Abstract
Pluripotent stem cells are an extremely powerful tool in modeling human diseases and hold much promise for personalized regenerative or cell replacement therapies. There is an increasing need for reproducible large-scale stem cell and differentiated progeny production, with minimal variation, rendering manual approaches impracticable. Here, we provide an overview of systems currently available for automated stem cell culture, and undertake a review of their capacities, capabilities, and relative limitations. With the merging of modern technology and stem cell biology, an increased demand and implementation of automated platforms for stem cell studies is anticipated.
Keywords
Introduction
The isolation of mouse1,2 and human embryonic stem cells (ESCs)3,4 has opened the possibility to study mechanisms involved in cell development, with particular interest in those that are dysregulated in disease pathology. The rapidly growing field of stem cell–based studies has accelerated significantly since the development of the cell reprogramming technique, which allowed the generation of induced pluripotent stem cells (iPSCs), from somatic cells, which have the ability to differentiate into all cell types of the body.5,6 Together with recent advances in methods for direct differentiation of iPSCs into a desired cell type, personalized regenerative medicine seems to be within our reach. However, a number of issues must be addressed before personalized therapies become available at scale. These include not only safety concerns and costs of potential treatment, but also the need to increase the efficiency and scale of iPSC generation, and to standardize the methodology of their production to reduce variability between cell lines.
Current Capabilities of Automated Systems
Various Automated Platforms for Cell Culture
Several companies manufacture automated platforms. These incorporate robotic arms for liquid and labware handling, Class II biosafety cabinets to ensure sterile conditions for cell handling, and incubators providing precise control over temperature and CO2 and O2 levels. Many of these systems are highly modular and may be easily upgraded with additional devices, such as automated cell counters, imaging systems, centrifuges, cell sorters, PCR thermocyclers, plate readers, and thermostats. Currently available robotic platforms may be distinguished based on the type of labware used, that is, microplates or cell culture flasks. For example, the Cellerity and subsequent Tecan Integration Group (TIG) customized Freedom EVO (Tecan, Zürich, Switzerland), Cellhost (Hamilton Robotics, Reno, NV), and BioCel (Agilent, Santa Clara, CA) systems are designed for microplate applications. Cell Culture Automatic Laboratory (CCAL) employs Petaka bioreactors (Celartia, Columbus, OH), Biomek Cell Workstation (Beckman Coulter, Brea, CA), and CELLSTAR Autoflasks (Greiner, Monroe, NC) for handling flasks, while SelecT and CompacT SelecT platforms (TAP Biosystems, Hertfordshire, UK) may utilize either microplates or T-flasks ( Table 1 ). Another liquid handler was recently reported for the maintenance and passaging of human pluripotent stem cells (PSCs). This smaller system is based on a self-contained Gilson’s pipette Max liquid handler, and can accommodate up to 96-well plates. 7 Some limitations include the absence of a connected incubator and the requirement of offline steps and substantial human contribution. Each system has a different capacity for labware handling, offering a high degree of flexibility for various applications, with the emphasis on large-scale studies that would otherwise be difficult to undertake.
Labware Capacities.
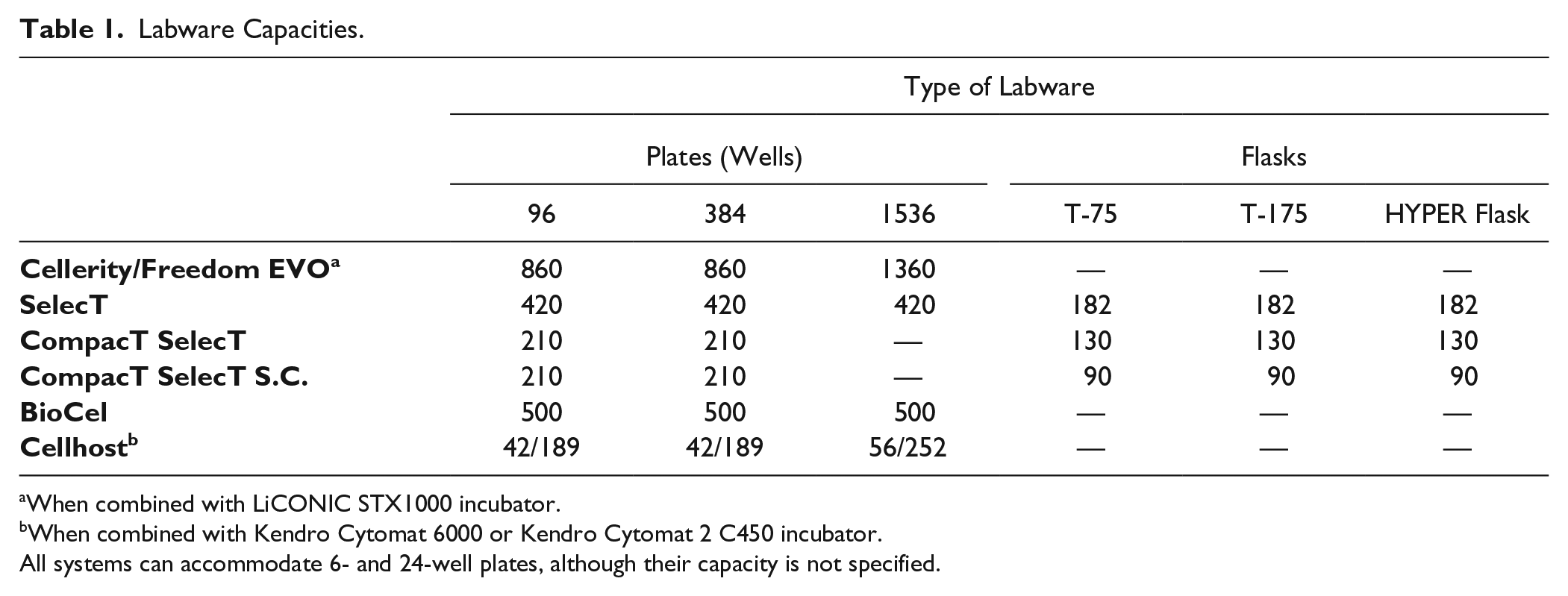
When combined with LiCONIC STX1000 incubator.
When combined with Kendro Cytomat 6000 or Kendro Cytomat 2 C450 incubator.
All systems can accommodate 6- and 24-well plates, although their capacity is not specified.
Current Usage
