Abstract
This article presents the design and development of a new hands-free ultrasonication robot for filamentous fungi homogenization. The platform was constructed with a modified inexpensive 3D printer, equipped with an upward-facing camera, a custom-designed wash station, and an add-on sonicator. While machine vision accomplished sample well screening based on image subtraction and color thresholding, it also determined the level of fungi homogeneity using color variance. Model fitting reveals that the process of filamentous fungi homogenization using ultrasonication included a period of significant exponential decay. Therefore, this procedure allowed for the rapid homogenization of the fungal samples during the initial stages of ultrasonication treatment followed by a deceleration in homogenization. Furthermore, a factorial experiment showed that higher sonicator power and temperature accelerated the homogenization process, while the cultivation time exhibited no effect on homogenization. In addition, the model parameters were varied between the wells, even when subjected to the same settings, meaning that the system cannot use the same asymptote of the homogeneity level to establish the termination time for different wells. Therefore, we used the standard deviation of the four most recent homogeneity level values to determine the termination time. This method was used for feedback control, forming a fully automated robot that did not require manual intervention during the experiment. A validation test on filamentous fungi demonstrated that the system was able to provide target quality of samples efficiently.
Keywords
Introduction
Microorganisms broadly serve as cell factories for the production of a variety of important high- and low-value products, such as lipids, biosurfactants, antibiotics, enzymes, therapeutic agents, exopolysaccharides, and bioplastics. It is possible to easily isolate some of these products following extracellular storage and excretion into the cultivation medium. However, others, such as lipids and polyhydroxyalkanoates, depend on intracellular production, necessitating their extraction for analysis via a process usually initialized by cell homogenization. 1 In addition, biochemical fingerprinting of microorganisms by high-throughput Fourier transform infrared (FTIR) spectroscopy, particularly in the case of filamentous fungi,2,3 uses homogeneous cell suspensions that require the application of ultrasonic homogenization.
Generally, it is possible to achieve cell homogenization by using chemical or mechanical/physical methods, 4 and recently, the development of a variety of cell homogenization procedures facilitated the efficient, low-cost, and effective release of intracellular products. 5 The effectiveness of these different homogenization techniques depends on the properties of the microbial cell, such as the physical strength of the cell wall and the location of the desired intracellular product. 5 Ultrasound disintegration—ultrasonication—is a common mechanical cell homogenization method based on the high shear force created by a high-frequency ultrasound (above 16 kHz), 5 which can achieve complete disruption of the microbial cell.5–7 Additional benefits of ultrasonication are that it reduces cell disruption processing time and energy consumption. 8 Unfortunately, however, the process also presents three main disadvantages. First, manual use of the ultrasonication system requires highly skilled technicians to hold, mix, identify, and monitor during processing. 9 Second, the process requires lengthy and repetitive operations, making it unsuitable for screening studies in which a large number of samples need to be sonicated within a short period of time. Finally, high-sound intensity has negative effects on human health, causing headaches, discomfort, and irritation. 10 It is, therefore, important that a platform be developed for the automation of manual ultrasonication operations.
Mancia and Love 11 presented a robotic ultrasound disintegrator with an electronically controlled sonicator probe mounted on a five-axis industrial robotic arm. Similarly, Almo et al. 12 developed an ultrasonication robot equipped with an EPSON robotic arm. In addition, previous work by Li et al. 9 used an ultrasonic module with a FESTO three-axis arm to prepare samples for analysis by FTIR spectroscopy. These systems simply used industrial robotic arms for ultrasonication manipulation without closed-loop control strategies for monitoring or automated control. The open-loop feature might, however, result in insufficient or excessive ultrasonication and, more important, might not always provide well-homogenized samples. Previous studies have provided neither automatic measurement methods for determining the degree of fungi homogeneity nor any mathematical models to quantify the phenomena of filamentous fungi homogenization by ultrasonication. Without appropriate sensing abilities, these machines continue to rely on the manual input of sample locations and technicians to monitor the process, adjust ultrasonication time, and so on.
In the laboratory automation field, many platforms with sensing systems have been developed to automate experiments and tests ranging from robotic sample preparation systems 13 to a machine vision-based system for sorting zebrafish embryos 14 and an automated system to test irrigation sprinklers. 15 Recently, the well-developed, open-source software for 3D printing technology allowed for rapid and low-cost prototyping. Based on a 3D printer, Nejatimoharrami et al. 16 presented a liquid-handling robot named EvoBot while also developing a 3D printer-based platform together with an imaging system for screening microscopy. 17
The main contributions presented in this article are the following:
We developed a novel, fully automated ultrasonication robot for the homogenization of fungal cells. The robot used machine vision to distinguish sample wells and blank wells.
We quantified the fungi homogenization process using model fitting, suggesting that homogeneity level to ultrasonication time of all the tested sample wells can be well fitted with exponential decay equations. A further factorial experiment showed temperature, sonicator power, fungal varieties, and well replicates have a significant influence on homogeneity level.
We developed a feedback control strategy that used the standard deviation of local homogeneity values to determine the ultrasonication termination time.
The proposed system was validated by the homogenization of filamentous fungi but might also be applicable to other types of microorganisms, such as yeasts, bacteria, and algae. The developed ultrasonication robot could be of particular use in processes that prepare filamentous fungi samples for biochemical phenotyping by FTIR spectroscopy, extraction of intracellular metabolites, and DNA from microbial cells.
Materials and Methods
System Design and Hardware Description
Figure 1 shows the hardware assembly of the developed platform. It comprises five functional modules: (1) an XYZ motion stage; (2) a control and vision module, including a controller and an upward-facing camera; (3) a washing module consisting of a pump and a newly designed wash station; (4) a sonicator module; and (5) the power supply. The overall size of the ultrasonication platform, excluding the water bottles, is 59 × 45 × 80 cm (length × breadth × height).
