Abstract
Metastasis is often critical to cancer progression and linked to poor survival and drug resistance. Early detection of metastasis, as well as identification of metastatic tumor sites, can improve cancer patient survival. Thus, developing technology to improve the detection of cancer metastasis biomarkers can improve both diagnosis and treatment. In this study, we investigated the use of nanodiamonds to develop a stimuli-responsive metastasis detection complex that utilizes matrix metalloproteinase 9 (MMP9) as a metastasis biomarker, as MMP9 increased expression has been shown to be indicative of metastasis. The nanodiamond–MMP9 biosensor complex consists of nanodiamonds functionalized with MMP9-specific fluorescent-labeled substrate peptides. Using this design, protease activity of MMP9 can be accurately measured and correlated to MMP9 expression. The nanodiamond–MMP9 biosensor also demonstrated an enhanced ability to protect the base sensor peptide from nonspecific serum protease cleavage. This enhanced peptide stability, combined with a quantitative stimuli-responsive output function, provides strong evidence for the further development of a nanodiamond–MMP9 biosensor for metastasis site detection. More importantly, this work provides the foundation for use of nanodiamonds as a platform for stimuli-responsive biosensors and theranostic complexes that can be implemented across a wide range of biomedical applications.
Introduction
Metastasis is the process by which cancer cells spread from their original site to distant tissues or organs and is the main cause of cancer-related deaths.1,2 Despite major advancements in diagnosis, surgical techniques, and adjuvant cancer therapies, the majority of cancer deaths are related to metastasis due to failure of treatments because of tumor resistance, direct organ damage, and other complications. 3 Since there is a strong correlation between poor patient outcome and the presence of metastases, early detection of metastases could potentially predict which patients will require additional therapeutic interventions after a successful resection of the primary tumor. 4 This will eventually lead to overall improved survival of the cancer patients.
A number of biomarkers have been identified for metastasis. While there are a number of methods available for the detection of cancer biomarkers, these biomarkers are most often identified as genes, microRNA, or proteins that are uniquely expressed or silenced in metastatic cancer.5–12 Such biomarkers include heat shock protein 70 (Hsp70), epithelial cadherin (E-cadherin), and death-associated protein kinase (DAPK).13–15 Many of these biomarkers were identified through increased understanding of the underlying molecular mechanisms of metastasis progression. Cancer metastasis is a complex process that involves tumor cell dissociation, epithelial–mesenchymal transition (EMT), intravasation, survival in the circulatory system, angiogenesis, and extravasation, with the ultimate establishment of secondary tumors at distant sites.16–19 Notably, EMT is an initial and crucial stage of metastasis, as it involves the detachment of cancer cells from the original tissue. Several proteins involved in EMT have been defined as metastasis markers, including the family of matrix metalloproteinases (MMPs).20,21
MMP proteins are a family of zinc-containing endopeptidases, also known as matrixins. 22 Under normal physiological conditions, MMP proteins are secreted and are responsible for tissue remodeling, inflammation, and tissue repair.23–26 However, in the context of cancer metastasis, MMP proteins mediate degradation of the extracellular matrix that forms physical constraint around tumors.27–30 MMP9 is a well-characterized member of the MMP protein family, and its overexpression has been reported in hepatocellular carcinoma (HCC), malignant lymphomas, prostatic adenocarcinoma, and gastric, glioma, and bladder cancers.30–37 Thus, developing improved methods for MMP9 biomarker detection may serve as a tool for improving metastasis detection.
As an endopeptidase, MMP9 cleaves specific peptide sequences. 38 Therefore, detection of MMP9 peptidase activity through cleavage of MMP-specific peptides may serve as a surrogate marker for detecting sites of cancer metastasis. One such substrate peptide sequence is LGRMRLPGK, which has been shown to be specific to MMP9 but not to other members of the MMP family, such as MMP2. 39 Although this MMP-specific peptide may serve as the foundation for an MMP9 biosensor, additional factors have to be considered before implementation of MMP9-specific peptides for clinical applications, in particular, the instability of the peptides under physiological conditions.40–43 In blood serum, nonspecific proteolysis can rapidly degrade peptides before they reach the target site. As such, good delivery methods to increase the stability of peptides are required for improved efficacy.
A wide range of approaches have been used to improve peptide stability and biosensor efficacy by nanomedical methods. This includes the formation of self-assembling peptide complexes, which can confer increased stability and efficacy. This, however, suffers from the challenges of controlling higher hierarchical structures while maintaining the desired functions. 44 Other common methods include the gold (Au) or titanium oxide (TiO2) nanoparticles functionalized with stimuli-responsive peptides.45–47 While these complexes can confer increased stability, improved delivery, and complementary properties due to their photoresponsive properties, Au and TiO2-based nanocomplexes are not biodegradable. Furthermore, Au and TiO2-based nanocomplexes have been linked to toxicity issues.48,49 As such, these complexes may be more suitable for enhanced, highly sensitive ex vivo biosensor diagnostic applications. 50 Detonation nanodiamonds (NDs) may serve as a useful platform for stimuli-responsive peptide delivery due to a number of favorable properties for drug delivery. NDs are ~3.5 nm truncated semioctahedral carbon structures with a number of physical properties, such as charged surface facets and chemical function groups decorated on their surface, that can impart enhanced functionality to their delivery cargo. As a result, our group and others have demonstrated that NDs are biocompatabile nanomaterials that can improve the delivery and efficacy of a wide range of therapeutic cargo for applications in both cancer treatment and diagnostics.51–61 Because of the versatility of NDs, NDs were evaluated in the development of a stimuli-responsive biosensor complex.
By utilizing a stimuli-responsive MMP9 cleavage system, we can quantify MMP9 activity by stimuli-specific cleavage signal and identify potential metastatic cancer sites. By delivering this stimuli-responsive system using an ND-based platform, we can improve stability and the potential of translating the peptide-based stimuli-responsive biosensor system into clinical use. Thus, the design and initial evaluation of an ND-MMP9 substrate peptide complex will provide evidence for use of this complex as a detection reagent for MMP9 biomarker-based metastasis detection.
Materials and Methods
Design of Stimuli-Responsive MMP9 Biosensor Using Nanodiamond Delivery Platform
The peptide sequence LGRMGLPGK was previously identified to be more specifically cleaved by MMP9 enzyme rather than the rest of the proteins of the MMP family. 39 Hence, this nine-amino-acid peptide sequence was selected as the MMP9 substrate in the MMP9-specific biosensor. In order to improve the stability and efficacy of naïve peptide, ND was used as the delivery vehicle ( Fig. 1A ). MMP9 activity through cleavage detection was designed to be quantified by using a pair of Förster resonance energy transfer (FRET) molecules, fluorescein isothiocyanate (FITC) molecule and 5-carboxytetramethylrhodamine (5-TAMRA), which were placed on either side of the peptide. The FITC molecule is quenched by TAMRA through energy transfer when the peptide is uncleaved. When MMP9 cleaves the peptide, the distance between FITC and TAMRA increases and there is no energy transfer to TAMRA. As a result, an FITC emission signal should be measurable ( Fig. 1B ).
