Abstract
Digital pathology is defined as the ability to examine digitized microscopic slides and to generate qualitative and quantitative data. The field of digital pathology is rapidly evolving and has the potential to revolutionize toxicologic pathology. Techniques such as automated 2-D image analysis, whole slide imaging, and telepathology are already considered “mature” technologies and have been used for decades in exploratory studies; however, many organizations are reluctant to use digital pathology in regulatory toxicology studies. Innovative technologies using digitized slides including high-content imaging modalities and artificial intelligence are still under development but are increasingly used in toxicologic pathology. While software validation requirements are already described, clear guidance for application of these rules to the digital pathology field are few and the acceptance of these technologies by regulatory authorities remains necessary for successful adoption of digital pathology into the mainstream of toxicologic pathology. This topic was discussed during a roundtable at the 2018 Annual Congress of the French Society of Toxicologic Pathology. This opinion article summarizes the discussion regarding the current questions and challenges on the integration of innovative digital pathology tools within a good laboratory practice framework and is meant to stimulate further discussion among the toxicologic pathology community.
*This is an opinion article submitted to the Toxicologic Pathology Forum and does not constitute an official position of the Society of Toxicologic Pathology or the journal
State of the Art/Context
Digital pathology is a timely topic for pathologists, and particularly toxicologic pathologists. This term covers a large array of technologies, which are generating both enthusiasm and concern at the same time. Digital pathology is defined by the Digital Pathology Association (https://digitalpathologyassociation.org/) as “a dynamic, image-based environment that enables the acquisition, management and interpretation of pathology information generated from a digitized glass slide.” The Digital Pathology Association was founded in 2009 and is a nonprofit organization comprised of pathologists, scientists, technologists, and industry representatives dedicated to advancing the field of digital pathology. The term “digital pathology” is often used interchangeably with “virtual microscopy.”
The field includes aspects of image analysis, whole slide digitization, viewing, handling and annotating, digital slide sharing, and remote analysis. Digital pathology also includes the technologies that may be applied to digital slides such as applications of artificial intelligence (AI) or in silico multiplexing (i.e., digital bright-field image overlay: use of computational tools to combine in one single digital image the results of 2 or more “simplex” bright-field staining protocols performed on serial sections of tissues; Lippolis et al. 2013). The storage, archive, and retrieval of digital slides and their accompanying information (metadata) are also part of the digital pathology field.
The role of a pathologist in a quantification team and an overview of digital pathology in the context of toxicologic pathology have been summarized elsewhere (McCullough et al. 2004; Aeffner et al. 2016; Saravanan et al. 2017). The Digital Pathology Association has also recently published a white paper about whole slide imaging providing more technical insights (Zarella et al. 2018).
This topic was discussed at the 2018 Annual Congress of the French Society of Toxicologic Pathology (SFPT: http://toxpathfrance.org/). This opinion article is based on the roundtable discussion animated by the speakers and summarizes the current questions and challenges on the integration of innovative digital pathology tools within a good laboratory practice (GLP) framework, aiming to stimulate further discussion among the toxicologic pathology community.
Innovative Technologies
The field of digital pathology is rapidly evolving. New equipment and imaging modalities are emerging from research and related biomedical fields such as mass spectrometry imaging (MSI). MSI encompasses a suite of technologies that enable label-free analysis of endogenous and exogenous molecules directly from tissue sections. Different ionization techniques and mass spectrometry may be combined and used within pharmaceutical research and development for a wide range of applications including assessment of drug and drug metabolite disposition, target engagement and/or toxicity biomarkers, and lipidomic or proteomic analyses. Thus, MSI can be used with classical microscopic examination to provide additional molecular information on the tissue. Digitization and associated processes of data mining are essential for interpretation of the complex images that result from this kind of innovative high-content imaging technology (Swales et al. 2018).
Tremendous innovations are also linked to the use of AI concepts and tools (Esteva et al. 2017; Litjens et al. 2016; Vandenberghe et al. 2017). There is an increasing demand for quantification of histological features, but histomorphometry is resource consuming and mostly limited to specialists. Partly or fully automated methods and application of machine learning principles open the door to automated morphology detection including detection of abnormalities or specific anatomic regions on a whole slide imaging (WSI). Major software companies and start-ups are entering the field, and the development of useful and complementary tools will require intense collaboration between pathologists, bioinformaticians, and software developers. The development of AI applications for digital pathology may be an opportunity to empower pathologists, improve accuracy and efficiency, and allow the pathologist to focus more on interpreting and integrating data and understanding patterns, thereby increasing the scientific content of their contributions.