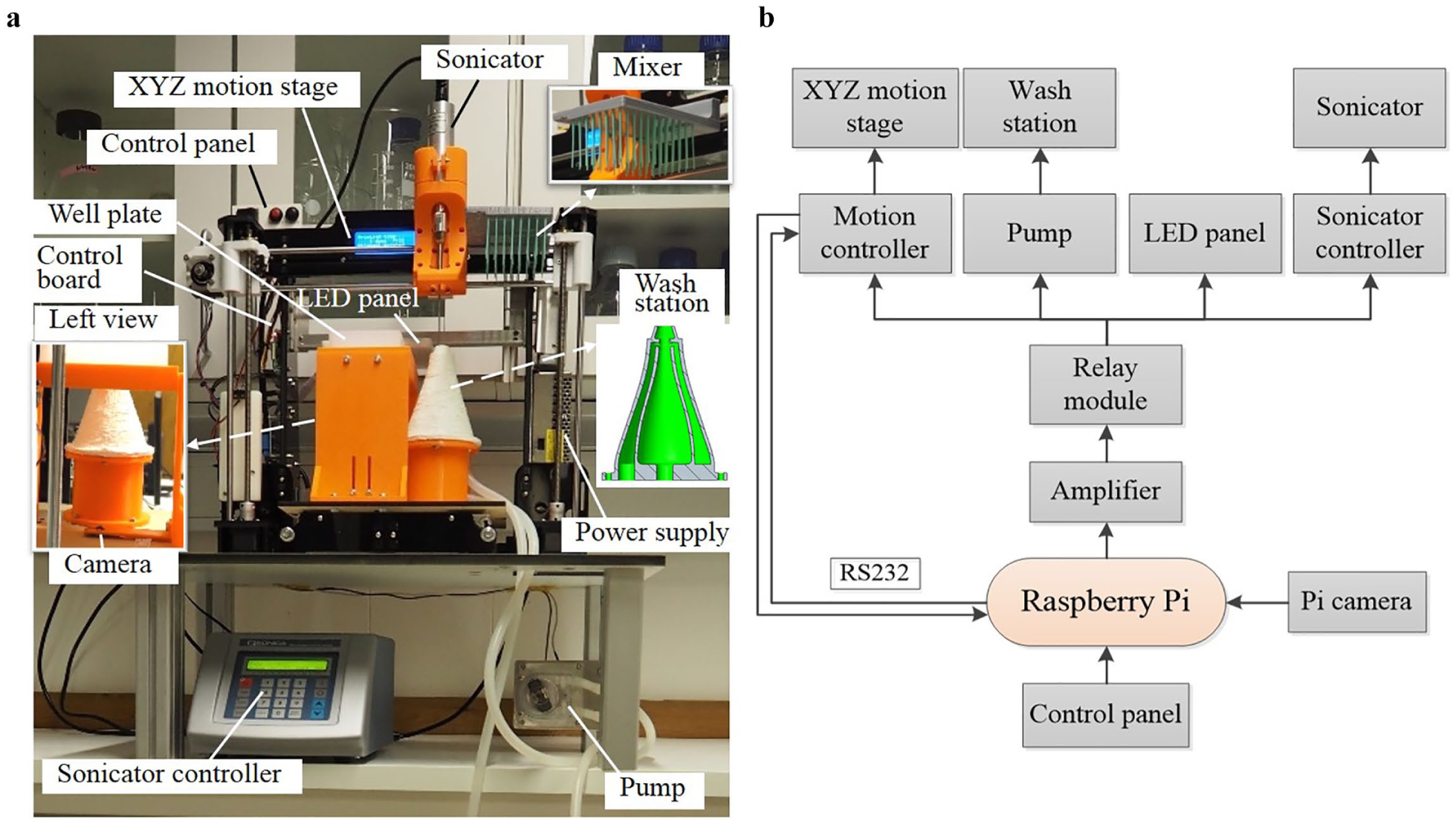
The ultrasonication robot: (
XYZ motion stage
Figure 1a
shows that the modification of a low-cost 3D printer formed the basis for the robot actuation module (Anet A8, Anet, China). This provided three degrees of motion for the sonicator, with an original working space of 220 × 220 × 240 mm. A 3D-printed sonicator holder with which to mount the sonicator on the robot arm replaced the extruder module used for 3D printing (
A laser-cut wooden board attached to a 3D-printed plate holder replaced the hotbed of the printer. Modifications to the original 3D printer controller facilitated motion control while being flashed with open-source firmware (0.92.5; Repetier, Willich, Germany) with a robust motion control algorithm that promoted external control through the G-code protocol. One disadvantage of the firmware is that it does not provide feedback upon completion of motion. However, this feedback is important in minimizing cycle time, since its utilization can trigger precisely sequential motions between the motion stage, pump, and sonicator. Therefore, we added a new function to the firmware to monitor when motions are complete and update the information to the upper controller. Since the robot arm is open-loop controlled, it requires calibration after each restart of the robot, as it is in 3D printing.
Control and vision module
The control board can be found on the left side of the platform. Selecting a Raspberry Pi 3 (Model B, Raspberry Pi Foundation, Cambridge, UK) as the main controller was due to its suitability for both image processing and electronics control. As shown in Figure 1b , the Pi controller has a serial port to communicate with the original 3D printer controller using G-code protocol for motion control. A centrally mounted, upward-facing camera (Raspberry Pi V2; Raspberry Pi Foundation) located underneath the plate holder provided a view of sample plates for image processing. The camera has a maximum resolution of 3280 × 2464 pixels; however, a lower resolution setting of 544 × 640 pixels facilitated fast processing. An LED panel consisting of several RGB LED units (5050; Golden Sun, Shenzhen, China), fixed on the center of the platform provided, a homogeneous light source for image processing. Customization of the LED panel shell allowed for easy attachment to the robot, and the lighting area (132 × 125 mm) was, therefore, large enough to pass light through the 96-well microtiter plate (128 × 86 mm) (CR1496; EnzyScreen, Heemstede, Netherlands).
