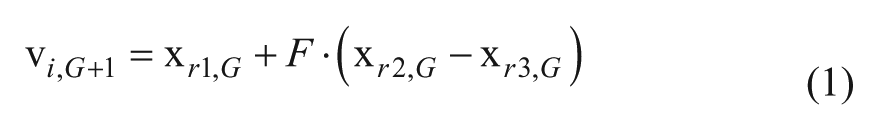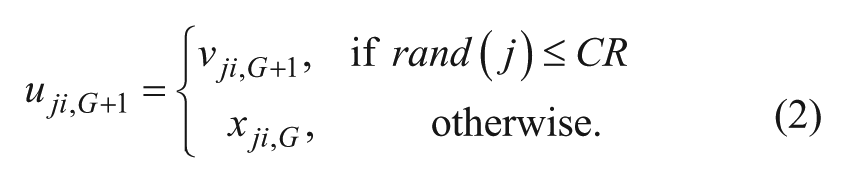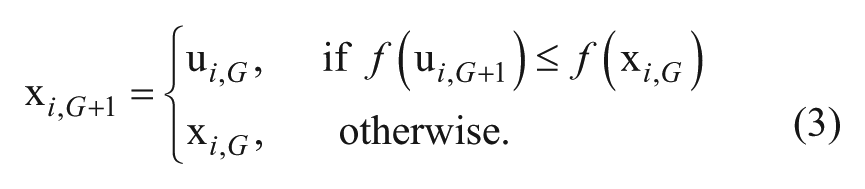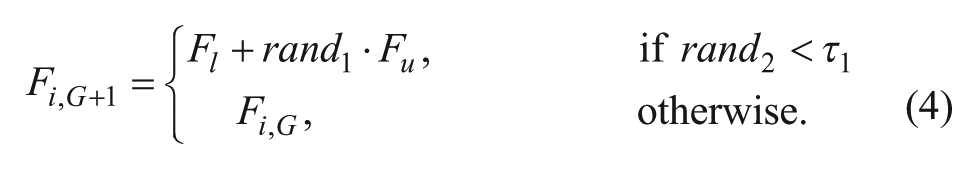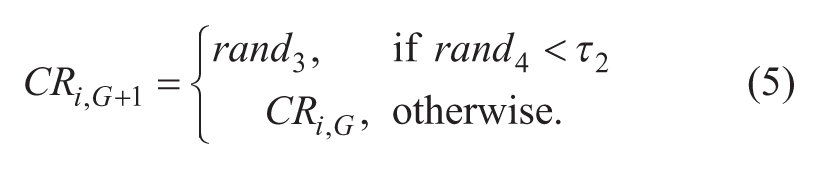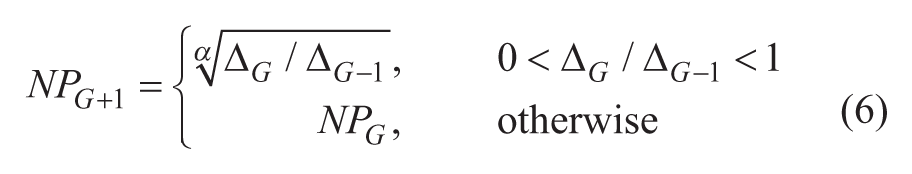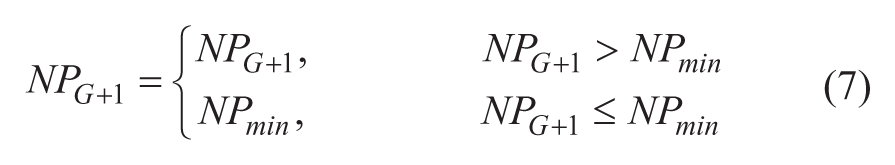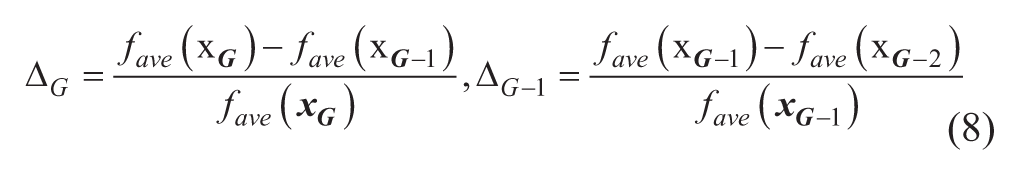Abstract
Differential evolution (DE) has been applied extensively in drug combination optimization studies in the past decade. It allows for identification of desired drug combinations with minimal experimental effort. This article proposes an adaptive population-sizing method for the DE algorithm. Our new method presents improvements in terms of efficiency and convergence over the original DE algorithm and constant stepwise population reduction–based DE algorithm, which would lead to a reduced number of cells and animals required to identify an optimal drug combination. The method continuously adjusts the reduction of the population size in accordance with the stage of the optimization process. Our adaptive scheme limits the population reduction to occur only at the exploitation stage. We believe that continuously adjusting for a more effective population size during the evolutionary process is the major reason for the significant improvement in the convergence speed of the DE algorithm. The performance of the method is evaluated through a set of unimodal and multimodal benchmark functions. In combining with self-adaptive schemes for mutation and crossover constants, this adaptive population reduction method can help shed light on the future direction of a completely parameter tune-free self-adaptive DE algorithm.
Keywords
Introduction
Differential evolution (DE) is a simple, reliable, and efficient population-based real-parameter function optimizer derived from the evolutionary algorithm (EA).1–4 A number of studies have pointed out that compared with other evolutionary-based algorithms, DE is more practical and exhibits much better performance on a wide variety of problems.5–9 Recently, the DE algorithm has been applied in combinatorial drug studies, in which multiple drugs with multiple dose levels are present and an optimal drug combination is required to develop desired treatment efficacy. To identify the optimal drug combinations from a large pool of drug candidates is traditionally considered a tedious and prohibitive task. With the advancement of technology in the emerging interdisciplinary field of combinatorial drug optimization, feedback system control strategies that were conventionally applied mainly in the engineering domain have gradually been adapted for biological complex systems. The hybrid of biological experimental advancements and engineering optimization platforms has paved the way for a paradigm shift for new drug development in various disease models.10–16 For instance, Ding et al. 15 applied DE to identify the optimal drug combination to treat herpes simplex virus type 1 (HSV-1). Nowak-Sliwinska et al. 16 demonstrated the application of DE to identify the optimal drug combination to treat cancer angiogenesis. Tsutsui et al. 17 adopted DE in their effort to identify the optimal drug combinations for long-term maintenance of human embryonic stem cells. Honda . et al. 18 proved DE could quickly provide a solution to control the osteogenesis process of human mesenchymal stem cells. The success of DE is found to be due to an implicit self-adaptation contained within the algorithmic structure, which makes it highly explorative at the beginning of the evolution and subsequently becomes more exploitative in the later stage of the optimization. 19
The simplicity, ease of implementation, and the great performance of DE in terms of accuracy, convergence speed, and robustness make it attractive for a diverse domain of real-world optimization problems. DE has been extensively applied in engineering fields such as robotics, 5 electronics, 20 engineering, 21 financial planning, 22 risk management, 23 and medical applications. 24 A review of DE applications has been presented by Plagianakos et al. 25
Although it is clear to the scientific community that DE is a powerful global optimization algorithm, there still remains considerable margin of improvement. This triggered extensive research efforts over the past decade, resulting in a wealth of variants designed to improve the performance of the original DE.5,6
The performance of the original DE algorithm is mainly affected by three control parameters: the mutation factor F, the crossover constant CR, and the population size NP. Because these values have to be tuned to fit specific optimization applications, a major problem that researchers face is the choice of a right set of control parameters. This problem is amplified when DE is employed for handling various difficulties in real-world applications such as high dimensionality and noisy optimization problems. In view of this, a wealth of DE variants has been proposed to improve the robustness and efficiency through the application of adaptive or self-adaptive techniques.26–30
Neri and Tirronen 5 suggested that, among a selection of DE variants, population size reduction, dynNP-DE, proposed by Brest and Maucec, 31 is one of the most effective algorithmic components in terms of robustness and efficiency, whereas the self-adaptive control parameter scheme (jDE) proposed by Brest and Maucec 32 is superior in terms of robustness and versatility over a diverse benchmark functions.
In this work, we propose a new population reduction method that can dynamically and continuously reduce the population size in response to the optimization progress. Our new method presents improvements in terms of efficiency and convergence over dynNP-DE. The method is DE structure independent, so it can be integrated into any type of DE variant.
