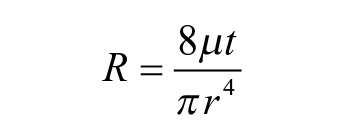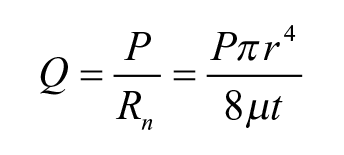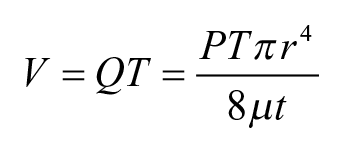Abstract
High-throughput enzyme screening for desired functionality is highly demanded. This paper utilizes a newly developed microfluidic pneumatic printing platform for high-throughput enzyme screening applications. The novel printing platform can achieve distinct features including a disposable cartridge, which avoids crosstalk; a flexible cartridge design, allowing for integration of multiple channels; and fast printing speed with submicroliter spot size. Moreover, a polydimethylsiloxane (PDMS)-based sandwich structure has been proposed and used during the printing and imaging, which can lead to better results, including reduced evaporation as well as a uniform light path during imaging. Using this microfluidic pneumatic printed PDMS sandwiched microdroplet array platform, we have demonstrated the capability of high-throughput generation of a combinatorial droplet array with concentration and volume gradients. Furthermore, the potential for enzymatic study has been validated by quantified cellulose reaction implemented with the printing platform.
Introduction
Enzymes have been important for applications in a variety of fields, including chemistry, biology, and the food industry. 1 The engineering of enzymes toward a specific property, such as better catalyst capability, chemical stability, or thermal resistivity, has gained largely increased interest in recent years, and directed evolution has been one of the most successful approaches to the engineering of these enzymes. 2 A typical directed evolution approach consists of two steps: first, genetic diversity is generated among the genotype, which results in diversified targeting phenotypes (i.e., enzymes) being displayed; second, a screening experiment is carried out to select the best phenotype in terms of a specific property. 3 During the screening procedure, high throughput has become one of the most important features, as one best phenotype needs to be identified among thousands or more mutants. Taking advantage of modern automation technologies, high-throughput screening allows for ultrafast evaluation of multiple mutants in parallel or streamline. 4 The evaluation procedure is often associated with chromogenic and fluorogenic methods, where the enzyme reaction takes place first, and the resultant chemical is labeled by markers, which can then be detected by an indirect substrate or sensor. 5 Prior to the screening, sample preparation, which includes mixing the reagents and labeling them with markers, requiring a large quantity of liquid handling procedures, is usually laborious due to the large scale of the screened library. In order to carry out enzyme analysis and screening more effectively, an automated, high-throughput method for liquid handling is needed. 6 Liquid handling has been one of the most important procedures to many large-scale screening applications. 7 With the advance of lab automations, many liquid handling techniques have emerged, such as automated pipetting, screen printing, microfluidic liquid delivery, and inkjet printing.
Automated pipetting utilizes robotics to automatically operate pipettes in order to deliver liquids according to program control. 8 The principle of automated pipetting is simple, and thus the method is robust. The method works well with traditional 96- or 384-well plates; however, due to the limitation of robotics precision and pipette precision, the minimum amount of liquid that can be handled by this method is usually in the range of 0.1 μL to 1 mL, which limits the density of the library. Moreover, given the loading volume of the pipette tip, during each delivery of liquid, the pipettes often need to move back to the liquid tank for refilling. Such frequent back-and-forth movement limits the throughput of the automated pipetting method. 7
Contact printing is usually based on an elastic pin or a pin array that, during the printing procedure, dips into the base of the liquid, allowing a certain amount of liquid transfer into the tip of the pin. Afterwards, the pins are brought into contact with the target substrate, to which the liquid is thus transferred.9,10 The material and geometric design of the tip, as well as the contact force between the tip and substrate, determines the amount of liquid to be transferred. Current development in fabrication allows for super-fine pin tips with micrometer diameter; thus, using an array of such pins with high density greatly enhances the throughput of this method, where thousands of spots can be printed at the same time. 11 However, in each delivery cycle, only one type of liquid can be printed. In order to deliver different liquids onto the same substrate, different patterns of pin arrays have to be made and applied for each type of liquid in a serial manner, which increases fabrication and time cost. Given these considerations, screen printing usually only works well for applications where fewer types of liquids need to be printed, such as DNA synthesis, where only four different nucleotides need to be printed.12,13
Microfluidic liquid delivery utilizes microfluidic chip designed to perform multiple liquid handling tasks, such as solution mixing, 14 gradient generation, 15 or breaking into droplets,16,17 under a submillimeter regime. Microfluidics provides a multifunctional platform for lab-on-a-chip analysis, and has been applied to many areas, including cell–drug interaction, PCR, and clinical diagnostics. 18 However, a microfluidic chip usually requires a sophisticated design and delicate fabrication, which greatly increases the cost, and such customized chips can usually serve only one specific purpose.
