Abstract
Induced pluripotent stem cells (iPSCs) are attractive for use in early drug discovery because they can differentiate into any cell type. Maintenance cultures and differentiation processes for iPSCs, however, require a high level of technical expertise. To overcome this problem, technological developments such as enhanced automation are necessary to replace manual operation. In addition, a robot system with the flexibility and expandability to carry out maintenance culture and each of the required differentiation processes would also be important. In this study, we established a platform to enable the multiple processes required for iPSC experiments using the Maholo LabDroid, which is a humanoid robotic system with excellent reproducibility and flexibility. The accuracy and robustness of Maholo LabDroid enabled us to cultivate undifferentiated iPSCs for 63 days while maintaining their ability to differentiate into the three embryonic germ layers. Maholo LabDroid maintained and harvested iPSCs in six-well plates, then seeded them into 96-well plates, induced differentiation, and implemented immunocytochemistry. As a result, Maholo LabDroid was confirmed to be able to perform the processes required for myogenic differentiation of iPSCs isolated from a patient with muscular disease and achieved a high differentiation rate with a coefficient of variation (CV) <10% in the first trial. Furthermore, the expandability and flexibility of Maholo LabDroid allowed us to experiment with multiple cell lines simultaneously.
Keywords
Introduction
Human induced pluripotent stem cells (iPSCs) represent attractive tools for investigating disease pathogenesis and screening drug candidates, and patient-derived iPSCs have accordingly been established from patients.1–14 For example, iPSCs derived from patients with inherited muscle and neurological diseases are used for drug discovery through phenotypic screening.1,3,5,12,13,15
Long-term culture of iPSCs and control of their differentiation require advanced technical skills. Several factors have been reported to affect reprogramming efficiency and iPSC performance, including culture-related stress. 16 Moreover, a lack of standardization in interlaboratory methodologies can lead to further variability. Automation has been reported to be effective in overcoming these problems. 17 In addition, some automated cell culture systems that improve the quality of iPSCs have been developed. Most robotic instruments, however, such as commercially available automated devices for iPSC culture, specialize in large-scale cell culture18–20 and are not suitable for cultivating multiple cell types. Because iPSCs are powerful tools in drug discovery research for modeling normal and disease phenotypes and their differentiation methods vary depending on the target cells and tissues, a robot system needs flexibility and expandability to perform phenotypic screening with various types of iPSC-derived models. Fleisher et al. built an automated system using a dual-arm robot that can be used for chemical analysis. 21 This system is suitable for performing liquid-handling processes, because the robotic arm has a human-like structure, allowing it to manipulate peripheral equipment typically used by researchers. Yachie et al. developed a dual-arm robot system named “Maholo LabDroid” as a part of efforts to establish a versatile laboratory automation system. 22 Considering the background above, we speculated that this dual-arm robot system, when combined with suitably customized tools and peripherals used by humans, could be used to realize various experimental steps.
Here, we established a platform to enable iPSC research using the Maholo LabDroid with reproducibility and expandability.
Materials and Methods
Culture of Human iPSCs
The protocols for this study were approved by the Astellas Research Ethics Committee of Astellas Pharma. Each donor signed an informed consent agreement. The human iPSC line GMP1 was established at the Astellas Institute for Regenerative Medicine. GMP1 was cultured using StemFit culture medium (cat. no. AK02N, Ajinomoto Healthy Supply, Tokyo, Japan); iMatrix-511 laminin-based cell culture substrate (cat. no. 892011 or 892012, Nippi, Tokyo, Japan); TrypLE Express Enzyme (1×), a phenol red–free cell exfoliation reagent (cat. no. 12604021, Thermo Fisher Scientific, Waltham, MA); CultureSure Y-27632 ROCK inhibitor (cat. no. 036-24023, FUJIFILM Wako Pure Chemical, Osaka, Japan); and D-PBS (−) (Dulbecco’s phosphate-buffered saline; cat. no. 045-29795, FUJIFILM Wako Pure Chemical), as previously described. 23 The human iPSC line FSHD-MyoD N#1 was established by Dr. Sakurai. 10 FSHD-MyoD N#1 cell was cultured using StemFit, iMatrix-511, and Y-27632 under the conditions described above for GMP1, with the exception that Accutase cell exfoliation reagent (cat. no. 12679-54, Nacalai Tesque, Kyoto, Japan) replaced TrypLE Express for cell dissociation and harvesting.
To induce myogenic differentiation of FSHD-MyoD N#1, cells were dissociated with Accutase, and 3000 cells each were added to the wells of 96-well plates (cat. no. 6055300, PerkinElmer, Waltham, MA) coated with Matrigel growth factor reduce, phenol red free (cat. no. 356231, Corning, Corning, NY). On the next day (day 1), the culture medium was replaced with primate embryonic stem (ES) cell medium (cat. no. RCHEMD001, ReproCELL, Kanagawa, Japan) containing 10 µM Y-27632. On day 2, the culture medium was replaced with primate ES cell medium containing 10 µM Y-27632 and 1 µg/mL doxycycline hyclate (Dox; cat. no. D5897, LKT Laboratories, Saint Paul, MN). Culture on day 3 was performed using medium containing 5% KnockOut Serum Replacement (KSR; cat. no. 10828010, Thermo Fisher Scientific), Alpha Minimal Essential Medium (α-MEM) culture medium (cat. no. 21444-05, Nacalai Tesque), Dox, and 2-mercaptoethanol (2-ME; cat. no. 21985-023, Thermo Fisher Scientific). Dox was removed on day 5, and 2-ME was removed on day 6. The cells were fixed and stained on day 8 (see below).
The human myogenic cell line Hu5/KD3 was provided by Dr. Hashimoto.24,25 Hu5/KD3 cells were cultured in myoblast medium [20% fetal bovine serum (FBS) defined (cat. no. SH30070.03, Cytiva, Grens, Switzerland), 2% Ultroser-G serum substitute (cat. no. 15950-017, Sartorius, Göttingen, Germany), 1% penicillin–streptomycin (cat. no. 15070063, Thermo Fisher Scientific), and 77% Dulbecco’s modified Eagle medium (DMEM) high-glucose culture medium (cat. no. D5796-500ML, Merck, Darmstadt, Germany)] on collagen-coated dishes (cat. no. 4020-010, AGC TECHNO GLASS, Shizuoka, Japan) or collagen-coated six-well plates (cat. no. 4810-010, AGC TECHNO GLASS). 0.25% Trypsin–EDTA (ethylenediaminetetraacetic acid) cell exfoliation reagent (cat. no. T4049-100ML, Merck) was used to detach cells from the collagen matrix. To induce Hu5/KD3 cells to differentiate into myotubes, the medium was replaced with myotube induction medium (1% FBS, 1% penicillin–streptomycin, and 98% high-glucose DMEM).