This article is organized as follows. The next section reviews the background of the original DE algorithm and surveys recent work on different variants of DE. The third section describes the mechanism of the self-adaptive control parameter DE scheme. The section also provides the detailed mechanism and properties of our new population-based method, which we termed the continuous adaptive population reduction (CAPR) method. The fourth section presents experimental results on the comparisons of the performance between our new method and the well-studied population reduction scheme dynNP-DE, by testing with well-known benchmark functions under a wide range of dimensions. The fifth section discusses the experimental results. The sixth section concludes this work.
Related Works
Most of the DE variants proposed in the literature are based on either adaptation or self-adaptation of the crossover and mutation rate, namely, F and CR.26,29,32–35 These variants of DE have been shown to greatly improve the robustness and efficiency by dynamically adjusting the control parameters.
Liu and Lampinen 33 proposed a fuzzy adaptive DE algorithm that uses fuzzy logic controllers to adapt the search parameters for the mutation and crossover operations. Brest et al. 27 proposed a self-adaptive scheme (jDE) in which the control parameters F and CR are assigned to each agent in the population. Better agents, resulted from better control parameters, are more likely to produce better offspring to survive the selection process. These offspring in turn enhance the likelihood for the better parameter values to propagate. Das et al. 36 modified the original mutation scheme in DE by involving both global and local neighbors, which are defined as the entire population and a portion of the nearby agents in the population, respectively. The combined contributions between these two parts are weighted self-adaptively. Neri and Tirronen 35 proposed a scale factor local search DE, which applied a local search with certain probability to the scale factor F, corresponding to the offspring with higher performance, during the generation of an offspring.
Besides the variants that focused on the adaptation and self-adaptation of the control parameters, others focused on the adaptation of various mutation strategies or hybridization with other strategies. Fan and Lampinen 37 proposed the trigonometric differential evolution (TDE) in which the mutation operation is replaced by a trigonometric mutation scheme with a prefixed probability. Qin and Suganthan 29 proposed a self-adaptive DE (SaDE) algorithm in which different mutation strategies are allowed to switch between each other while F and CR are allowed to self-adapt without any predefinition. Rahnamayan et al. 38 proposed an opposition-based DE that applies the logic of opposition points to generate initial population. The method improves the convergence properties of DE on noisy and large-scale problems. Mezura-Montes et al. 39 conducted a comparison study on the different DE variants.
Besides controlling the mutation and crossover parameters, it should be obvious that varying the population size during the optimization process could also be a highly rewarding approach. However, population size has so far not been studied as much as the other two control parameters. In contrast, in the context of EAs, population-sizing schemes have long been investigated in a number of works, such as Genetic Algorithm with Varying Population Size (GAVaPS), 40 Adaptive Population size Genetic Algorithm, 41 Random Variation of Population Size GA, 42 and Population Resizing on Fitness Improvement GA. 43 Lobo and Lima 44 surveyed a variety of population-sizing schemes for EAs.
The effect of population size NP on the DE algorithm is analogous to the EAs in which too small a population could cause premature convergence and too large a population could lead to stagnation. 45 In the context of DE, there exist a few works that focused on the population size. Teo 34 made the first attempt by self-adapting NP using an absolute and relative encoding scheme which was improved by Teng et al. 46 Neri et al. 24 proposed an integration of fitness diversity into the DE structure where population size and the other parameters encoded within each agent are self-adapted based on a measurement of the fitness diversity. Brest and Maucec 31 proposed a population size reduction scheme in which the population size is reduced in a stepwise fashion during the optimization process, so as to avoid DE stagnation with high dimensional problems. The idea is shown to significantly improve robustness for large-scale problems.31,47
In addition, several new variants of population-resizing mechanisms have been proposed recently in the framework of DE. Zhang et al. 48 recently incorporated the lifetime and extinction mechanisms of GaVAPS to regulate the population size in an adaptive manner. Dziedzic 49 proposed a clustering-based population size reduction method in which only the best specimens from each of the resulting clusters are transferred to the next generation. Wang and Zhao 50 proposed a self-adaptive population-resizing mechanism, SapsDE, in which the population size can be adjusted dynamically by eliminating a fixed amount of poorest performing population based on some online solution-search status. Zamuda et al. 51 proposed a structured population size reduction algorithm in which dynNP-DE is combined with multiple population size–dependent mutation strategies using a structured population.
DE and Related Algorithms
DE is a stochastic, population-based algorithm developed for real-valued function optimization problems.
2
It operates by having a population of agents (
A brief description of the original DE algorithm 2 iteration procedures is as follows:
Generate a population of D-dimensional target vectors
Generate a trial vector Mutation: Generate a mutant vector
where r1, r2, r3 ∈{1, . . . , NP} are random indexes and are mutually different from each other and i. F is the amplification factor ∈[0, 1], which controls the amplification of the differential variation ( Crossover: Perform binomial crossover:
where CR is the crossover control parameter. Selection: Perform selection between the trial vector and target vector using a greedy selection criterion:
Output the results of the best target vector.
Self-adaptive DE algorithms eliminate the manual tuning of F and CR for different types of functions. To facilitate the sole comparison with an existing dynamic population reduction algorithm, we used the same self-adaptive schemes and parameter settings as presented in Brest et al. 52 This self-adaptive scheme has been shown to provide enhanced robustness and great performance for large-scale problems, 48 constrained optimization, 53 and multi-objective problems 54 and has been extensively discussed.5,6
The method allows a self-tuning of the control parameters, F and CR, of each individual agent in the population in each iteration. Fi,G+1 and CRi,G+1 are updated according to the following scheme:
where randj, j∈{1, 2, 3, 4}, are uniformly distributed random values between 0 and 1; τ1 and τ2 are constant values, 0.1, which represent the probabilities that the control parameters F and CR are updated, respectively. Fl and Fu are constant values, 0.1 and 0.9, respectively, which represent the minimum and maximum values that F can take. This allowed Fi,G+1 and CRi,G+1 to take a value from [0.1, 1.0] and [0, 1], respectively, in a random manner. These parameters are renewed before each trial vector generation and are used during the mutation, crossover, and selection steps during the creation of each offspring xi,G+1.
In this section, we introduce our CAPR scheme. We have mentioned in the previous section that two control parameters, F and CR, can be self-adapted within each individual agent during each iteration run, through a self-adaptive DE scheme (jDE). 27 To further improve the efficiency of the DE algorithm, we propose a new dynamic population size reduction scheme that eliminates the manual tuning of the third control parameter, NP. The method gradually reduced the population size according to the change of gradient of the fitness value. A new NP is calculated after one cycle of function evaluations for all agents in the population:
where
After one cycle of evaluation of all agents in the population (i.e., NPG function evaluations), the evaluated function values of all agents in the population are averaged to be
The design behind our population reduction scheme was inspired by dynNP-DE proposed by Brest and Maucec. 31 In their method, population reduction was performed in a stepwise fashion a few times throughout the whole optimization process. The numbers of function evaluations are distributed evenly for each population size. More specifically, the algorithm was predefined to reduce the population by pmax number of times. After each size reduction, the new population size is always equal to half of the previous population size. During the population reduction, one individual from each half of the current population is compared based on their fitness values, and the better one becomes the survivor for the next generation. For example, if the total number of function evaluations is predefined as 100,000, pmax = 4, and initial population = 200, the population size for each generation will be NPi ∈ {200, 100, 50, 25}, and each generation will last for 100,000/pmax/NPi = {125, 250, 500, 1000} number of function evaluations. This stepwise population size reduction mechanism allows us to wonder about the possibility of a continuous, instead of stepwise, population size reduction scheme that may more efficiently and interactively reduce the population size.
In CAPR, the population size reduction rate is dependent on the gradient of optimization performance. Compared with dynNP-DE, in which the population is always reduced by half of the previous population for a predefined number of times, the population reduction size in CAPR is not a constant but a variable based on the performance of the current and previous iterations. The population is reduced only if the gradient of the current generation, Δ G , is lower than the gradient of the previous generation, Δ G-1 . As is for all population-based evolutionary algorithms, a large pool of diverse populations is required at the beginning of the optimization process for exploration reasons. In the later stages of the evolutionary process, exploitation should take place to improve the best-so-far individuals. Exploitation should start only when exploration starts to slow down. Therefore, an increase in the gradient simply means the optimization process is still in the exploration phase, and therefore, reduction of population size should not be carried out. For the same reason, if the current gradient has an opposite sign of the previous gradient, which means the performance of the current iteration is worse than the previous iteration, the population will also not be reduced.