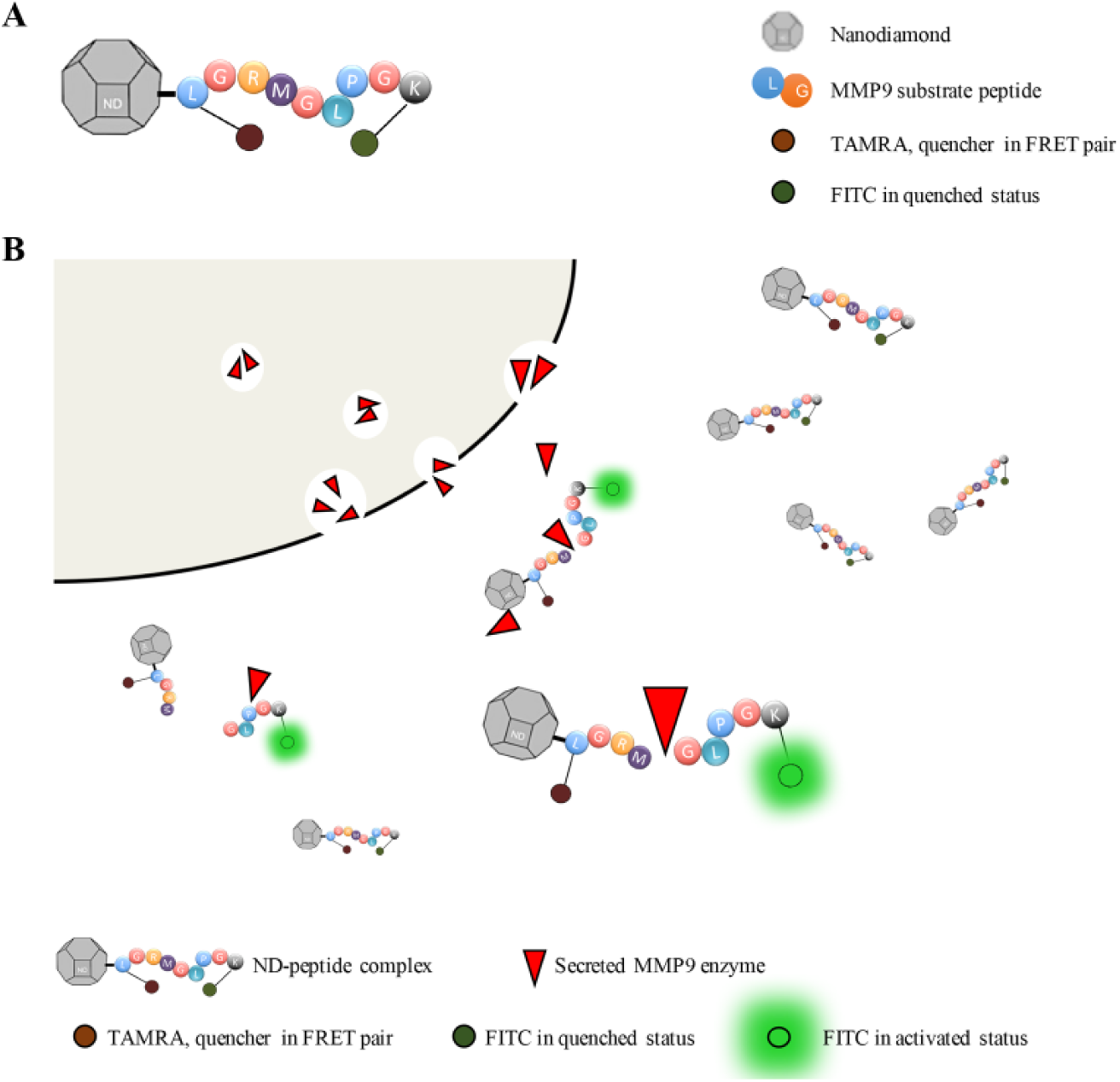
Schematic illustration of ND–peptide cleavage model. (
Preparation of Nanodiamond–MMP9 Substrate Peptide Complex
NanoAmando Soft Hydrogel (NanoCarbon Research Institute, Nagano, Japan) was freeze-dried to obtain ND powder. Carboxyl-modified NDs (ND-COOH) were synthesized according to the previous studies. 62 Briefly, ND powder was dispersed in a mixture of concentrated sulfuric acid and nitric acid (3:1, v/v), followed by stirring and reacting in a reflux device at 70 °C for 3 days. The resulting ND-COOH was then washed by deionized water until the suspension pH reached neutral. Subsequently, ND-COOH was freeze-dried and the powder was stored at room temperature until further use.
To obtain an ND-MMP9 substrate peptide complex, one anhydrous reaction was performed. ND-COOH powder was suspended in DMSO (10 mg/mL), and sonicated well before conjugated with peptide. A certain amount of dicyclohexylcarbodiimide (DCC) and 4-dimethylaminopyridine (DMAP) was added into ND-COOH suspension to activate the carboxyl groups in ND-COOH. After 10 min of activation at room temperature, MMP9 substrate peptide (with a sequence of LGRMGLPGK, Mimotopes, Melbourne, Australia) was then added into the reaction suspension at a weight ratio of 1:1 (MMP9 peptide to ND-COOH). Following reaction at room temperature overnight, the reaction mixture was centrifuged at 15,000 rpm for 15 min. Subsequently, the precipitates (ND-MMP9) were resuspended in deionized water. Unreacted peptides, together with DCC and DMAP, remained in the supernatant and were collected for quantifying conjugation efficiency.
Characterization of ND-MMP9 Complex
ND-COOH and ND-MMP9 were suspended in deionized water (0.2–0.3 mg/mL, ND concentration). The particle size and ζ-potential were determined by the dynamic light scattering (DLS) method using a Malvern Zetasizer Nano ZS (Malvern Instruments, Malvern, UK) at 25 °C. All measurements were performed in triplicate.
Fourier transform infrared spectroscopy (FTIR) was performed by a PerkinElmer FTIR Spectrum 2000 (PerkinElmer, Waltham, MA). Samples were dried overnight using R-210 Rotavapor (Buchi, New Castle, DE). Five milligrams of each sample was mixed with 0.1 g of KBr powder using mortar and pestle before pressing the sample to a thin film, from which the spectra were taken.
