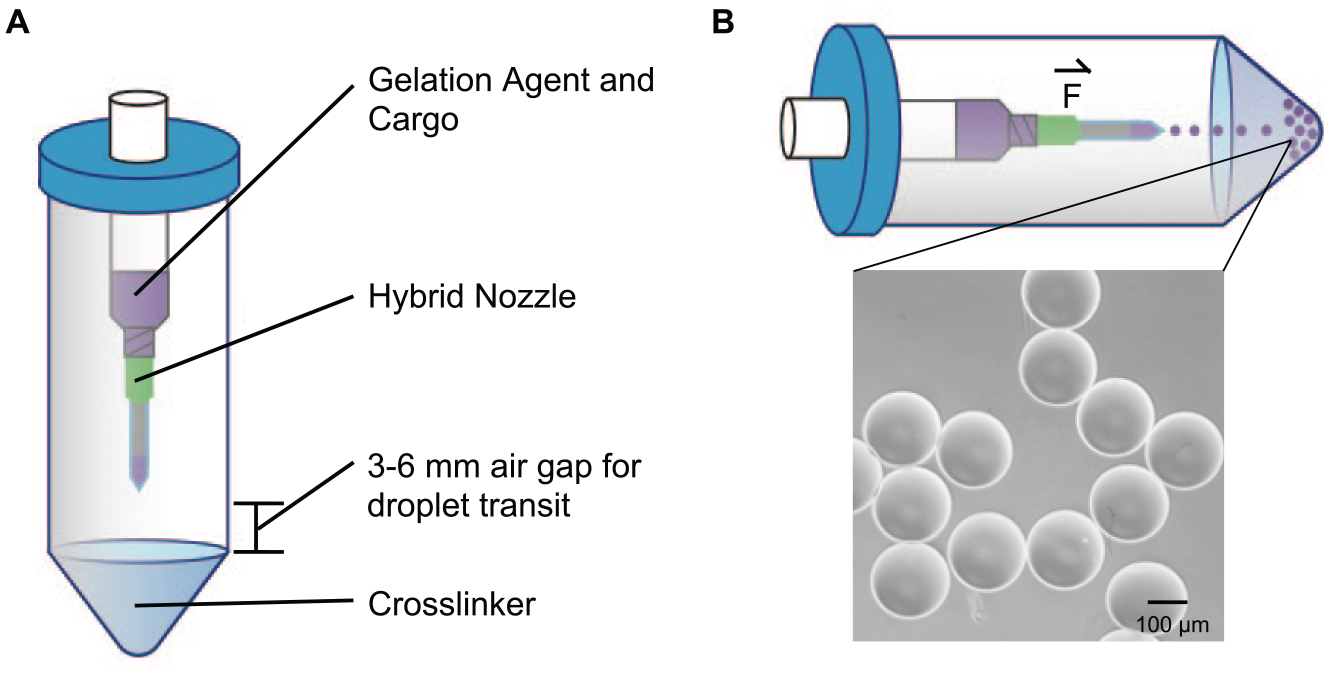
Schematic of the device design and operation. (
Figure 2A
presents microsphere diameters measured as a function of centrifuge rotor speed (RPM) as well as by the corresponding acceleration (
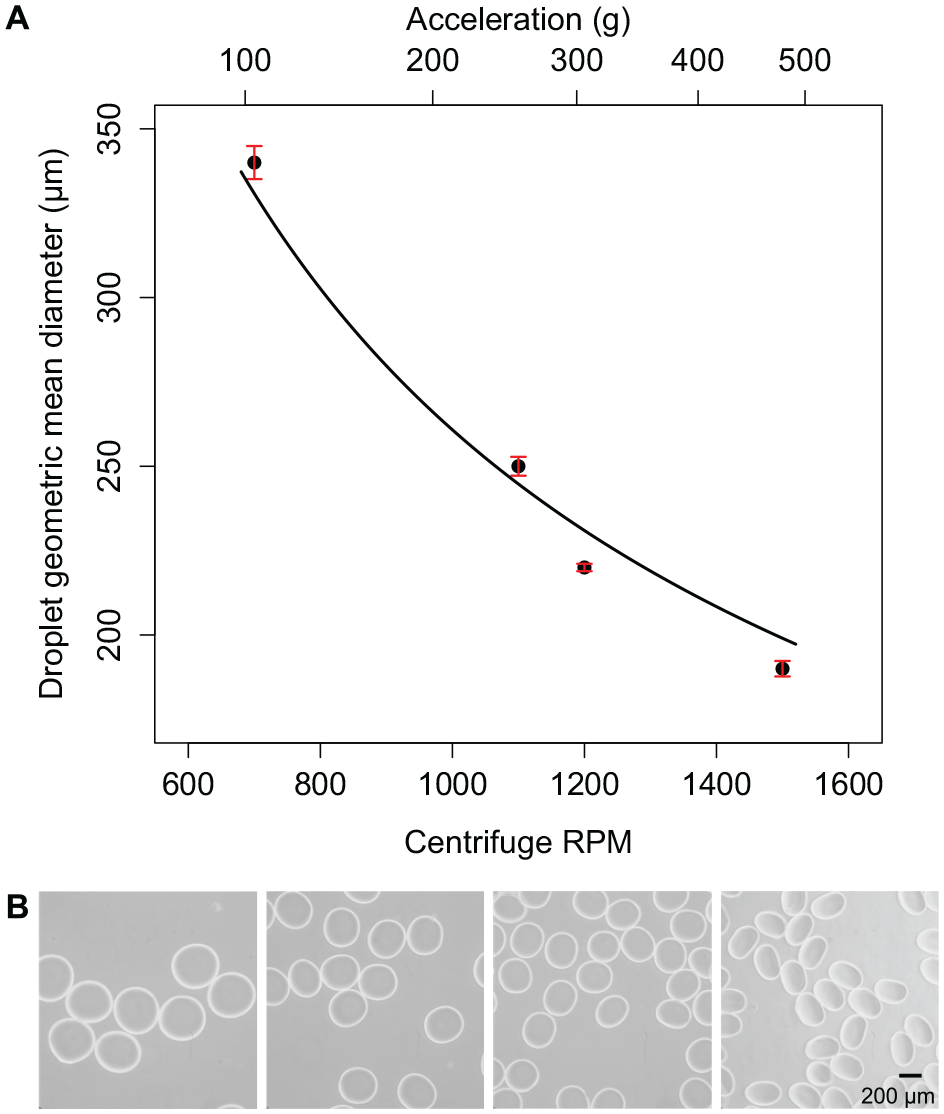
Calibration of centrifugal droplet generation with hybrid nozzles. (
To demonstrate modulation of microsphere size, we generate microspheres using the new hybrid nozzle design (diameter 50 µm) and the centrifuge tube assemblies described above (
Fig. 2B
). The devices are operated in pairs across a range of centrifuge speeds from 700 to 1500 RPM (corresponding to accelerations from 104 to 478
We explore the generation of droplets with a range of cell occupancies from 100 to 500 cells/droplet using a 50 µm diameter hybrid nozzle with centrifuge tube assemblies spun at 1250 RPM (332
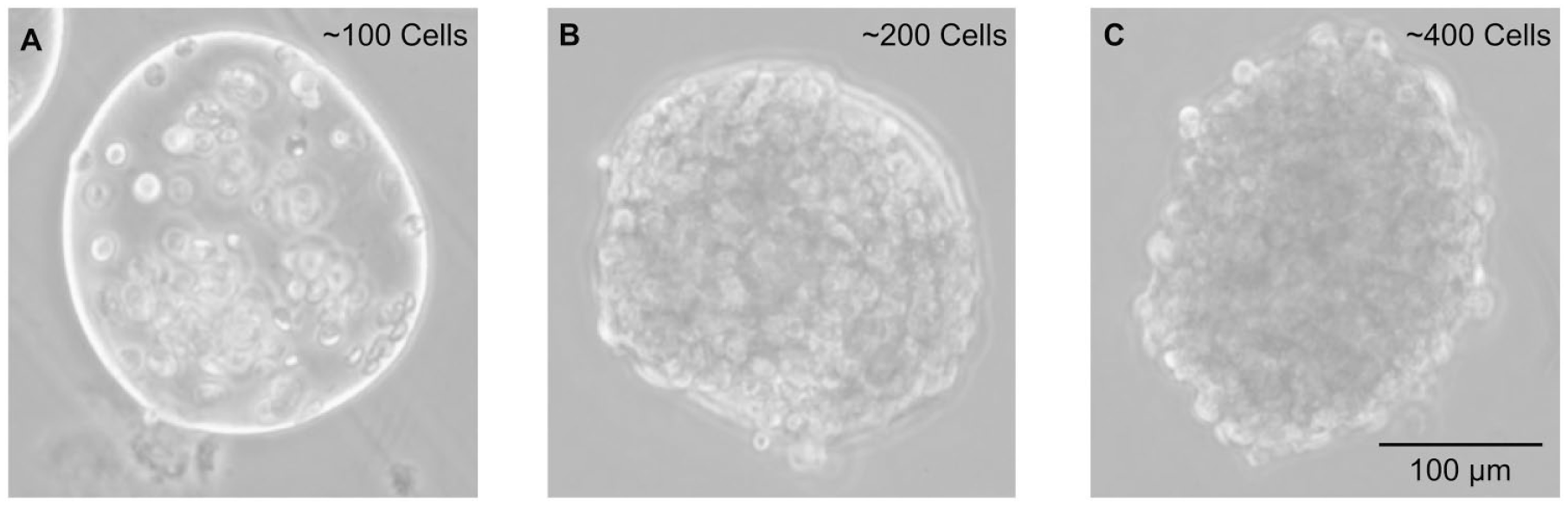