On the top-left corner of the platform is a user-friendly control panel with one power switch and two buttons (start and pause), which connected directly to the GPIO pins of the Pi controller. An intentionally positioned low-cost four-channel relay module (SainSmart, Lenexa, KS) together with a transistor array (ULN2003A; Texas Instruments, Dallas, TX), segregated the Pi controller from the 3D printer main board. As shown in Figure 1b , the relay module is used to control the 3D printer controller, pump, LED panel, and sonicator controller for on/off operations. The Pi controller commands the relay module with current amplification through the transistor.
Sonicator module, washing module, and power supply
A sonicator (Q125; Qsonica, Newtown, CT) combined with a standard probe facilitated the sonication of fungal cells. The washing module consists of a peristaltic pump (WPL 810; Williamson, Southwick, UK), a newly designed wash station (middle right of
Fig. 1a
and
Image Processing
Measurement of fungi homogeneity level
Accurate measurement of the fungi homogeneity level is essential to close the loop of ultrasonication control. Since the level of fungi homogenization in a well plate varies from well to well, analysis transpired via single-well-based image processing. This necessitated the segmentation of the wells and the identification of their centroid locations in the image pixel coordinates. One simple method was first to calculate the real-world coordinates (
Filamentous fungi have a multicellular structure, where hyphae cells form a complex structure called mycelium. When filamentous fungi were homogenized by ultrasound sonication, we observed disruption of mycelium structure, and fungal cells were homogenized. With time, the ultrasonication treatment reduced the size of single or several large pieces of fungal mycelia in each well. Hence, the first measurement method attempted to calculate the fungal area for each well based on color thresholding. However, because of differences in the fungal biomass quality, amount, and possible fungal varieties in each well, the thresholds varied from well to well. Furthermore, during the ultrasonication, in some cases, the whole pieces of fungal biomass split into several small pieces, making area calculation difficult. Homogenization of the filamentous fungal mycelia disrupted the mycelia into smaller pieces, causing the sample suspension to become more homogeneous. Homogeneity level, on the opposite side, also means variance level. Therefore, this study proposed a robust method based on the color variance within each single well, which calculated and combined all of the three basic color channels, R (red), G (green), and B (blue), ranging from 0 to 255. The total variance
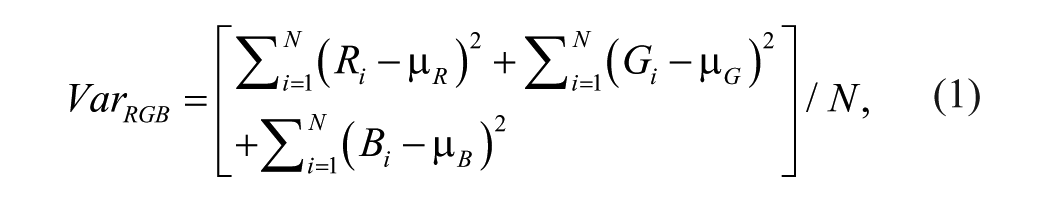
where
Sample screening