Inkjet printing is traditionally used in the office, where inks of different colors are printed on papers, combining to form text or pictures.19,20 Nowadays, it has begun attracting people’s attention to replace traditional inks with chemical or biological reagents.21–25 Such methods have been applied to many biochemical fields, including combinatorial synthesis,26–28 drug screening,29,30 chemical dilution, 31 and gene analysis. 32 The advantages of inkjet printing include its high-throughput, flexibility and high resolution. A well-modified inkjet printer can deliver nanoliter-scale liquids into any desired pattern, with a speed of more than 1000 droplets per second. Many different types of inkjet printers, including piezoelectric, 33 thermal electric, 34 and acoustic printers, 35 have been well developed and are starting to be applied in the field of biochemistry. Of most printers, the cartridges where the “inks” are stored are usually integrated with an actuation piece, either a piezoelectric material or a thermal electric material, which generates the energy for droplets to shoot out during operation. Such a piece is relatively expensive compared with the rest of the cartridge, resulting in a higher cost. In such cases, cartridges are often recollected and washed for repeated usage, which could potentially introduce contamination concerns for biological or chemical applications.
We have developed a new type of inkjet printer, referred to as a microfluidic pneumatic printer (MPP), as a modification to our previously developed microfluidic impact printer,30,36 in order to reduce the cartridge cost. The MPP printer head is composed of a microfluidic cartridge, where the liquids are contained; a pneumatic driving source; and an array of valves that control the pneumatic source. Driven by compressed air, no piezoelectrical or thermal–electrical piece is required for droplet generation, while the pneumatic source and valves can be reversibly detached from the cartridge. In this way, the cartridge can be made from low-cost polymeric materials and thus be disposable, which solves the high cost and potential contamination issues that exist among traditional inkjet printing methods. Moreover, with the easy plug-and-play design of the printer head assembly, the loading of the solution becomes much easier, while the cartridge design allows for easy customization of droplet size (1–100 nL) and integration of multiple channels (up to 24 channels per cartridge).
By applying the MPP platform, high-density droplet arrays with flexible patterns were able to be created. The MPP-enabled droplet dispensing not only increases the throughput but also reduces chemical waste. As a proof-of-concept study, the platform was applied to enzyme reaction analysis with a Glucose Colorimetric/Fluorometric Assay Kit and an enzymatic assay of cellulose. To further facilitate the result acquisition, a three-layer structure was developed by sandwiching the droplet array with polydimethylsiloxane (PDMS), which not only prevents evaporation during printing but also provides a uniform light path during the imaging procedure. Combined with an optical imaging and analysis technique, we have developed a novel, high-throughput, quantitative analysis platform for a minute quantity of reagents. The results demonstrate the great potential of the printing platform for enzyme reaction analysis.