A recently founded Irish start-up (Deciphex: https://www.deciphex.com/) has, for example, developed a novel approach using AI for toxicologic pathology. They are using control tissues within each study to develop classifiers, to which the test tissue is compared, yielding an abnormality score and color-coded overlay, highlighting both the location and the intensity of the abnormality. Such a tool could facilitate integration of image analysis data with other types of data and open new research fields uncovering features that have not yet been identified by humans. However, the challenges for the required validation of these advanced technologies seem to be even more critical than for conventional image analysis tools.
Quality Expectations
The emergence of digital pathology and the massive use of electronic data create a challenge to ensure the quality and reliability of data and their compliance to GLP. The concept of data integrity should fully apply to the data generated using digital pathology tools. The World Health Organization (WHO) defines data integrity as “the degree to which data are complete, consistent, accurate, trustworthy and reliable and that these characteristics of the data are maintained throughout the data life cycle. The data should be collected and maintained in a secure manner, such that they are attributable, legible, contemporaneously recorded, original or a true copy and accurate. Assuring data integrity requires appropriate quality and risk management systems, including adherence to sound scientific principles and good documentation practices” (
WHO Expert Committee 2017). Data means all original records and certified true copies of original records, including source data and metadata and all subsequent transformations and reports of this data, which are recorded at the time of the GxP activity and allow full and complete reconstruction and evaluation of the GxP activity. Data should be accurately recorded by permanent means at the time of the activity. Data may be contained in paper records (such as worksheets and logbooks), electronic records and audit trails, photographs, microfilm or microfiche, audio- or video-files or any other media whereby information related to GxP activities is recorded. (
WHO Expert Committee 2017) the data and their metadata (defined as the associated data describing the main data, e.g., date and device used for the acquisition of digital slides) shall be inseparably and dynamically linked; the data integrity requirements apply throughout the entire life cycle of data and information, from creation to disposal; and the efforts and resources to maintain data integrity must be commensurate to the risks, particularly the risk of data integrity loss.
The risk of data integrity loss covers multiple aspects that are commonly agreed upon by the scientific community and health authorities and comprise: key quality attributes (e.g., ALCOA+: attributable, legible, contemporaneously recorded, original or a true copy, accurate, complete, consistent, enduring, and available) requirements not fulfilled; loss of data/metadata; unintentional alteration of data; and intentional alteration of data (fraud).
FAIR principles are also applicable to data since the goal of pathology assessment is to create knowledge. FAIR stands for:
The general prerequisites for computerized systems apply to “any computerized system that creates, records, processes, reviews, analyses, reports transfers, stores and retrieves data related to GxP activities” ( WHO Expert Committee 2017) and, therefore, they apply to digital pathology tools (see a summary of general prerequisites for computerized systems in Table 1).
Quick Card: General Prerequisites for Computerized Systems.a

a Example of standard quality audit questionnaire that summarize the general prerequisites for computerized systems extracted from the oral presentation of Alain Piton’s “Digital Pathology, Quality expectations and regulatory requirements to ensure data quality and data integrity” at the Society of Toxicologic Pathology Meeting (2018) in Montpellier, France.
Several challenges must be overcome when using digital pathology tools in a regulatory framework. First of all, the pathologist must justify that the following technical constraints are adequately managed: Information technology infrastructures are in place and secured: high-resolution digitization takes time and digital images use memory space. Storage and management of digital slides and associated metadata require large storage capacities and an adequate data security process. The resolution of the digital slides should be adequate for pathology assessment and/or peer review. The computer screen must be of sufficient quality (color fidelity/uniformity, bit depth, luminance, resolution/pixel pitch, and refresh rate/frequency) to permit correct visualization of the fine structures and diagnostic features.
There are currently no guidelines that define, from a practical point of view, the adequate conditions for whole slide digitization in toxicologic pathology, for example, level of scanning magnification, acceptable focus, use of multiple stacks, and if multiple stacks are used, number and spacing used. All of the elements that must be included in the digital slide should be defined properly: label, the tissue of interest or the whole tissue and elements present on the glass slide including potential defects such as artifacts like air bubbles, debris, or glue spots.
