Abstract
A single-bead-based, fully integrated microfluidic system has been developed for high-throughput CD4+T lymphocyte enumeration at point-of-care testing. Instead of directly counting CD4+T lymphocytes, CD4+T lymphocyte enumeration is achieved by quantitatively detecting CD4 antigen from the lysed blood sample with a functionalized polycarbonate single bead based on chemiluminescence. To implement the sandwiched chemiluminescence immunoassay with reduced nonspecific binding, a streamlined microfluidic chip with multiple reaction chambers is developed to allow each reaction step to be completed in an independent chamber where reagent is pre-stored. With simple magnetic control, the single bead with an embedded ferrous core can be consecutively transported between each of two adjacent chambers for different reactions. Meanwhile, enhanced mixing can be achieved by moving the single bead back and forth inside one chamber with magnetic actuation. High-throughput detection can be performed when a linear actuation stage is adopted to introduce synchronous magnetic control to multiple single beads in parallel microfluidic chips. A sensitive charge-coupled device (CCD) camera is adopted for high-throughput chemiluminescence detection from multiple single beads. Experimental results show that with the fully integrated microfluidic system, easy-to-operate, accurate, low-cost, immediate, and high-throughput CD4+T lymphocyte enumeration can be successfully achieved at resource-poor settings.
Keywords
Introduction
As a serious global infectious disease, human immunodeficiency virus (HIV) extraordinarily threatens public health with the increasing number of HIV-infected patients. 1 Because HIV-infected patients cannot be cured right now, improving their living status becomes an important goal of clinical treatment. It has been proved that CD4 cells, being one of the important lymphocytes in the body’s immune system, are attacked by HIV viruses, which as a result will weaken the immune system of HIV-infected hosts. 2 Without treatment, CD4 levels will gradually decrease, and eventually HIV-infected hosts will be diagnosed as having acquired immune deficiency syndrome (AIDS) when their CD4 level drops lower than a critical threshold (e.g., 200 cells/µL). 3 It has been confirmed that periodically monitoring CD4 levels has important clinical significance, for example the decision to start antiretroviral therapy (ART) when the CD4 level is lower than a threshold value (e.g., 350 cells/µL). 4
Because of its heavy reliance on infrastructure, for example electricity, reagent, refrigerator, and deionized water, CD4+T lymphocyte enumeration with flow cytometry must be performed in central labs with complicated procedure and high cost. 5 As a counterpart, with a microscope and hemocytometer, one can perform manual CD4 counting with relatively low cost, although it is not a reasonable solution for accurate and efficient detection. 6 Therefore, to perform CD4 counting at point-of-care testing, it is highly desired to develop new methods to implement convenient, easy, low-cost, and efficient CD4 detection. 7 Microfluidic systems for different purposes, which consist of microvalves, micropumps, microchannels, and microchambers, are intensively studied to enable bioanalytical processes to be completed in a simple, rapid, automatic, and efficient way with disposable microfluidic chips.8–10 “Lab-on-a-chip,” or a microfluidic system, has been regarded as a promising solution to provide simple, rapid, low-cost, and accurate diagnosis at resource-poor settings.11–13
Different methods for CD4 counting based on a microfluidic system have been developed based on the chemical, physical, or bioreaction characteristics of CD4+T lymphocytes.14,15 In whole, there are two different strategies to perform CD4+T lymphocyte enumeration: one is to identify and count CD4 one by one, and the other is to quantitatively detect the physical or chemical property related to the amount of a batch of CD4. For the first strategy, how to isolate and recognize each CD4 cell are major tasks with microfluidic chips.16,17 For example, CD4 cells are first captured by a functionalized surface or an array of cell-sized microholes within a reaction chamber to isolate them from other cells, and then each CD4 cell is recognized based on geometry or fluorescence labeling.18–20 In an alternative way, a specific cell-sized microfluidic channel is designed to allow each CD4 to separately go through the detection point where the fluorescence, optical, or impedance signal from CD4 can be recorded for CD4 counting.21–23 For the second strategy, how to quantitatively detect the physical or chemical property of one batch of CD4 related to its amount is the major challenge with microfluidic chips.24,25 For example, semiquantitative CD4+T cell counting is achieved by detecting the monolithic chemiluminescence signal from specifically captured CD4+T lymphocytes in a microfluidic chip with densely arranged microfabricated traps. 26 Or, alternatively, CD4+T lymphocyte enumeration is achieved through detection to the impedance change of the chemically lysed CD4+T lymphocytes. 27 In an even more concise way, a CD4 cell count can be directly figured based on their geometry when they are orderly accumulated together.28,29 Compared to the first strategy, with the second strategy, CD4+T lymphocyte enumeration can be performed with less complicated microfluidic chips and less elaborate companion instruments because there is no need for single-cell CD4 manipulation.
Here, we describe a single-bead-based, fully integrated microfluidic system for high-throughput CD4+T lymphocyte enumeration based on sandwiched chemiluminescence analysis in POC testing. A functionalized single bead with a high surface-to-volume ratio is fabricated to specifically capture CD4 antigen from the lysed blood sample, and its chemiluminescence signal is quantitatively detected for CD4+T lymphocyte enumeration. A streamlined microfluidic chip, which consists of multiple reaction chambers, is developed to allow the single bead to separately react with the sample lysate, wash buffer, horseradish peroxidase (HRP) enzyme, and mixed substrate buffers at different chambers. In addition, the single bead and the required reagents are pre-stored on the fully integrated microfluidic chips; this allows CD4 counting to be thoroughly competed on a disposable chip after loading the lysed blood sample. Actuated by the external magnetic field in one dimension, the single bead with an embedded ferrous core can not only be moved back and forth inside one reaction chamber for enhanced mixing, but also be transported between two adjacent reaction chambers for continuing reaction. High-throughput CD4+T lymphocyte enumeration can be conveniently achieved with a linear actuation stage through synchronous magnetic control to multiple single beads belonging to different microfluidic chips. Chemiluminescence analysis is adopted to reduce the signal background in quantitative CD4 antigen detection. In high-throughput detection, a chemiluminescence signal from multiple single beads in different microfluidic chips is simultaneously collected by a sensitive charge-coupled device (CCD) camera. The critical performance of the developed microfluidic system is validated and confirmed with different experiments.