Culture of Human iPSCs Using Maholo LabDroid
Maholo LabDroid was used to seed iPSCs and change medium. Details of this automated iPSC culture are described in the Results section.
Three-Germ-Layer Induction
To investigate the differentiation ability of GMP1 cultured by Maholo LabDroid, three-germ-layer induction was performed by embryoid body (EB) formation. Details are described in the Results section.
Immunocytochemistry (ICC)
To fix the GMP1, the supernatant was discarded from the culture wells, and 4% paraformaldehyde (PFA)–phosphate-buffered saline (PBS) solution (4% PFA/PBS; cat. no. 163-20145, FUJIFILM Wako Pure Chemical) was added. After incubation at room temperature for 15 min, the cells were washed three times with D-PBS (−). To fix Hu5/KD3-derived myotubes, fixation buffer HC containing 90% Histochoice tissue fixative (cat. no. H2904-100ML, Merck), 25 mM (volume: 6.25%) calcium chloride (cat. no. 07055-08, Kanto Chemical, Tokyo, Japan), and 3.75% glyoxal cell fixative reagent (cat. no. 078-00905, FUJIFILM Wako Pure Chemical) was used instead of 4% PFA/PBS. The supernatant was removed, and ICC-blocking buffer containing 5% normal goat serum (cat. no. S-1000, Vector Laboratories, Burlingame, CA), 0.3% Triton X-100 (cat. no. 35501-15, Nacalai Tesque), and 94.7% D-PBS (−) was added. The cells were then incubated at room temperature for 20–30 min, the supernatant was removed, and the 1st Ab mix ( Suppl. Table S1 ) was added.
To fix FSHD-MyoD N#1–derived myotubes, 4% PFA/PBS was added to an equal volume (100 µL/well) of the culture supernatant. After incubation at room temperature for 30 min, the cells were washed three times with D-PBS (−). ICC-blocking buffer-2 containing 0.4% Triton X-100 and 99.6% Blocking One blocking reagent (cat. no. 03953-95, Nacalai Tesque) was added, and the cells were then incubated at room temperature for 1 h. The supernatant was removed, and the 1st Ab mix ( Suppl. Table S1 ) was added.
After treatment with the 1st Ab mix, the cells were washed three times with D-PBS (−) or phosphate-buffered saline with Tween 20 (PBST) pH 7.4 (×10) (cat. no. 163-24361, FUJIFILM Wako Pure Chemical) diluted 10-fold with distilled water (1× PBST). The supernatant was removed, the 2nd Ab mix ( Suppl. Table S1 ) was added, and the cells were incubated at room temperature for 1 h and subsequently washed three times with 1× PBST and D-PBS (−).
Imaging of GMP1 was performed using a Biorevo fluorescence microscope BZ-9000 (Keyence, Itasca, IL), EVOS FL Auto 2 Imaging System (Thermo Fisher Scientific), or Gel Imager AE-6911CX (Atto, Tokyo, Japan). Images of Hu5/KD3 cells were acquired using an IN Cell Analyzer 6000 (Cytiva). Images of FSHD-MyoD N#1–derived myotubes were acquired using a CQ-1 (Yokogawa Electric, Tokyo, Japan).
Quantitative PCR (qPCR) Analysis
Total RNA from cultured cells was collected and purified using a RNeasy Plus Mini Kit (cat. no. 74136, Qiagen, Hilden, Germany). Reverse transcription of messenger RNA (mRNA) was performed using the Super Script III First-Strand Synthesis System (cat. no. 18080-051, Thermo Fisher Scientific). PCR was performed using Power SYBR Green PCR Master Mix (cat. no. 4368706, Applied Biosystems, Waltham, MA), and detection was performed with a 7900HT Fast Real-Time PCR System (Applied Biosystems). The sequences of primers are shown in
Supplemental Table S2
. The temperature and time of the PCR were 95 °C, 5 min; 95 °C, 10 s; 62 °C, 30 s; and 72 °C, 30 s for 40 cycles. Relative expression of the target gene was normalized with its relative expression of the internal control, β-actin (
Results
Figure 1A shows the Maholo LabDroid system constructed for the purpose of performing various experiments using iPSCs. Maholo LabDroid is a general-purpose humanoid robotic system for life sciences developed by the Robotic Biology Institute (hereinafter, RBI, Tokyo, Japan). The Maholo LabDroid robot system used in this study comprises a MOTOMAN-CSDA10F robot (YASKAWA Electric, Kitakyushu, Japan), pipettes, centrifuges, microscopes, and other routine equipment. In particular, because long-term culture is required for differentiation of iPSCs, we selected the dedicated robotic CO2 incubator SCALE120 (RORZE Lifescience, Ibaraki, Japan). The central MOTOMAN-CSDA10F robot was programmed to perform operations such as transporting plates and tubes, pipetting, opening and closing the covers of laboratory equipment, and accessing peripheral research equipment. Experimental protocols were visually organized using the Protocol Maker software developed by RBI. The aspirating and dispensing speed, angle, and aspirated and dispensed volume can be individually and freely set. Maholo LabDroid was operated on a class 100 clean bench. Figure 1B shows examples of movements of Maholo LabDroid.