Materials and Methods
Printing Platform
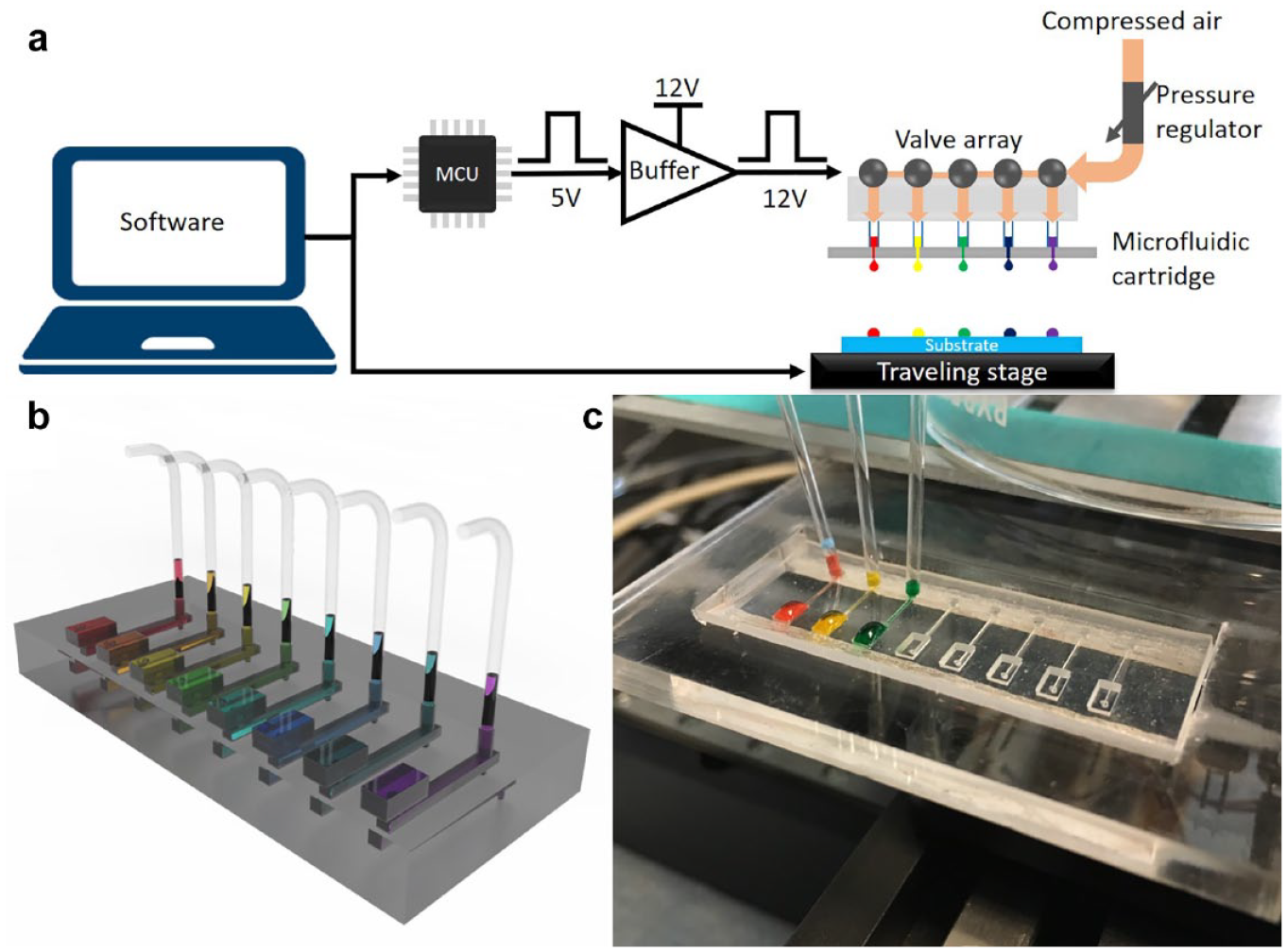
The MPP platform setup is shown in Figure 1a , which includes a printer head, a two-dimensional traveling stage (Speed Limit 300 mm/s; Thorlabs Inc., NJ, USA), and peripheral control circuits and software. Software is customized so that the user can design a printing pattern in Excel and import it into the software, which then directs the movement of the traveling stage and printer head. By coordinating the printer head dispensing and traveling stage movement, different patterns in an array format can be generated.
(
The printer head is an assembly of a controlling valve array and a multichannel microfluidic cartridge, as seen in
PDMS Sandwich Structure Preparation
Prior to printing, a PDMS-coated slide (2 × 3 inches) was prepared by spin coating (at 9000 rpm, 30 s) 100 µm of PDMS on a glass slide. A 250 µm PDMS layer was directly purchased (Rogers BISCO, CT, USA) and laser cut (Universal Laser Systems) into a rectangular frame shape and plasma bonded with the PDMS-coated slide. Such a two-layer structure was used as substrate for the following droplet array printing, after which another PDMS-coated slide was prepared in the same manner and covered on top of the printed structure, forming a three-layer PDMS–droplet–PDMS sandwiched structure. In order to prevent evaporation, two rows of droplets were printed around the area of interest before the actual droplet array was printed. The sacrificial droplets can increase the vapor pressure of the surrounded array, which served for humidity control purposes.
Glucose Colorimetric/Fluorometric Assay
Glucose, a Glucose Colorimetric/Fluorometric Assay Kit (Sigma Aldrich, MO, USA), and buffer (PbS core-type quantum dots; Sigma Aldrich) were loaded into three channels of a cartridge, respectively. The droplet volume of each channel was first characterized and adjusted uniformly to 10 nL/drop by choosing the corret nozzle size (100 µm thickness, 100 µm diameter) and pulse duration (2 ms) and fine-tuning the air pressure (at 2 psi). Zero, 2, 4, 6, 8, and 10 drops of glucose (concentration: 1 nmol/μL) were printed into a column array format, followed by printing 10, 8, 6, 4, 2, and 0 drops of buffer correspondingly in order to maintain the uniformity of the total volume. Afterwards, 10 drops of assay mix were printed onto every spot. The same experiment was repeated five times along different columns, which resulted in a 6 × 5 array with a total volume of 200 nL on each spot. While such an array provides a study toward the influence of different concentrations with consistent volume, a similar procedure was carried out for studying different volumes with a consistent concentration, where 0, 2, 4, 6, 8, and 10 drops of glucose and 0, 2, 4, 6, 8, and 10 drops of buffer were printed correspondingly, followed by printing 10 drops of staining solution on every spot.