Transmission electron microscopy (TEM) was performed using a JEOL JEM-2010 transmission electron microscope (JEOL, Tokyo, Japan). ND-COOH and ND-MMP9 were dispersed in deionized water at a concentration of 0.1 mg/mL, and the suspension was dropped on the copper mesh (coated with carbon film), and then dried overnight at room temperature prior to imaging.
Cell Culture
Human HCC cell lines Huh7 and SNU398 were obtained from the Japanese Collection of Research Bioresources (Tokyo, Japan) and American Type Culture Collection (Manassas, VA), respectively. Huh7 cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM, BioWest, Kansas City, MO), whereas SNU398 cells were cultured in RPMI-1640 (BioWest) medium. Both media were supplemented with 10% (v/v) fetal bovine serum (Gibco, Grand Island, NY) and 1% penicillin/streptomycin (Gibco). All the cells were cultured in a 5% CO2-humidifed chamber at 37 °C.
Protein Extraction and Western Blot
Huh7 or SNU398 cells were washed twice with phosphate-buffered saline (PBS) and lysed in RIPA buffer consisting of 0.5% sodium deoxycholate, 1% NP-40 detergent, 0.1% sodium dodecyl sulfate (SDS), 150 mM NaCl, 10 mM Tris-HCl (pH 7.4), protease, and phosphatase inhibitors (Roche, Basel, Switzerland). The lysed cells were then sonicated on ice for 20 s (10 times, 2 s each) and centrifuged at 10,000g for 15 min to remove cell debris. The protein content of each sample was then determined by Bio-Rad Protein Assay (Bio-Rad Laboratories, Hercules, CA) according to the manufacturer’s instructions. Equal amounts of sample protein were then separated by 8% SDS–polyacrylamide gel electrophoresis (SDS-PAGE), and transferred to polyvinylidene difluoride (PVDF) membranes. Membranes were blocked by incubation in TBST (Tris-buffered saline, 0.1% Tween-20) containing 5% bull serum albumin (BSA) for 1 h, and probed with primary antibodies. Thereafter, membranes were processed using standard procedures. Antibody binding was visualized by an imaging system ImageQuant LAS 500 (GE Healthcare Life Sciences, Pittsburgh, PA). The following antibodies were used: anti-human MMP9 (1:500, R&D Systems, Minneapolis, MN) and anti-β-actin (1:10,000, Sigma, St. Louis, MO).
The extracellular expression level of MMP9 (or the secreted MMP9) was determined by Western blot as well. Briefly, equal numbers of these two human HCC cells were seeded and equal volume of culture media were collected after culturing for 3 days. Total proteins in the culture media were concentrated and collected by centrifuging at 2000g for 20 min using Corning Spin-X UF concentrators (Sigma). Equivalent volumes of these two concentrated proteins underwent standard Western blot procedures as described above.
Immunohistochemistry
Two cases of human HCC and matched adjacent noncancerous tissues from patients who had undergone treatment at the National University Hospital of Singapore were collected. Tissues were collected from a single HCC lesion, along with surrounding normal tissue. All HCCs were clinically and pathologically confirmed. Written consents were obtained from all patients before sample collection. The study was approved by the National University of Singapore Institutional Review Board (NUS-IRB).
HCC patient tissues were fixed using 4% (w/v) paraformaldehyde (PFA) and embedded with paraffin. The paraffin blocks were then cut into sections of 4.0 μm thickness, dehydrated in graded Histo-Clear, and rehydrated in graded ethanol. For antigen retrieval, slides were placed in antigen retrieval solution (pH 9.0, Dako, Carpinteria, CA) and boiled for 5 min in a microwave (Milestone, Milan, Italy). Slides were then allowed to cool for 10 min, followed by a 5 min wash in water. Before immunohistochemical staining, the slides were washed twice (5 min each) in Phosphate Buffered Saline with Tween 20 (PBST), and blocked in blocking buffer (Protein Block Serum-Free, Dako) at room temperature for 1 h. Afterwards, the sections were incubated overnight at 4 °C in a humidified chamber with primary antibodies. For primary antibodies, anti-human MMP9 antibodies (1:500, R&D Systems) were used. The slides were then washed three times in PBST and incubated for 1 h in anti-mouse HRP-conjugated secondary antibodies (1:10,000, donkey). Slides were washed in PBST, developed with 3,3′-diaminobenzidine (DAB), and counterstained with hematoxylin. The slides were then washed with deionized water, followed by dehydration in graded ethanol and graded Histo-Clear. The coverslips were mounted using mount medium, and the slides were visualized on a Zeiss Axioplan 2 fluorescence microscope (Zeiss, Oberkochen, Germany).
In Vitro Cell-Free Cleavage Assay
Recombinant human MMP9 protein (R&D Systems) was first activated by p-aminophenylmercuric acetate (APMA, Sigma) in assay buffer (50 mM Tris, 10 mM CaCl2, 150 mM NaCl, 0.05% Brij-35 (w/v), pH 7.5). Fluorophore-modified MMP9 substrate peptide and the corresponding ND-MMP9 complex were added separately into activated MMP9 buffers containing different MMP9 protein concentrations. The time-course cleavage signal was analyzed and recorded by an Infinite 200 PRO microplate reader (Tecan, Zurich, Switzerland). Measurements were conducted in a black/transparent fluorescence-block 96-well plate (Falcon, BD, San Jose, CA) in triplicate.
To better simulate the physiological conditions in vivo, the cleavage of both MMP9 peptide and ND-MMP9 in human serum was detected. MMP9 peptide and ND-MMP9 (10 μM) were added into a certain volume of human serum, followed by a time-course cleavage signal assay, as described above.
In Vitro Cell Culture Cleavage Assay
Huh7 and SNU398 cells were seed into black/transparent fluorescence-block 96-well plates. After 24 h of incubation, culture media were collected for detecting the cleavage extracellularly. MMP9 peptide and ND-MMP9 were added into the culture media of either Huh7 cells or SNU398 cells. For cleavage intracellularly, MMP9 peptide and ND-MMP9 were diluted using serum-free media and added into each well with either Huh7 or SNU398 cells. Time-course cleavage assays were performed using an Infinite 200 PRO microplate reader, as above. To better elucidate the correlation between MMP9 expression level and the cleavage signal, cells were seeded in various numbers (representing different amounts of MMP9). The cleavage assay process was the same. To demonstrate the specific cleavage between the MMP9 and ND-MMP9 complex, an MMP9 enzyme inhibitor (ab142180, Abcam, Cambridge, UK) was used. Cells were incubated with the MMP9 inhibitor for 1 h (50 μM). The cleavage signal of the inhibiting group was recorded continuously for 7 days. For the control group, no inhibitor was used.