Material and Methods
Fabrication of Magnetically Actuated Single Beads
A direct way to perform immunoassay-based detection in microfluidic chips is to specifically capture the target through the action between antibody and antigen with the immobilized antibody or antigen on the surface of the microchamber. To improve the detection sensitivity, one solution is to fabricate the specific structure within the reaction chamber to increase the surface-to-volume ratio. For example, functionalized pillar arrays can be fabricated on the surface of the microchamber to improve the chip performance through a high surface-to-volume ratio. Here, in a simple way, a single bead with 2 mm diameter, which is made of polycarbonate, is adopted to capture CD4 antigen from sample lysate through its functionalized surface. The single bead with a high surface-to-volume ratio can be conveniently integrated into a microfluidic chip to cooperate to achieve sensitive CD4+T lymphocyte enumeration. To easily actuate the single bead, two iron balls with a diameter of 0.6 mm are embedded in the single bead as the ferrous core, as shown in Figure 1 .
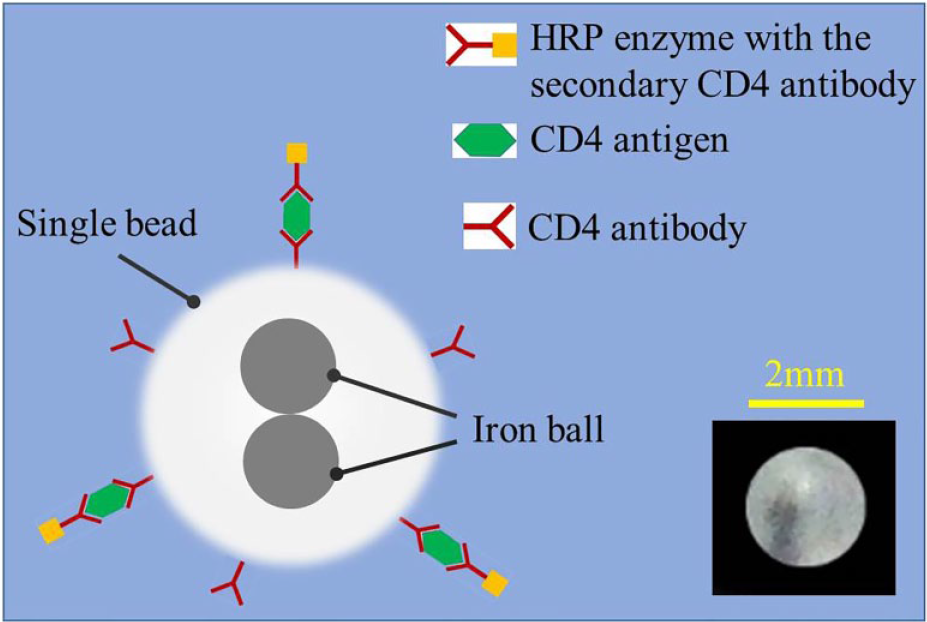
The illustration of the chemiluminescence analysis with the single bead. The bead is immobilized with CD4 antibody first, then the CD4 antibody reacts with the surrounding CD4 antigen in the lysate sample. Finally, horseradish peroxidase (HRP) enzyme is specifically captured by the single bead through the secondary CD4 antibody before it speeds up the chemiluminescence with the mixed substrate buffers. The inset is a real picture of the single bead.
The ferrous core allows the single bead inside the microfluidic chip to be magnetically actuated by the outside controllable magnetic field. The single bead (shown in Fig. 1 , inset) is first drilled by a 0.6 mm drilling bit to a depth of 1.5 mm, then two iron balls are pushed into the blind hole before it is sealed with hot melting. After that, the surface of the single bead is functionalized with CD4 antibody for specific CD4 antigen capturing based on physical adsorption. In principle, the CD4 antibodies will be effectively immobilized on the surface of the polycarbonate bead when a consistent hydrophobic bond is formed between CD4 antibodies and the polycarbonate bead whose surface is naturally hydrophobic. In detail, first, the single bead is incubated with CD4 antibodies overnight at 4 °C. Second, the single bead is washed multiple times with the surfactant wash buffer. Third, the single bead is incubated with casein in the blocking buffer overnight at 4 °C. Finally, the single bead is dried overnight at room temperature. With efficient blocking, the nonspecific binding of the secondary antibody to the single bead is effectively avoided. As shown in Figure 1 , a sandwiched chemiluminescence immunoassay is implemented through the functionalized single bead for CD4+T lymphocyte counting. To detect CD4 antigen, the blood sample is first chemically lysed, which is also helpful to avoid interference from a large quantity of red blood cells, especially when the whole blood sample is tested. Then, CD4 antigens from the sample lysate are specifically captured by CD4 antibodies immobilized on the surface of the single bead. Finally, HRP enzyme specifically captured through the secondary CD4 antibody triggers the intensive chemiluminescence for quantitative detection.