Initially, automated cell culture systems were developed to validate their capabilities to replicate tasks that were principally performed by human operators. The successful outcome of such studies and rapid progress in the iPSC field have prompted the establishment of large-scale repositories of patient-specific iPSC lines derived from both disease-affected individuals and healthy controls. The overarching goal of these banks is to provide researchers with a reliable source of cells in order to facilitate studies of numerous conditions, including diabetes and cardiovascular and neurodegenerative diseases. McKernan and Watt recently described both existing banks and those under development. 8 Several of these initiatives plan to utilize robotic systems to achieve their goal. The New York Stem Cell Foundation (NYSCF) established in 2005 aims to generate 2500 human iPSC lines using a self-designed robotic platform composed of three connected platforms using STAR liquid handling systems (Hamilton Robotics).8,9 This system remarkably demonstrated its potential in the automation of the entire process of iPSC production, including the automated culture of fibroblasts, reprogramming, the selection of iPSCs, maintenance, passaging, differentiation, thawing, and the assessment of pluripotency, differentiation, and karyotype. 9 The California Institute for Regenerative Medicine (CIRM), in collaboration with Cellular Dynamics International, Inc. (CDI), plans to generate 9000 iPSC lines from 3000 individuals from the United States with nonintegrating episomal vectors. 8 Several consortiums in Europe also aim to generate large biobanks of iPSCs. The European Bank for induced pluripotent Stem Cells (EBiSC) intends to bank more than 10,000 lines covering a wide range of diseases. 8 The StemCellFactory initiative’s objective is to automate production of iPSC lines, and their subsequent characterization and differentiation into neural and cardiac derivatives with a self-built system, 10 while I-Stem is an initiative that aims to automate the screening of compounds with therapeutic potential and subsequent disease modeling using the CompacT SelecT platform. Moreover, the AUTOSTEM Consortium is undertaking a project to expand the automated platform developed for the StemCellFactory project and utilize it for the production and banking of therapeutic stromal cells. 11 Finally, our group aims to establish a large repository of patient-specific iPSCs and differentiated cells for ophthalmic research utilizing a Tecan Freedom EVO platform.12–14
Automated Processes
Maintenance
The manual maintenance of iPSCs introduces several limitations for transition into large-scale experiments. First, the maintenance of the iPSC culture to retain pluripotency and for directed differentiation protocols requires highly trained and experienced staff. Moreover, technician variability and human error pose major limitations when high numbers of samples are being processed in parallel. This variability also contributes to significant differences between cell lines generated and maintained in various laboratories,15,16 which are exacerbated by factors affecting the reprogramming efficiency itself. These include the tissue of origin, 17 the transcription factors employed in the reprogramming cocktail, their stoichiometry, 18 the method of reprogramming, the genetic background, and the stress that cells are exposed to during their reprogramming and subsequent culture. 19 Overcoming these limitations by moving toward an automated process will allow the handling of a greater numbers of cells, which will in turn facilitate the optimization of existing methods and generation of new protocols for cell maintenance and directed differentiation. Furthermore, it will enable standardization of liquid handling and incubation times, which are two factors underlying the volatility of the manual approach. Therefore, the use of fully automated platforms for iPSC derivation, expansion, and differentiation may be key in transitioning to large-scale cell cultures. Cells derived from such cultures would provide a means for thorough disease modeling in vitro, as well as a broad range in toxicity screening of new chemical compounds, and pave the way for therapeutic cell replacement.3,4
Several groups have already adopted an automated approach and thoroughly tested its suitability for everyday use in various applications. Initially, research in the field of automation was focused on assessing whether employing the automated platforms for culture maintenance would have any effect on cellular characteristics, for example, cell morphology or gene expression, comparing them with cultures that relied solely on manual handling. Joannides et al. explored the capability of automated mechanical passaging (AMP) to expand human ESCs. 20 Their analysis revealed that AMP-expanded cells were more uniform in colony shape and colony numbers than the manual control. The stem cells also retained both morphological features and pluripotency marker expression, characteristic of pluripotent cells.