In Vitro 3D Metastasis Cell Culture Chip Biosensor Analysis
SNU398 spheroids were first formed by seeding SNU398 cells on ultra-low-attachment culture dishes (Corning, Corning, NY) in serum-free spheroid medium. DMEM/F-12 serum-free medium (3:1, v/v) contained 1% penicillin/streptomycin, 1% w/v minimum essential media nonessential amino acids, 1% w/v sodium pyruvate, 2% B27, 1% v/v insulin transferrin selenium, 4 μg/mL heparin, 10 mM nicotinamide, 20 ng/mL epidermal growth factor, and 10 ng/mL fibroblast growth factor 2.
SNU398 spheroids were collected and stained using CellMask Deep Red plasma membrane stain (C10046, Invitrogen, Thermo Fisher Scientific, Waltham, MA) at 37 °C for 15 min. Thereafter, the spheroids were suspended in serum-free spheroid medium, and the spheroid suspension was then mixed with Matrigel Matrix (Corning), followed by seeding into a three-dimensional (3D) Cell Culture Chip (AIM Biotech, Singapore). After culturing for 24 h, equivalent (1 μM) MMP9 peptide and ND-MMP9, both diluted with serum-free spheroid medium, were added separately to the AIM Chip wells. For the control group, spheroid medium without peptide was added to a separate well. To better trace the substrate peptide cleavage by MMP9, images were taken using an Olympus FV1000 confocal fluorescence microscope (Olympus, Center Valley, PA) at different treatment time points. Cy5 (633 nm/664 nm) and fluorescein (488 nm/519 nm) were applied as the detection channels. Finally, the raw data were evaluated by FV10-ASW 4.2 Viewer Software (Olympus).
Statistical Analysis
All experimental data were performed at least in triplicate, and described as the mean ± standard deviation. Unpaired Student’s t test and one-way analysis of variance were used to analyze the results.
Results
Expression of MMP9 in HCC Patient Samples
Previous studies of MMP9 have demonstrated its key role in promoting tumor invasiveness and metastatic characteristics in HCC.63,64 In addition, reduced expression of E-cadherin has been shown to portend a poorer prognosis in HCC patients. 65 To determine the levels of MMP9 expression and its metastatic potential in HCC, we first analyzed the protein expression of MMP9 and E-cadherin in primary HCC patient specimens (HCC141015 and HCC240216) via immunohistochemistry. We showed that HCC patient samples expressed higher levels of MMP9 and reduced levels of E-cadherin compared with the normal adjacent tumor tissues (NAT240216) ( Fig. 2 ). These data suggest that MMP9 could potentially be used as a biomarker for HCC, in particular for metastatic HCC tumors.
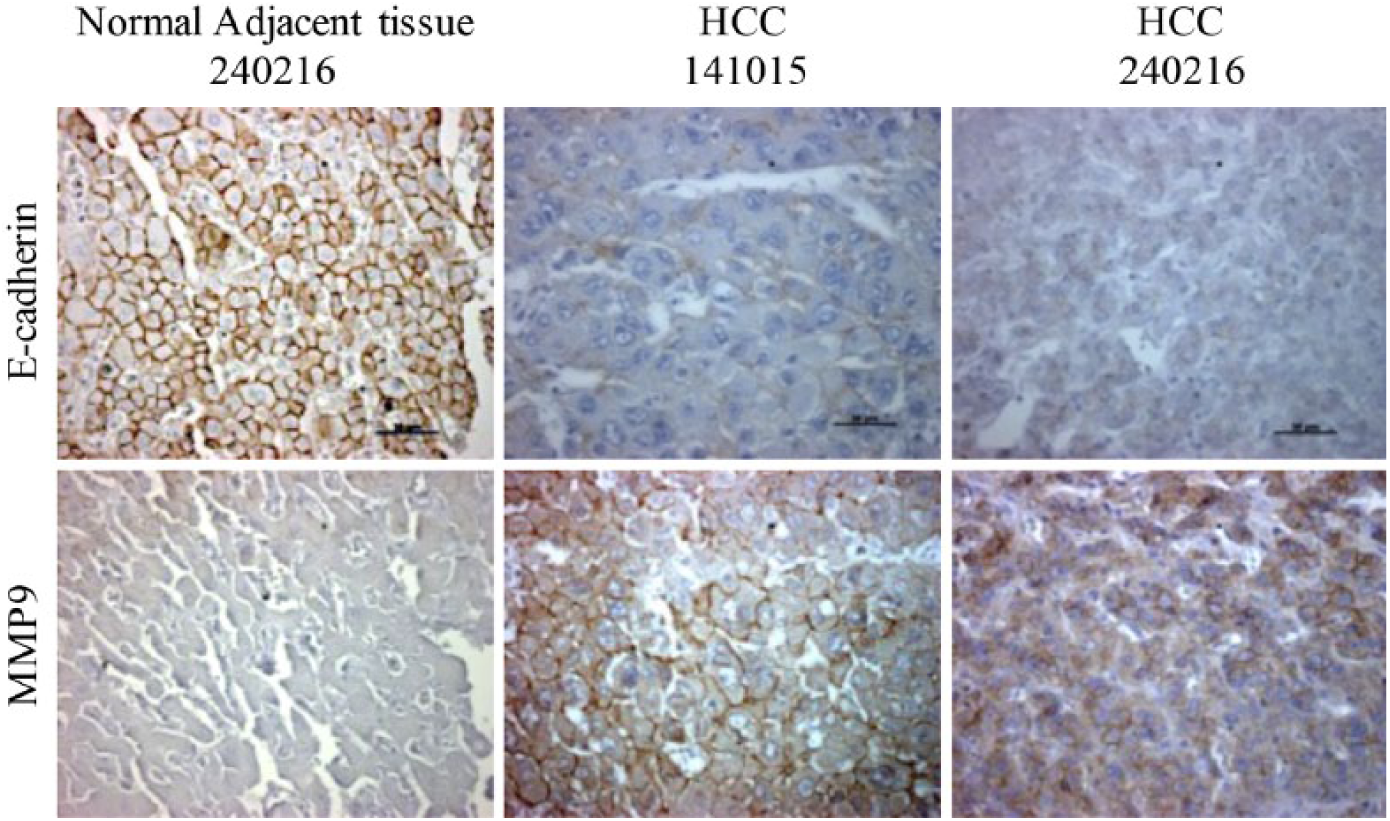
MMP9 expression in HCC patients. Representative pictures showing immunohistochemistry staining of E-cadherin and MMP9 in HCC patient samples and normal adjacent tissue. Brown-colored edges around cells denote the positive staining of E-cadherin and MMP9. Scale bar (black), 50 μm.