Fully Integrated Microfluidic Chip with Pre-stored Single Bead and Reagent
A fully integrated microfluidic chip with pre-stored reagent is developed to implement CD4+T lymphocyte enumeration with the fabricated single bead. As shown in Figure 2A , the microfluidic chip consists of a reaction module (41×35×8 mm) and an auxiliary module (50×30×16 mm), and both of them are fabricated by polymethyl methacrylate (PMMA) with acetonitrile-based solvent bonding after laser ablation. The reaction module not only provides multiple reaction chambers for the sandwiched chemiluminescence immunoassay, but also pre-stores the required reagent in the same chambers. As shown in Figure 2A , once all the required reagents have been pre-stored in multiple reaction chambers, the reaction module is sealed with aluminum foil from the outside before use. As shown in Figure 2A , similar to liquid reagent, the functionalized single bead is also pre-stored in one reaction chamber. As shown in Figure 2B , before use, the aluminum foil is removed from the reaction module, and then the reaction and the auxiliary modules form a functional chip consisting of multiple reaction chambers with pre-stored reagent. As shown in Figure 2B , the reaction chamber located on one end of the functional chip is responsible for single-bead storage and the CD4 antigen-capturing step, the subsequent four for wash buffer storage and the wash step, the next one for HRP enzyme storage and the HRP labeling step, the subsequent four for wash buffer storage and the wash step, and the final one for substrate storage and the chemiluminescence step. As shown in Figure 2A , especially for the final chemiluminescence reaction, the related reaction chamber is separated into two independent subchambers by an isolation part to allow two substrate buffers to be separately stored before reaction. Once the sample lysate is manually loaded into the reaction chamber with the single bead, CD4+T lymphocyte enumeration is performed by sequentially transporting the single bead between each of two adjacent reaction chambers until it reaches the final reaction chamber for chemiluminescence. Especially after the single bead reaches one of the subchambers for substrate buffer storage, the middle isolation part is manually removed, and the mixing between the two substrate buffers is significantly enhanced when the single bead is moved back and forth by the outside magnet. There is no need for any waste storage chamber because the waste reagent is always left behind the single bead once it is transported to the adjacent chamber for the next reaction step.
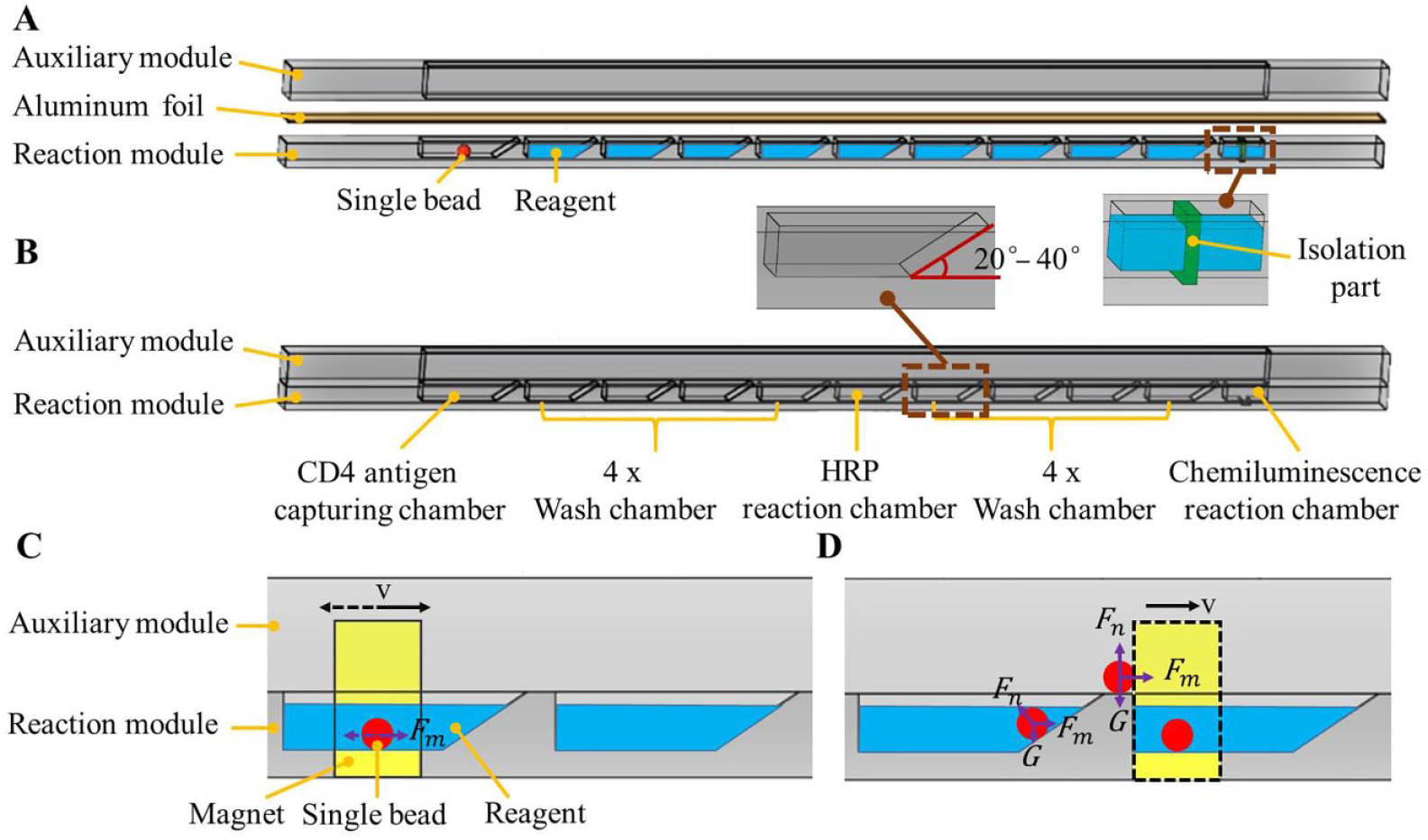
Microfluidic chip with a single bead for CD4+T lymphocyte enumeration. (A) The detailed structure of the reaction and auxiliary modules. (B) Assembled chip with multiple reaction chambers. (C,D) Mixing and transport of the single bead with magnetic control.