Successive groups examined both ESCs and iPSCs of human and rodent origin maintained on automated platforms in feeder-free conditions9,14,21–23 or on a mouse embryonic fibroblast (MEF) feeder layer.20,24–26 Paull and colleagues are the only study that report on the entire process of human iPSC generation, from fibroblast culture to iPSC maintenance. 9 They carefully examined gene expression profiles, revealing that automatically processed cells retained their round-shaped morphology and pluripotent characteristics, such as markers of pluripotency; for example, OCT4, TRA-1-60, TRA-1-81, NANOG, and SSEA-4 were expressed at the same or higher level compared with manual controls.9,20,21,24,26 Stem cells were also tested for any karyotypic anomalies that could be introduced during the cell culture itself—most groups reported that no anomalies occurred during the time of their experiments,20,21,24,26 while one 9 identified aneuploidy in 11% of lines tested, that is, the acquisition of chromosome 22. This is not surprising given that chromosomal abnormalities are to be expected with reprogramming and culturing of PSCs. 27 The variation observed in the number of chromosomal anomalies is most likely a reflection of the size of sample analyzed, or time of culture prior to analysis or the sensitivity of the analysis itself. For instance, Konagaya and colleagues analyzed one iPSC line by G-banding, 26 Thomas et al. reported on two human ESC lines by G-banding, 21 and Paull et al. analyzed 38 iPSC lines using a NanoString karyotype assay, with improved sensitivity compared with traditional G-banding. 9 Of importance, three of the four abnormal lines tested by Paull and colleagues originated from the same fibroblasts and shared the same anomalies, suggesting that those originated within the fibroblasts and passed on, but did not arise from the automated reprogramming or maintenance. 9 The high-resolution analysis of copy number variant in iPSCs maintained manually or on the automated platform revealed a similar amount of variation, hence suggesting that continued assessment of cell genetic stability is essential. 9
Several automated platforms are also able to be adapted for processing other stem cell types. Some examples are detailed below. Thomas et al. successfully shifted the manufacturing of current good manufacturing practice (cGMP)–compliant human neural stem cell line CTX0E03, from manual to automated processes, maintaining the cell number and transcription profile according to strict quality control specifications, that is, level of expression of neural markers, together with cell survival and expansion rate. 28 The CTX0E03 cell line is currently in Phase II clinical trials for the treatment of patients with ischemic stroke (NCT02117635). On the other hand, Kato et al. evaluated the possibility of expanding primary cell lines, that is, human fibroblasts and bone marrow stromal cells (BMSCs), using a benchtop platform with a footprint not exceeding 0.5 m2. 29 Cells obtained using the automated system displayed characteristics similar to those of manually cultured BMSCs and the ability of bone formation when transplanted into nude mice. Furthermore, Liu et al. tested the human osteosarcoma (HOS) cell line to identify sources of variation during cell expansion and the influence of residual trypsin on cell viability after passaging. 30 Recently, a Biomek Cell Workstation was successfully used to cultivate and passage suspension cultures from four different cell lines, demonstrating the feasibility of this platform for automation of floating cell culture. 31
Passaging
Enzymatic Passaging
Passaging is one of the crucial steps in PSC culture. It is essential both for the maintenance of cells in an undifferentiated state and for their expansion to the sufficient numbers required for downstream applications. Cell passaging can be performed enzymatically or mechanically. Enzymatic passaging is widely used and efficient. 32 However, it has occasionally been reported by several groups that enzymatic passaging of human PSCs increases the risk of introducing karyotypic abnormalities directly correlating with the duration of culture and, consequently, with the passage number.33–36 This issue can be addressed with the use of either the recombinant enzyme TrypLE Express 37 or an enzyme-free cell dissociation buffer, such as Versene. However, studies report the successful long-term maintenance of enzymatically passaged human PSCs without induction of chromosomal abnormalities: human ESCs were passaged with trypsin EDTA for >30 passages, 21 and iPSCs with a combination of trypsin and collagenase and knockout serum replacement (CTK) 26 or with accutase 9 for ~20 passages. These should reassure us of the feasibility of maintenance of karyotypically normal human PSCs with enzymatic passaging, and on automated platforms.