Characterization of ND-MMP9 Complex
The chemical conjugation of peptide and ND was accomplished through connection of the carboxyl group on acid-washed ND (ND-COOH) and the amine group on a modified C-terminus of the peptide. The synthesized ND–MMP9–peptide complex (ND-MMP9) showed slightly increased diameter (33.36 ± 4.28 nm) compared with naïve ND particle (23.37 ± 2.13 nm), as well as the ND-COOH (30.57 ± 3.20 nm) ( Fig. 3A ). Meanwhile, the surface net charge of ND also changed postconjugation, dropping from 53.15 ± 0.92 mV to 27.03 ± 0.25 mV ( Fig. 3B ). However, the lattice structure on the surface of nanoparticles was clearly observed on both ND and ND-MMP9 ( Fig. 3C ). The FTIR assay, which identifies the active functional groups on the nanoparticle surface, indicated that the typical functional groups in both ND (C=C stretch, O–H stretch) and peptide (C–N stretch, N–H wag) were detected on the surface of ND-MMP9 ( Fig. 3D ), demonstrating the successful conjugation of MMP9 peptide onto the ND-COOH surface.
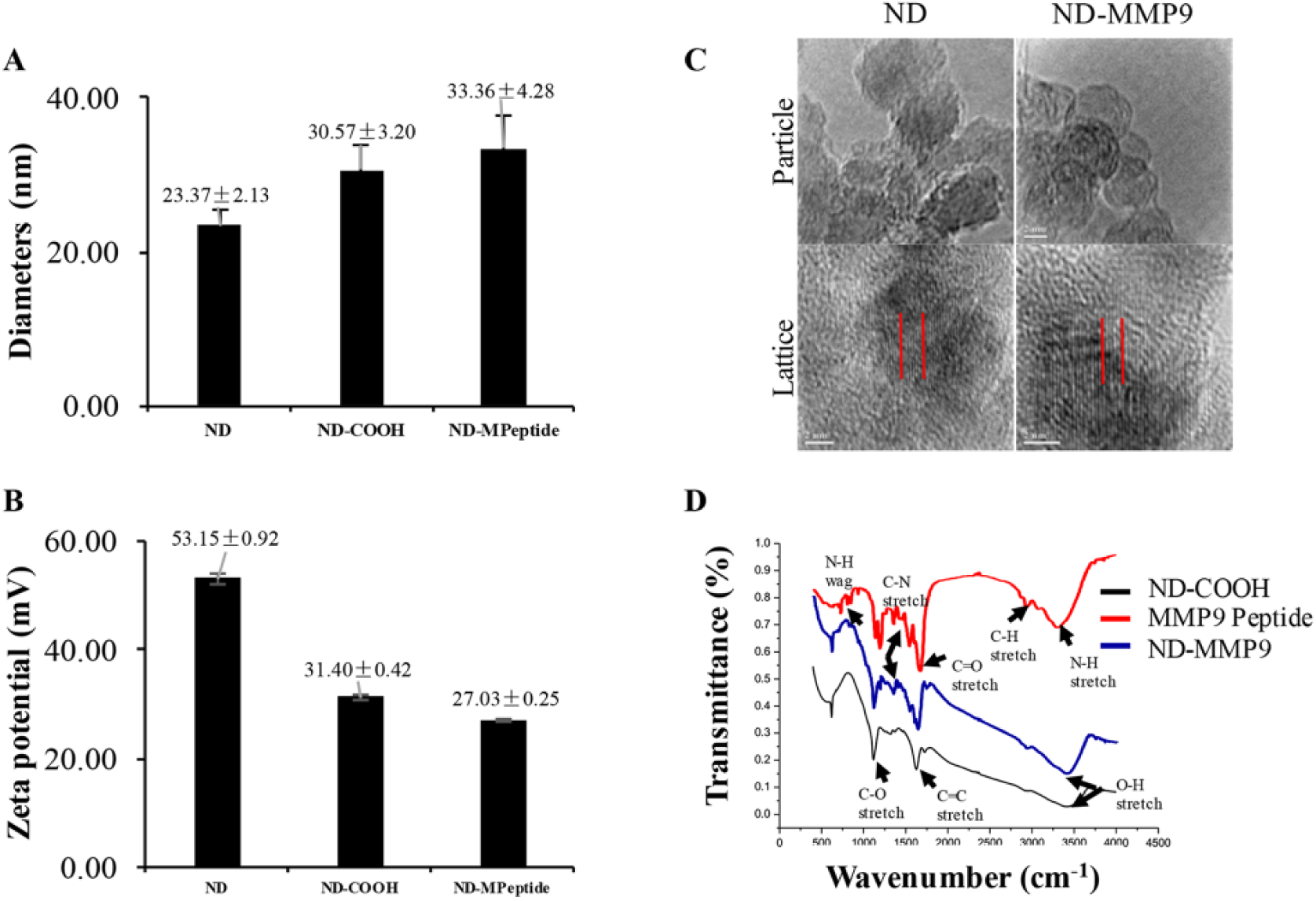
ND-MMP9 peptide characterization. (
ND-MMP9 Responded to MMP9 Enzyme but Remained Stable in Serum
Upon successful conjugation of peptide onto ND, we first tested whether the substrate peptide on the ND surface was still feasible for MMP9 cleavage. The results showed that both the peptide and ND-MMP9 were able to respond to purified MMP9 enzyme, and the cleavage level was dependent on the concentration of purified MMP9 in the reaction system ( Fig. 4A , B ). Nonspecific substrate peptide cleavage was also evaluated. When treated with human serum under physiological conditions (37 °C, pH = 7.4), the naïve MMP9 substrate peptide showed significantly stronger degradation than peptide conjugated on ND ( Fig. 4C ). These results indicated that ND-MMP9 is able to respond to specific MMP9 cleavage and also protect against nonspecific degradation.