As shown in Figure 2C , a permanent magnet, which is mounted on a linear moving stage, is used to horizontally move the single bead back and forth with magnetic control for enhanced mixing when it reacts with the lysate sample, HRP enzyme, mixed substrate solution, or wash buffer in one of the reaction chambers at different times. As shown in Figure 2D , after the reaction in the previous chamber is completed, the single bead is transported into the adjacent reaction chamber via magnetic control when the magnet keeps moving horizontally toward the next chamber. As shown in Figure 2B , once the chip is assembled, an integral wall is formed above the reaction chambers that provides a path for transport of the single bead. The single bead, which is actuated by the outside magnet, is transported by first following a sloping wall (e.g., with an angle of inclination between 20 and 40°, as shown in Fig. 2B ) and then moving along the integral wall formed by the auxiliary module until it reaches the next reaction chamber. As shown in Figure 2D , when the magnetic force (Fm) is large enough to actuate the single bead by overcoming the resisting forces from both the sloping wall (Fn) and the gravity (FG), the single bead will continually move forward until it finally drops into the next chamber. With magnetic control, the single bead can be conveniently actuated by one magnet to achieve enhanced mixing as well as smooth transport between different reaction chambers. As shown in Figure 2B , instead of relying on one single reaction chamber, multiple independent reaction chambers are incorporated into the developed chip to reduce nonspecific binding by allowing each chamber for one reaction step. Without complicated flow control, for example pumping or valving, the developed microfluidic chip is able to perform CD4+T lymphocyte enumeration in a simple, robust, and low-cost way.
Instrument for High-Throughput CD4+T Lymphocyte Enumeration with Microfluidic Chips
As shown in Figure 3 , a portable instrument is developed to perform high-throughput CD4+T lymphocyte enumeration with microfluidic chips. As shown in Figure 3A , only one linear actuation module, which consists of a step motor, a belt, a tray, and a custom positioning strip with multiple thin slots, is required to perform high-throughput detection on multiple microfluidic chips at one time. As shown in Figure 3A , permanent magnets mounted on a tray are used to actuate single beads when they are driven by the step motor through the belt. The movement of each magnet can be flexibly controlled when each of its specific positions is detected through one of the thin slots from the positioning strip, which as a result allows the single bead to be moved back and forth for enhanced mixing, or transported between each of two adjacent chambers. As shown in Figure 3A , a total of three magnets mounted on a tray is required to implement high-throughput detection with at most four microfluidic chips at one time. In principle, more magnets can be easily added to allow high-throughput detection with more microfluidic chips. For three magnets, two of them are used to respectively actuate two single beads in two separate chips, and the third one (sitting in the middle) is shared by another two separate chips. As shown in Figure 3A , based on synchronous magnetic control to multiple single beads, convenient high-throughput detection can be achieved within multiple chips with the streamlined structure.
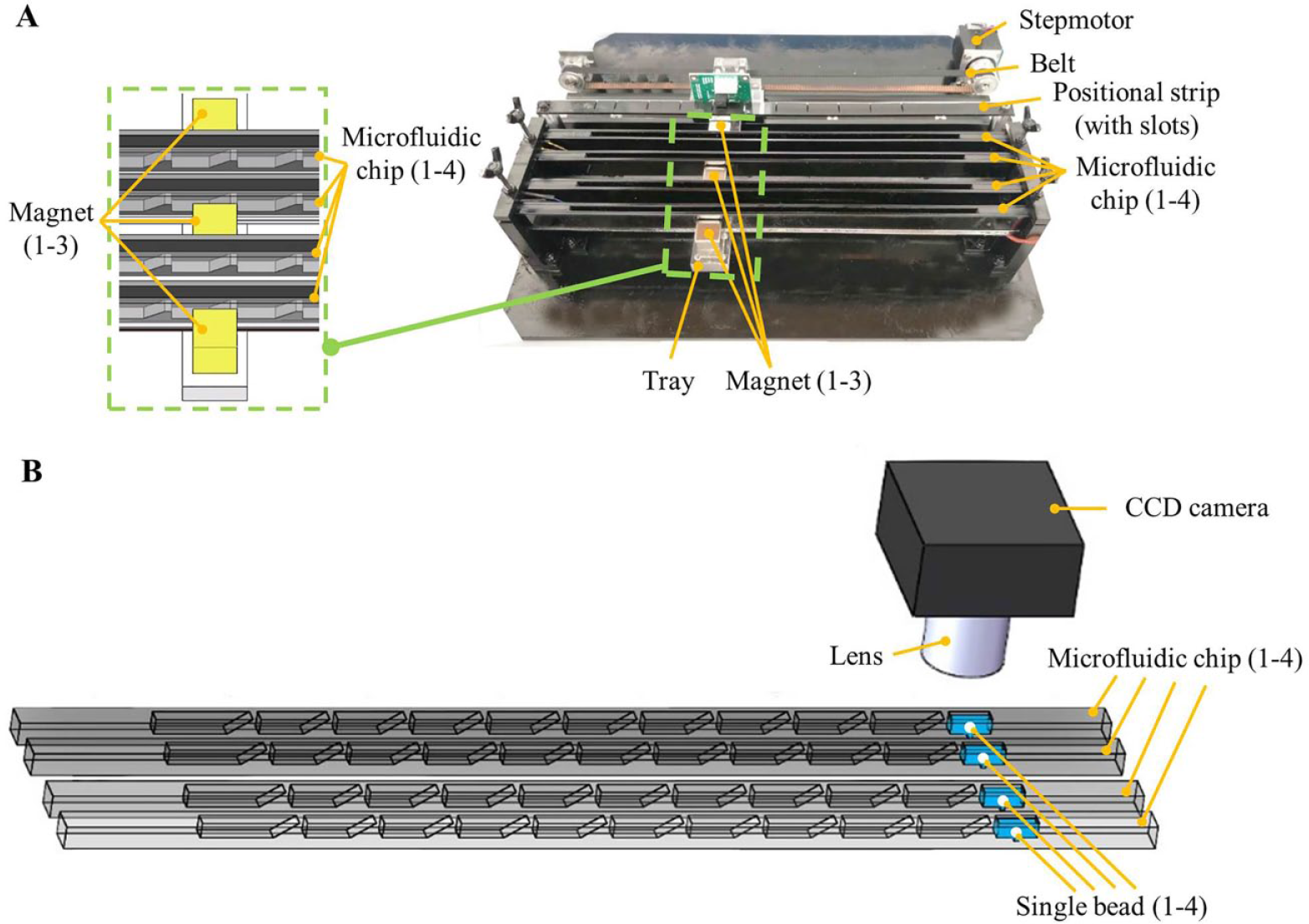
Instrument for high-throughput CD4+T lymphocyte enumeration with microfluidic chips. (A) Picture of the high-throughput detection system with one linear actuation module. (B) Schematic of a collection of chemiluminescence signals from four separate single beads with a charge-coupled device (CCD) camera.