Ranges of cell concentrations are encapsulated. (
Finally, we demonstrate microsphere stability as well as cell proliferation over a 7-day time course in suspension culture using an initial population with 200 cells/droplet, which provides sufficient space for cell proliferation. For this assay, the encapsulation is performed using 1300 RPM (359
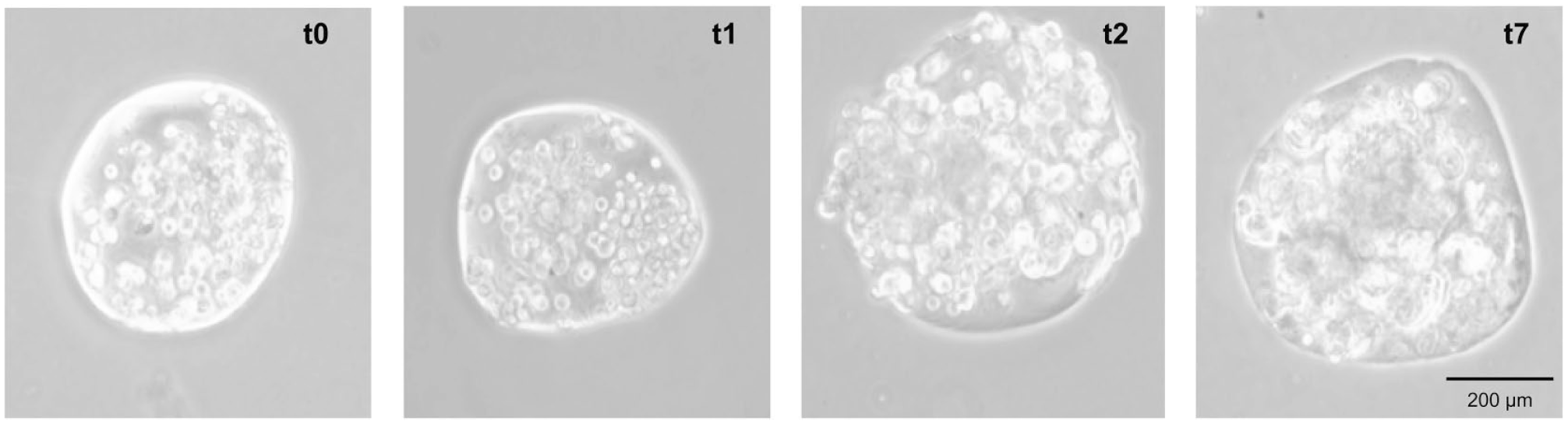
Cell-encapsulating suspension culture. Photomicrographs of microspheres initially encapsulating 200 cells/microsphere (
Discussion
The results presented here demonstrate the implementation of a simple-to-construct droplet-generating device for the production of 3D cell cultures. Our device fits into the context of previous work as an approach that combines direct (droplet formation in air) and indirect (spontaneous droplet formation in microfluidic channels) device configurations. Recent studies, also combining direct and indirect approaches, demonstrate centrifuge devices that produce water/oil emulsions for droplet-based digital PCR and droplet-based hydrogels for cell encapsulation using custom-built centrifuges.18–22 In other device designs, centrifugation of specially designed microwell plates is used to trap multiple cells 23 or to grow cultures in hypergravity conditions, 24 forcing the aggregation of cells in submicroliter volumes that would otherwise not form cell aggregates. These approaches, although useful for design, implementation, and understanding of centrifugation-based devices, are more complicated to construct and use than the approach reported here. In general, the advantages of our work in comparison with others include the control of nozzle diameter, the simplicity of device construction and operation, and the ease in creating relatively large, monodispersed populations of droplet-based 3D cell cultures across a range of cell concentrations.