Moreover, because of the fast population reduction rate of CAPR, we do not perform any sorting or comparison, which keeps only the best individuals in the reduced NP population. Instead, we simply randomly truncate the excessive agents in the population. This prevents the method from always picking the best individuals, which may trap the optimization into local optimum. This also avoids any addition on the time complexity to the algorithm.
We have limited the minimum population size, NPmin, to be the same as the dimension. This is higher than the minimum requirement of standard DE, which is 4 (r1, r2, r3, and i itself), for the crossover step. We found that the higher NPmin is helpful to our algorithm. As we will notice in the later sections, our algorithm indeed reduces the population quite fast ( Figs. 1 and 2 ), and thus a higher NPmin could help to reduce the chance of being trapped in local optimum.
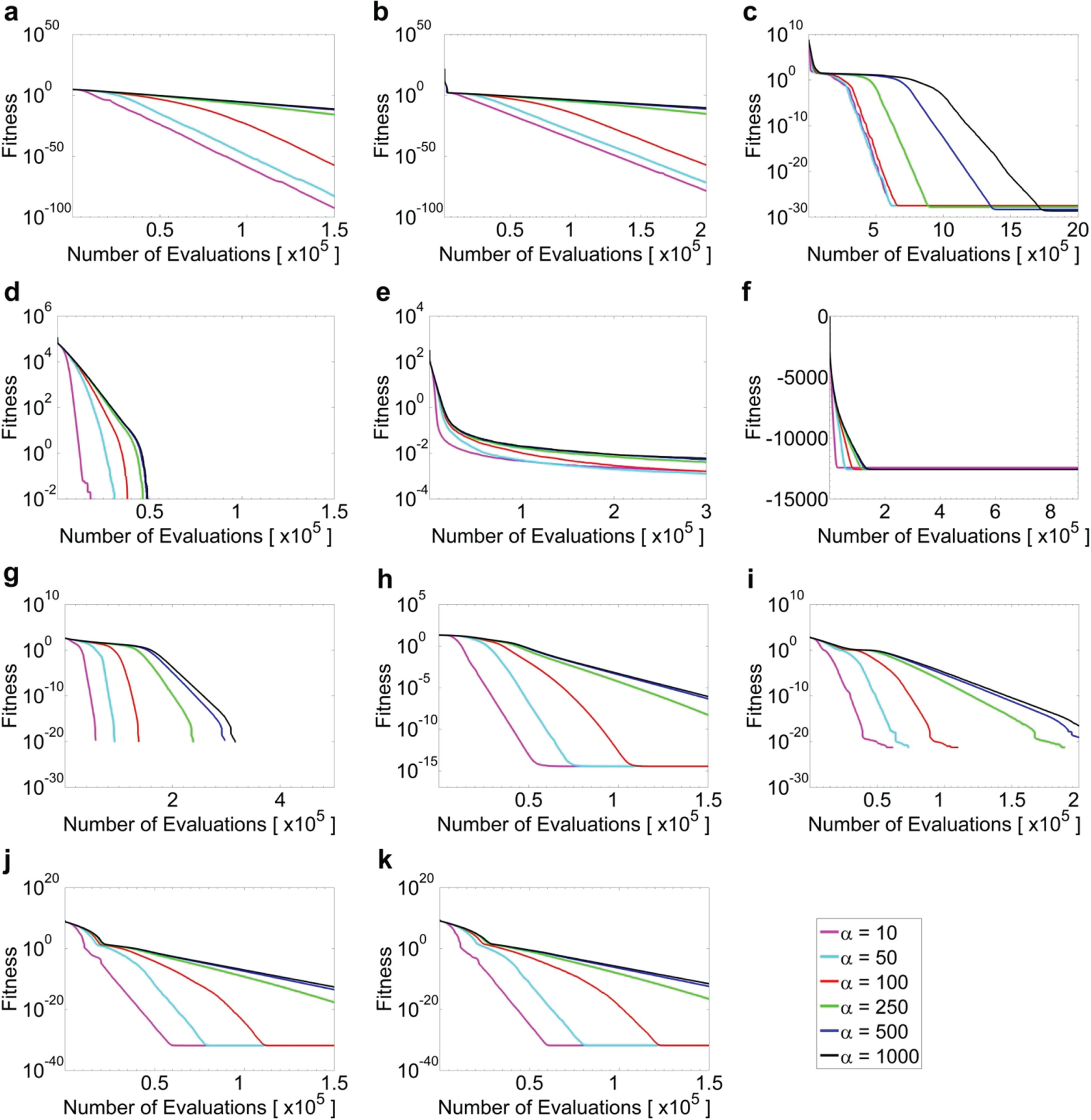
Average fitness value of all performing agents averaged over 100 independent experiments for f1 (a), f2 (b), f5 (c), f6 (d), f7 (e), f8 (f), f9 (g), f10 (h), f11 (i), f12 (j), and f13 (k) at D = 30 at different α values (see legends).
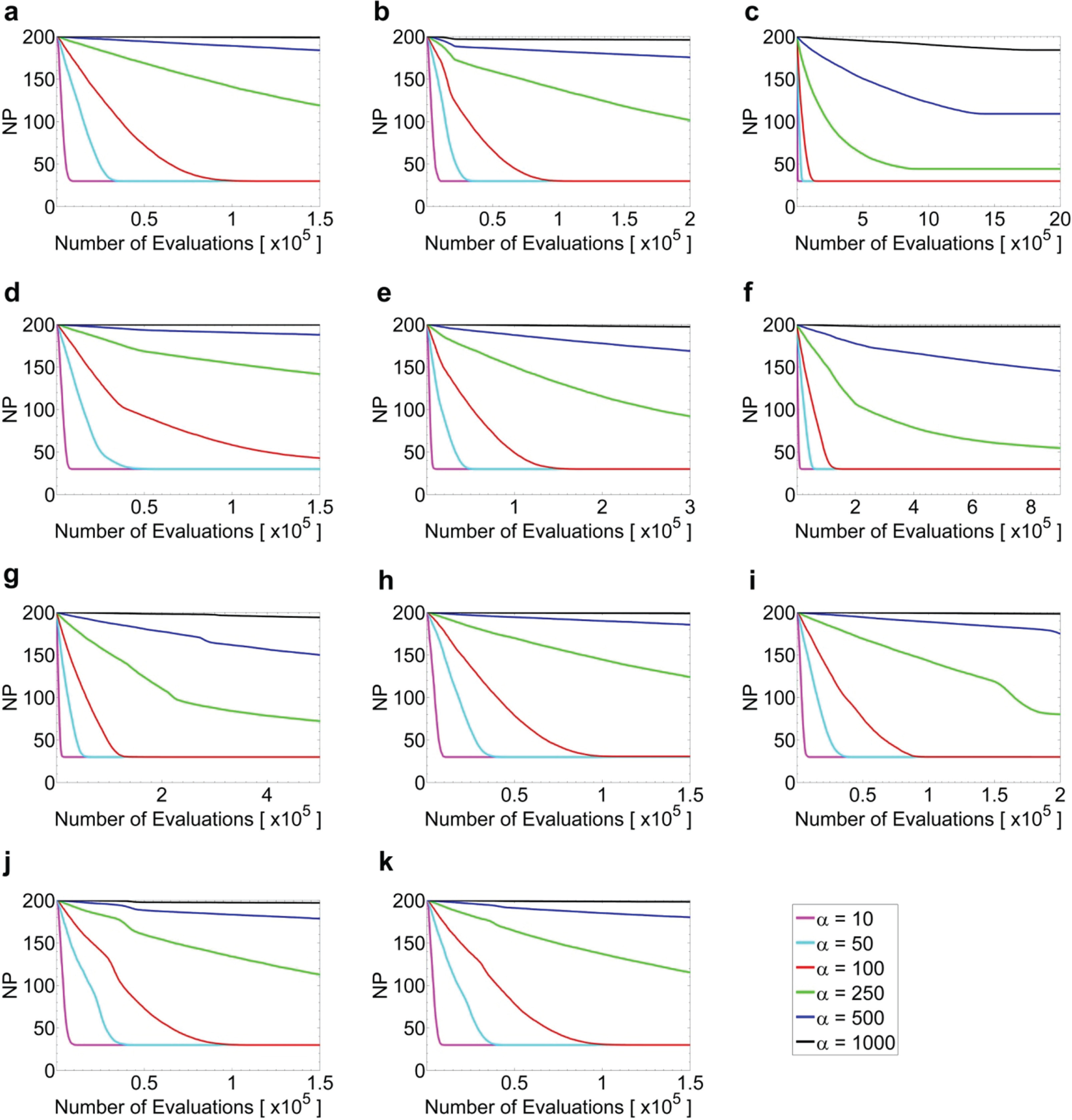
Average population size of all performing agents averaged over 100 independent experiments for f1 (a), f2 (b), f5 (c), f6 (d), f7 (e), f8 (f), f9 (g), f10 (h), f11 (i), f12 (j), and f13 (k) for D = 30 at different α values (see legends).