As shown in Figure 3B , for high-throughput detection with four microfluidic chips, a 16-bit CCD camera (ST-1603, SBIG), which is located above the four parallel reaction chambers, is used to take the chemiluminescence picture from four single beads. Custom software developed with Visual Studio 2010 is used to control the CCD camera with a laptop, for example to take pictures or set the exposure time. Meanwhile, a custom image-processing algorithm was developed to automatically analyze the chemiluminescence signal intensity for each of the single beads in high-throughput detection. The procedure of the sandwiched chemiluminescence immunoassay performed on the developed microfluidic chip system is quite similar to the traditional enzyme-linked immunosorbent assay (ELISA) except for the difference in signal detection. As has been done with ELISA, 30 the reaction efficiency of the enzyme-based immunoassay can be improved by heating the reaction chamber to a proper temperature. Therefore, to further improve the detection sensitivity with reduced incubation time, all the microfluidic chips are simultaneously heated to 37 °C by a heating block from the bottom when single beads react with sample lysate or another reagent. To avoid negative effects from outside environmental light, the mechanical actuation subsystem, the electrical control subsystem, and the detection subsystem with microfluidic chips are all enclosed in a small box.
Chemiluminescence Analysis in Streamlined Microfluidic Chips
First, 30 µL test sample is mixed with 210 µL lysis buffer outside the chip, and then a total of 240 µL sample lysate is manually loaded into the reaction chamber with a single bead and incubated for around 10 min. It has been proved that 10 min is enough for a reasonable detection signal to be achieved even with low positive samples based on the systematical experimental results (data not shown here). Meanwhile, the single bead is moved back and forth within the reaction chamber with a speed of 1.5 mm/s through the magnetic control to improve the binding efficiency of CD4 antigen from the sample lysate with enhanced mixing. In the following steps, the single bead is always moved back and forth (with a speed of 1.5 mm/s) through the magnetic control when it is incubated with other reagents in different reaction chambers. After that, the single bead is transported into the next wash chamber with 260 µL wash buffer and incubated for 1 min. There are four adjacent wash chambers to repeat the wash step one time in each chamber; it has been proved that four wash times are necessary to reduce nonspecific bonding as a compromised solution based on the systematic experimental results (data not shown here). Second, the single bead is transported into the next reaction chamber with 160 µL HRP and incubated for 10 min for specific enzyme binding. After that, the single bead is transported into the next wash chamber with 260 µL wash buffer and incubated for 1 min. There are four adjacent wash chambers to repeat the wash step one time in each chamber; again, the wash is repeated four times to reduce nonspecific bonding as a compromised solution based on the systematic experimental results. Finally, the single bead is transported into the reaction chamber with 100 µL of two mixed substrate buffers (50 µL each), and the chemiluminescence with the mixed substrate buffers around the single bead is significantly enhanced by the HRP enzyme due to catalysis effect. CD4+T lymphocyte enumeration can be achieved by quantitatively analyzing the intensity of the chemiluminescence signal of the single bead collected by a CCD camera. With the improved mixing efficiency, the incubation time for each reaction step is significantly reduced, which allows detection with the developed microfluidic chip to be completed within just 30 min. Moreover, the chemiluminescence analysis with the streamlined microfluidic chip can drastically reduce nonspecific binding and reaction based on the rule of one chamber for one reaction step.
Results and Discussion
A calibration model between the concentration of CD4 antigen in the sample lysate and the corresponding CD4+T lymphocyte count in the blood sample is studied and developed with flow cytometry. It is found that 1 RU/µL (RU/µL is a relative unit calibrated by Beijing Wantai Biological Pharmacy Enterprise Co., Beijing, China) of CD4 antigen concentration in the sample lysate corresponds to 1 cell/µL of CD4+T lymphocyte count in the blood sample. Therefore, based on the quantification model, the number of CD4+T lymphocytes in a unit volume can be determined from the concentration of CD4 antigen that is achieved with the developed microfluidic system. Significant signal reflection is found from the inside wall of the detection chamber when the microfluidic chip is made from transparent PMMA. Therefore, to achieve low and clear background around the single bead in detection, black PMMA is used for chip fabrication to achieve the desired signal-to-noise ratio.
Comparison of Protocols with Different Wash Times
To optimize the protocol for CD4+T lymphocyte enumeration with the developed microfluidic chip, two protocols with different wash times were compared for optimal chemiluminescence analysis. For protocol A, after the single bead reacts with the sample lysate or the HRP enzyme, it is washed four times. In contrast, for protocol B, each wash is repeated only twice after the single bead reacts with the sample lysate or the HRP enzyme. In the experiment, lysate of white blood cells was used as the test sample, and a CCD camera (exposure time: 1 min) was used to take pictures from each single bead for chemiluminescence signal collection. As shown in Figure 4 , the detection result with protocol A (each wash repeated four times) is much better than that with protocol B (each wash repeated twice). The experiment was repeated three times, and similar results were attained.