Mechanical Passaging
Automated Approach of Mechanical Passaging
The mechanical passaging of undifferentiated PSCs allows for a precise selection of undifferentiated parts of PSC colonies. This is an obvious advantage of mechanical passaging, which, however, remains a tedious process and requires trained personnel to perform, hence rendering this approach difficult to adapt to well-standardized high-throughput experiments. 38 Once automated, mechanical passaging would become a method of choice for PSC expansion, significantly reducing the risk of unwanted genetic alterations. Several groups attempted to adapt an automated approach of mechanical passaging,20,38,39 including the modified tissue chopper of Joannides et al. 20 One of the most significant drawbacks of this system is the lack of imaging options, which forces colony picking by the operator and does not allow fully automating the passaging process. The company AVISO (Gera, Germany) solved this issue by developing the CellCelector system. This system comprises an inverted microscope with a motorized stage, and a robotic arm to pick colonies, which uses either autoclavable capillaries (glass or metal) or a scraping module for cell harvesting. Colony selection and cell harvesting are done without any manual intervention, while images of processed colonies are stored for further reference. 38 CellCelector may also be fitted with a laminar cabinet AVISO FlowBox, thus enabling precise control of environmental conditions (temperature and CO2 and O2 levels) (manufacturer’s promotional materials). Cells passaged with CellCelector (human and mouse ESCs and iPSCs) maintained expression of pluripotency markers (e.g., Oct-4, Nanog, Tra-1-60, Tra-1-81, Rex1, or Ssea-1 [mouse]) and were able to differentiate into embryoid bodies (EBs).38–41 Furthermore, CellCelector was used to remove pluripotent colonies from feeder cells without compromising the differentiation potential of the mouse ESC line tested 39 and for selective isolation of hybridoma colonies producing high quantities of monoclonal antibodies with survival rates corresponding to colonies that were processed manually. 42 The ability of isolating single cells for further experiments with CellCelector offers the possibility to work with homogenous populations of cells of interest. This may be beneficial for high-sensitivity assays for which contamination with unwanted cells could have detrimental effects on the results and lead to errors in data analyses. 39 The need for isolation of single cells is not a new concept, and numerous methods have been established for tissue microdissection. 43 However, all are time-consuming and labor-intensive, thus requiring highly skilled staff members, making them unsuitable for large-scale applications.
Laser-Based Passaging for Cell Ablation
In 1976, Meier-Ruge et al. reported a laser-based microdissection technique that allowed excision of single cells from frozen tissues. 44 This procedure was later expanded to the isolation of rat renal tubules from freeze-dried tissue 45 and to the development of the laser capture microdissection (LCM) system. With the LCM, cells of interest adhere to a temperature-sensitive film activated by a laser beam pulse, allowing their separation. 43 The LCM is, however, not contact-free, and the presence of residual cells postdissection has been reported in tissue isolated this way. 46 Interestingly, an unexpected observation provided a solution to this issue. Schutze et al. noticed that cells isolated by LCM are catapulted from the processed tissue and could be collected in a separate tube without any external intervention, in effect dramatically reducing the risk of contaminants. The phenomenon of laser pressure catapulting (LPC) was then coupled to LCM, resulting in a novel method named laser microdissection and pressure catapulting (LMPC). 47 LMPC is a very flexible technique that proved its ability to be applied in various fields, for example, analyses of genomic aberrations, proteomics, botany, forensics, and manipulations of living cells (reviewed in Schutze et al. 46 ). It utilizes pulses of the UV-A laser to cut out either single cells or larger fragments, which are then captured on a suitable tube cap as a result of the process of cold ablation. The wavelengths of maximal absorption for DNA and proteins do not overlap with those of the UV-A laser; thus, cells subjected to LMPC should not display any abnormalities on either the genetic or protein level. LMPC was tested with tumur cell lines as well as stem cells, and no changes in morphology, gene expression pattern, or genome integrity were observed in LMPC-treated cells.32,48–51 Moreover, it is also possible to perform microdissection on differentiated cells derived from PSCs as a way to purify the cell population, which may be essential to generate large numbers of cell types of interest with high efficiencies. 51 The LMPC procedure using the PALM MicroBeam system can be fully automated with integration of the RoboMover robotic unit. It can also be expanded by the addition of a small incubator that provides defined conditions during the cutting process, hence offering a high-throughput and contact-free method of sample isolation from the biological material (promotional materials of Carl Zeiss Microscopy GmbH, Jena, Germany).