Digitization of glass slides and on-screen assessments are only one part of a global process from the original specimen to the data interpretation and reporting. The validation of algorithms used for automated measurements and/or diagnoses should be performed comparing the machine’s results with: the morphology and expected results of the examined specimen, and the results obtained by an experienced pathologist on the same tissues with a standard method.
Two Examples of Approaches That May Be Used to Ensure Data Integrity in Digital Pathology
The following two case studies are proposed here to illustrate a possible validation strategy.
Use of a Risk Analysis Approach in GLP-compliant Implantation Studies
Two intradermal implantation studies in the rabbit were requested specifically by the China Food and Drug Administration (FDA) in the context of the registration dossier of new hyaluronic acid–based gels. These studies aimed to assess the
Digital slide acquisition via the scanner: At each timepoint and before each series of scans, a universal calibration glass slide (PS20 pattern, Pyser Optics, United Kingdom) comprising 13 different predefined image areas was acquired. Selected areas on the calibration slide were measured with Definiens® and the actual areas were compared with the theoretical values to ensure reliability of the scanner over time.
Segmentation quality control: Automatic segmentation was reviewed at 100 percent by a study pathologist and/or study scientist. Conformity, manual change, and reason for change were documented on a dedicated form, and all images including segmentation were stored in a dedicated shared drive with limited access (initial and after manual changes if applicable).
Analysis script on Definiens® Algorithm integrity: Analysis of one digital slide from the first timepoint (D0) was performed again as a reference for the further timepoints to ensure the algorithm integrity over the whole study duration. In addition, the data export function was validated prior to the study phase start. All results exported in Excel® (version 2010) sheets were printed, signed, and kept in raw data.
Statistical analysis and archiving: Full validation of SAS® PC and of the archiving function in IMS® was performed prior to study phase start.
Use of a Digital Pathology System in the Context of OECD Guideline 443-compliant Studies
The European regulation Registration, Evaluation, Authorisation, and Restriction of Chemicals that has been enforced since 2007 requires all manufacturers and importers of chemicals to identify and manage risks linked to the substances they produce and market in Europe. In quantities exceeding one metric per year and per company, a registration dossier must be submitted to the European Chemicals Agency that in turn evaluates compliance with regulations and eventually grants authorization with or without restrictions. Since 2015, the European Commission uses an extended one-generation preclinical reproductive toxicity study (OECD test guideline 443). Notably, neurotoxicity is investigated to detect early effects in pups (cohort 2B) and late effects in adult animals (cohort 2A). The neurohistopathology assessment for the cohort 2B is done in pups at weaning postnatal day (PND) 21 or PND22; the brain maturation in PND21 rats corresponds to brain maturation of children about 2 to 3 years old (Semple et al. 2013). Histomorphometry is mandatory in addition to standard examination of different areas of the brain and includes linear and/or areal measurements on representative areas of the brain. At least three consecutive sections should be taken at each level section, and the pathologist or histologist selects the most homologous and representative sections. The guideline stresses the importance of preanalytical precautions and gives recommendations on the reporting. The report should include statistical analyses, details on the morphometric procedures and on the device, images of representative microscopic findings, and low-magnification images to show that the sections were homologous. If needed, historical data should also be included. The use of a digital pathology system is the easiest way to fulfill all requirements.
The trimming, embedding, sectioning, and staining of brain sections were performed according to Garman et al. (2016) to comply with the OECD 443 guideline. All slides were scanned with the Leica AT2 whole slide scanner and the associated software: ScanScope (version 12.3.2.5030) which drives the scanner; eSlide Manager (eSM, version 10.2.0.4.6): the online repository where the digital slides can be identified manually or by a bar code, visualized, arranged into different folders, and accessed through a login and a password, even remotely from third parties such as sponsors; and ImageScope (version 12.3.2.5030), the image processing and analysis software.
The “compliance module” from Leica provides 3 sets of functionality:
“Change validation”: any change of data entry in the system must be accompanied by a “reason for change.” The system administrator can create a predefined list of reasons for change (e.g., “spelling error,” “manual correction”);
“Electronic signatures” for any changes of data entry in the system. This can be used in conjunction with “reason for change”;
“System auditing,” which allows designated persons to extract data or events that occurred over a given time frame. This functionality (“audit trail”) can be configured, including the “reason for change.”
The qualification/validation of the systems for their use in the study workflow described was performed according to the GLP consensus document no. 17 and to the principle described in the GAMP®5. In our example, the computerized systems consisted of the scanning device itself (equipment), the hardware (computer), the software (programs), the network, all the standard operating procedures pertaining to the activity, and also the users, who must be properly trained (see a schematic representation of a computerized system in Figure 1). The validation included the qualification of the equipment and the validation of the process and software.