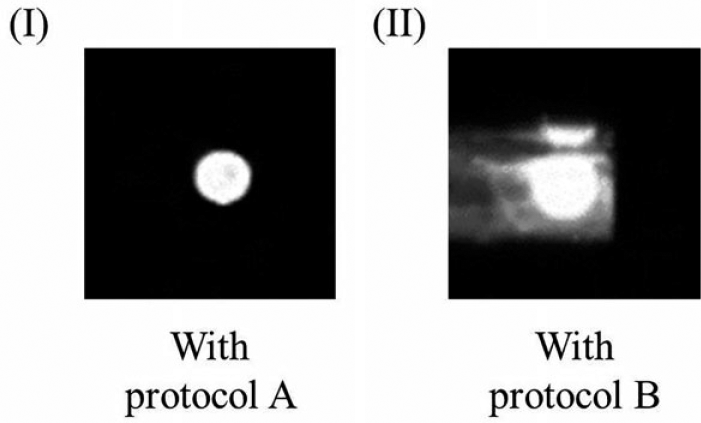

Chemiluminescence image of the single bead in the reaction chamber, which is achieved with (A) protocol A and (B) protocol B.
As shown in Figure 4B , witnessing the extraordinarily high and complicated background around the single bead caused by nonspecific binding and reaction, it is possibly not sufficient to wash the single bead just twice (with protocol B) after it reacts with the sample lysate or the HRP enzyme. In contrast, as shown in Figure 4A , with protocol A, detection with low and clear background around the single bead can be achieved when the single bead is washed four times after each reaction. Therefore, to achieve desired chemiluminescence analysis with a high signal-to-noise ratio, protocol A is chosen to perform single-bead-based CD4+T lymphocyte enumeration.
Sensitivity Evaluation of Single-Bead-Based CD4+T Lymphocyte Enumeration
It has been found that after 2 min with chemiluminescence reaction, the signal intensity around a single bead will gradually drop with the elapsing of time because of both the decreasing concentration of the mixed substrate buffers and the continuing HRP consumption. Therefore, to achieve reliable and stable detection, the mean value of the chemiluminescence signal intensity readings from the CCD camera (exposure time: 1 min) in the first 2 min are adopted as the detection result.
In the experiment, lysate of white blood cells was used as the test sample, and a CCD camera was used to take pictures from the single bead for chemiluminescence signal collection. For the positive test, the test sample with CD4 antigen concentration of 75 RU/µL, which corresponds to 75 cells/µL of CD4 count in a blood sample, was used to evaluate the detection sensitivity. Meanwhile, lysis buffer was used in a negative control test. The incubation time for the single bead to react with the sample lysate or the HRP enzyme was set to 10 min in the experiment. Experiments with both positive and negative samples were repeated at least three times. As shown in Figure 5 , the chemiluminescence signal intensity of the single bead subtracted by the background with the positive test (586) is significantly higher than that with the negative test (18).
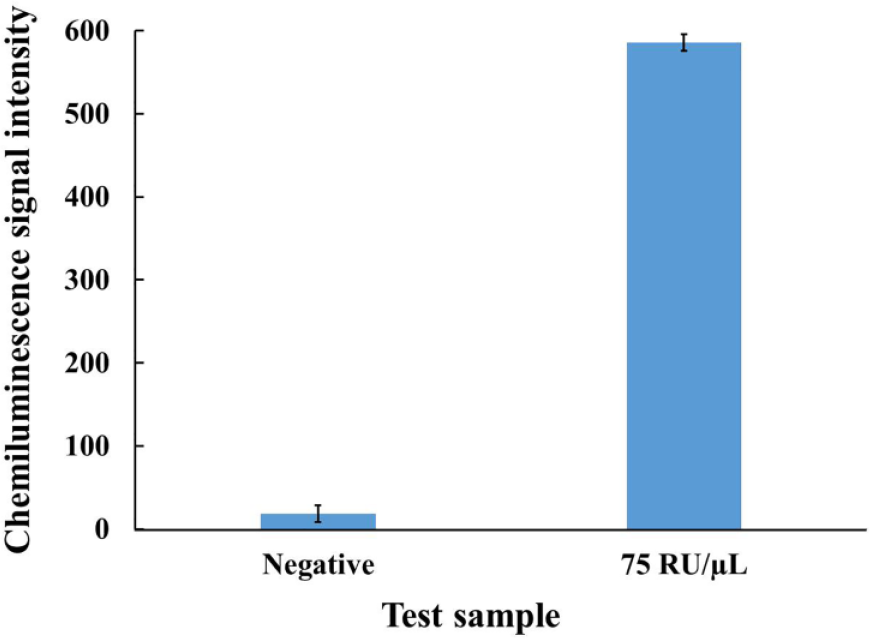
Chemiluminescence signal intensity of the single beads with negative sample and positive sample (75 RU/µL), respectively.
In another way, for the positive or the negative test, the ratio of the signal amplitude between the single bead and the background is, respectively, 1.53 or 1.02, which ensures that the positive test with a low concentration sample (75 RU/µL) can be reliably differentiated from the negative test. Therefore, it is sufficient for the single bead to react with the sample lysate or the HRP enzyme within 10 min to achieve the desired limit of detection (LOD) of 75 RU/µL. Although with an even lower concentration than 75 RU/µL, the detection signal of the single bead is still higher than that of the background (data not shown here), 75 RU/µL is regarded as an acceptable and reliable LOD with the single-bead-based microfluidic system for CD4+T lymphocyte enumeration. Therefore, reliable and rapid CD4+T lymphocyte enumeration with acceptable sensitivity can be completed within just 30 min, which is beneficial to POC testing at resource-poor settings.
Dose–Response Curve of High-Throughput CD4+T Lymphocyte Enumeration
The dose–response curve of CD4+T lymphocyte enumeration, based on the developed high-throughput microfluidic chip system, is established with different test samples with low to high concentrations, including 75, 150, 300, 600, and 1200 RU/µL. In the experiments, lysate of white blood cells was used as the test sample. Meanwhile, lysis buffer was used for the negative control test. Each time, four parallel experiments with four independent microfluidic chips can be performed simultaneously. Experiments with each concentration were repeated three times. The dose–response curve of CD4+T lymphocyte enumeration with the developed high-throughput microfluidic system is shown in Figure 6 . In Figure 6 , each data point is accompanied with an error bar, which represents mean ± SD (standard deviation) and n = 3 (each experiment is repeated three times).