With hybrid nozzle centrifuge tube assemblies ( Fig. 1 ), we find that the most important parameters for producing spherical alginate droplets, in agreement with previous work, 17 are the viscosity of the solution, the diameter of the nozzle, the height of the air gap between the nozzle tip and catch solution, and the acceleration of ALG out of the nozzle. We adjust these parameters to demonstrate the generation of a range of monodisperse and controlled microsphere sizes from 150 to 350 µm in diameter. The inclusion of cells over a range of concentrations did not alter the mean size or homogeneity of the microspheres.
To standardize the measurement, we report both RPM of the centrifuge and the acceleration at the nozzle in terms of multiples of
Control over droplet size is demonstrated to be improved using a 50 µm diameter hybrid nozzle when compared with using the smallest-diameter commercially available needles as the nozzles (
Fig. 2B
in comparison with
Suppl. Fig. S2
, summary in
Suppl. Table S1
) over a range of centrifuge spin speeds. It is observed that as centrifugal acceleration increases, the GMD of the collected microspheres becomes smaller. Although both hybrid nozzles and needles operate within the same range of accelerations, the microspheres generated with the hybrid nozzle have a smaller GMD (
Achieving a single-cell suspension is important for standardized cell culture conditions as well as for homogeneity in the microsphere population. A well-mixed single-cell suspension helps avoid cell clumping that can cause clogs in the nozzles, especially at high cell concentrations. In addition, it helps ensure that each microsphere will have effectively the same numbers of cells when initiating cell assays like proliferation experiments. We establish cell processing procedures for encapsulation that use buffers without Ca2+ to count, wash, and introduce single-cell suspensions into ALG precursor solution without prematurely crosslinking the gel, and enabling a high degree of control over the concentration of cells per droplet. The 100 and 200 cells/droplet conditions ( Fig. 3A,B ) yield cell-encapsulating microsphere populations with cells somewhat spaced out and lacking initial cell–cell contacts. In contrast, the 400 cells/droplet condition ( Fig. 3C ) produces microspheres that are densely packed. Immediately after generation, these droplets have a cell density of ~25% of the cell density of a fully mature nontemplated spheroid. 6 For the highest concentration of 500 cells/droplet (not pictured), no microspheres were produced due to nozzle clogging and a high multiplicity (cell clumping even after filtering through a 40 µm cell strainer) in the cell suspension.
The stability of the encapsulated cell culture is investigated over a 7-day period to demonstrate the feasibility of the approach for use in spinner flask bioreactor suspension culture. A 200 cells/droplet encapsulation experiment is performed using 1300 RPM (359
The improved method for droplet-based 3D cell encapsulation by centrifugal droplet generation reported here demonstrates an alternative technology that addresses the limitations of microfluidic and droplet generation in-air platforms. A single centrifuge tube assembly with a hybrid nozzle has a moderate throughput on order of 1000 droplets/s depending on the operating parameters, which in itself is a significant improvement to microfluidic throughputs that are generally on the order of 10 droplets/s. As mentioned previously, this moderate throughput in centrifugal synthesis can enter the high-throughput regime by operating up to 16 centrifuge tube assemblies in parallel (depending on the centrifuge). Additionally, it takes a few hours to form a templated 3D cell culture population that has excellent uniformity in size and control over the cell concentration using the centrifugal synthesis method described in this work. The centrifuge tube assemblies, in a way, are to be included in both the droplet generation in-air and the microfluidic droplet generation categories as the flow out of the hybrid nozzle and droplet formation under centrifugal force occur in an air gap of 3–6 mm in height, and the hybrid nozzle is similar in construction to a glass capillary microfluidic nozzle. As an aside, degradation in performance with much larger and highly nonspherical particles is observed for a larger air gap of 9 mm. The centrifugal droplet generation approach is less cumbersome than classical microfluidics and uses simple aqueous gelation chemistries, further agreeable to using materials that may be more biomimetic than alginate.