One should notice that our population reduction scheme is designed only for reducing the population and not increasing the population. It is certainly possible to increase the population size by a certain amount if the evolutionary process is not able to improve the optimization after a long quiescent period. In this present work, we did not investigate this aspect and would like to focus solely on the population size reduction mechanism.
Results
Our algorithm was tested with 21 benchmark functions shown in
Tables 1
Unimodal and Multimodal Benchmark Functions f1 to f13.

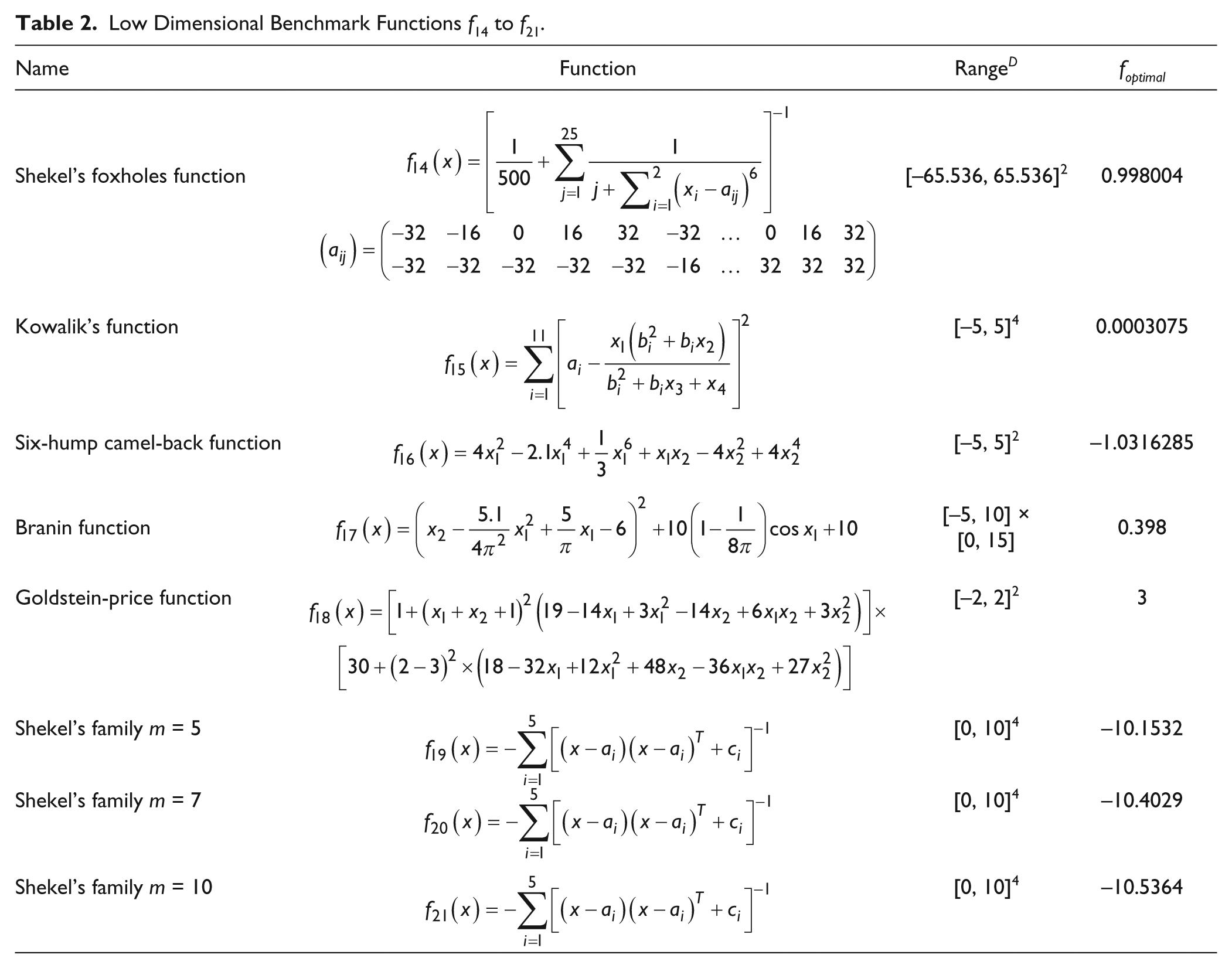
Low Dimensional Benchmark Functions f14 to f21.

Values of ai and ci for f19 to f21.
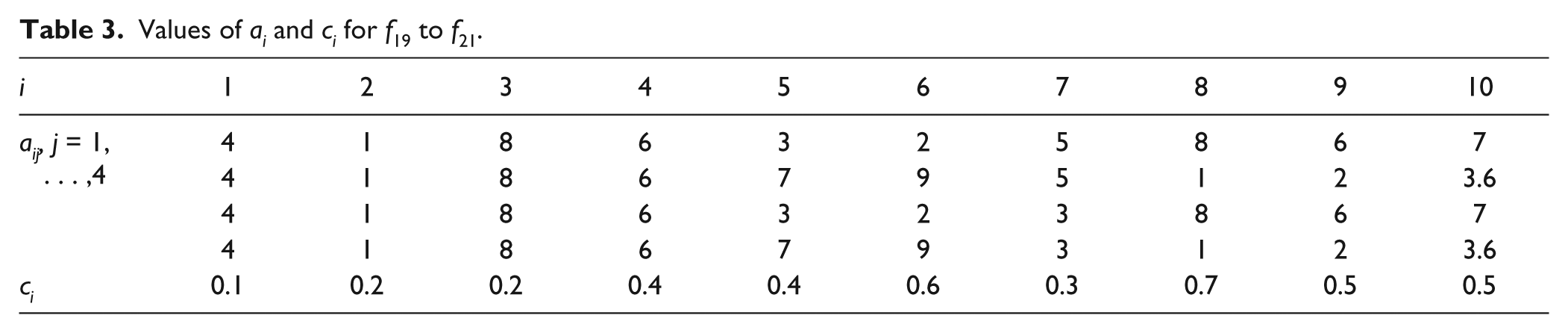
Table 4 presents the experimental results of our algorithm in comparison to dynNP-DE and the original DE algorithm with dimensions D = 30. To illustrate the sole effect caused by the population reduction algorithm, and to facilitate comparison with the dynNP-DE algorithm, we used the same self-adaptive schemes and parameter settings used in Brest and Maucec. 31 Initial values of the self-adaptive control parameters, F = 0.5 and CR = 0.9; initial NP value, NPinit = 200; and α = 100 were used in most experiments in this work, unless stated otherwise. The minimum NP value is set to NPmin = 30, which is the same as the dimension size D.