Interestingly, increasing the homogeneity of the in vitro culture is also achievable by using another laser-based method. Contrary to previously described cold ablation techniques, photothermolysis uses high temperatures and the phenomenon of thermal confinement to eliminate unwanted cells from mixed cell populations.52,53 The key components of this system are either iron oxide microparticles or gold nanoparticles that robustly absorb light within the visible spectrum. These particles are conjugated with antibodies against the antigen of interest to label cells that are subsequently irradiated with pulses of laser in the nanosecond range, with the duration depending on the size of the particle used. The energy from laser pulses is then absorbed by particles, swiftly raising their surface temperature, leading to elimination of specific cell types from cultures by membrane damage of a particle-labeled cell. 54 The photothermolysis offers a high level of precision, highlighted by reports of the absence of damage to neighboring cells, which maintained their ability to proliferate and differentiate 53 and did not undergo apoptosis. 52 Despite several advantages of laser-based purification methods, these systems are cost-prohibitive.
Magnetic-Activated Cell Sorting for Cell Enrichment
Many of the mechanical passaging methods described above have not yet been integrated to fully automated platforms, and scalability must be taken into consideration, as the feasibility of passaging must be maintained with an increasing number of cells. Ideally, an automated method for selective colony isolation should be relatively simple, allow the high-throughput processing of biological material at a low cost, and result in a potent enrichment of the cell population of interest without a negative influence on cell viability. Magnetic-activated cell sorting (MACS) offers these benefits. It uses nanoparticles conjugated with an antibody against the antigen of choice, together with a magnet to generate a magnetic field and capture labeled cells within a separation column. After removal of the column from the magnetic field, labeled cells can be easily eluted for further processing. The idea of MACS was introduced in 1990, 55 and since then, it has been employed in more than 20,000 studies for the isolation of various cell populations (promotional materials of Miltenyi Biotec, Bergisch Gladbach, Germany). MACS allows an efficient and straightforward enrichment of cells expressing TRA-1-6056–59 and SSEA-4,57,59 markers of human ESCs and iPSCs, for the selection of bona fide iPSCs after reprogramming.56,58 This approach is elegantly exemplified in Paull et al. 9 with the selection of incompletely reprogrammed cells, which resulted in 26-fold enrichment of TRA-1-60-positive cells following enzymatic dissociation. We also used this approach for the selection of reprogrammed iPSCs, as well as for cell enrichment following passaging on the automated platform. 14 Automating and integrating cell ablation and magnetic-based sorting techniques into robotic platforms will allow purification of heterogeneous cell populations, and hence is likely to increase the efficiencies of differentiation protocols. This will enable generation of the cell types of interest from PSCs in a large quantity, which could be used to elucidate mechanisms underlying the progression of several diseases and may ultimately lead to developing a cure for their efficient treatment. Furthermore, the elimination of undifferentiated stem cells will significantly reduce the risk of teratoma formation, which is one of the biggest concerns for cell transplantation studies using PSCs. The importance of addressing this issue is highlighted by the observation that only two mouse ESCs are sufficient to initiate malignant transformation, 60 and thus the method for efficient elimination of undifferentiated cells must be developed before PSC-based therapies will become common treatment for a variety of illnesses.
Differentiation
Having established that the maintenance and passage of PSCs by automated platforms does not lead to changes in cellular morphology, gene expression, or karyotype anomalies, it is important to assess their retention of pluripotency and their capacity to differentiate. Several teams have analyzed the ability of rodent and human PSCs maintained on automated platforms to differentiate into the three germ layers. Each group observed that EBs generated by automation express differentiation markers from the three germ layers,9,21,22,24–26,38 while Soares et al. confirmed this ability in an adherent setting. 23 Moreover, Kowalski et al. optimized an EB formation method in a 384-well microplate format for high-throughput cardiomyocyte differentiation, achieving an efficiency exceeding 40%. 61 The positive outcome of these experiments prompted further experiments differentiating cells into various lineages in monolayers. Automated platforms have been utilized to successfully differentiate stem cells into neural lineages, 22 including midbrain dopaminergic neurons;9,26 neural stem cells and oligodendrocytes; 9 endodermal derivatives, such as hepatocytes, definitive endoderm, 9 and pancreatic islet cells; 26 and cell types of mesodermal origin, such as cardiomyocytes and metanephric mesenchymal cells. 9 It is noteworthy that the efficiency of differentiation experiments with automated platforms was comparable to those performed manually,9,26 while the intersample variation was noticeably lower.9,22 Broader use of automation will facilitate optimization of differentiation protocols to increase the homogeneity of produced cells and also increase statistical power, allowing researchers to analyze low-penetrance phenotypes.