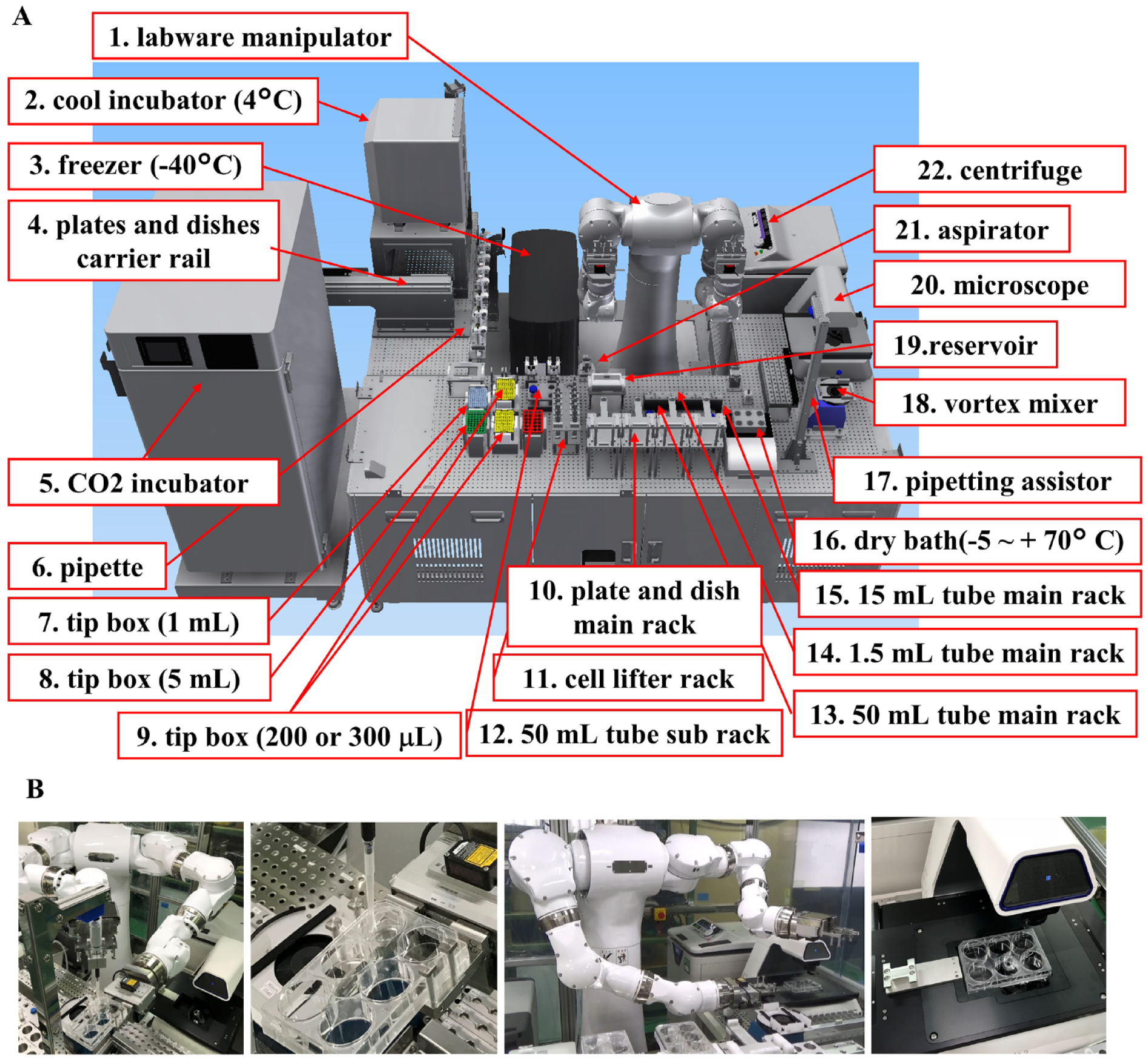
Maholo LabDroid optimized for cell cultivation. (
The experimental procedures performed by Maholo LabDroid can be customized using Protocol Maker. An example of a medium change protocol for a six-well plate is shown in Figure 2A , in which the tube containing medium was incubated in an aluminum block bath for 10 min and then moved to the main rack containing 50 mL tubes (Steps 1–3). Next, the plate was moved from the CO2 incubator to the main rack of plates, culture medium was aspirated, fresh medium was added from the 50 mL tube on the main rack, and the plate was returned to the CO2 incubator (Steps 4–6). These steps were repeated for the next plate (Steps 7–9). Finally, the aspirator was washed with ethanol (Steps 11 and 12). The operator can precisely control the configuration of the system, such as the volume of pipetting and the time, speed, angle, and height of addition or aspiration. Figure 2B shows the schedule of weekly tasks performed by Maholo LabDroid. Here, five independent experiments could be simultaneously performed, including a medium change of six- and 96-well plates and cell passage from one 6-well to five 6-well plates. These user-friendly systems enabled the researchers to use Maholo LabDroid for many types of cell culture, such as undifferentiated iPSCs, primary cultured skeletal cells, and iPSC-derived skeletal muscle.
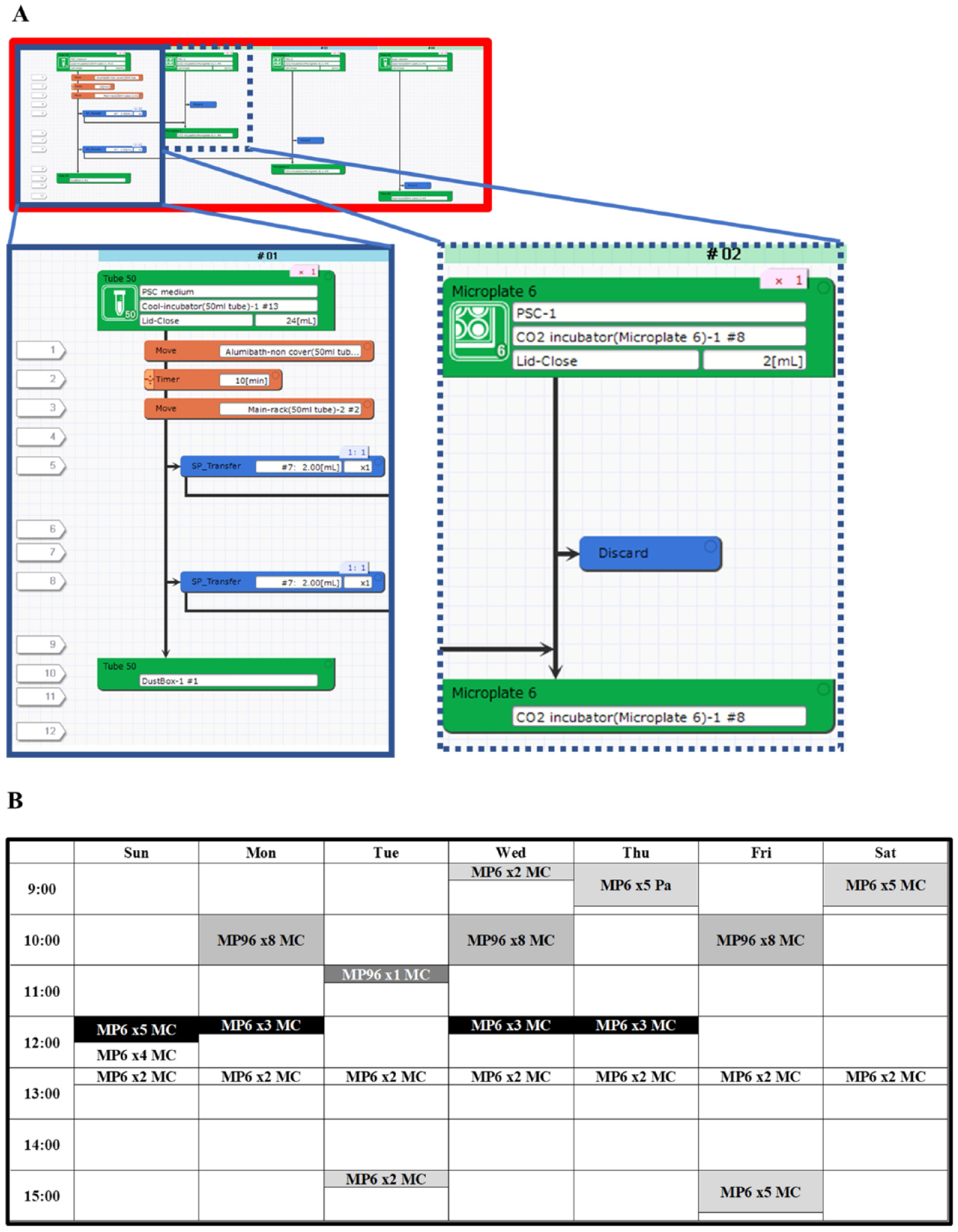
Maholo LabDroid’s weekly schedule and protocols. (
First, we optimized the movement of Maholo LabDroid for performing maintenance culture of undifferentiated iPSCs ( Fig. 3 ). A skilled researcher who has a wealth of experience tilts the plate to 20° and aspirates the medium to avoid leaving old medium. Accordingly, Maholo LabDroid was programmed to tilt the plate to 20° to mimic the motions of a researcher. The remaining amount of supernatant was equivalent to that when the medium change was performed by a researcher (data not shown). We cultivated the iPSCs for 7 days ( Fig. 3A ). Unexpectedly, the distribution of colonies in the well was biased in one direction on day 7. The area without cells tended to be on the top of the tilted plate, and the size of this area tended to be increased in the order of removal. Therefore, medium exchange of iPSCs was performed daily for 7 days with the plate tilted at either 0° or 20°. When Maholo LabDroid tilted the plate 0° to remove the medium, there was no area without cells ( Fig. 3B ). In contrast, when Maholo LabDroid tilted the plate 20°, areas without colonies tended to occur at the top of the tilted