Comparison between CAPR, dynNP-DE, and the Original DE in Experimental Results, when D = 30. a
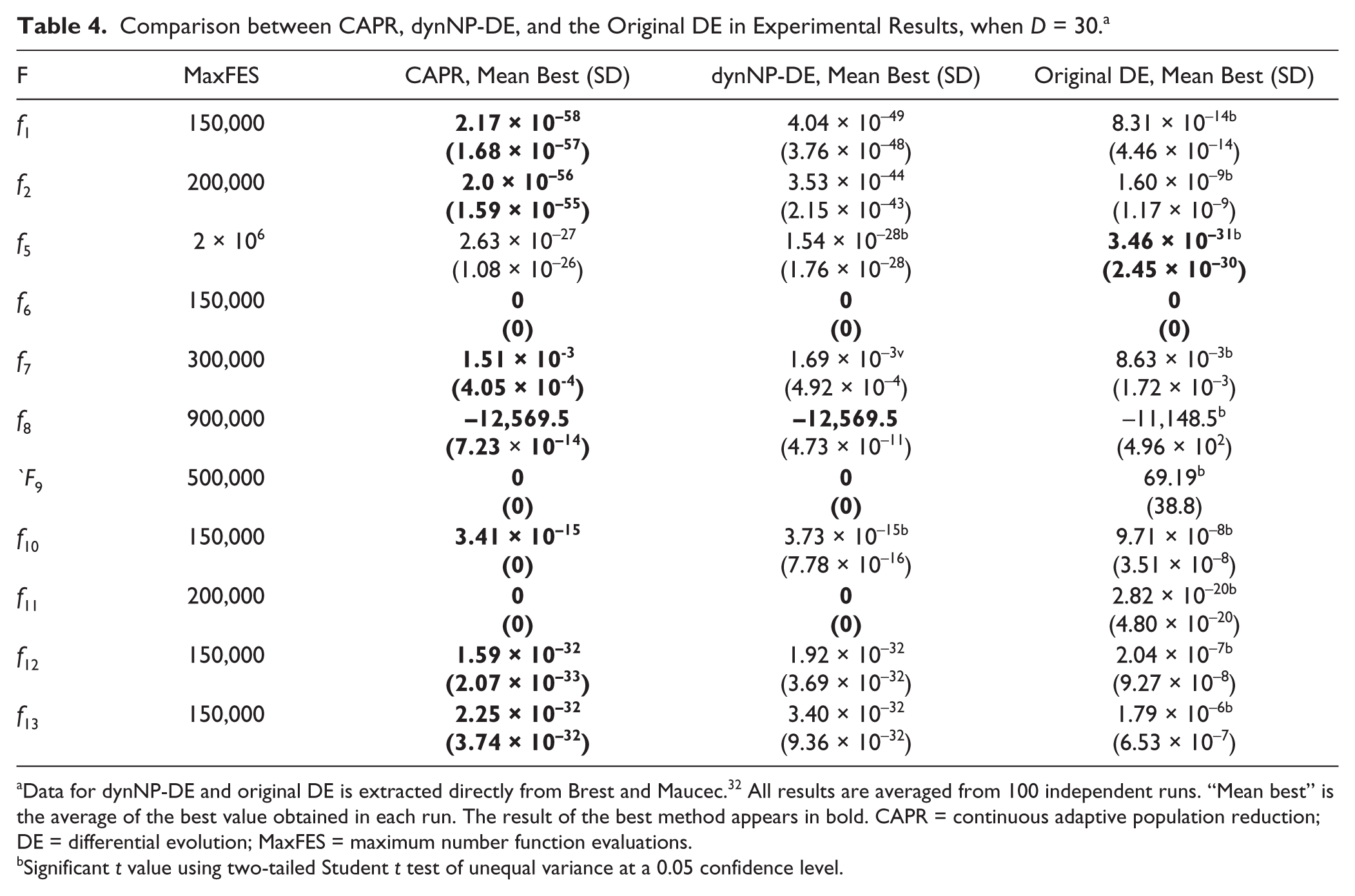
Data for dynNP-DE and original DE is extracted directly from Brest and Maucec. 32 All results are averaged from 100 independent runs. “Mean best” is the average of the best value obtained in each run. The result of the best method appears in bold. CAPR = continuous adaptive population reduction; DE = differential evolution; MaxFES = maximum number function evaluations.
Significant t value using two-tailed Student t test of unequal variance at a 0.05 confidence level.
The optimal results from 100 independent runs of each function were averaged and are listed in Table 4 as “mean best.” For each function, the result of the best method is highlighted in bold. All mean best results between CAPR and other methods were statistically compared using a two-tailed Student t test of unequal variance with a confidence level of 0.05. As shown in Table 4 , CAPR presented better results for all functions compared with the dynNP-DE, except f5. Functions f1 and f2 presented a couple orders of magnitude improvement over the dynNP-DE algorithm, and f10 showed 0 standard deviations. For function f5, both our algorithm and the dynNP-DE algorithm outperformed the original DE algorithm, most likely because of the population reduction, which decreases the diversity in the population. However, the results in Table 4 show that CAPR has the best overall performance compared with the dynNP-DE and original DE algorithm.
A parametric study of the CAPR method was investigated with different values of NPinit ∈ {200, 400} and α ∈ {10, 50, 100, 250, 500, 1000}. MaxFES remained the same as shown in
Table 4
. The average of the best (mean best) and standard deviation of the average of the best, as well as the number of successful tests that did not get trapped in local optimum, are presented in
Table 5
. Fitness value considered as trapped in local optimum is at least 1010 higher than the mean best value. For most functions with α = 10, the population size NP shrunk too fast and therefore easily got trapped in the local optimum. This can be seen from the fitness and population size plots (
Figs. 1
Parametric Study of CAPR when D = 30. a
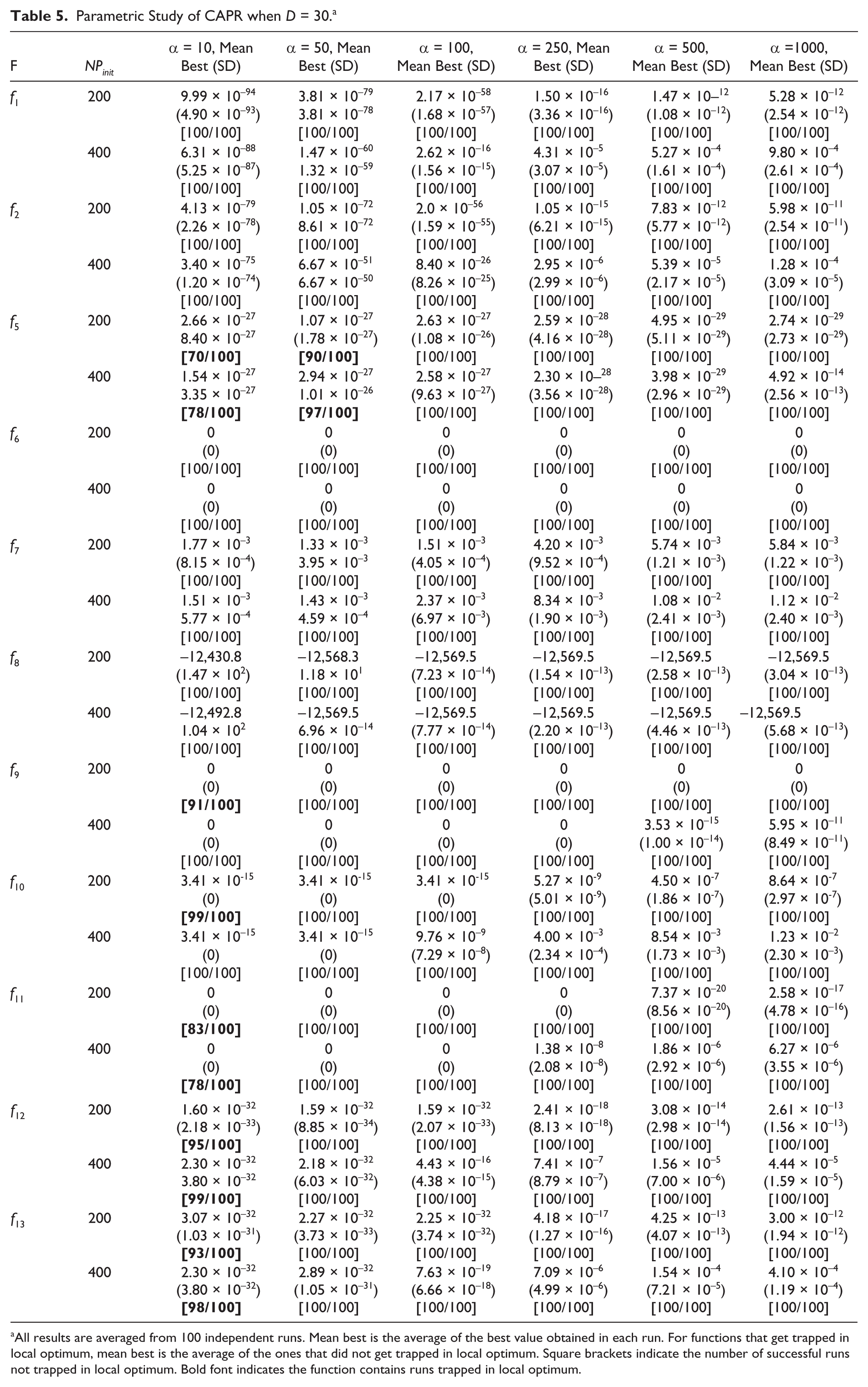
All results are averaged from 100 independent runs. Mean best is the average of the best value obtained in each run. For functions that get trapped in local optimum, mean best is the average of the ones that did not get trapped in local optimum. Square brackets indicate the number of successful runs not trapped in local optimum. Bold font indicates the function contains runs trapped in local optimum.