The resulted PDMS sandwiched droplet array was then incubated in an incubator (35 °C) for 10 min. The fluorescence was then imaged by a gel imager at 587 nm wavelength according to the glucose assay protocol. The fluorescence value was calculated by taking the accumulation of each pixel’s fluorescence intensity in the droplet area. The droplet area was selected automatically by the gel imager software (Bio-Rad Image Lab, Bio-Rad, CA, USA), which can autodetect fluorescence and generate reports with quantified intensity.
Enzymatic Assay of Cellulose
The experimental setup is the same as that in the glucose colorimetric assay, only the glucose solutions have been replaced by carboxymethylcellulose (CMC) sodium solution and cellulase solution. Specifically, CMC sodium solution, cellulase solution, buffer, and assay mix were loaded into four channels, respectively, and droplet volumes were characterized and adjusted to 10 nL/drop for consistency. Ten drops of CMC (excess) were printed along each column, while 1–10 drops of cellulase solution were printed along each row. The total volume in each spot was compensated by printing buffer accordingly. Afterward, the droplet array was covered by the PDMS-coated slide as before and incubated for 10 min. The results were obtained by the gel imager (GelDocXR; Bio-Rad, CA, USA) and processed in the same way.
Results and Discussion
MPP Platform Setup
In the current work, we have implemented the MPP platform by integrating the pneumatic printer head with the previously developed control system.
Figure 1
shows the systematic setup of the MPP platform, as well as its key component, the microfluidic cartridge. The MPP system comprises customized software, a control circuit, a printer head, and a traveling stage (
Droplet Size Characterization
Droplet size has been one of the most important characteristics of printing technologies. To achieve fine resolution and flexibility, technology seeks a minimal droplet size with a tunable range. In our MPP system, the droplet size can be controlled by changing the geometry of the cartridge, including the nozzle diameter and thickness, as well as the supply pulse duration of the compressed air. A microfluidic model, based on flow resistance, has been developed for the calculation of the droplet size. Given the scale of the microfluidic cartridge, the liquid performs as laminar flow, which results in flow resistance of the nozzle as
where µ is constant and t and r are the nozzle length and diameter, respectively. Thus, the flow rate (for approximation, assume the pressure given is constant) can be calculated as
where P is the supply pressure. Thus, the resulted droplet size is
In this model, the droplet size is proportional to the fourth power of the nozzle diameter, as well as to the pulse duration and inverse of the nozzle length. To verify the model, three sets of droplets are printed with three sets of different parameters, and the result is shown in Figure 2 . The droplet size is measured by taking pictures of the printed droplets on the PDMS substrate and measuring the droplet diameter, and then calculating the droplet volume based on diameter. In future work, measuring the droplet by weight with a fine-resolution scale would provide better accuracy. For each set of data, three devices were tested, with 20 droplets measured for each device, resulting in 60 droplets per data point. Figure 2c shows that the smallest droplet we managed to print was 2 nL. As can be seen in the figure, the experimental results match well with the linear regression curve, which proves the linear relationship as the equations indicate. The error bar, which represents the standard deviation of multiple measurements (n = 50), shows that droplet volume variation is less than 5%. Further calibration of the droplet size can be done by fine-tuning the printing pressure to ensure that all channels are printing droplets of the same size.