The technical ability to generate uniform large-number populations of scaffold-supported 3D cell cultures by droplet-based methods has the potential to enable widespread use of 3D culture in place of less realistic alternatives. Generating large and uniform populations of these types of cell-encapsulating constructs is important for applications in high-throughput drug screening as well as cancer research. Culturing the templated cell-encapsulating microspheres produced here uses the same techniques as conventional spinner flask approaches and thus is fully compatible with downstream processing. For example, the microsphere concentration in a spinner flask is easily volumetrically counted by microscopy and microspheres can be transferred by pipetting into a well plate for high-throughput drug discovery applications using conventional sterile cell culture practices. These 3D cell culture populations are more realistic models of a tumor and enable high-throughput drug screenings that are amenable to personalized medicine applications because of the quick processing. Many other applications are possible, and here we discuss some directions related to cancer research. In such research, treatments have been discovered that enable a drug compound to be directed to a specific type of cell. Often, these approaches are developed by using model cell line screening processes or in vivo xenograft models that identify unique genetic biomarkers (usually because of a single mutation). 26 However, these models have poor clinical translation because of a systematic inability to mimic key pathophysiological features. For example, 2D tissue culture conditions are entirely unrealistic in that tissue culture media as well as incubator conditions are designed to sustain mammalian cells on a flat surface rather than to mimic conditions of an in vivo primary tumor.27–29 In comparison with the human systems that xenograft models are designed to mimic, the mouse has a higher metabolic rate, is inbred, has a short lifespan, is reared in sterilized environments, and in immunodeficient mice, transplantable tumors grow to treatment size over weeks (not years, as is usual in humans). 30 In addition, the use of monogenic human tumor cell lines in xenograft models is lacking the genomic variability of a primary cancer and is one of the main reasons why xenotransplantation data often fail to translate to human clinical results. 31
Although recapitulating all of the pathophysiological aspects of an in vivo tumor may never be fully realistic, the work presented here makes new, more realistic, 3D tissue culture practices accessible, allowing new assay formats that mimic more of the essential aspects of heterogeneity in human tumors. For example, this new technique could be used to culture patient-derived primary cells in 3D with a drug challenge experiment. There is also opportunity for more complex co-culturing experiments where mixed cell line populations (e.g., immunological cells with cancer cells) within identical 3D microenvironments could be measured. Adaptation of this technology for other advanced cell assays may require further protocol development and optimization. However, since the methods described here rely on readily available materials that most biomedical research labs will have in house, the implementation of centrifugal droplet-generating technologies in other labs interested in using 3D cell cultures to investigate advanced cancer research questions is feasible.
Supplemental Material
Supplemental_Material_for_Centrifugal_Generation_of_Droplet-Based_3D_Cell_Cultures_by_De_Lora,_et_al – Supplemental material for Centrifugal Generation of Droplet-Based 3D Cell Cultures
Supplemental material, Supplemental_Material_for_Centrifugal_Generation_of_Droplet-Based_3D_Cell_Cultures_by_De_Lora,_et_al for Centrifugal Generation of Droplet-Based 3D Cell Cultures by Jacqueline A. De Lora, Jason L. Velasquez, Nick J. Carroll, James P. Freyer and Andrew P. Shreve in SLAS Technology
Footnotes
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: A.P.S. discloses financial interest in BennuBio Inc., a company developing high-speed flow cytometry instrumentation. J.P.F. is currently employed by BennuBio Inc.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: We acknowledge the University of New Mexico for partially funding this work. A.P.S. and J.A.D. were, in part, supported for work on development of technology for centrifugal droplet formation by the Photosynthetic Antenna Research Center (PARC), an Energy Frontier Research Center funded by the U.S. Department of Energy, Office of Science, Office of Basic Energy Sciences under award no. DE-SC0001035. J.A.D. was supported for the cell culture and encapsulation research reported in this publication by the National Cancer Institute of the National Institutes of Health under award no. 5 F31 CA189682.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