Microfluidic Platforms
Numerous groups have highlighted that successful implementation of cell-based therapies in the clinic requires thorough and high-throughput quality control testing to ensure efficacy and safety (e.g., Yamanaka, 62 Gonzalez et al., 63 Andrews et al., 64 Seifinejad et al. 65 , and Miura et al. 66 ). Microscale technology or microtechnology offers multiple improvements over the standard-sized tissue culture dishes. With microtechnology, it is possible to miniaturize assays and use microfluidic platforms to conduct multiple experiments simultaneously. Parallelization of the analysis decreases variability between samples, improving the consistency of acquired data. It also significantly reduces the amount of reagents required to conduct an experiment, and shortens analysis time, leading to obtaining results faster, for example, allowing the detection of HIV and syphilis infections within 20 min. 67 Moreover, it provides an unprecedented precision in the control of the in vitro microenvironment. 68 Indeed, as the in vitro environment is artificial, it is essential to optimize the settings to achieve near-native conditions. Microfluidics gives the ability to meticulously define cell seeding density to achieve the cell number required and to control tissue-specific spatiotemporal changes in concentrations of small molecules, which can be delivered to cells in femtoliter volumes. 69 It is also feasible to organize cells into a characteristic three-dimensional structure to reproduce tissue complexity, for instance, using synthetic scaffolds and coating with extracellular membrane proteins to facilitate adhesion. 70 Microfluidic platforms can be easily automated, and thus could eventually be used for the long-term cell culture without human intervention. This would be an advantageous feature over a conventional approach, as it would standardize the cell culture process. Furthermore, microfluidic platforms are characterized by low reagent consumption, which would be beneficial for experiments involving continued medium changing, often required with long-term cell cultures and differentiation experiments. 71 Microfluidic platforms could also enable the production of high-resolution live images of cell cultures, a useful feature not only for tracking cells throughout the experiment, but also for observing the effects of external stimulation with drugs or toxins on their survival.70,72 Although these are key advantages of microfluidics, and some aspects of stem cell culture can be performed using them (reviewed in Wu et al. 73 ), most of the current platforms remain experimental, and none have yet been successfully used for the long-term maintenance of stem cells.
Applications
A schematic description of novel applications or facilitated by automation is described in Figure 1 .
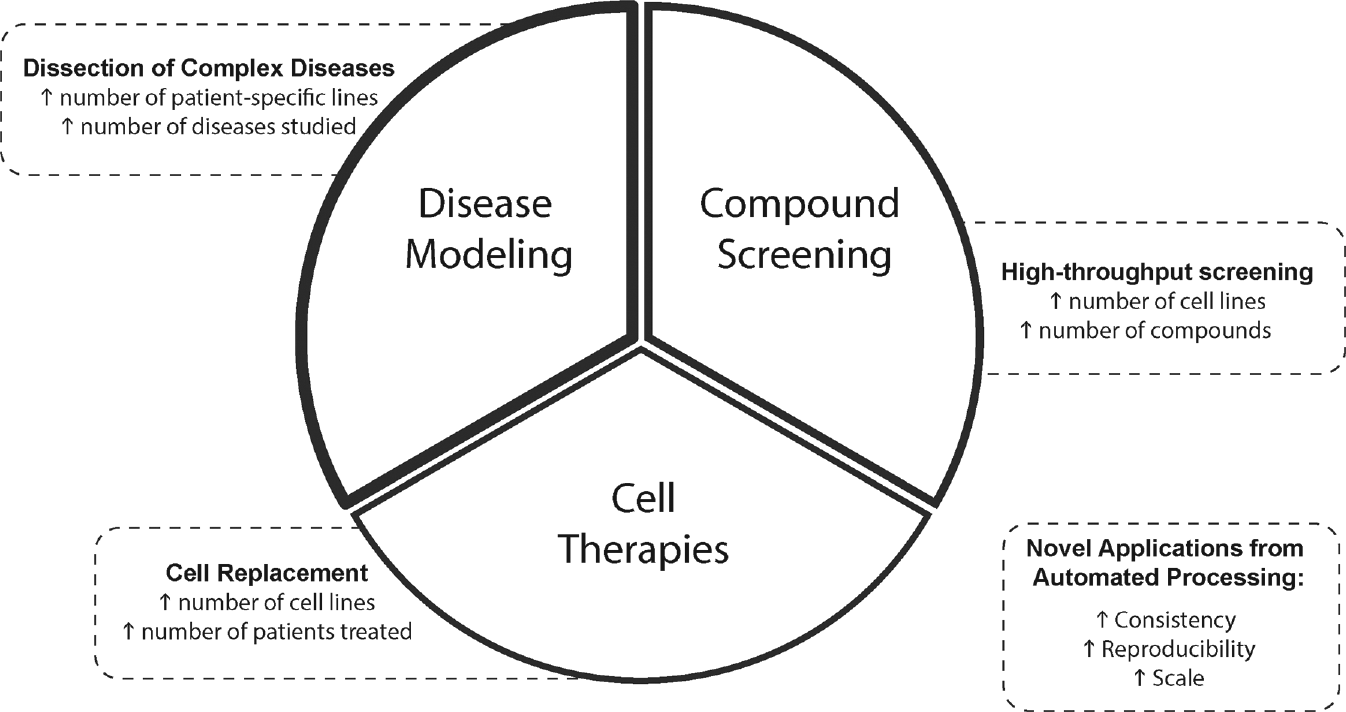
Schematic representation of novel or facilitated applications for hPSCs by automation.