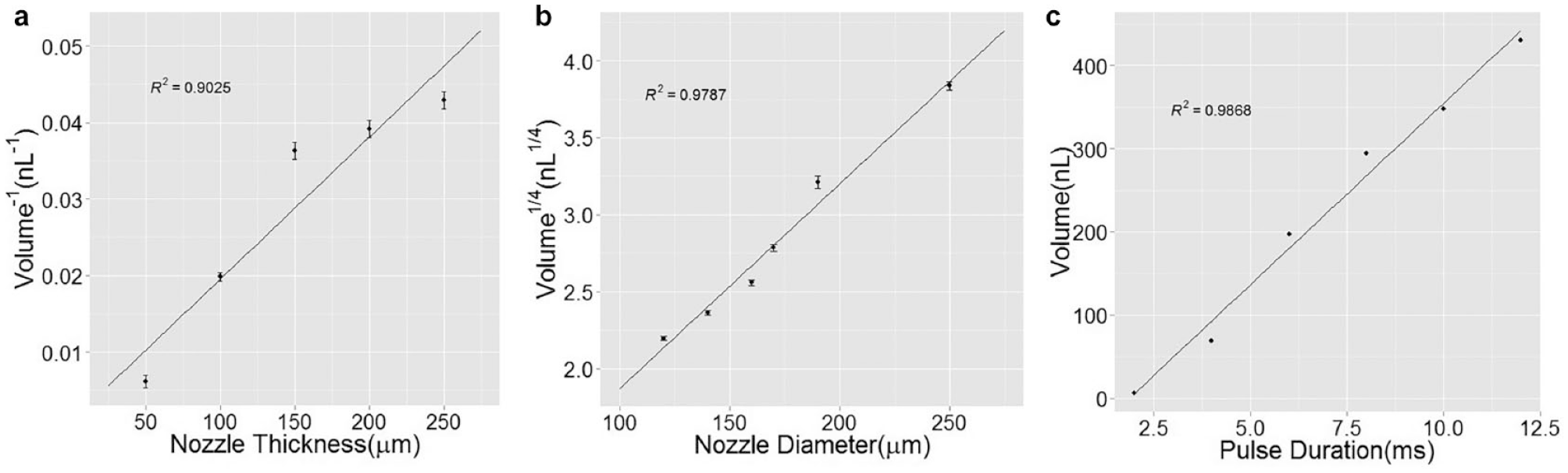
Droplet volume in relation to cartridge geometrical design including (
To demonstrate the capability of the printing platform to generate uniform droplet sizes with different channels, a multicolor droplet array was generated by printing a 1:1 mixture of propylene glycol and deionized water. As a result, a Tetris pattern is printed as shown in Figure 3 . We have measured 100 droplets in the Tetris pattern across three different experiments, and the size distribution histogram can be seen as in Figure 3b . The measured droplet size averages 260 µm in diameter, which guarantees a high droplet density. As can been seen, multichannel printing can be implemented with a single cartridge. A different pattern was also created and is shown in Figure 3c , where three food dyes, red, yellow, and blue, were printed, each in a rectangular array format, and then overlapped, forming different colors. The droplet volume of each spot in a single-color array was around 10 nL. Note that the measure deviation for this result was 7%, which was slightly larger than what was obtained in the droplet size characterization experiment, which is mainly due to the usage of multiple channels, which increased the error. This also shows that the droplets can be printed with minimal misalignment.
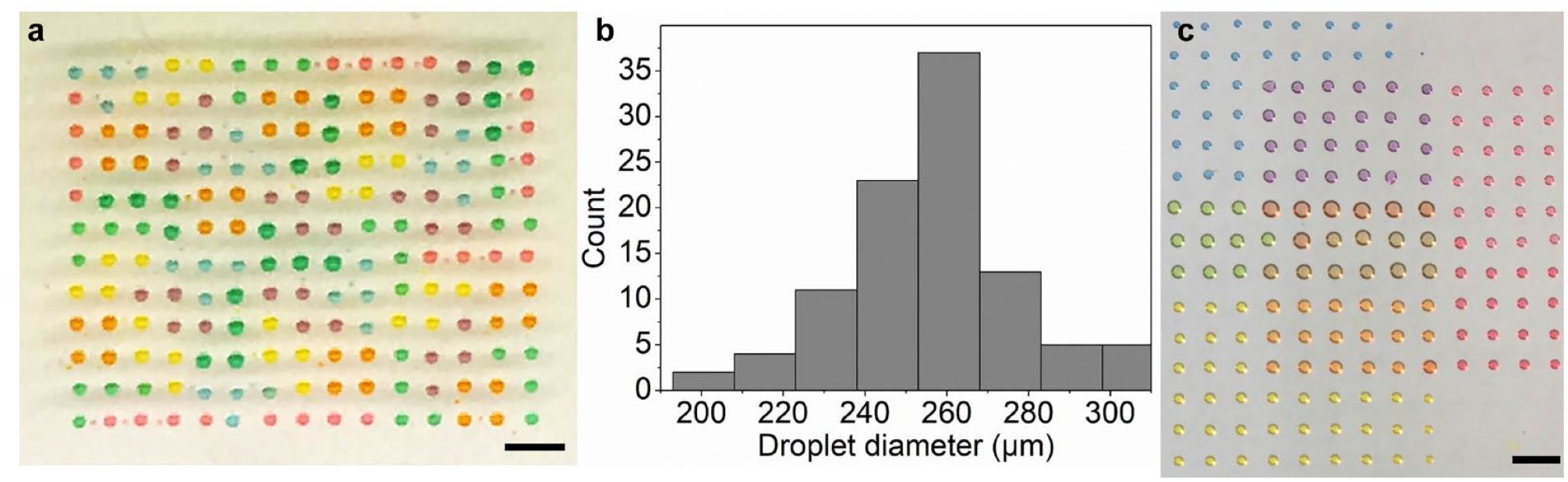
(
Sandwiched Structure of Droplet Array
In order to improve the optical detection uniformity, a cylindrical droplet is preferred over a hemisphere as the flat top and bottom surfaces ensure a uniform optical light path in the whole area of interest.