plate ( Fig. 3B ). Compared with the case at 0°, when the plate was tilted 20° by Maholo LabDroid, the variation between wells was large, and the number of collected cells was small. Furthermore, when the plate was not tilted, about 1.5 times the number of cells was recovered compared with the case with human researchers ( Fig. 3C ). When the plate was tilted at 0° by Maholo LabDroid, the remaining Y-27632 concentrations were 1.5 µM from 30 h to 48 h and 225 nM from 48 h to 72 h after seeding because of the residual medium (300 µL/well) in each well (data not shown). When Maholo LabDroid changed the medium without tilting the plate, the undifferentiated property of the cells was maintained to the same extent as when a skilled researcher changed the medium ( Fig. 3D ). We therefore decided that Maholo LabDroid should be programmed to change the medium of maintenance cultures of undifferentiated iPSCs without tilting the plate.
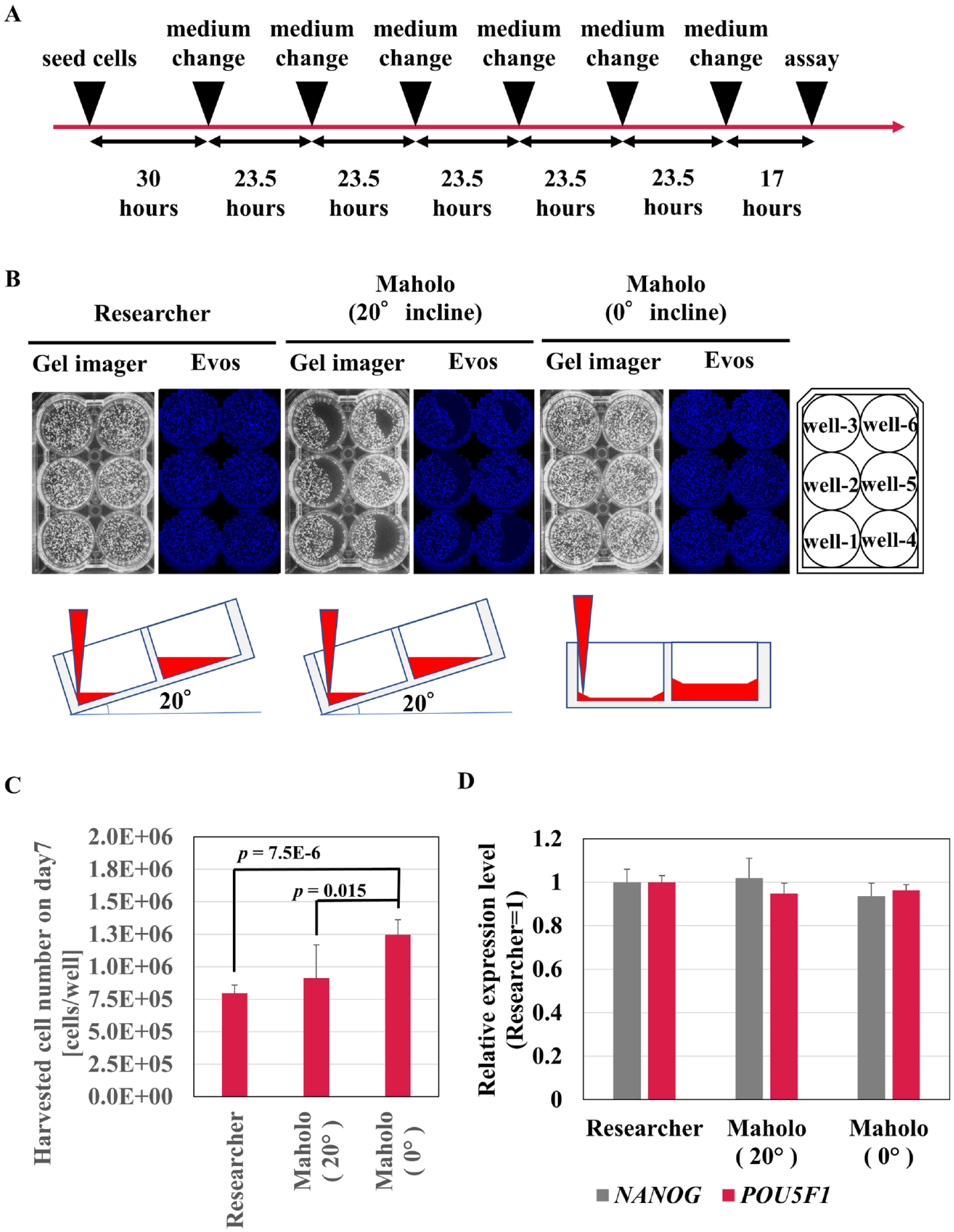
Optimization of maintenance techniques of induced pluripotent stem cells (iPSCs) using Maholo LabDroid. (
We next investigated the long-term cultivation of iPSCs using Maholo LabDroid. The daily medium change was optimized as described above, and weekly cell passaging using Maholo LabDroid was conducted by the following method ( Fig. 4A ). Washing with D-PBS (−) was performed with the plate tilted 20° to efficiently remove the medium. After washing with D-PBS (−), 0.5× TrypLE Select (1 mL/well; Thermo Fisher Scientific) treatment was performed for 10 min at 37 °C. Maholo LabDroid added 5 mL StemFit and recovered cells into a 15 mL tube containing 5 mL StemFit using a 5 mL pipette (total: 11 mL). The suspension was centrifuged, and the cells were resuspended with StemFit 10 mL. 150 µL of the cell suspension and 15 mL of the StemFit medium were mixed to make the diluted cell suspension. 1 mL/well of the diluted cell suspension was added to a new plate pre-dispensed with 1 mL of StemFit medium containing Y-27632 (final: 10 µM) and iMatrix (final: 0.25 µg/cm2). The weekly diluent ratio was 167-fold, and the growth rate was constant during this study ( Fig. 4B ). Maholo LabDroid took longer to operate than researchers. Maholo LabDroid took 1 h and 30 min to pass the undifferentiated iPSCs from one plate to two plates of a six-well plate. Maholo LabDroid took 38 min to change the medium in four plates in a six-well plate. The waste of media by Maholo LabDroid was considered to represent the same consumption as the researcher because Maholo LabDroid used the same experimental equipment as the researcher.