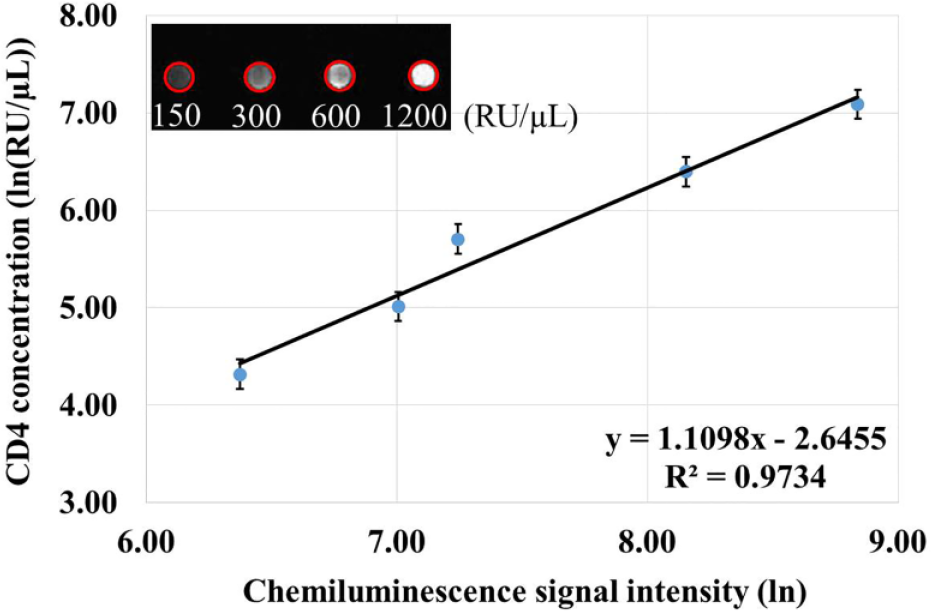
Dose–response curve with test samples of different concentrations (from 75 to 1200 RU/µL). (Inset) A chemiluminescence image of high-throughput detection from four separate single beads (marked with red circles).
For high-throughput detection, a chemiluminescence signal from a total of four separate single beads belonging to different microfluidic chips can be simultaneously collected with a CCD camera. Based on the custom intelligent image-processing algorithm, the chemiluminescence signal intensity of the single bead subtracted by the background is attained and adopted as the detection result for quantitative analysis. As shown in Figure 6 , with the increasing concentration of the test sample, the signal intensity of the single bead increases accordingly. In Figure 6 , the inset is a chemiluminescence image of high-throughput detection from four separate single beads that react with different test samples (150, 300, 600, and 1200 RU/µL). Also in Figure 6 , with the dose–response curve, a desired linear fitting model with an exponential logarithm (the coefficient of determination R2 = 0.9734) between the CD4 antigen concentration or the CD4+T lymphocyte count and the chemiluminescence signal intensity can be properly established. Two test samples with known CD4 antigen concentrations of 200 RU/µL and 800 RU/µL were used to verify the accuracy of the linear fitting model, and their test results were, respectively, 189 RU/µL and 894 RU/µL, with an average detection error of 8.63%. For monitoring of CD4 cells in HIV-infected patients, there are two clinically critical thresholds, 200 and 350 cells/µL. The lower one is regarded as a clinical decision point of CD4 monitoring, and the higher one is regarded as an optimal point to start ART. Therefore, with the developed single-bead-based microfluidic system, accurate CD4+T lymphocyte enumeration with a reasonable detection range between 75 and 1200 RU/µL can be successfully achieved for effective CD4 monitoring of HIV-infected patients at POC testing.
Evaluation of Single-Bead-Based CD4+T Lymphocyte Enumeration with Whole Blood Sample
The performance of single-bead-based CD4+T lymphocyte enumeration with the developed microfluidic system was further evaluated with a total of six whole blood samples (provided by Beijing Wantai Biological Pharmacy Enterprise Co.). In each experiment, 30 µL whole blood sample was first mixed with 210 µL lysis buffer for 10 s outside the microfluidic chip, then 240 µL sample lysate was manually loaded into the reaction chamber with a single bead. Six whole blood samples were also detected with flow cytometry for parallel testing. Each experiment was repeated three times. The correlation curve between the developed method and the flow cytometry is shown in Figure 7 .
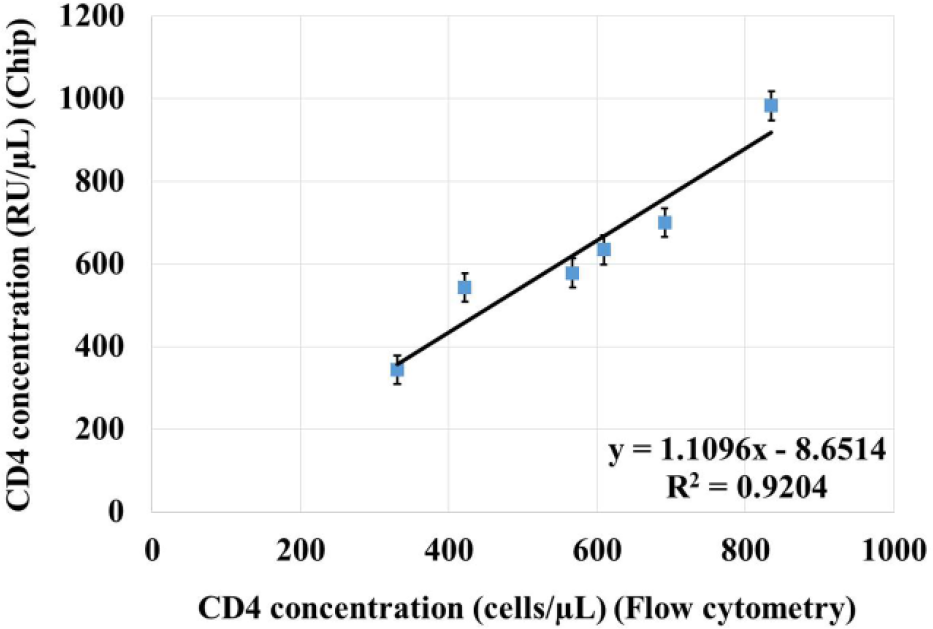
Correlation curve of CD4+T lymphocyte enumeration between the single-bead-based microfluidic system and flow cytometry.