The purpose of sample screening was to extract the sample wells (filled with filamentous fungal mycelia) from the well plate so that the treatment could skip the blank wells (empty wells) without any preinput information, a feature that is essential for hands-free operation.
Figure 2a
shows the original picture of a well plate where a few sample wells are randomly located. To identify the sample wells, hue, saturation, and value (HSV) color thresholding was considered, since it is widely used for object segmentation.
19
As mentioned above, due to the different angles at which the camera photographs the wells, the light saturation detected by the camera is uneven, and there are also shadows caused by the wells. This means that fixed global color thresholds cannot work well on different wells in the original picture, as verified by these tests. In addition, different fungal species of samples have different colors and thus require different color thresholds. To make the vision system robust to different wells and types of samples, we used a picture
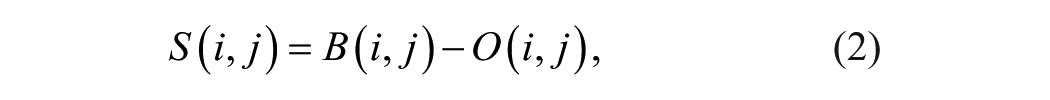
where
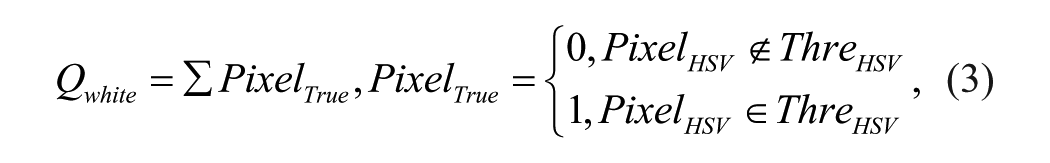
where
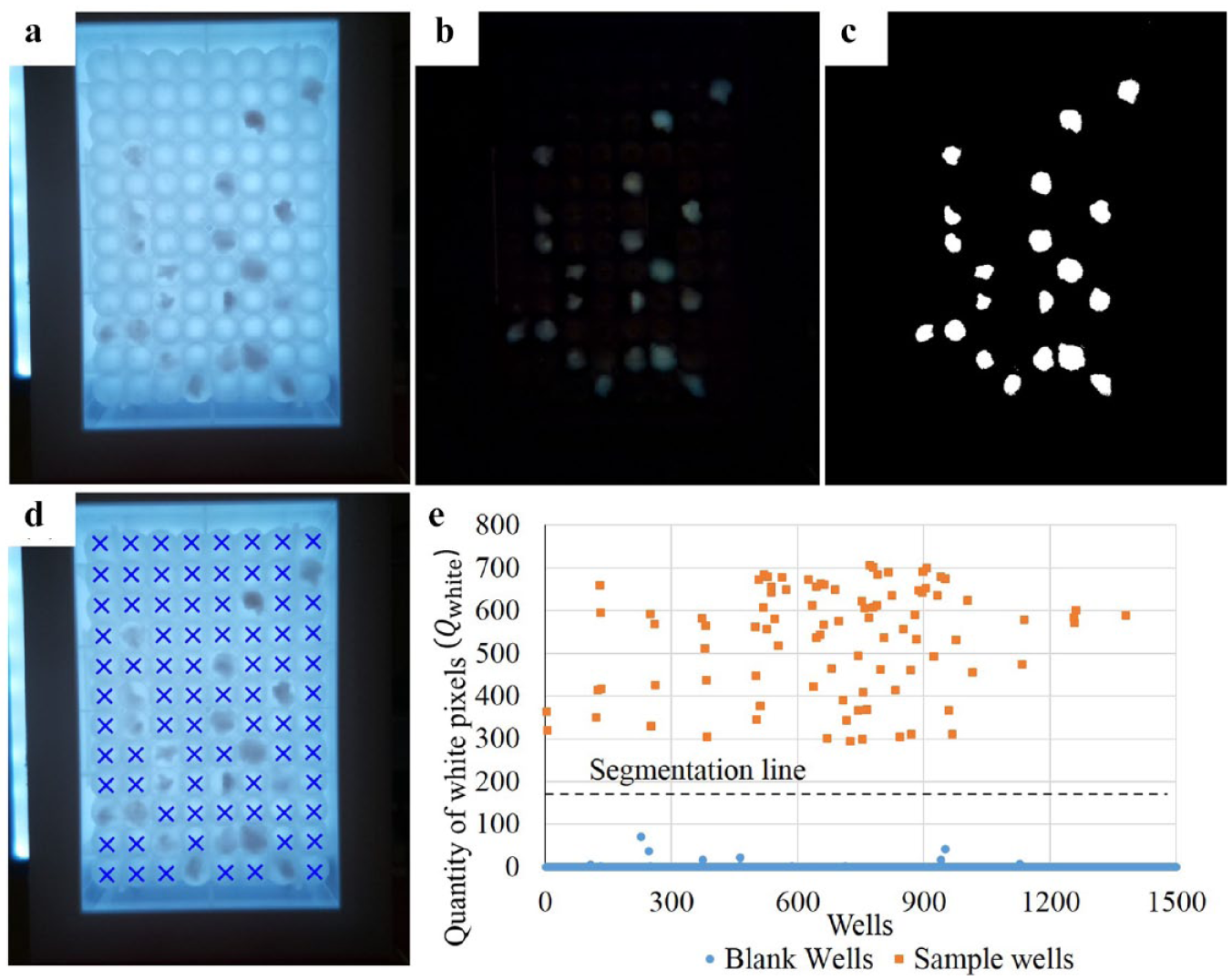
Sample screening approach: (
Furthermore, an experiment performed on fourteen 96-well plates (1344 wells) with different sample well distribution and different types of filamentous fungi determined the extraction threshold of
Filamentous Fungi Samples for the Ultrasonication Experiment
The modeling and control tests involved two varieties of filamentous fungi—namely,
Fungi were cultivated in 800-µL liquid medium malt extract broth (Oxoid, Basingstoke, UK) in 96-square polypropylene deep-well plates using the Duetz-MTPS (EnzyScreen) microtiter plate system for 2 to 3 days at 25 °C. After cultivation, the filamentous fungal mycelia were washed with distilled water in three centrifugation steps. After each centrifugation step, the mycelia formed a pellet at the bottom of wells of the microtiter plate, and supernatant above the mycelia was removed. Thereafter, the mycelia were transferred into new 96-square polypropylene deep-well plates filled with 2 mL of deionized water to homogenize fungi. We used distilled water according to the microbiology protocol for washing mycelia from the growth medium. Cells of mycelia already appeared in swallowed form before washing, since the fungi grew in a liquid medium.
Modeling and Control
Modeling of the Filamentous Fungi Homogenization Process
The method used to measure the fungi homogeneity level was also the technique employed to investigate fungi homogenization changes during ultrasonication treatment. Knowledge of the exact relationship between the fungi homogeneity level and the ultrasonication treatment time could reveal the characteristics behind ultrasonication homogenization of filamentous fungi, which denoted a possible approach to precisely control ultrasonication procedures.