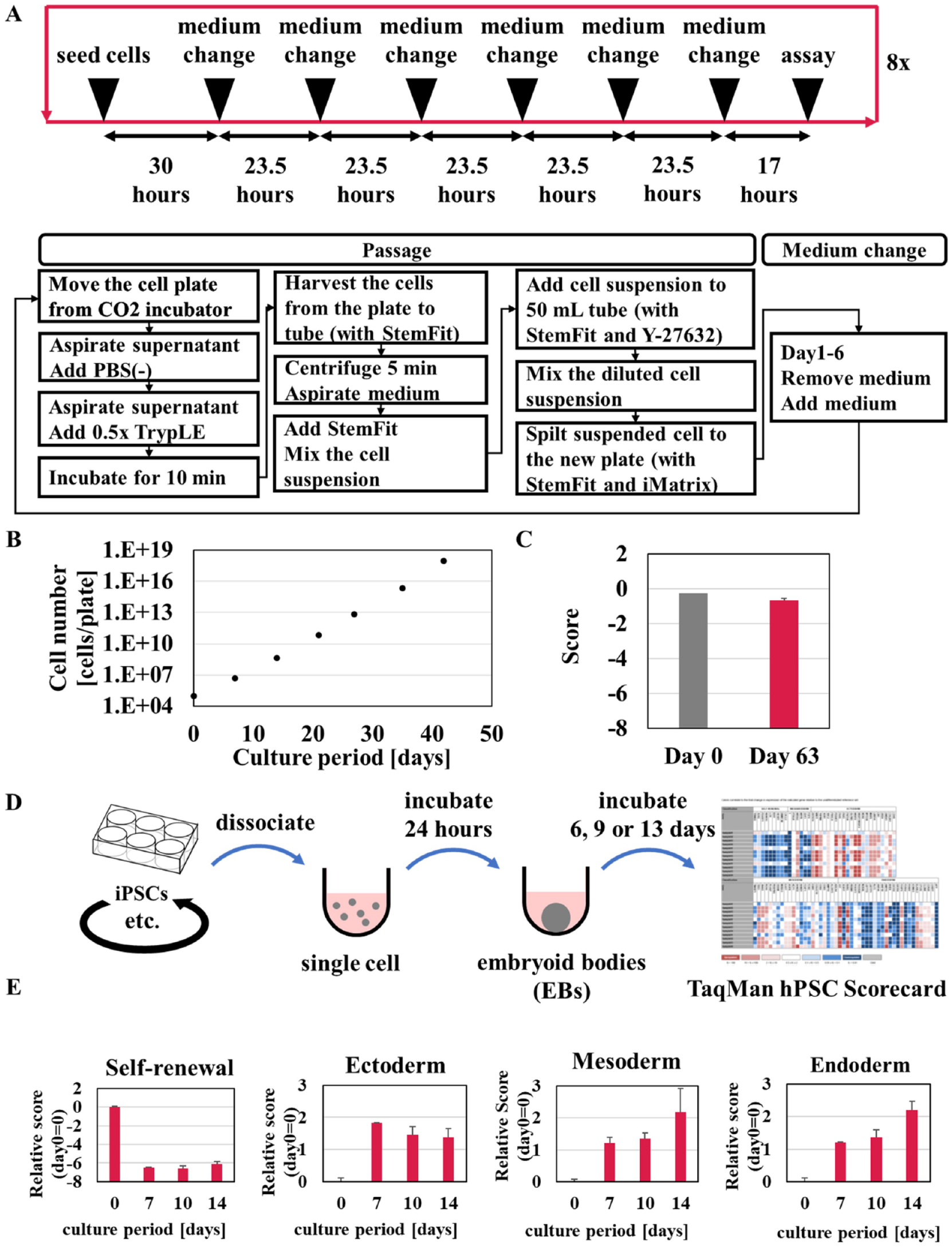
Optimization of long-term culture of induced pluripotent stem cells (iPSCs) using Maholo LabDroid. (
The target period of cell culture was set to continuous culture of 60 days or more, with reference to a report that culture of undifferentiated iPSCs could be automated and the culture could be continued for 60 days or more. 19 These operations by Maholo LabDroid for long-term cell culture were automatically repeated for 63 days. Maholo LabDroid was able to exceed the 60-day target. Human iPSCs possess the capabilities of self-renewal and differentiation into multiple cell types. The TaqMan hPSC Scorecard (cat. no. A15872, Thermo Fisher Scientific) enables verification of the potential for self-renewal and trilineage differentiation of pluripotent stem cells (PSCs) using a qPCR assay that consists of 94 genes. 26 On days 0 and 63, the cells were harvested to perform an assay with the TaqMan hPSC Scorecard. Results showed that the “self-renewal” scores were similar on day 0 and day 63 ( Fig. 4C ). To estimate the differentiation ability of these long-term cultured iPSCs, we performed induction of three germ layers with embryoid body (EB) formation of 14 days ( Fig. 4D ). Results showed that the expression levels of markers of ectoderm, mesoderm, and endoderm dramatically increased ( Fig. 4E ), and it was proved that iPSCs cultured with Maholo LabDroid for 63 days maintain pluripotency.