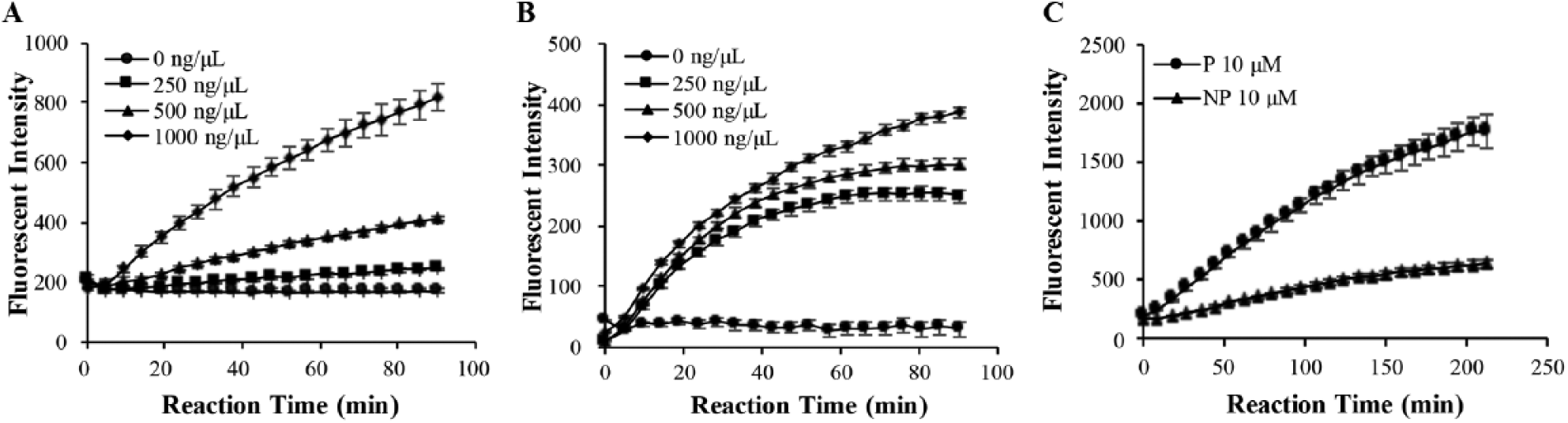
ND-MMP9 responds to MMP9 enzyme cleavage and shows stronger serum stability. (
ND-MMP9 Cleavage Signal Correlates with MMP9 Expression Level
In this study, we chose to chemically conjugate a stimuli-responsive peptide biosensor onto an ND surface. In order to track the peptide, as well as develop a stimuli-responsive quantifiable sensor, a FRET pair consisting of FITC and TAMRA was attached on either terminal of the peptide. 66 Energy is transferred from FITC to TAMRA under the FITC excitation wavelength, so FITC will be quenched when the peptide maintains its full length. When the peptide is cleaved by MMP9 from ND, this energy transfer is terminated due to the increasing distance between FITC and TAMRA; hence, the FITC emission can be observed under FITC excitation. By inspecting the FITC emission signal, the substrate peptide cleavage by MMP9 can be quantitatively measured.
In order for ND-MMP9 to function as an MMP9 biosensor, there should be a correlation between the cleavage signal and MMP9 expression level. Hence, the MMP9 protein expression level in HCC cell lines was measured, both intracellularly and extracellularly. MMP9 protein was primarily secreted extracellularly into culture medium, with the Huh7 cell line expressing higher levels of MMP9 protein than the SNU398 cell line ( Fig. 5A ). Both intracellular and extracellular MMP9 activities against ND-MMP9 were evaluated ( Fig. 5B ). After quantifying the MMP9 expression in these two cell lines as, well as the end-point cleavage signal level, a correlation between these two readouts was determined ( Fig. 5C ). To elucidate this correlation more accurately, the ND-MMP9 cleavage level under designated amounts of MMP9 protein was further explored ( Fig. 5D , E ). ND-MMP9 cleavage signal increased according to the intensity of the MMP9 protein with linear correlation ( Fig. 5F ). Furthermore, inhibition of the MMP9 protein triggered a corresponding decrease in ND-MMP9 cleavage activity ( Fig. 6 ). These data demonstrate that the cleavage of ND-MMP9 correlates with the MMP9 protein expression level and its activity.
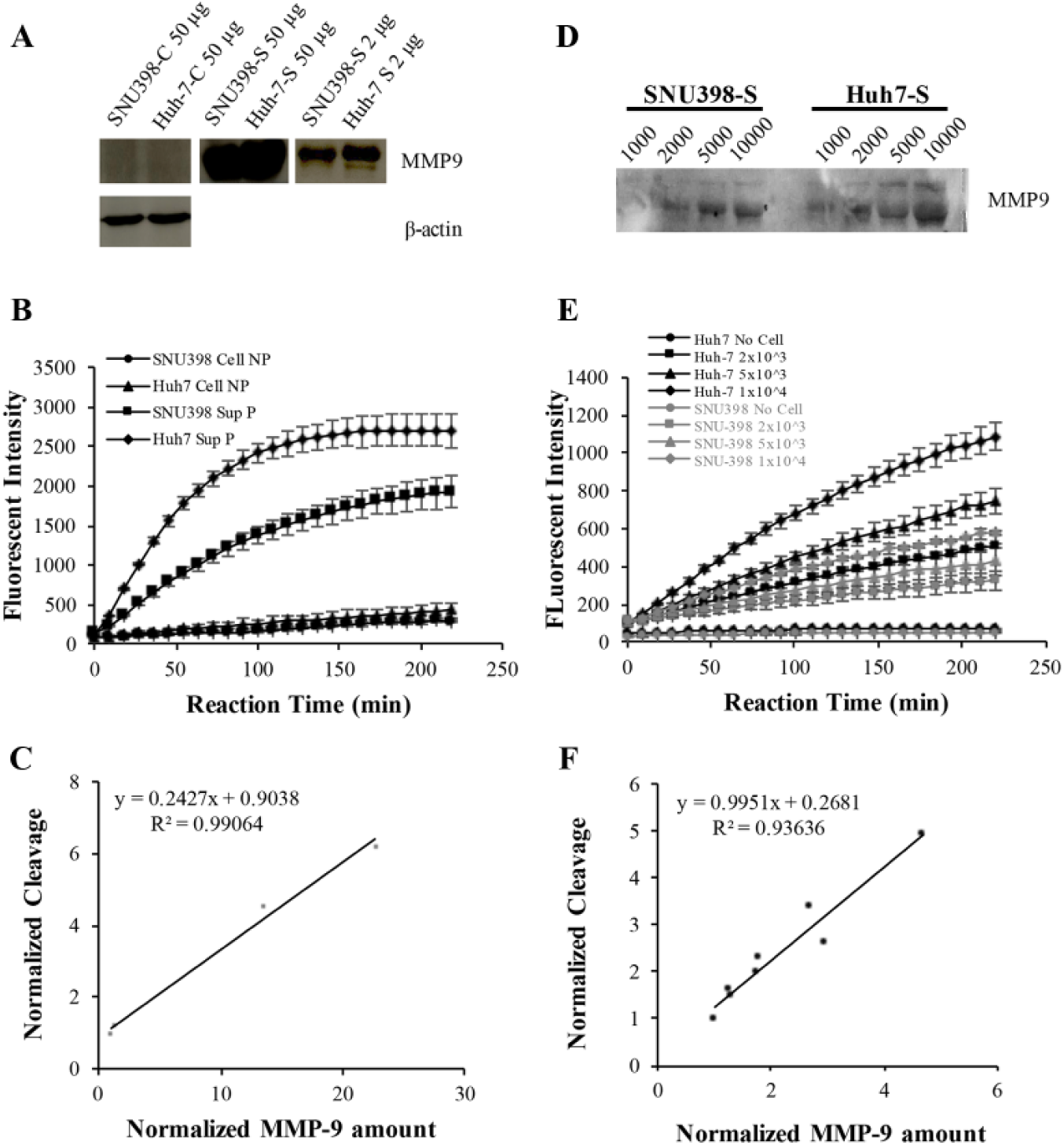
ND-MMP9 cleavage is correlative with MMP9 expression level in HCC cell lines. (