Disease Modeling
Direct examination of human cells affected in disease is not always feasible due to their anatomical location; thus, there is often reliance upon isolation from postmortem tissues. A significant drawback of this is the fact that it generally allows observing the phenotype at the terminal stages of the disease, after aging and environmental impacts, which are difficult to control. Thus, postmortem tissues cannot be readily employed to investigate the development of disease and progression of disease pathophysiology. 74 Human iPSCs offer a solution to overcome this problem, as these cells represent an unlimited source of cellular material for analysis. Furthermore, iPSCs have a genetic background identical to that of the person they were derived from; thus, generating iPSCs from those with genetic diseases or susceptibilities, and subsequently differentiating them to the pathological cell types, offers the possibility of directly examining dysfunction in cells with disease-specific genotypes and phenotypes. Utilizing such disease-affected cells to track molecular changes may give insight into genetic interactions and/or mechanisms that bear a disease risk or contribute to disease development. Indeed, numerous groups have successfully reported derivation of patient-specific iPSCs from monogenic disease or multifactorial disorders. 75 However, modeling of the multigene disorders remains a great challenge because of the interplay between genetics and the environment. Additionally, the tissue-specific growth conditions and differentiation efficiency must be optimized first to ensure that cellular phenotypes displayed in vitro are in concordance with those observed in vivo. 76 The automated approach will greatly facilitate optimization of both processes, allowing researchers to streamline experiments and thus test multiple combinations of conditions at the same time.
It is worth noting that modeling late-onset diseases using iPSCs may require additional modifications of cell culture conditions in order to more closely recapitulate the age-related disease phenotype. Cell reprogramming erases epigenetic marks of somatic cells, and at the same time, pluripotency-associated marks are created.77,78 However, several groups have shown that reprogramming is a continuous process, and iPSCs at an early stage retain an epigenetic memory of the somatic cell of origin, which is gradually lost upon passaging.79–81 Nevertheless, iPSC reprogramming results in the generation of rejuvenated cells that lost age-associated characteristics, such as shortened telomeres, decreased mitochondrial activity, and senescence markers,82–84 and that also require a prolonged maturation to a functional state. 85 To address this issue, fully differentiated iPSCs could be exposed to senescence-associated compounds, such as progerin 85 or ceramide, 86 to simulate aging in vitro. Interestingly, the use of progerin reactivates the expression of age-related markers that were lost during reprogramming. 85 This strategy might not be sufficient to recapitulate all features of aging, but could allow capturing specific cell characteristics that would not be observed with fetal-like iPSC-derived progeny, hence improving our understanding of the mechanisms involved in the progression of late-onset diseases.
In addition, disease modeling with patient-specific iPSCs faces other obstacles. First, analysis between cell lines and between early and late passages may introduce variability and hinder the comparison of results. Second, many current studies employ samples from healthy individuals as controls for iPSC disease modeling. 76 This approach may introduce unwanted variability between cells derived from different individuals. Therefore, recently there has been a strong push in the stem cell field to incorporate isogenic controls for iPSC disease modeling. This involves the use of genome editing technologies to correct mutations in the patient-derived iPSCs 87 with the use of programmable endonucleases, including zinc-finger endonucleases (ZFNs), 88 TALENs, 89 or CRISPR/Cas9. 90
Drug Screening
Human PSCs and their progeny could be used for drug screening, but also for toxicity assessment, using cells differentiated into those affected in disease, as well as from other tissues, in order to assess secondary side effects and toxicity. The large-scale use of patient cells for this type of approach would greatly benefit from automation of cell culture. For instance, McLaren et al. performed a medium-throughput screening of 1,000 compounds using human iPSC–derived neuroepithelial-like stem cells and identified 24 compounds promoting proliferation/survival. 91 Microfluidic platforms would be particularly useful, as those can capture single-cell responses to external stimuli and allow visualization behavior of rare cell populations that otherwise would be missed from analysis. 92 With the advantages offered by combining an automated approach with microfluidics, iPSCs could be used to conduct large-scale adsorption, distribution, metabolism, elimination, and toxicity (ADMET) studies of newly developed drugs. 93
Despite many advantages, the two-dimensional system commonly used in the in vitro culture cannot fully recapitulate the process of development and cell differentiation occurring during organ development in vivo. The main reason behind this phenomenon may be a lack of proper cell–cell interactions that are normally required for tissue patterning during organogenesis. However, this issue may be addressed thanks to recently developed three-dimensional systems and use of organoids, 94 which are composed of multiple cell types that self-organize into an organ-like structure. 95 The first PSC-derived organoids were established by Sasai’s team, who generated three-dimensional cortical tissues from both mouse and human ESCs. 96 Since then, organoids have been developed for multiple organs, including the brain, liver, heart, lung, pancreas, and kidney (reviewed in Huch and Koo 94 and Lancaster and Knoblich 95 ). Organoids are likely to mimic organ responses to external stimuli more accurately than monolayer cell culture, and therefore could prove beneficial for toxicity screening. 95 Moreover, they have great potential to help unravel the mechanisms governing the development of human organs, such as the brain 97 and the retina, 98 and also malignant transformation. 99 The use of automation would offer greater consistency of generated organoids, resulting in the increased reproducibility of this method, and help to elucidate the mechanisms involved in tissue development that may also play a role in disease progression.