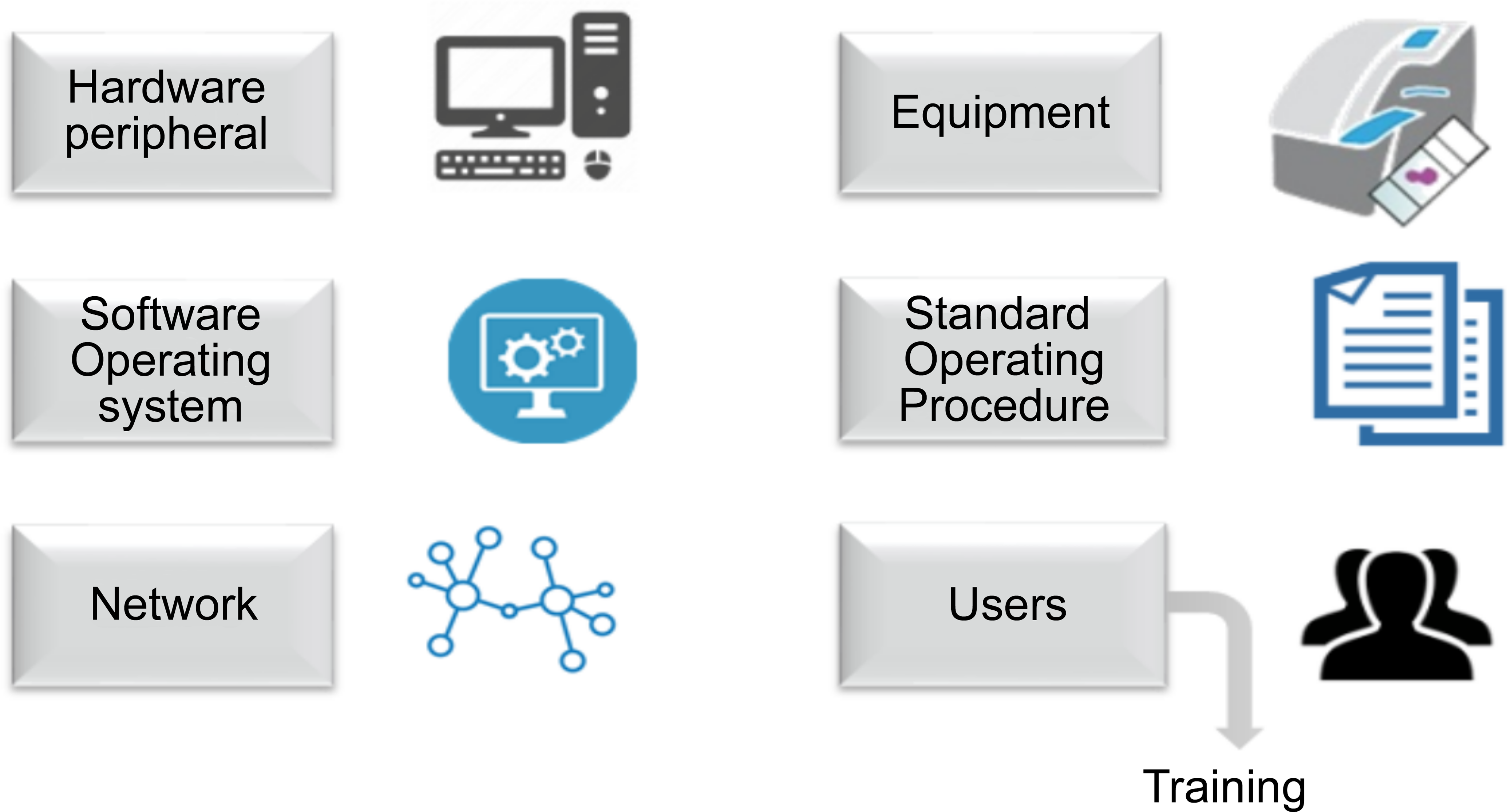
Schematic representation of a computerized system.
A validation plan that defined the scope and goals of the validation project was issued. It included the deliverables, personnel involved, time lines, acceptance criteria, compliance requirements, and risk analysis. The validation process consisted of tests that fell into three categories according to the 3Qs approach: installation qualification (IQ), operational qualification (OQ), and performance qualification (PQ).