As shown in Figure 7 , the detection results with the developed microfluidic system are quite close to those achieved with flow cytometry. A satisfied correlation model (R2 = 0.9204) between the single-bead-based microfluidic system and flow cytometry is properly developed, and the average error for all the test samples is around 9.6%. Because CD4 is also expressed on monocytes, for most of the existing microfluidic systems, monocytes are normally isolated from CD4+T lymphocytes for accurate counting. Although there are different methods for monocyte separation, the complexity and cost of the microfluidic system will be significantly increased when monocytes are isolated, with elaborate and complicated chip design. However, for POC diagnosis at resource-poor settings, a desired way is to make a reasonable compromise between precision and simplification to achieve relatively accurate detection with convenience, ease, and low cost. Therefore, instead of separating monocytes from CD4+T lymphocytes, both of them are lysed in the whole blood sample simultaneously, then CD4 antigen is detected with the developed microfluidic system. Principally, the detection error caused by the unexpected small amount of CD4 antigen from monocytes can be regarded as a systematic error in CD4+T lymphocyte enumeration, which can be partly compensated by the calibration model between CD4 antigen concentration and CD4 count.
With the developed single-bead-based high-throughput microfluidic system, simple, efficient, accurate, low-cost, and immediate CD4+T lymphocyte enumeration can be successfully achieved within just 30 min at POC testing. Compared to other existing methods with flow cytometry or complicated microfluidic systems, as a proof of concept, it has been demonstrated that high-throughput CD4+T lymphocyte enumeration can be performed conveniently and easily with the developed, much-simplified microfluidic system.
Conclusions and Outlook
A fully integrated microfluidic system with a single bead has been developed to perform high-throughput CD4+T lymphocyte enumeration at POC testing. A single bead of 2 mm in diameter with an embedded ferrous core, which is immobilized with CD4 antibody on the surface, is used to capture CD4 antigen and react with different reagent for CD4+T lymphocyte enumeration. A fully integrated microfluidic chip, which consists of multiple reaction chambers where different reagents are pre-stored in advance, is developed to perform the sandwiched chemiluminescence immunoassay with the single bead. Before use, the microfluidic chip consisting of a reaction and auxiliary modules made from black PMMA is assembled to form the interconnected wall between each two adjacent reaction chambers. Assisted by magnetic actuation, the single bead can be consecutively transported between each of two adjacent reaction chambers along their interconnected wall. Meanwhile, to improve detection sensitivity with reduced reaction time, enhanced mixing can be achieved by moving the single bead back and forth inside each reaction chamber with magnetic actuation. Assisted by transport, the single bead is able to alternately react with lysate sample, wash buffer, HRP enzyme, and mixed substrate buffers in different relative reaction chambers according to the predefined protocol for CD4 antigen detection, which is especially helpful to decrease nonspecific binding.
Because of the streamlined structure with the developed microfluidic chip, high-throughput detection can be conveniently achieved by introducing synchronous magnetic control to multiple single beads belonging to different microfluidic chips with a linear actuation stage. With the developed high-throughput microfluidic system, four experiments can be performed in parallel when four single beads are simultaneously actuated by three shared magnets. In principle, more experiments can be performed in parallel with the developed high-throughput microfluidic system when more magnets are incorporated. To accommodate high-throughput detection, a sensitive CCD camera is used to collect chemiluminescence signals from multiple single beads by taking pictures with a preset exposure time. A custom image-processing algorithm is developed to quantitatively analyze the detection signal for each single bead. Different protocols with changed numbers of washings (wash times) are compared to ensure that desired detection with low and clear background can be properly achieved. The acceptable detection sensitivity is confirmed, and a proper limit of detection with CD4+T lymphocyte enumeration at POC testing is reliably achieved. A dose–response curve established with high-throughput detection shows that a reasonable quantification model with the single-bead-based microfluidic system can be successfully developed for sensitive and accurate CD4+T lymphocyte enumeration. Moreover, the performance of the developed microfluidic system is further evaluated with whole blood samples, and it is demonstrated that accurate and immediate CD4+T lymphocyte enumeration can be successfully achieved by quantitatively detecting CD4 antigen from the sample lysate with the single-bead-based microfluidic system within just 30 min, which is especially appealing to HIV/AIDS diagnosis in resource-poor settings.
In summary, as a proof of concept, an easy-to-operate and fully integrated microfluidic system with pre-stored reagent, which consists of a disposable microfluidic chip with a functionalized single bead and a low-cost companion actuation instrument with one motorized linear stage, is developed for high-throughput CD4+T lymphocyte enumeration. With the advantage of reagent pre-storage, CD4 counting can be automatically completed within the fully integrated microfluidic chip after first loading the lysed sample. Compared to other existing microfluidic systems for CD4+T lymphocyte enumeration, rapid, simple, and low-cost detection can be achieved without complicated actuation and elaborate detection, which is beneficial for CD4 counting in resource-poor settings. A reasonable high-throughput platform is proposed to allow the single bead and the developed microfluidic chip to properly cooperate to achieve sensitive detection with low complexity. In principle, to extend its applications, other different diagnoses can be potentially implemented on the developed high-throughput microfluidic platform by custom making specific beads for different purposes, as well as flexibly rearranging the one-dimensional array of the reaction chamber for different protocols.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work was supported by the National Natural Science Foundation of China (no. 81371711) and the research fund to the top scientific and technological innovation team from Beijing University of Chemical Technology (no. buctylkjcx06).