Differentiation of iPSCs is often induced in a 96-well plate format. We examined the movements that Maholo LabDroid was required to perform to seed iPSCs in a 96-well plate at the same cell density per plate (
Fig. 5A–5C
). Maholo LabDroid harvested iPSCs from a six-well plate and added them to 96-well plates using an eight-channel pipette. Maholo LabDroid started sequentially adding cells to columns 1–12 (
Fig. 5C
). Cells were initially added at 20,000 cells/well to investigate their distribution in the well. The cells were fixed and stained with Hoechst dye immediately after seeding. The dissociated single cells were uniformly distributed in the well (
Fig. 5B
). The number of cells per well decreased gradually, however, in the order of columns 1–12 (
Fig. 5C
, seeding condition 1). To improve the uniformity of cell numbers in the plates, we optimized the pipetting speed and number of times the cell suspension was mixed in the reservoir. We determined that Maholo LabDroid could seed equal numbers of cells into 96-well plates (
Fig. 5C
, seeding condition 2). Next, we investigated whether the quality of the iPSCs was sufficient when cultured in the 96-well plates. Maholo LabDroid seeded the cells at 1000 cells/well and maintained them for 7 days (
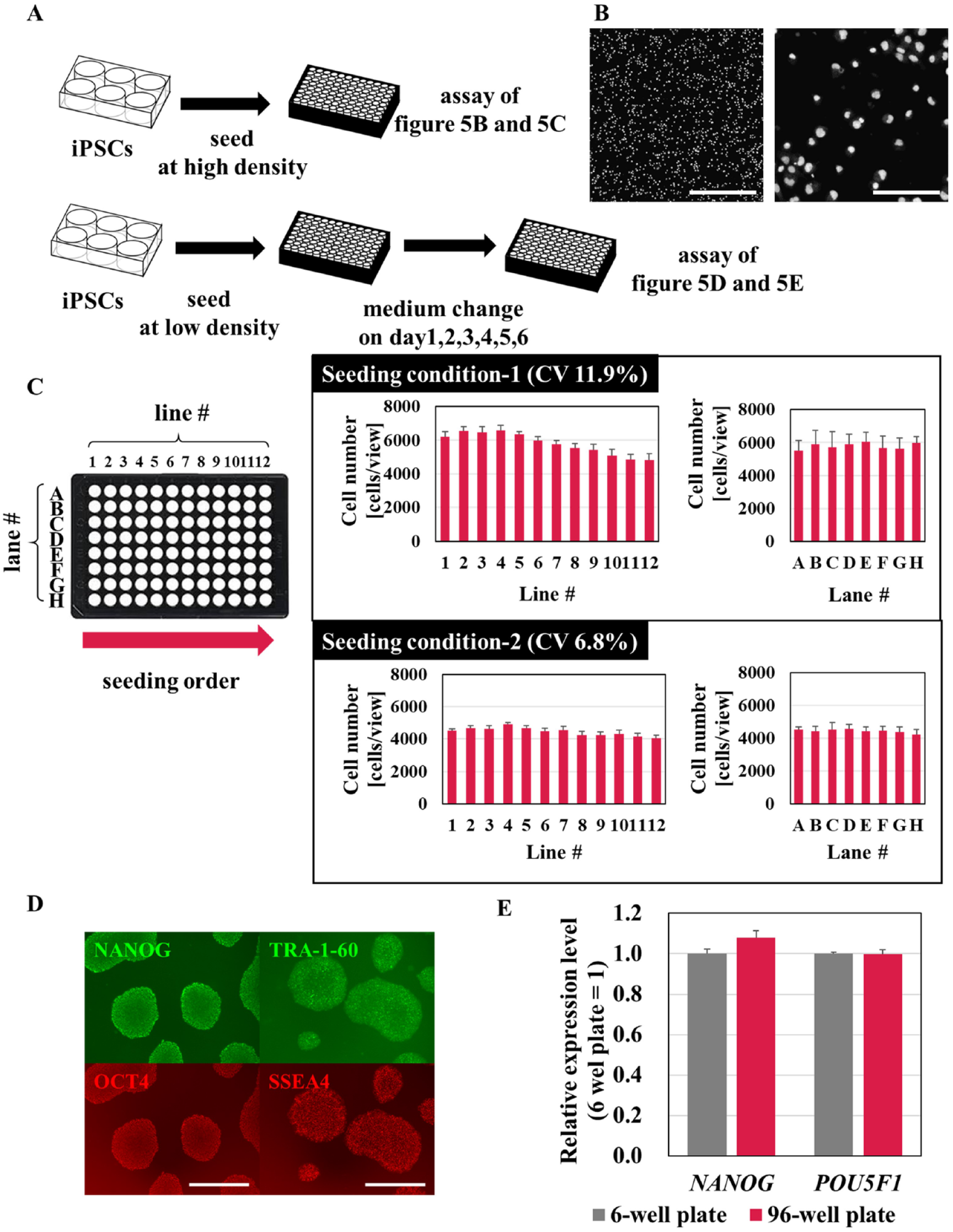
Optimization of seeding induced pluripotent stem cells (iPSCs) into 96-well plates using Maholo LabDroid. (
We examined the movement of Maholo LabDroid when performing the steps required using Hu5/KD3 cells in six-well plates as follows: cell harvest, seeding 96-well plates, induction of differentiation, fixation, and immunostaining ( Fig. 6A ). Furthermore, we also evaluated inter- and intraday variability in differentiation to assess Maholo LabDroid’s cell-culturing capabilities ( Fig. 6B ). We selected the area of immunostained myosin heavy chain (MyHC) as an index for myogenic differentiation; this area is often used in our multiple-evaluation system using Hu5/KD3 cells. A representative image of MyHC-positive myotubes is indicated in Figure 6C . The uniformity of myogenic differentiation conducted by Maholo LabDroid was demonstrated using heat-map analyses of Hoechst-positive areas and MyHC-positive areas in each plate ( Fig. 6D ). Statistical analyses of the MyHC-positive area in each vertical or horizontal lane showed reasonably high uniformity of myogenic differentiation in 96-well plates ( Fig. 6E ). The coefficient of variation (CV) values were <10%, except plate 1 ( Fig. 6F ), indicating high accuracy with low variability.