Therapeutic Cell Replacement
The most anticipated application of human PSCs is their widespread use for cell replacement therapies. Both ESCs and iPSCs are considered potential sources of cells for transplantation;100–102 however, patient-specific iPSCs have the advantage to match the genetic background of the potential patient. This holds great promise for regenerative medicine, as it would theoretically significantly reduce the risk of immune rejection. 103 iPSC-based therapies have already been examined in rodent models of Parkinson’s disease 104 and sickle cell anemia, 105 where in vitro–derived cells engrafted into host tissues offered significant improvement of observed disease symptoms. Yet, several challenges need to be addressed before iPSC-based therapies will be successfully implemented in humans. First, the generation of iPSCs for clinical applications needs to be thoroughly assessed for safety, as the viral delivery of reprogramming factors poses a risk of malignant transformation of such cells. 65 However, Sendai virus (SeV)–based vectors are nonintegrative and, at the same time, can transfect cells with high efficiency. 106 The recent advances in technology have enabled virus-free generation of iPSCs with the use of proteins,107,108 chemical compounds, 109 nonintegrating episomal vectors, 110 or miRNAs. 111 Moreover, much progress has been made to increase the efficiency of in vitro differentiation into the cell type of interest, but it is still far from obtaining a pure, homogenous cell population. Optimizing the efficacy of iPSC generation, their differentiation, and integrating methods for the removal of undifferentiated cells from the cell culture to cGMP standards will ultimately allow us to obtain high-quality biological material for safe and durable transplantation.
Conclusion
Human PSCs have opened new avenues for disease modeling, drug screening, and insight into the fundamental mechanisms underlying cell biology, in addition to the development of cell replacement therapies. Considering the tremendous potential of human PSCs, the large quantity of cells needed, the extensive manual labor, and the time-consuming procedure in their reprogramming and the maintenance and differentiation of cells, automated systems offer the possibility of reproducibility, standardization, and a robust throughput of samples. Despite slight variations in configuration and specification between these platforms, the capabilities of each system are generally fairly uniform. Consideration to automate should take into consideration the ability of a platform to increase the scale of cultures, from the generation of iPSCs; stem cell maintenance and passaging; and differentiation to selected progeny, but also the ability to perform quality control of these steps on a high scale. Costs associated with automation and the large-scale production of cells are not negligible, and should also be considered in order to keep experiments financially feasible. For instance, conductive sterile tips are expensive. Further, cultures plates, volumes, and components of culture media, as well as passaging solutions and subsequent quality control checks, will also increase as the scale of the cultures increases. Additionally, maintenance contracts for automated platforms can also represent a further financial constraint if they are to be added to a research budget. Another important challenge of transferring manual stem cell culture to automated system-based production is to merge the current technology with stem cell culture practices, and to combine the necessary hardware requirements with adequate software to ensure a smooth running of automated platforms. With the ongoing refinement of technology, automated facilities will become more versatile and user-friendly, offering scalable supply, decreased variability, and a high quality of human PSCs for research and clinical applications.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the Ophthalmic Research Institute of Australia and Retina Australia. Further support was provided by an UROP-Stem Cells Australia scholarship (RH), an International Postgraduate Award Scholarship (MD), an NHMRC Fellowship (AWH), an Australian Research Council (ARC) Future Fellowship (AP, FT140100047), a MAWA fellowship (RCBW), and University of Melbourne and Operational Infrastructure Support from the Victorian government.