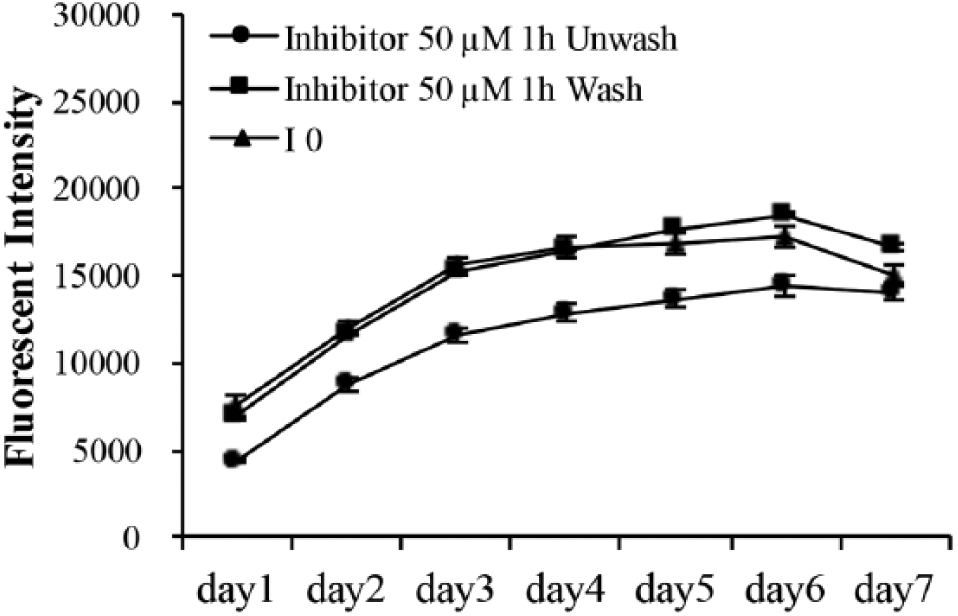
ND-MMP9 peptide cleavage was inhibited by MMP9 enzyme inhibitor. Fluorescent intensity change within reaction period of ND–peptide (NP) cleaved by Huh7 cells with or without pretreating with 50 µM MMP9 inhibitor. “Wash” and “Unwash” denote the washing steps performed 1 h after inhibitor treatment.
Evaluation of ND-MMP9 in In Vitro 3D Metastasis Model
Metastatic tumors do not reside in cell-free liquid solutions, but rather in complex microenvironments where secreted MMP9 degrades the surrounding extracellular matrix and allows tumor cell movement to new local and distal sites. 3D spheroid cultures and microfluidic-based devices can be utilized to better analyze metastatic tumors in the context of these complex environmental conditions.67–72 We next explored if ND-MMP9 was capable of emitting a fluorescent signal in the surrounding tumor microenvironment of a metastatic tumor by using a 3D spheroid culture system within a multichannel microfluidic chip system. ND-MMP9 efficacy was evaluated using a 3D Cell Culture Chip. The 3D Cell Culture Chip is a three-channel microfluidic chip that is capable of recapitulating spheroid tumor growth and metastasis potential. A central channel serves as the site of tumor spheroid growth within an extracellular matrix scaffolding. The other two parallel channels can serve to represent nearby blood vessels to determine the effect of drugs, proteins, or in this case, a biosensor on detection, survival, extravasation, and/or intravasation of metastatic tumors. Following seeding of SNU398 spheroids in the central channel, 1 μM of either MMP9 peptide or ND-MMP9 was delivered into the central channel by directional flow from one outer channel to the other. Biosensor activity was then visualized and measured under a confocal fluorescence microscope. The FITC signal of peptide cleavage was compared with the Deep Red staining that outlined spheroid cellular structures to determine peptide cleavage location, which primarily occurred extracellularly ( Fig. 7 ). Our data suggest that secreted MMP9 enzyme is the primary mediator of ND-MMP9 cleavage, and that ND-MMP9 can fluorescently paint the surrounding area of metastatic cancer sites based on MMP9 activity.