Figure 4
shows a comparison of a regular droplet array versus a sandwiched droplet array. The static contact angle of aqueous droplets on our PDMS surface is around 90° (
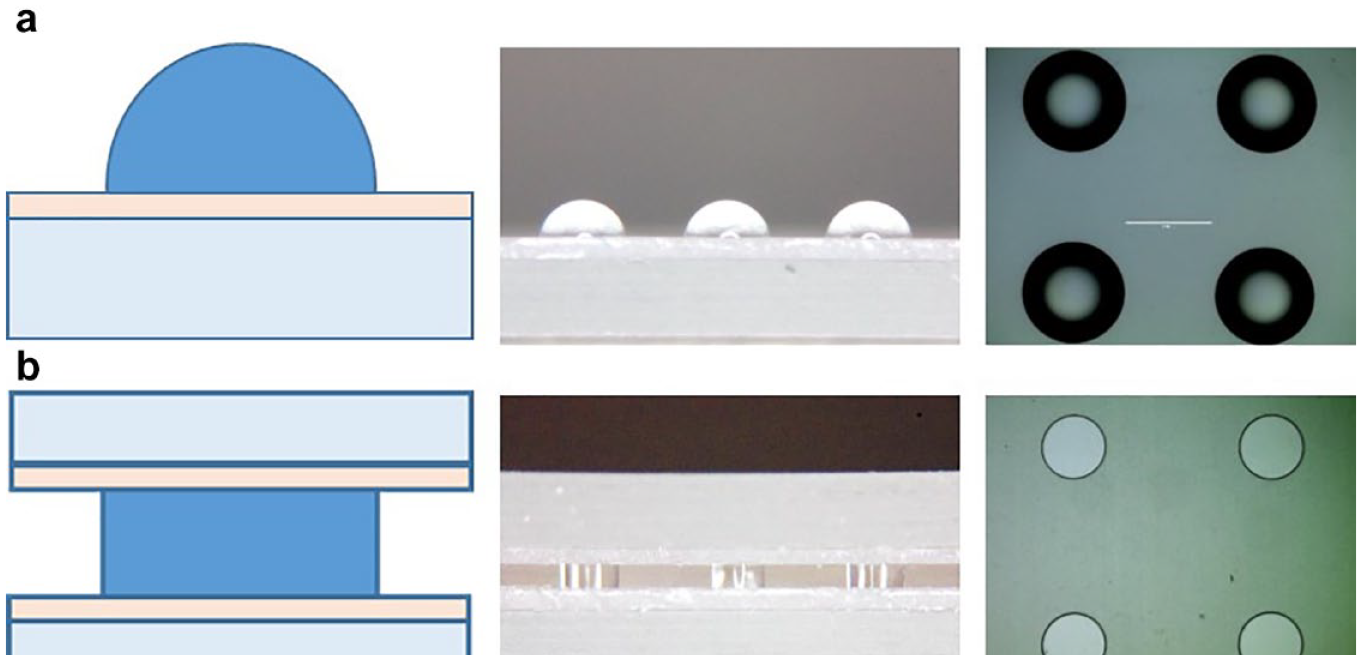

(
Characterization of Light Intensity versus Concentration and Volume
The setup and procedure for the glucose test assay were used for the initial characterization and calibration for intensity against different concentrations and droplet volumes. First, for concentration calibration, a droplet array with a glucose concentration ranging from 50% to 0% (including the assay mix) was created in a droplet array column using the printing platform, while being repeated along each column five times, resulting in a 6 × 5 array. The throughput of the printing procedure is on average ~0.8 s per spot, depending on how limited it is by stage movement. We are currently developing a faster “print-on-the-fly” mode where the printing is performed while the stage is moving. We believe that in the future this will greatly improve the throughput. As can be seen in
Figure 5
, 10–20 droplets were printed at each spot, with all droplets converging into one. This proves the printing accuracy with no misalignment. The fluorescence was then imaged by the gel imager at 587 nm. In this study, the gel imager was applied for the results analysis, allowing for visualization and quantification of the droplet array in a parallel and high-throughput manner. To the best of our knowledge, this is the first time that a gel imager has been used for imaging of a droplet array in a parallel manner.