The first experiment intended to observe the
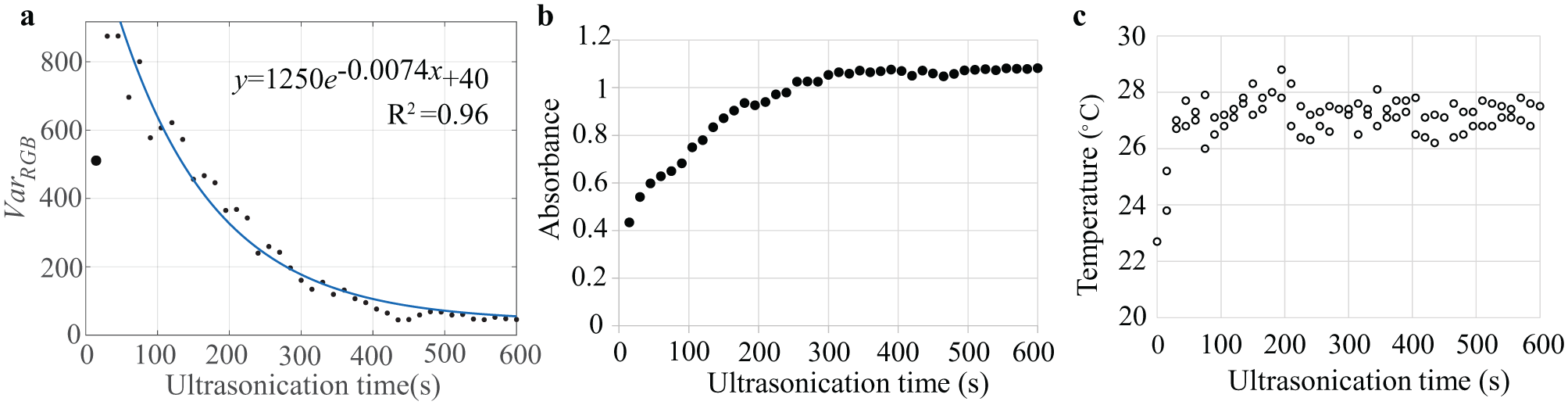
Figure 3
displays the three parameter changes during ultrasonication of a
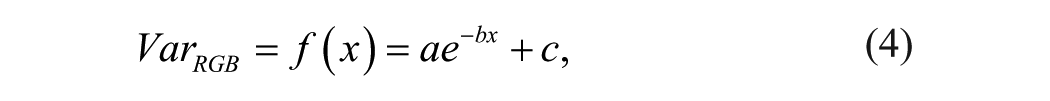
where
To identify whether factors such as temperature, sonicator power, fungi varieties, cultivation time, or even the well positions affect changes in the
This study used a statistical software (JMP 13, SAS, Medmenham, UK) to perform factor effect analysis on all the data using one-way analysis of variance (ANOVA). In this analysis, the response was
System Control
Termination time
Since the timeous termination of ultrasonication ensures efficiency, it is essential to be familiar with the target homogeneity level regardless of the control method employed. However, according to the results shown in
Then we used the standard deviation
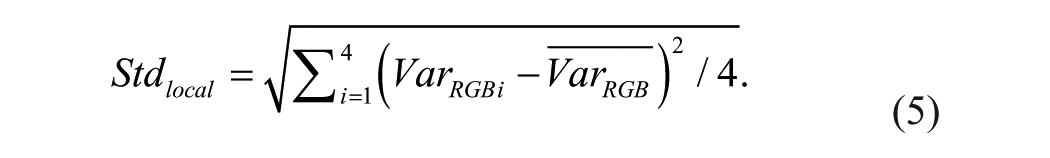
Figure 4
shows the
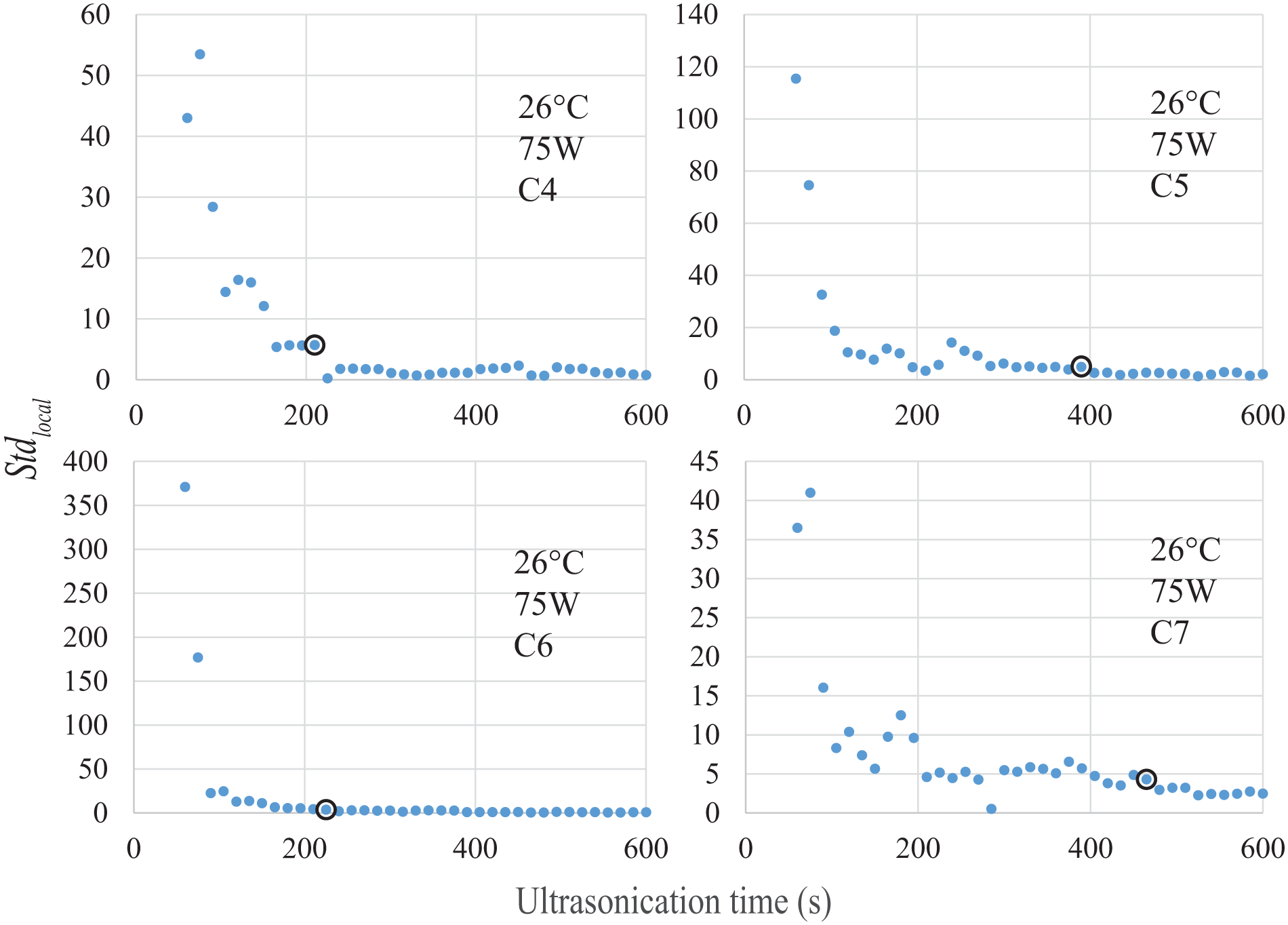
Sequential control
Equation (4) shows promise in model-based control for the ultrasonication process. The idea is that, after an initial period of ultrasonication (e.g., 30 s), the system uses a model from equation (4) to estimate the remaining treatment time based on the difference between the target and current
Due to the limitations of ultrasonication, this system used a screening-based control method for the automation. Similar to the modeling process, the robot treated each well with 15 s of ultrasonication at each interval and monitored the termination time for all wells. When reaching the termination time, the well was passed.