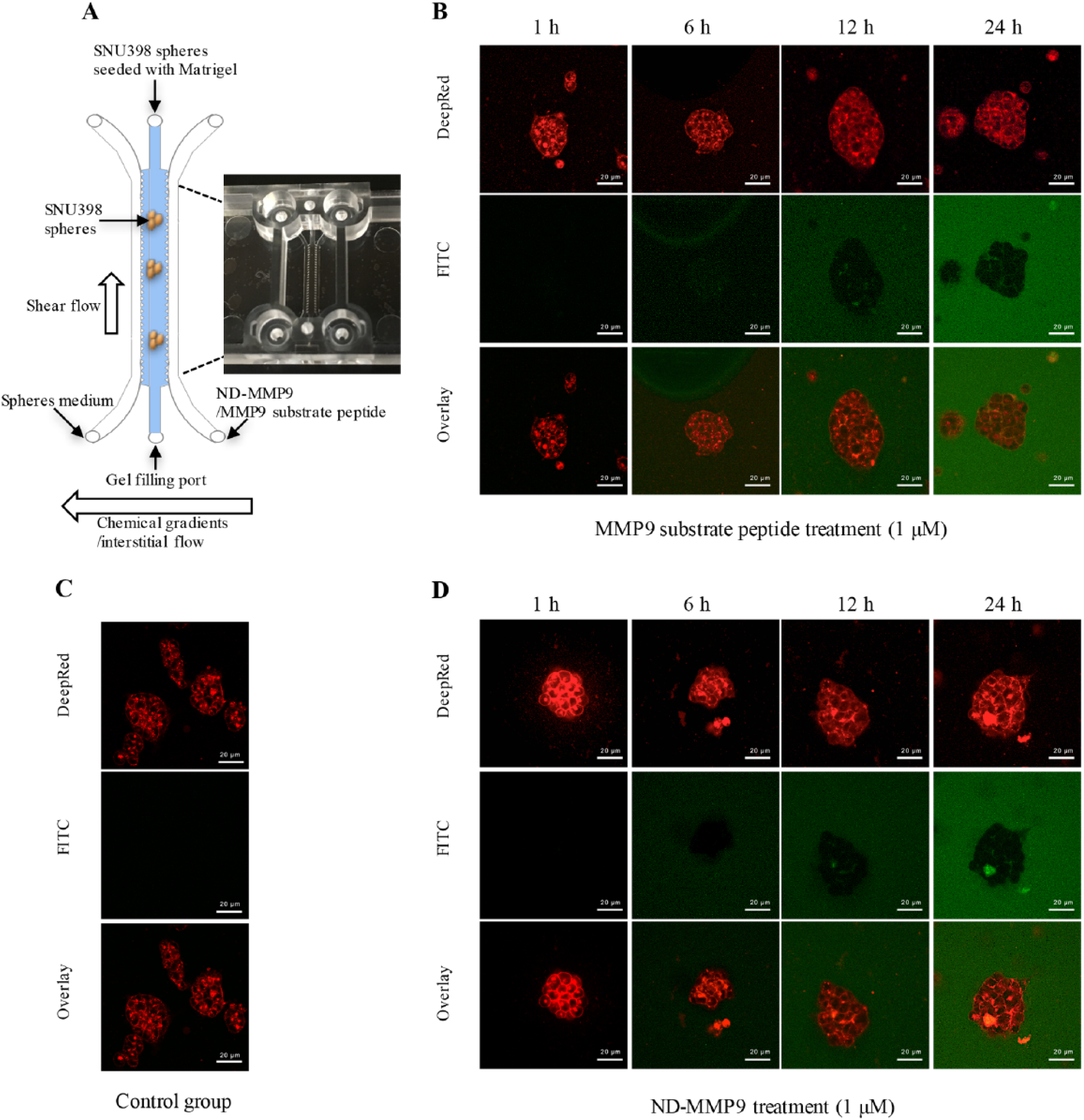
Visualization of ND-MMP9 cleavage by secreted MMP9 enzyme from SNU398 spheroids. (
Discussion
A nanoparticle-based biosensor requires several key properties before translating toward preclinical and clinical development. In evaluating the potential of an ND–peptide-based stimuli-responsive biosensor, it is first important to understand how chemical conjugation of peptides to NDs affects the overall size and charge of the delivery complex, as this will influence the biodistribution, cellular uptake, and biocompatibility. Previous studies that utilized reversible physical adsorption of drugs on the surface of NDs resulted in an aggregate complex diameter increase of approximately 400%. In this study, we observed less than a 4 nm increase in diameter following chemical conjugation of peptide to the NDs. This could be due to the differences in conjugation methods. During physical adsorption, it is possible that the drug may form one or more drug layers around the ND that cover the charged facets of the ND. A complete covering of ND facet surfaces by these drug layers would result in a more dramatic change in net charge that would also likely affect aggregate size. For chemical conjugation of peptides, peptides were connected to carboxyl groups presented on the ND surface. This likely serves to create some space between the peptide and the charged facets, as well as permitting loading of peptide to the ND without covering the charged facets. As a result, the change in net charge (31.40–27.03 mV) was not strong enough to generate the formation of new aggregates. This suggests that the diameter of the final complex could be manipulated through different methods of synthesis for various applications. Furthermore, the relatively small size of the ND-MMP9 aggregate complex ensures that the complex fits the physical characteristics for proper biodistribution and metastatic tumor delivery.
Peptide-based biosensors must overcome a number of challenges in order to be successful for in vivo applications. These challenges include protein hydrolysis in the blood stream, the shredding force in caterpillar circulation, and proteases/peptidases in the tissue matrix. Although the cleavage signal of ND-MMP9 is not as strong as naïve peptide in both in-tube assay and cellular cleavage, the ND-MMP9 complex maintained a clear and quantifiable cleavage signal while also significantly improving the stability of substrate peptide and protection against nonspecific proteases. Through enhancing the stability of the peptide, a greater concentration of peptide biosensor at tumor sites should be achievable. Meanwhile, the minimization of random degradation can help to decrease the possibility of false positives in in vivo applications. Improved peptide stability, paired with the demonstration that ND-MMP9 can quantitatively measure MMP9 expression level, suggests that ND-MMP9 biosensors may serve as a useful reagent in detecting MMP9-positive metastatic tumor sites. While this initial study utilized the FITC-TAMRA FRET pair, other more clinically relevant FRET pairs can be explored for the quantification of MMP9 activity in vivo. Additionally, other imaging modalities, such as DNA barcode tagging or isotope labeling, can also be incorporated to further improve the diagnostic sensitivity.
While this study focuses on the use of an ND–peptide biosensor platform for the detection of MMP9 activity, this ND–peptide system has potential applications beyond MMP9 activity detection of metastatic tumor sites. This stimuli–response biosensor system can be applied to a number of other disease and biological functions where specific enzyme–substrate reactions take place, for instance, in the detection of glutathione (GSH) and cathepsin B (CTSB), whereby both enzymes have unique expressions in tumor sites compared with normal tissues.73–75 Beyond applications in diagnostics, this ND-mediated peptide delivery platform can also be used to deliver therapeutic peptides, as well as in the development of theranostics or drug combinations. For the development of ND-based drug combinations, however, experimental-based rational design of optimal drug combinations must be done in order to identify the most appropriate therapeutics to include in more complex multifunctional ND-based theranostic devices.76–82 Overall, this study can serve as the foundation for an ND–peptide-based approach to developing stimuli-responsive biosensors for the detection of disease-specific enzymatic activity that can serve to help clinical diagnosis and the treatment of a wide range of cancers and other diseases.
Footnotes
Acknowledgements
The authors thank Dr. Chiu (Department of Pharmacy, NUS) for her kind help in DLS measurements, as well as Professor Phillip Koeffler (Cancer Science Institute of Singapore, NUS) for providing the Tecan Microplate Reader.
Authors’ Note
The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Cancer Institute or the National Institutes of Health.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors are supported by the National Research Foundation Cancer Science Institute of Singapore RCE Main Grant, Ministry of Education Academic Research Fund (MOE AcRF Tier 2 [MOE2015-T2-2-126, Seed Fund Grant T1-BSRG 2014-05]).