The IQ was performed to ensure that all documents pertaining to the equipment were available and to check that the equipment was properly installed and configured. The OQ was done to verify that software was functioning according to its operational specifications in the selected environment. Specifically, during the OQ of eSlide manager and ImageScope, specific tests scripts were prepared and executed, addressing the key potential issues such as logging, allocation of slides into studies and identification of the slides, measurements of lengths using a gauge/calibration standard, measurements of surfaces, Ki-67-positive nuclei counting, data/results processing, analysis report processing, backup/restore, and archiving. Stress case scenarios were also included.
The PQ was finally done to verify that software consistently performed to the specification for its day-to-day use. The presence of accurate quality documentation and training processes was also checked at this step. The validation report included the deviation reports (description, cause, corrective actions) as well as a conclusion and the formal authorization from the test facility management to use the system in GLP studies.
The pathologist in this validation procedure selected the most homologous and representative sections, and low-magnification images were included in the report. This is the reason why the decision was made to use specific forms documenting the selection of those sections. The images including the measurements were printed, signed, and retained in the study raw data. The drawback of this strategy was to maintain two types of data, that is, electronic and paper data.
Discussion
Digitalization brings value to toxicologic pathology and the innovative technologies that are currently under development have the potential to improve pathophysiologic understanding, safety assessment, and efficiency. Digital toxicologic pathology shows promise in the long run to enable toxicologic pathologists to save time on labor-intensive and repetitive tasks, allowing them to have more daily time to qualify the changes, deepen interpretations, and propose relevant investigative studies. Indeed, the images become shareable records; they are searchable and may allow interstudy or interproject correlations when properly curated and linked to adequate metadata. Moreover, digitization is a mandatory step for image analysis, bringing quantification to the pathologist’s report and opening the road toward the use of AI-based analysis.
The examples presented in this article demonstrated that digital pathology tools may be used in a regulatory framework but require workflow integration and major validation efforts. The risk of divergent quality expectations exists, however, as guidance adapted to digital pathology is lacking. International efforts have already been made in the field of human pathology such as the four-year international project Academia and Industry Collaboration for Digital Pathology that focused on developing efficient and innovative products for digital pathology (http://aidpath.eu/). In addition, validation approaches have been proposed (Pantanowitz et al. 2013; Parwani et al. 2014) and the recent FDA approval of Philips IntelliSite Pathology Solution (PIPS) shows that progress is being made particularly in the field of human surgical pathology (press release April 12, 2017: https://www.fda.gov/NewsEvents/Newsroom/PressAnnouncements/ucm552742.htm). Approval of this digital pathology solution was based on the results of a clinical study of approximately 2,000 surgical pathology cases showing that the interpretation made based on the PIPS images was comparable to those made using glass slides.
However, with regard to safety assessments conducted by the pharmaceutical, chemical, and agrochemical industries, a lack of a clear framework of acceptance by regulatory authorities has resulted in a reluctance to engage further in this type of work.
All roundtable participants agreed that emerging research technologies are bringing innovation to the understanding of mechanisms of action and that high-content imaging modalities are inseparable from computerized systems.
Nevertheless, it was recognized that within the field of toxicologic pathology, economic and resource constraints have the potential to impede the use of a fully digital workflow and the acceptance of digital pathology tools could also face resistance from pathologists who may prefer using a microscope to examining tissues on a screen. As a result, some colleagues have been reluctant to promote the use of automatized diagnosis whereas others have deplored the extensive validation effort that would be necessary to cover all aspects of digital pathology. Based on the roundtable discussion, it was apparent that many toxicologic and experimental pathologists preferred to physically evaluate glass slides with a microscope and provide scientifically sound interpretations rather than spending time testing and validating new systems, particularly since these technologies are continuously and rapidly evolving.
Other limitations are related to the available equipment and software. Most of them were initially developed and implemented in non-Good pratices (GxP) context, therefore compliance to 21 CFR Part 11 and a true integration of all the systems together was and remains a challenge. Although some vendors are now proposing modules specifically designed to address issues related to GLP-requirements, software providers may not understand all regulatory constraints in the field of toxicologic pathology.