Figure 5
shows the entire control sequence of the robot. The codes in the Pi controller were written in Python 2.7, and the image processing was based on the
Sample screening: After calibration of the arm, the vision system distinguished between blank wells and sample wells. The locations of the sample wells were recorded for ultrasonication, while the blank wells were disregarded.
Feedback control: Once the sample wells had been screened out, the feedback control step was initiated, starting with ultrasonication treatment for a duration of 15 s. During ultrasonication, the sonicator tip in the well moved in a compound manner (circling and up-down) to encourage more homogeneous fungi homogenization, which resembled the way human operators would apply the device. Furthermore, between processing any two wells, the sonicator tip was cleaned thoroughly in the wash station to avoid contamination. As shown in
Figure 6c
, fresh water passes through the outer hollow space of the wash station to clean the sonicator tip and then flows through the hollow inner space into the waste bottle. To avoid droplets from contaminating other wells, the sonicator was powered on for 1 s to remove the droplet from the tip of the sonicator probe after withdrawing it from the well (
Fig. 6b
). In addition, when extracting the sonicator probe following the washing process, the robot touched the inner top of the wash station to remove the droplet (
Fig. 6d
). The top of the wash station has a conical surface for better scraping of the droplets. Subsequently, measurement of the fungi homogeneity levels of all sample wells occurred online using the image-processing system. After completion of the first four treatments, the system began to calculate the
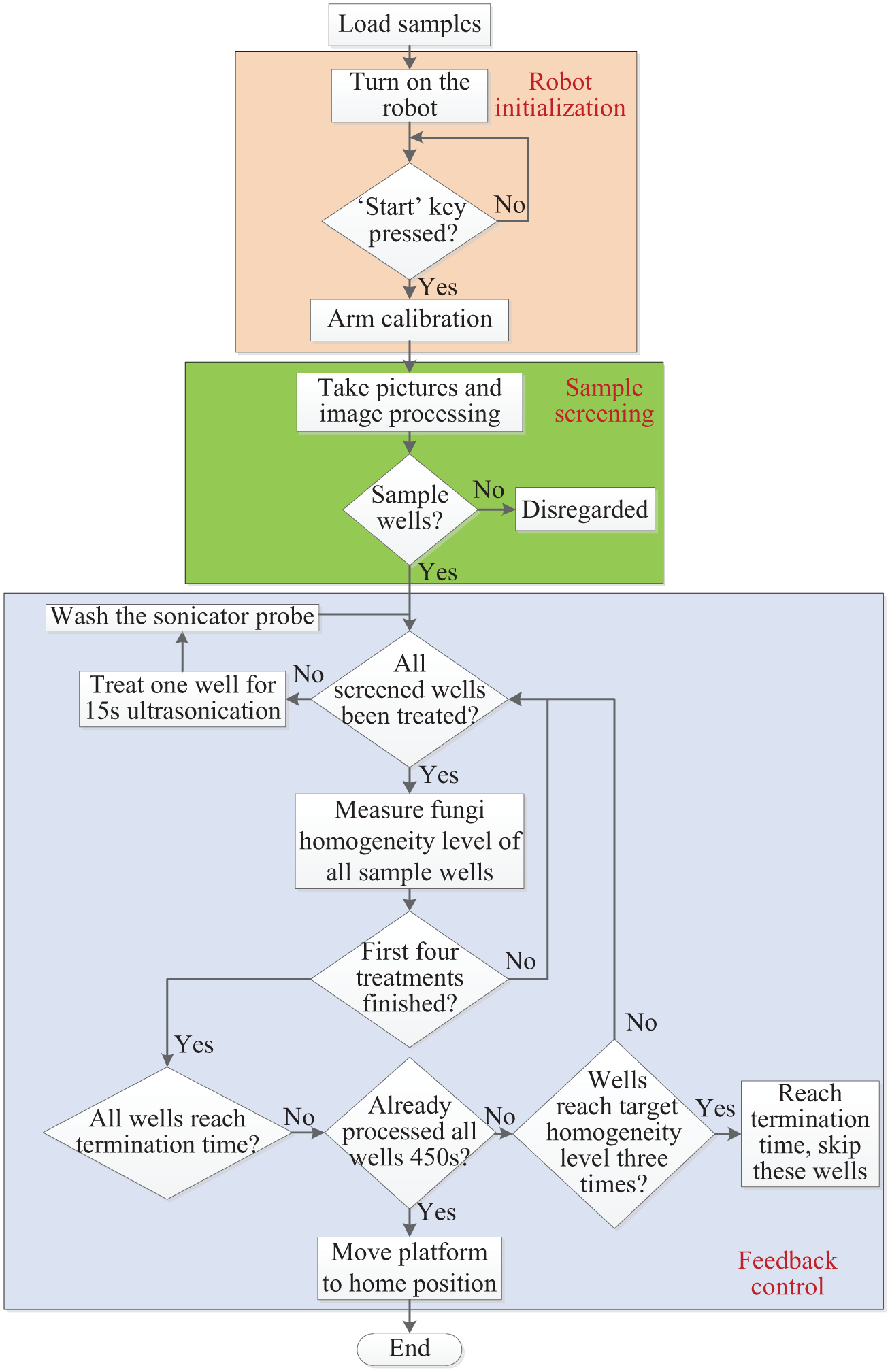
Workflow of the robot control sequence: the ultrasonication sequence can be divided into three main steps: robot initialization, sample screening, and feedback control. The system measured the fungi homogeneity levels of all sample wells after each 15-s ultrasonication and screened them again for a new cycle of ultrasonication until the termination requirements were satisfied or the wells had completed a 450-s processing cycle.