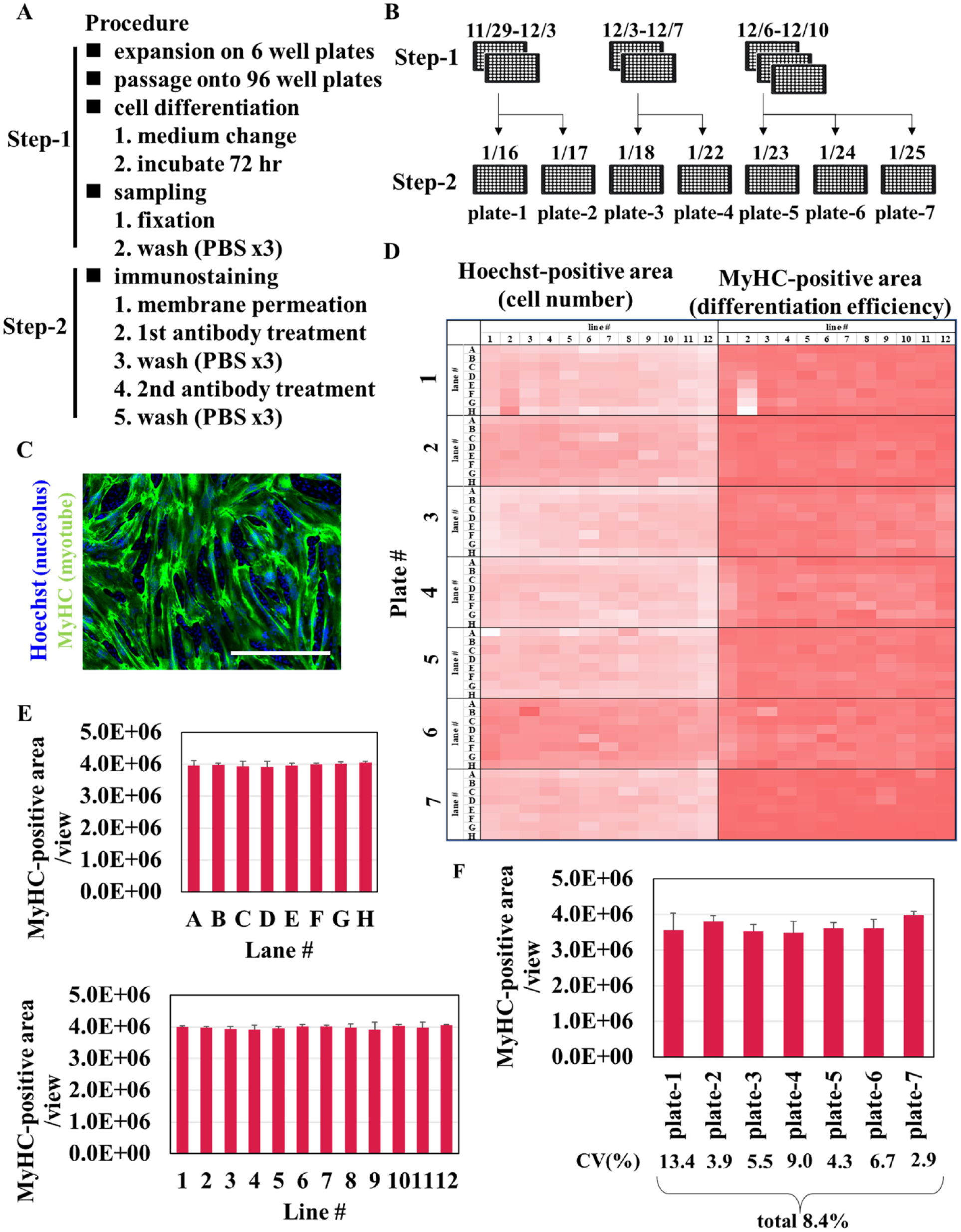
Differentiation of Hu5/KD3 cells in 96-well plates using Maholo LabDroid. (
In addition, the same evaluation was performed using FSHD-MyoD N#1. In this study, Maholo LabDroid conducted all differentiation and ICC steps except those for cell harvest and cell fixation. Specifically, Maholo LabDroid could not count the number of cells, and the handling of PFA must be done under local or external exhaust ventilation, so these steps were carried out by the researcher (
Fig. 7A
). Cell differentiation was induced using a published method.
10
Immunostaining was performed by Maholo LabDroid (
Fig. 7B
), and the count (cells/view) and ratio (%) of MyHC-positive cells were evaluated (
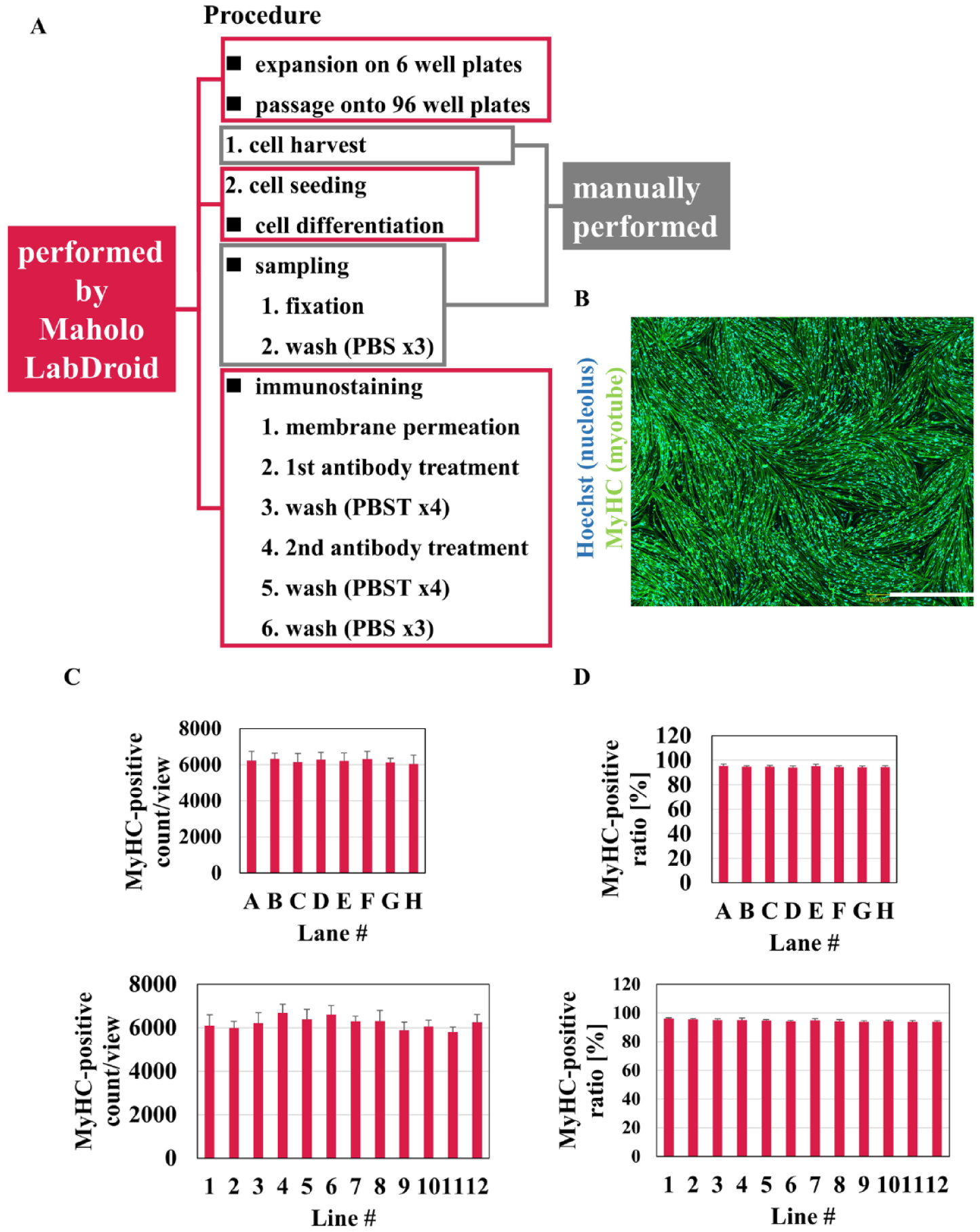
Differentiation and immunostaining of FSHD-MyoD N#1 on 96-well plates using Maholo LabDroid. (
Discussion
Here, we optimized Maholo LabDroid and established a platform for the long-term culture of undifferentiated iPSCs and for a cell-based assay with complicated steps. Maholo LabDroid showed flexible experimental capabilities combined with scheduling capabilities that allowed the simultaneous culture of multiple cell types.