Significant improvements in convergence speed can be seen when dimension D is increased to 60 and 90 ( Table 6 ; Fig. 3 ). Comparing this to the results of dynNP-DE, the overall average numbers of function evaluations (FES) needed for CAPR to reach VTR are significantly lower than those needed by dynNP-DE. At D = 60, we tested two initial population sizes, NPinit ∈ {200, 400} for both CAPR and dynNP-DE. For dynNP-DE, pmax = 4 is used when NPinit = 200 and pmax = 5 is used when NPinit = 400. These are the experimental conditions recommended by Brest and Maucec. 32 Other control parameter settings remain unchanged, as in the experiments in Table 4 , except for MaxFES, which is doubled. At both NPinit values, CAPR showed significant convergence speed improvement for all functions. When NPinit = 200, CAPR can reach VTR in less than half of the FES needed by dynNP-DE for f2, f5, and f7 to f11. Similar results are found when NPinit = 400, where CAPR needs less than half of the FES of dynNP-DE to reach VTR for f2, f5, f8, and f9. From Table 6 , it seems that for both methods, a lower initial population size would lead to a faster convergence, but it should be noted that a low population sometimes could lead to a local optimum trap. Similar to dynNP-DE, f5 is still found to be a challenging function that may be trapped in local optima. When D is increased to 90, similar improvement is found. Here, pmax = 5 is used for dynNP-DE. CAPR is found to require less than half of what is needed for dynNP-DE to reach VTR for f2 and f7 to f10 and less than one-third of what is needed for dynNP-DE to reach VTR for f8 and f9.
Comparison between CAPR, dynNP-DE, and Original DE in Average FES to Reach VTR when D = 60 and 90. a
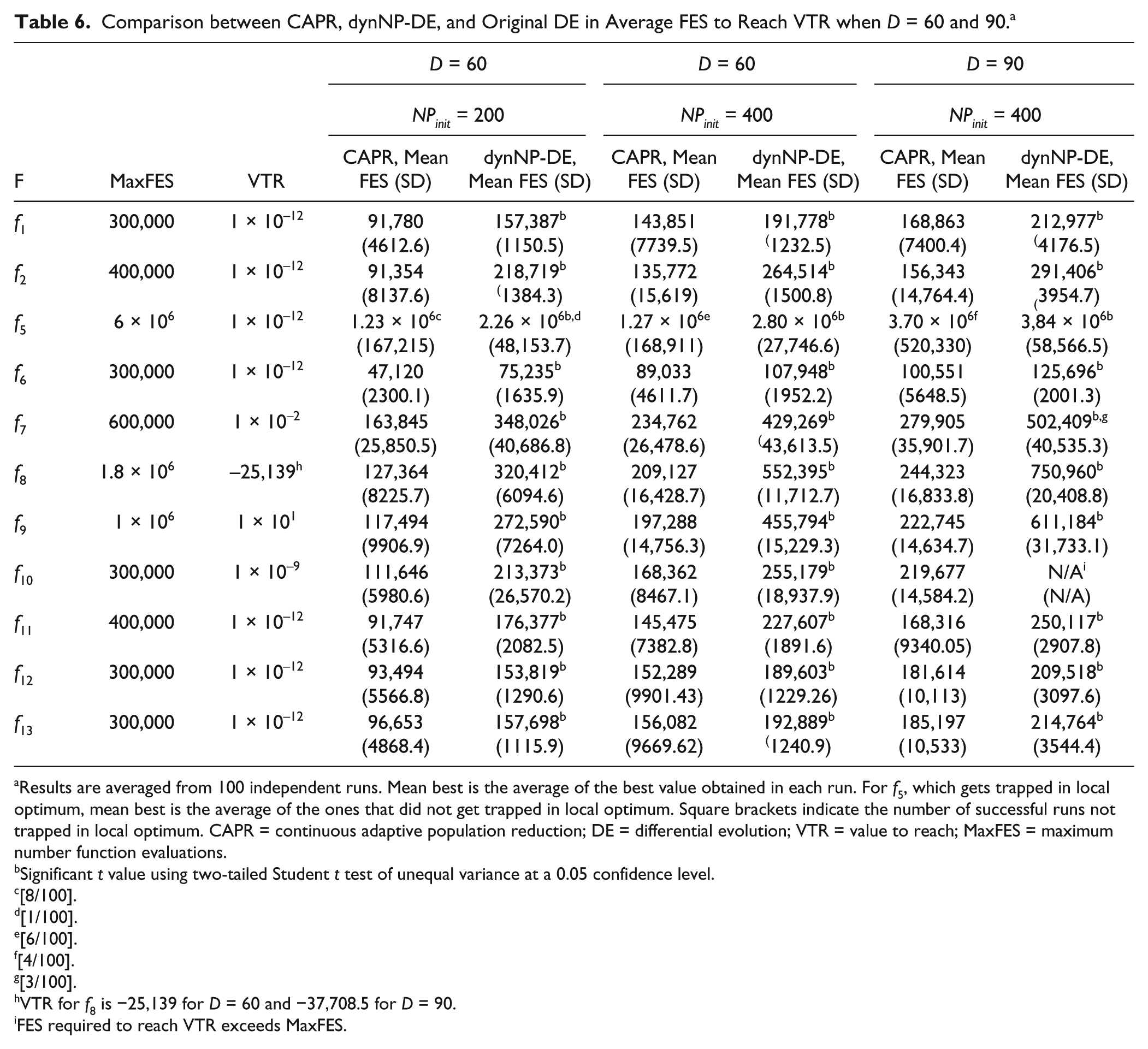
Results are averaged from 100 independent runs. Mean best is the average of the best value obtained in each run. For f5, which gets trapped in local optimum, mean best is the average of the ones that did not get trapped in local optimum. Square brackets indicate the number of successful runs not trapped in local optimum. CAPR = continuous adaptive population reduction; DE = differential evolution; VTR = value to reach; MaxFES = maximum number function evaluations.
Significant t value using two-tailed Student t test of unequal variance at a 0.05 confidence level.
[8/100].
[1/100].
[6/100].
[4/100].
[3/100].
VTR for f8 is −25,139 for D = 60 and −37,708.5 for D = 90.
FES required to reach VTR exceeds MaxFES.