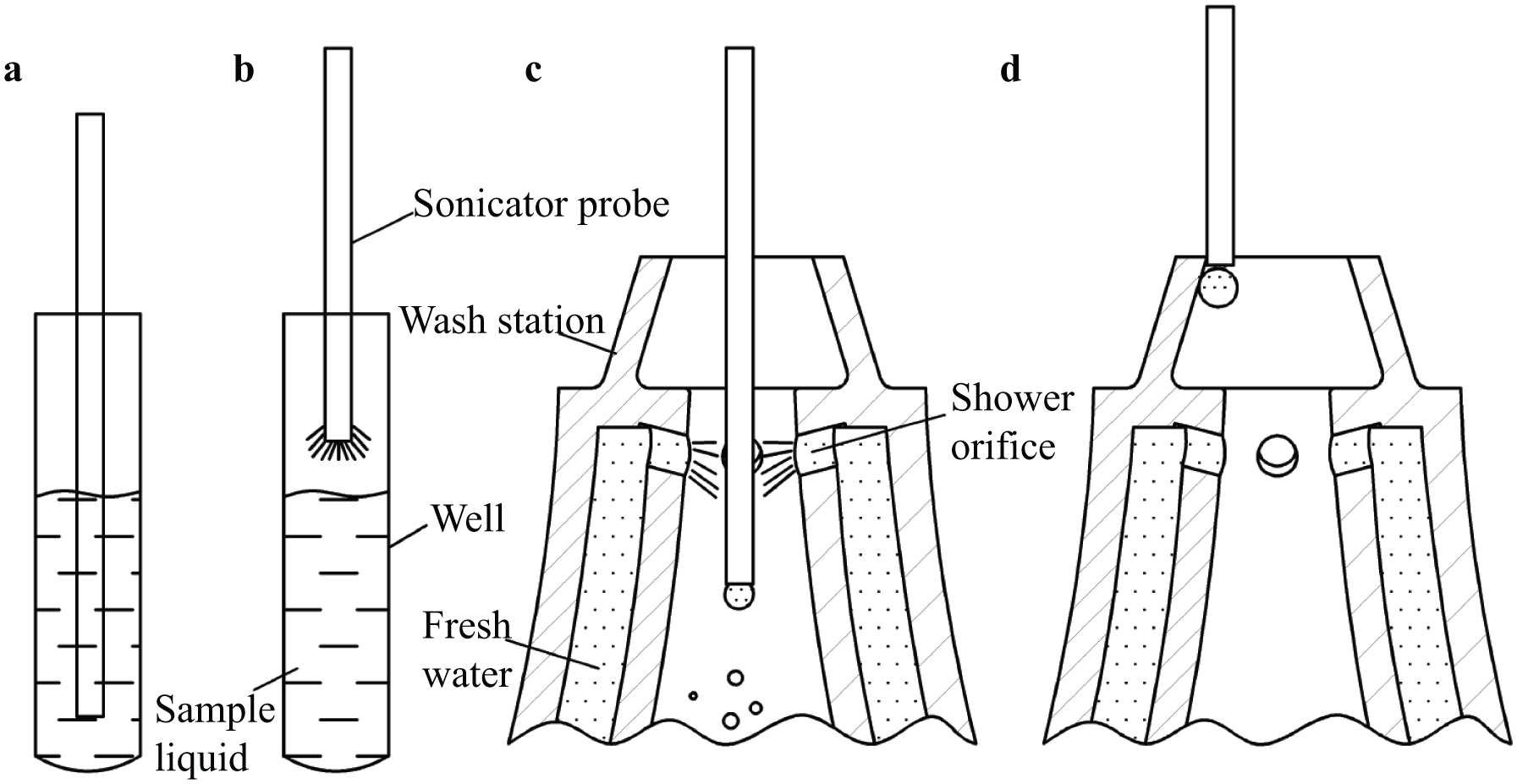
Schematic of ultrasonication and washing steps: (
This screening-based control is simple, is robust, and ensures high homogenization quality. Consequently, the vision-based feedback control system allowed for the full automation of the machine without requiring manual intervention during the experiment.