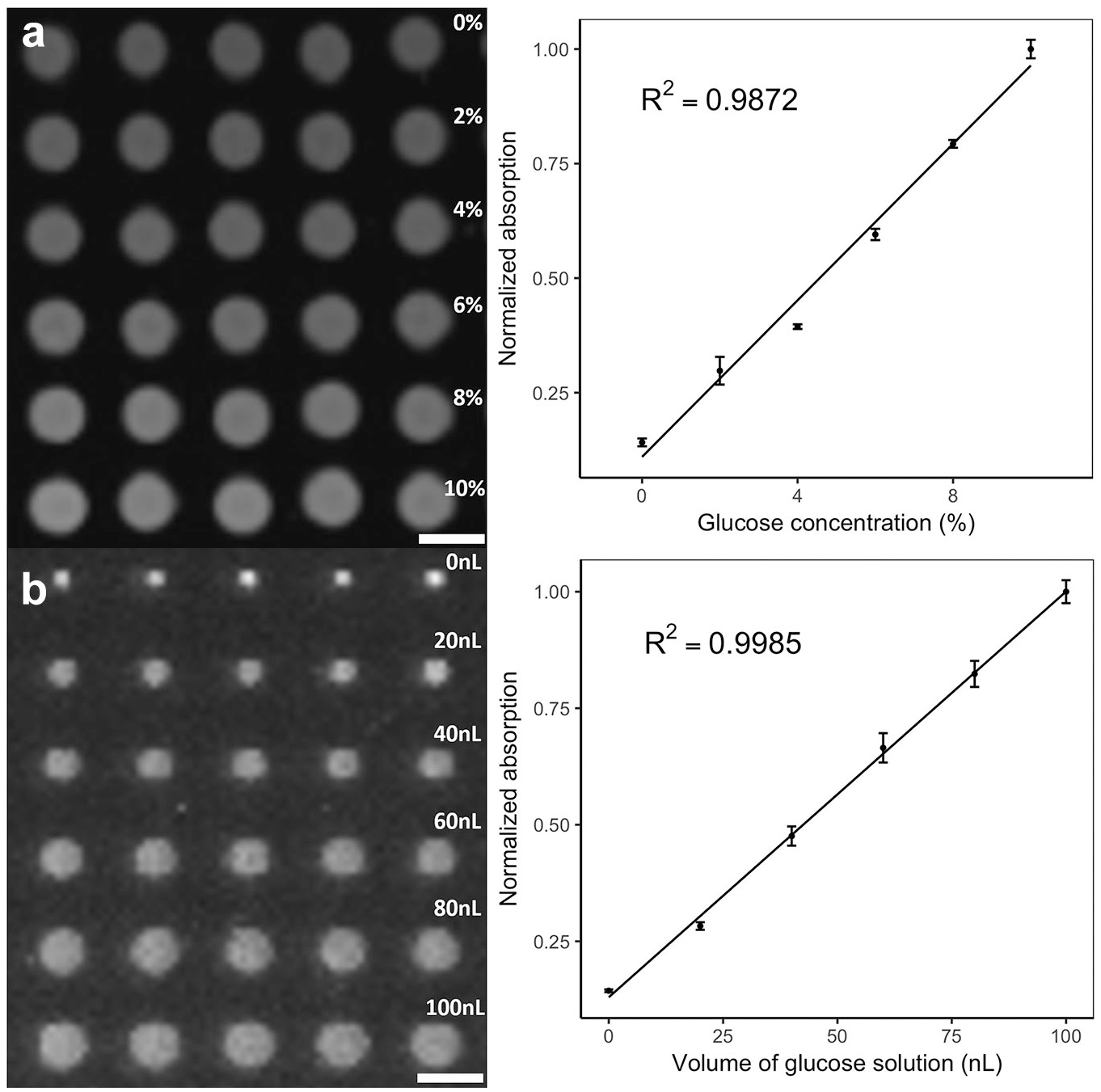
Printed droplet array with concentration gradient and volume gradient. The results are quantified by measuring the normalized absorption versus the (
Analysis of Cellulose Reaction
Given a valid method to evaluate glucose amount with the printing platform, we have gone a step further and applied the platform to the evaluation of enzyme activity. As a proof of concept, cellulase with different concentrations was used to react with the same concentrations of CMC sodium solution. The resulting hydrolyzed glucose level can then be used to reflect the cellulase concentration. Figure 6 shows the result of the cellulose reaction. From the figure, the fluorescence intensity gradient can be observed as the concentration of the cellulase solution decreases. At each cellulase level, as the cellulase concentration decreased, the fluorescence intensity also decreased, indicating a lower level of glucose being produced. This indicates the effectiveness of the cellulose enzymatic reaction analysis. The MPP platform-based enzymatic reaction analysis provides an approach to evaluate different enzyme activities if the same concentration of enzyme solution were being used and different amounts of glucose produced. As the printing platform can generate a high-density droplet array with high frequency using a minute amount of solution, such a high-throughput, low-cost approach shows great potential in the quantitative analysis of an enzyme reaction, and even high-throughput enzyme screening in the future.