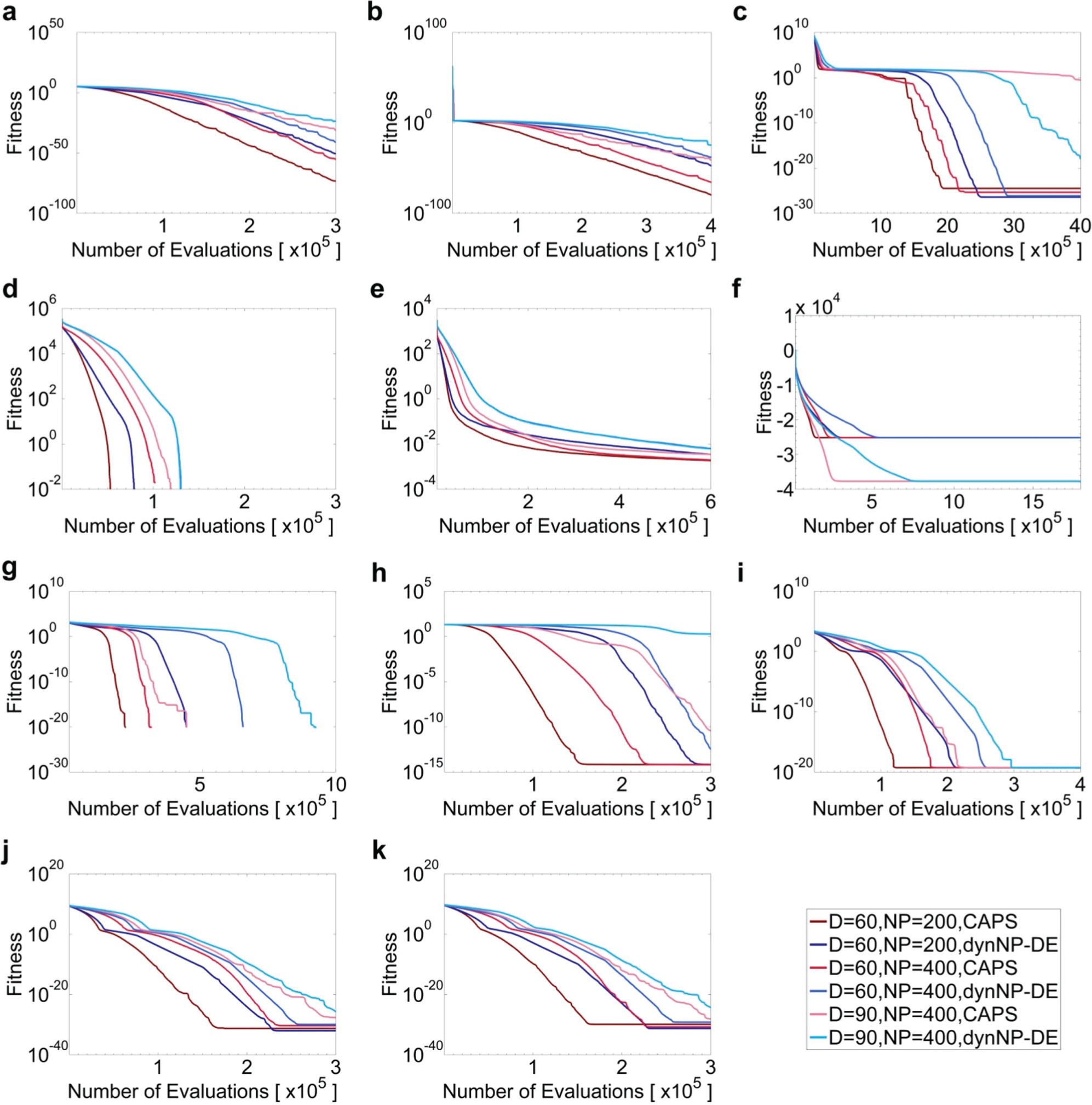
Comparison of average fitness values of all performing agents averaged over 100 independent experiments for f1 (a), f2 (b), f5 (c), f6 (d), f7 (e), f8 (f), f9 (g), f10 (h), f11 (i), f12 (j), and f13 (k) between CAPR (red color–based lines) and dynNP-DE algorithms (blue color–based lines; see legends).
Graphs plotted in
Figures 1
Figure 3 presents the comparison between CAPR and dynNP-DE on the iteration dynamics of the average fitness and NP values at D = 60 and 90 with NPinit ∈ {200, 400}. Both the curves of best fitness and NP values are plotted. Same as Table 6 and Figure 3 , pmax = 4 is used for NPinit = 200 and pmax = 5 is used for NPinit = 400 for the dynNP-DE algorithm. The other control parameter settings remain unchanged as the previous experiments. For all functions, CAPR reaches optimal fitness faster than dynNP-DE when comparing to figs. 15 to 25 in Brest and Maucec. 31
We also tested some low-dimensional functions, f14 to f21, which have only a few local minima. 54 Because the dimension D is much lower than the previous experiments, we lowered NPinit to 50 as suggested previously, where NPinit should be chosen to be about 5 × D to 10 × D2. Moreover, NPmin is also lowered to 4. Table 7 shows the comparison of performance of CAPR with dynNP-DE and the original DE.
Comparison between CAPR, dynNP-DE, and Original DE in Experimental Results, when D = 30. a .
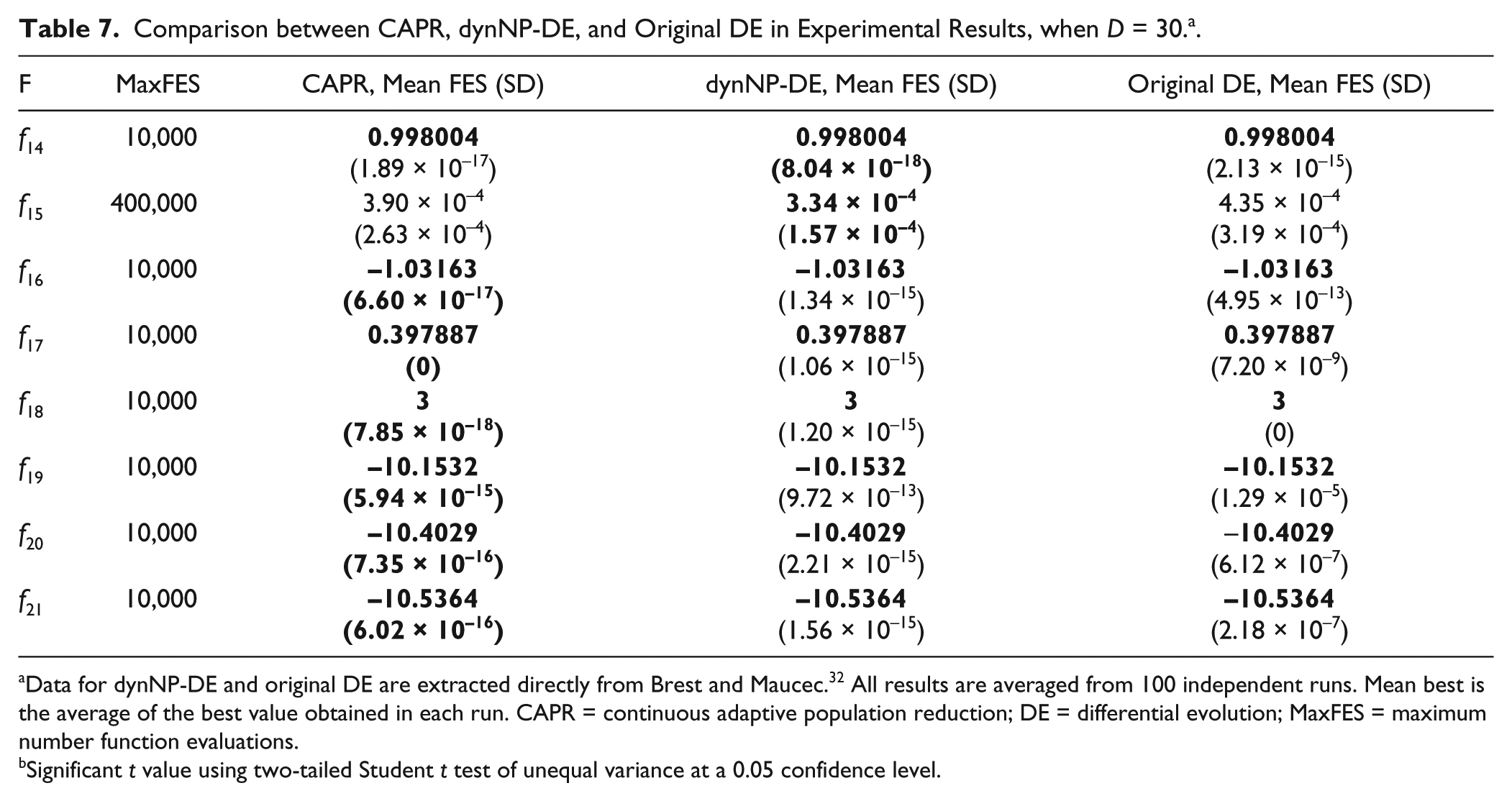
Data for dynNP-DE and original DE are extracted directly from Brest and Maucec. 32 All results are averaged from 100 independent runs. Mean best is the average of the best value obtained in each run. CAPR = continuous adaptive population reduction; DE = differential evolution; MaxFES = maximum number function evaluations.
Significant t value using two-tailed Student t test of unequal variance at a 0.05 confidence level.
CAPR shows slightly better performance than dynNP-DE for most functions, except f14 and f15. Other than f15, all three methods can reach the same global optimal values with CAPR showing the best in terms of standard deviations. Considering the fact that even the original DE can reach the global optimal values, which is not the case for f1 to f13, it is hard to realize the full capacity of CAPR because of the simplicity of these low-dimensional functions.
Moreover, we performed a characterization of the population truncation methods during the population reduction process. One is by random truncation, which eliminates the excessive number of agents randomly, and this is the scheme we have been using in all of the previous experiments. The other one is through a sorted truncation, which eliminates the excessive number of the poorest-performing agents from the population. Parameter settings are the same as those in the experiments shown in Table 4 and Table 7 , where NPinit = 200 and α = 100. As shown in Table 8 , the random truncation scheme results in relatively better convergence than the random truncation scheme for most functions, except f1, f2, and f7. This is most likely because keeping a more diverse population could prevent the optimization process from getting trapped in local optima, thus enhancing exploration.