Results and Discussion
Measurement Precision and Accuracy
To evaluate the precision of the fungi homogenization measurement, 11 wells with different homogeneity levels were measured continuously without any treatment and repeated 25 times. The results, illustrated in
To evaluate the measurement accuracy, this study utilized fully homogenized samples to fill the well plate. As shown in
System Control Result
To validate the control method, an experiment was performed with six
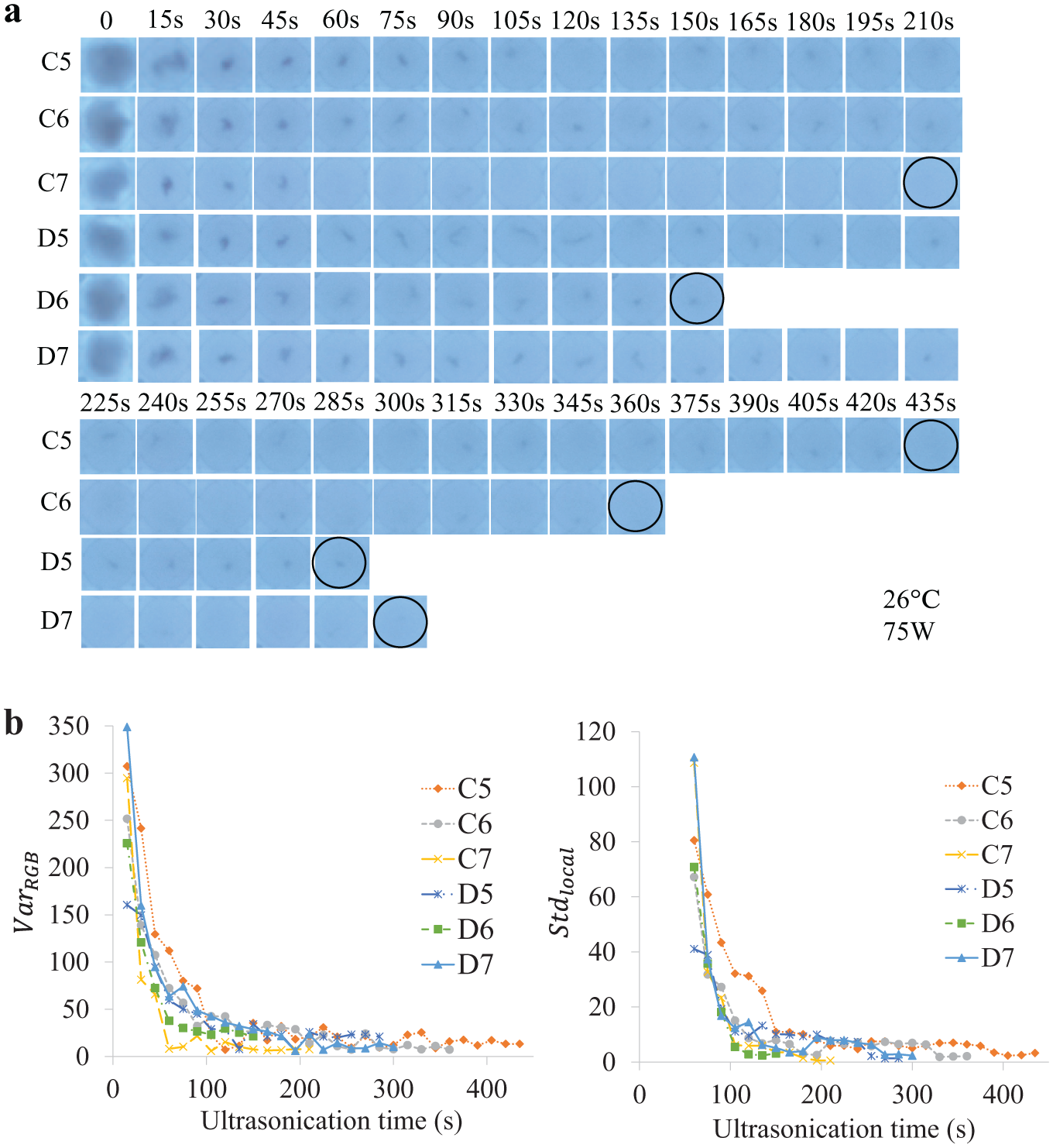
Validation test results: (
Consequently, sufficient homogenization of all the fungi samples transpired and reached similar states comparable to the modeling process (refer to the circled wells in
Limitations and Suggestions
Since the homogeneity measurement method depended on an upward-facing camera, the bottom mycelia primarily determined the color variance calculation. Therefore, the system was not sensitive to the mycelia that remained suspended in the well. On some occasions, the mycelia did not fall to the bottom before the images were captured, which resulted in the acquisition of a smaller
However, based on our tests, for some filamentous fungi, such as
In addition, the
We tested the system with filamentous fungi and found that filamentous fungi homogenization by ultrasonication might be an exponential decay process. However, this theory has not been verified by testing various fungi. Further work should include testing of other types of fungi or even other microorganisms.
In conclusion, this study developed a novel inexpensive platform to automate filamentous fungi homogenization using ultrasonication and constructed the platform hardware by modifying an inexpensive 3D printer with various custom-designed parts and additional electronics. While machine vision distinguished between sample wells and blank wells based on image subtraction and color thresholding, it also measured the level of fungi homogeneity using color variance. The utilization of model fitting indicated that all the tested fungal homogenization using ultrasonication caused significant exponential decay. This process allowed for the rapid homogenization of fungal samples during the initial stages of ultrasonication treatment followed by gradual homogenization. In addition, one-way ANOVA analysis denoted that higher sonicator power and temperature accelerated the homogenization process, while the cultivation time exhibited no effect on homogenization. Moreover, the model parameters varied between the wells, even when subjected to the same settings, meaning that the system cannot use the same asymptote of the homogeneity level to establish the termination time for different wells. Therefore, we used the standard deviation of the four closest homogeneity level values to determine the termination time. This method allowed for feedback control, forming a fully automated robot that did not require manual intervention during the experiment. A validation test on filamentous fungi demonstrated that the system was efficient and able to provide target quality of samples.
Supplemental Material
DS_TECH861361 – Supplemental material for A Laboratory-Built Fully Automated Ultrasonication Robot for Filamentous Fungi Homogenization
Supplemental material, DS_TECH861361 for A Laboratory-Built Fully Automated Ultrasonication Robot for Filamentous Fungi Homogenization by Ya Xiong, Volha Shapaval, Achim Kohler and Pål Johan From in SLAS Technology
Footnotes
Acknowledgements
We thank Julie Eymard for helping to prepare filamentous fungi samples.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by Norwegian University of Life Sciences.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