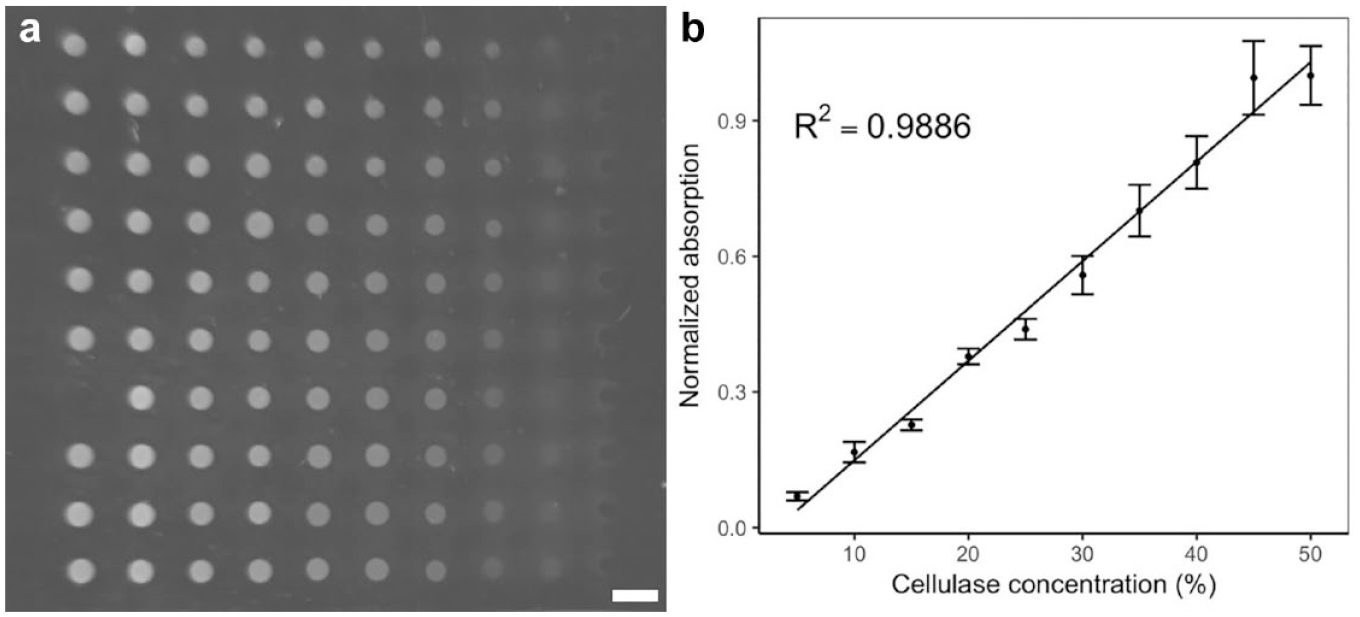
(
Conclusion
This paper reports a novel microfluidic pneumatic printing platform, which is demonstrated to show great potential in application for high-throughput enzyme screening. The MPP platform possesses several distinct features: (1) The detachable cartridge greatly reduces cartridge fabrication cost and avoids contamination, (2) The customized design allows for high flexibility and multiplication. (3) Fast printing speed and small droplet size can reduce chemical waste and increase the throughput. (4) The screening method assisted by the PDMS sandwich structure provides better results in terms of a uniform light path and reduced evaporation. The printing platform was used to carry out a test assay of glucose concentration, as well as an enzymatic assay. The results demonstrate the feasibility and advantage of the printing platform for the application of enzyme screening. With the above-described advantages, the MPP platform can be expected to play an important role in future high-throughput enzyme screening and other fields, such as drug screening and combinatorial synthesis.
Supplemental Material
supplemental_material_JL_et_al_-_revision – Supplemental material for Microfluidic Pneumatic Printed Sandwiched Microdroplet Array for High-Throughput Enzymatic Reaction and Screening
Supplemental material, supplemental_material_JL_et_al_-_revision for Microfluidic Pneumatic Printed Sandwiched Microdroplet Array for High-Throughput Enzymatic Reaction and Screening by Yang Sun, Gang Wang, Jingting Liang, Jiajie Sui, Jinzhen Fan and Jiannan Li in SLAS Technology
Footnotes
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work has been supported in part by National Science Foundation Awards ECCS-0846502 and DBI-1256193 and National Institute of Health Awards NCI-R21CA173243 and NIEHS-P42ES004699, as well as the preparation of highly active modified antimicrobial peptides, national key research and development program, 2017YFD05010000, key technological research on feed utilization of corn straw, major tendering project of science and technology in Jilin province, 20170203009NY, key technological research on recycling utilization of corn straw resource, key technology research and development project in Jilin province, 20180201067SF. J.L. acknowledges the assistance from Paolo Disano for chip fabrication.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.