Comparison of Experimental Results between Random Truncation and Sorted Truncation for CAPR and dynNP-DE at D = 30. a .
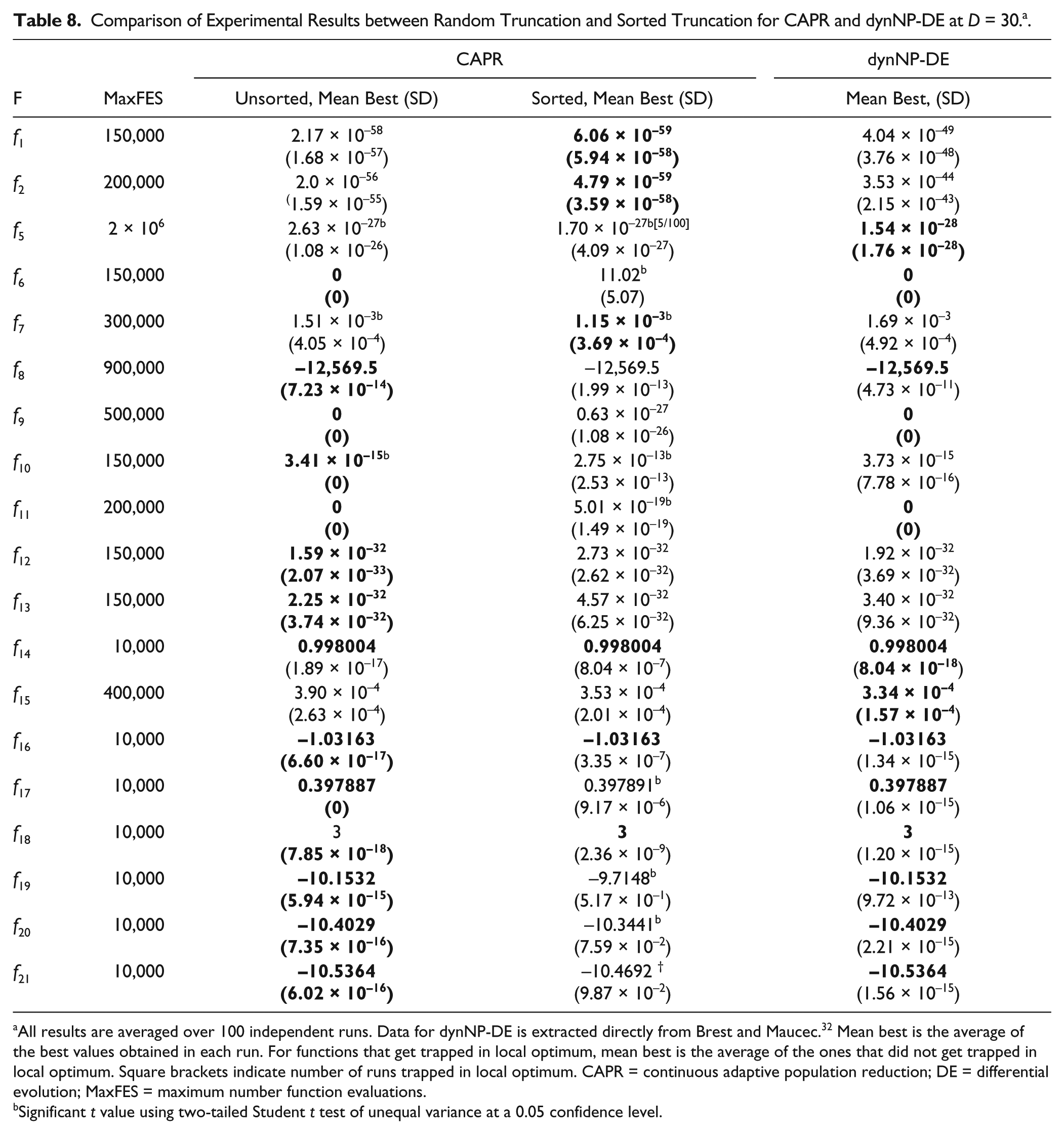
All results are averaged over 100 independent runs. Data for dynNP-DE is extracted directly from Brest and Maucec. 32 Mean best is the average of the best values obtained in each run. For functions that get trapped in local optimum, mean best is the average of the ones that did not get trapped in local optimum. Square brackets indicate number of runs trapped in local optimum. CAPR = continuous adaptive population reduction; DE = differential evolution; MaxFES = maximum number function evaluations.
Significant t value using two-tailed Student t test of unequal variance at a 0.05 confidence level.
Discussion
As inspired by the dynamic population reduction scheme (dynNP-DE), our proposed dynamic population reduction algorithm is designed to interact dynamically with the optimization space to more effectively distribute the population. Our population reduction scheme works by distributing the number of populations in response to the average fitness value in a continuous fashion, in contrast to a stepwise population reduction scheme of dynNP-DE. The study shows that updating the population size adaptively to the exploration-exploitation stage in the evolutionary process is an effective way to enhance the DE convergence efficiency.
One important feature of our population reduction scheme is that we limit the population reduction to happen only when the gradient ratio (Δ G /ΔG-1) is between 0 and 1. This limits exploitation (refining the best population) to occur only when improvement starts to slow down. If the gradient ratio of the current iteration is greater than 1 (much better performance than previous iteration) or worse than that in the previous generation, no population reduction will take place.
It is always desirable to have an optimization algorithm be both effective and robust. CAPR improves, besides the final fitness values, significantly on the speed of convergence. Such improvements are significant when the dimension is D = 60 and 90, where the average number of FES required for CAPR to reach the predefined VTR can be as low as half of those needed by dynNP-DE.
According to our parametric study, we found that a higher initial population may not always be helpful to the optimization problem. As shown in Table 5 , a higher NPinit is helpful only when the population reduction is too fast (when α = 10) by lowering the chance of being trapped in local optimum. However, when population reduction is slower (α ≥ 50), all functions perform better when NPinit is indeed smaller, except f5. We believe that the performance of a DE algorithm truly relies on the proper balance between exploration (requires a diverse population) and exploitation (requires a refined population). A similar situation is also found on selecting α, which controls the population reduction rate. At the two extremes, α = 10 and 1000, DE either reduces the population too fast or reduces the population too slow, which greatly affects the final results. Similar observations have recently been discussed by Neri and Tirronen. 5 This observation is in contrast to the common consensus that population-based evolutionary algorithms always desire a large pool of diverse population at the beginning. Although. currently, we do not have a self-adaptive mechanism of providing the optimal balance for all functions, this study proposed some best-parameter settings for CAPR, which could yield better overall results for most functions: NPinit = 200 and α = 100 for D ∈ {30, 60}.
One important process that surprises us is the agent elimination step during population reduction. The random truncation method is relatively better than the sorted truncation method. The random truncation method prevents the algorithm from always picking the best individuals during the evolution, which may trap the optimization into local optimum. Random truncation also does not need any extra sorting process, which will increase the time complexity compared with the unsorted truncation method.
Conclusions
This article proposes a population reduction mechanism for the DE algorithm, namely, CAPR. The performance of the method is evaluated through the use of a number of unimodal and multimodal benchmark functions. The new method improved the convergence speed compared with the stepwise population dynNP-DE scheme. We believe the improvement is due to the fact that our population reduction scheme is continuously responding to the progression of the optimization process, which more effectively and adaptively controls the population size during the evolutionary process, in accordance with the exploration and exploitation stages.
The article also presents parametric analysis on the control parameters involved in the continuous population reduction scheme, namely, α and NPinit, which control the population reduction rate in response to the optimization progress and the initial population size, respectively. The plotted curves of the fitness and NP values illustrate the evolutionary dynamics of CAPR and provide us information on the boundaries on the choice of these control parameters. Because the method does not require any additional sorting or comparison of the agents in the population, no additional cost is introduced, except the calculation for a new NP value after one cycle of evaluations of all agents.
An extension of our population reduction method is to include the population increment method to further improve DE. Furthermore, because there is still much less work available on the self-adaptive population control method in contrast to the self-adaptive F and CR control method, future investigations into a completely parameter tune-free self-adaptive DE algorithm will be greatly beneficial.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by National Natural Science Foundation of China (81301293) and National Science and Technology Major Projects for “Major New Drugs Innovation and Development” (2014ZX09507008).