Maholo LabDroid performed continuous cell culture of iPSC for 63 days, while maintaining the undifferentiated phenotype and the ability of the cells to differentiate into the three embryonic germ layers ( Fig. 4E ). These results are comparable to those of an automated cell culture system designed based on the video analysis of expert culture operations described previously. 19 Maholo LabDroid was able to achieve the long-term culture of undifferentiated iPSCs without error thanks to the high motion accuracy of the MOTOMAN-CSDA10F (0.1 mm increments) and subsequent confirmation steps. In the case of the medium change of six-well plates, Maholo LabDroid usually required a verification procedure that was intuitively performed by a human operator as follows: (1) checking whether the chip was removed from the aspirator, (2) correcting the orientation of the tube containing the medium, (3) opening the lid, and (4) mounting the tip on the pipettor. These confirmation steps helped Maholo LabDroid to operate continuously without error and to perform the long-term culture required for the differentiation of iPSCs.27,28 Maholo LabDroid, however, took a longer time to operate than researchers in terms of pipette operation and its confirmation steps. Our preliminary results showed that replacing the manual pipette with an electric pipette with continuous dispensing function reduced the working time.
When Maholo LabDroid dispensed undifferentiated iPSCs into 96-well plates, the number of cells per well decreased in the order of columns 1–12 ( Fig. 5C ). When researchers performed this task, in contrast, no decline in cell numbers was observed. With optimization of the movement used to mix cell suspensions in the reservoir, Maholo LabDroid seeded equal numbers of cells in 96-well plates; and once the parameters were determined, Maholo LabDroid moved with the same precise motion required to add equal numbers of cells.
The edge effect is one of several factors that increase the noise and variability of cell-based assays. 29 We chose the SCALE120 incubator for use with Maholo LabDroid. We selected the SCALE120 incubator due to its ability to control changes in environmental conditions due to robot transfer and its ability to achieve accurate temperature and humidity distribution with an independent heater panel. In addition to the experimental accuracy provided by Maholo LabDroid, we believe that the high performance of the incubator is also explained by the absence of an edge effect.29–31
Maholo LabDroid maintained Hu5KD3 in six-well plates and established the following processes required for assay in 96-well plates: seeding, differentiation, fixation, and immunostaining processes. Intraday and day-to-day variations of CV values in the reseeding of cells from maintenance culture, their induction of differentiation, and immunostaining experiments were all within 10%, except plate 1 ( Fig. 6F ): The second row of this plate showed an increase in the number of cells from row A to row H ( Fig. 6D ). Therefore, the CV value was not <10%, because the number of cells was not suitable for differentiation. The CV value after excluding the second row of plate 1 was 4.3%. The CV values of other plates were <10%, and no similar phenomenon was observed. These results suggest the possibility of plate quality issues rather than a Maholo LabDroid motion problem.
Maholo LabDroid was then applied to myogenic differentiation of FSHD-MyoD N#1 using the same workflow as that established for Hu5KD3 cells. The established workflow brought good CV values for FSHD-MyoD N#1 without fine-tuning of movement. As indicated in the Introduction, iPSCs are used in disease models through phenotypic screening,1,3,5,12,13,15 which requires flexible operations using numerous protocols that require different media. To support the multiple operations necessary to differentiate iPSCs into various cell types, a robot system needs flexibility, such as the ability to handle various culture vessels and media. These operations—including seeding, harvesting, and agitating cells—are challenging even for a skilled researcher. To obtain the results with good reproducibility, it is necessary to thoroughly optimize the movement of the robot. The motion of the MOTOMAN-CSDA10F, which is optimal for cell culture, was not always precisely the same as that of human researchers in some cases. Maholo LabDroid’s medium replacement method was potentially damaging to cells (
Figure 2 shows that the LabDroid scheduler ran five different iPSCs and different protocols each during a week. The expandability and flexibility of Maholo LabDroid allowed us to experiment with multiple cell lines simultaneously. Maholo LabDroid consists of equipment used by researchers, so we can freely change the system configuration. The MOTOMAN-CSDA10F, being a humanoid robot, is easy to perform what humans are doing. In addition, the MOTOMAN-CSDA10F has a seventh joint, which allows it to work in tight spaces.
Judging from publicly known information, most robotic instruments, such as commercially available automated devices for iPSC culture, are not suitable for cultivating multiple cell types.18–20 Maholo LabDroid has the flexibility to add peripherals and devices to execute new protocols. Therefore, we believe that the Maholo LabDroid will accelerate the use of iPSCs for drug discovery research by overcoming the difficulties in their use.
Supplemental Material
sj-pdf-1-jla-10.1177_24726303211000690 – Supplemental material for Establishment of a Robust Platform for Induced Pluripotent Stem Cell Research Using Maholo LabDroid
Supplemental material, sj-pdf-1-jla-10.1177_24726303211000690 for Establishment of a Robust Platform for Induced Pluripotent Stem Cell Research Using Maholo LabDroid by Miho Sasamata, Daisuke Shimojo, Hiromitsu Fuse, Yohei Nishi, Hidetoshi Sakurai, Tatsutoshi Nakahata, Yukiko Yamagishi and Haruna Sasaki-Iwaoka in SLAS Technology
Footnotes
Acknowledgements
We thank the members of the Robotic Biology Institute Inc. for collaboration and support in developing the robot system technology.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was partly supported by grants to H.S. from the Acceleration Program for Intractable Diseases Research Utilizing Disease-Specific iPSC, provided by the Japan Agency for Medical Research and Development (AMED).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