From a practical point of view, although scanning and screen quality have markedly improved, using digital slides for toxicologic pathology interpretation instead of glass slides remains time-consuming. Useful tools are integrated in most available viewers, such as the ability for side-by-side comparisons, annotations, light boxes to visualize several thumbnails on the same screen, metadata search capabilities, and more. Some image management systems provide figure makers that allow the automatic preparation of image-based documents. Several pathologists during the roundtable at the 2018 French SFPT meeting pointed out the need for innovative interfaces such as touch screens and/or joysticks resembling the microscope handling experience to improve the screening speed. Using a mouse and simple screen panning options are currently inadequate to allow a pathologist to perform the efficient assessment of hundreds of slides or tissues per day, as can be done with a microscope. This issue must be solved by suppliers with guidance from pathologists in defining the specific requirements needed to achieve an optimal system for the needs in toxicologic pathology. A variety of input devices for digital slide navigation were reviewed by Molin, Lundström, and Fjeld (2015), and currently many of the roundtable participants are gaining further experience with different devices.
Remote peer review of digital slides for exploratory and GLP studies appears to be more and more common in pharmaceutical, chemical, and agrochemical companies, as well as in contract research organizations. The definition of the regulatory status of images in pathology and its consequences have been discussed ten years ago in a “best practices” article from Tuomari et al. (2007) where the authors made a clear distinction between images used for illustrative purpose or used for diagnosis. The definition of the regulatory status of digital slides remains nevertheless a key point and the practical conditions and validation scheme to be applied should be better defined in order to ensure harmonization and regulatory acceptance. If there is now a clear consensus on the acceptance of diagnoses made from a digital slide in the OECD region, there is also a clear regulatory requirement to archive the original glass slide. Indeed, the glass slides, as per OECD document no. 16, are considered as study specimens derived from the test system. Therefore, the glass slides and the corresponding blocks are needed for the reconstruction of the histopathology phase of the study and consequently must be archived in addition to the digital slides regardless of the validation and qualification efforts made to obtain a digital “true copy” of the tangible specimen. A “true copy” or “certified copy” is “a copy (irrespective of the type of media used) of the original record that has been verified (i.e., by a dated signature or by generation through a validated process) to have the same information, including data that describe the context, content, and structure, as the original.” This definition has been given in the Integrated Addendum to ICH E6(R2) on Good Clinical Practice (2016), but no guideline to date has considered the practical consequences of this definition for digital slides.
AI tools are even more complex and controversial. Algorithm validation should include a comparison of the results obtained using the automated diagnosing system with a standard method. However, this will not be as easy as comparing the checksum of two files. Designing an unbiased scenario to perform the PQ of such systems will be challenging and would need close collaboration between pathologists and software providers but also, at some point, endorsement by regulatory authorities. Moreover, the performance of machine learning processes is, per definition, not frozen in time. The question may be raised about the nature and type of documentation to record the “training” of machines in addition to the initial validation of algorithms and processes.
Overall, digital pathology systems are subject to the same requirements as any other GLP-computerized systems including fully validated systems and methods, ALCOA+ characteristics of data, adequately trained staff, appropriate procedures, and qualified hardware and infrastructure. Digital pathology tools require a higher level of specific qualification and validation due to the complexity of equipment and software, number of components, and algorithm management. Several papers have begun this process (Tuomari et al. 2007; Long et al. 2013; Zarella et al. 2018), but, to move forward and engage in a true digital evolution of toxicologic pathology, it is time now to have an international “best practices” paper on digital pathology validation giving a practical toolbox for equipment requirements and software validation. This position paper should give accurate definitions and provide a comprehensive summary of the current issues and risks with recommendations on how to validate digital technologies within the field of toxicologic pathology. The profession needs to embrace these new technologies and include them in existing processes.
Footnotes
Acknowledgments
The authors thank the board of the French Society of Toxicologic Pathology (SFPT), Erio Barale-Thomas (Janssen R&D, Beerse, Belgium), and Jean-François Gallas (Sanofi, Montpellier, France) for their review. The authors would also like to thank the participants to the 2018 SFPT Annual Meeting (May 24 and 25, 2018, Montpellier, France) for their valuable input during the roundtable.
Author Contributions
Authors contributed to conception or design (BG, FG, GH, DO, AP, VS); data acquisition, analysis, or interpretation (BG, FG, GH, DO, AP, VS); drafting the manuscript (BG); and critically revising the manuscript (FG, GH, DO, AP, VS). All authors gave final approval and agreed to be accountable for all aspects of work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Declaration of Conflicting Interests
The author(s) declared no potential, real, or perceived conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
