Abstract
There is a critical need to develop high-throughput assays to identify compounds that offer therapy for individuals suffering from neurodegenerative diseases. Most brain disorders, including neurodegenerative diseases, share the common neuropathology of mitochondria dysfunction, which can lead to apoptosis of neurons, overproduction of reactive oxygen species (ROS), and other cellular neuropathologies characteristic of these diseases. Human induced pluripotent stem cells (iPSCs) with a stable genomic insertion of the neurogenin-2 transcription factor under the control of the TetOn promoter can be differentiated into excitatory human neurons (i3Neurons) within 3 days of exposure to doxycycline. These neurons have been used to develop and validate a live-cell assay for parameters of mitochondrial dynamics and function using two compounds known to promote mitochondrial elongation in mouse neurons, 4-hydroxychalcone and 2,4-dihyrdroxychalcone. The assay involves plating the neurons in 384-well microtiter plates, treating them with known or unknown substances, and then capturing morphological information for the neuronal mitochondria using a lentivirus vector to express a mitochondrial-targeted fluorescence reporter. The i3Neuron cultures exposed to these two compounds for 24 h exhibit significantly decreased circularity and significantly increased length compared to controls, two morphological parameters correlated with increased mitochondrial health. The assay is rapid, with results obtained after a one-week-long i3Neuron culture or one month if neurons are co-cultured with astrocytes. This live-cell, mitochondrial phenotypic assay can be used for high-throughput screening or as an orthogonal assay for compounds obtained via other high-throughput screening campaigns.
Introduction
The progress in bringing potential therapeutics for neurodegenerative diseases from the research bench to the clinic has been extraordinarily slow. In fact, only 8% of drugs targeted to treat CNS diseases become approved after clinical trials. 1 Compared to drug discovery for other metabolic or autoimmune diseases, these diseases present difficult target identification and validation, animal models with poor face validity, and a lack of biomarkers.1–4 Phenotypic high-throughput compound screens, however, offer promise by providing an unbiased method for uncovering novel targets for neurodegenerative diseases. 5
Mitochondrial dysfunction is a hallmark of most neurodegenerative diseases, including Alzheimer’s disease (AD), Parkinson’s disease (PD), Huntington’s disease (HD), and amyotrophic lateral sclerosis (ALS).6,7 Mitochondria perform numerous biochemical functions in cells beyond the typically recognized functions of adenosine triphosphate (ATP) generation and calcium buffering, including the regulation of apoptosis, generation of reactive oxygen species (ROS), lipid and pyrimidine biosynthesis, and others.8–11 When mitochondria become compromised in neurons due to aging or other factors, the precise regulation of these functions leads to the pathologies observed in many neurodegenerative diseases. 7 With the advent of induced pluripotent stem cell (iPSC) technology, 12 we can now use human-derived neurons in high-throughput screens to search for compounds that improve mitochondrial health and function.
iPSC-derived human neurons are now being used effectively in high-throughput screening platforms. Two recent examples include the search for compounds that reactivate the silenced Fmr I gene in Fragile X and for compounds that promote neurite outgrowth and branching for potential nerve regeneration.4,13,14 iPSCs exposed to the neurogenin-2 (Ngn2) transcription factor via lentivirus transduction differentiate into glutamatergic neurons. 15 Fernandopulle et al. 16 improved on this system by stably integrating the Ngn2 gene under TetOn promoter 17 regulation in a human iPSC line, so that virtually all cells treated with doxycycline differentiate into excitatory human neurons (i3Neurons). The i3Neurons express classic cortical neuron markers such as MAP2, TAU, and Tuj1 (beta-III tubulin) after differentiation. 16 When the i3Neurons are co-cultured with astrocytes, they exhibit spontaneous excitatory currents that can be inhibited by glutamate receptor antagonists. 16
Here, we have combined the advantages offered by i3Neurons with a novel and effective high-content assay we developed to extract morphological features of mitochondria in mouse neurons as proxies for parameters of mitochondrial health and function. 18 The structure and function of mitochondria are highly related. For instance, when mitochondria become defective, they assume a small and rounded morphology that can be measured by a circularity index. Moreover, 90% (45/50) of the compounds identified to promote mitochondrial length in mouse neurons in this high-content assay caused a corresponding increase in ATP generation. 18 i3Neurons are generated rapidly and in large batches that are ideal for reducing variability in high-throughput assays and screens. In addition, virtually all of the iPSCs differentiate into excitatory neurons, which reduces variability from the multiple cell types found in primary neuron cultures established from brain tissue. The assay generates results in a week’s time from the plating of cryopreserved i3Neurons to imaging, allowing for the rapid accumulation of data. The assay as described for measuring mitochondrial features in neurons can be adapted to any cellular target for which a fluorescent reporter can be fashioned.
Materials and Methods
iPSC Maintenance, i3Neuron Differentiation, and Cryopreservation
All reagents were obtained from Thermo Fisher Scientific (Waltham, MA) unless otherwise noted. iPSCs and i3Neurons were maintained and differentiated according to protocols outlined in Fernandopulle et al. 16 In short, iPSCs were thawed and plated in Essential 8 media with 1× RevitaCell Supplement in a six-well dish coated with 0.5 µg/cm2 vitronectin, and then incubated at 37 °C with 5% CO2. Media were changed daily, using only Essential 8 media after the first day. iPSCs were clump-passaged twice with 0.5 mM ethylenediaminetetraacetic acid (EDTA) in 1× Dulbecco’s phosphate-buffered saline (DPBS) before being plated in a 15 cm2 dish coated with a 1:100 dilution of Growth Factor Reduced Matrigel (Corning, Corning, NY) in Dulbecco’s Modified Eagle Medium: Nutrient Mixture F12 (DMEM/F12). When the 15 cm2 dish was 70% confluent, the iPSCs were single-cell-passaged with 7 mL of Accutase. Ten million iPSCs were plated per 15 cm2 dish coated with Matrigel in induction media: 1× N2 supplement, 1× non-essential amino acid (NEAA), and 1× L-glutamine with a DMEM/F12 and HEPES base supplemented with 1× RevitaCell Supplement and 2 µg/ml doxycycline (Millipore Sigma, Burlington, MA). We consider this as cell day 0 for all experimental timelines described below. On days 1 and 2, 100% media changes were performed, with induction media supplemented with only doxycycline (minus 1× RevitaCell Supplement). On day 3, i3Neurons were cryopreserved in CTS PSC Cryomedium and stored at −80 °C. In all succeeding experiments with i3Neurons, each plate is derived from an individual cryovial of previously differentiated cells and can therefore be considered to be independent of other plates used.
i3Neuron Lentiviral Multiplicity-of-Infection (MOI) Titration Assay
i3Neurons, now considered to be at cell day 4, were revived and plated in cortical neuron media: 1× B27 supplement, 10 ng/mL brain-derived neurotrophic factor (BDNF; PeproTech, Rocky Hill, NJ), 10 ng/mL NT-3 (PeproTech), and 1 µg/mL laminin with a BrainPhys Neuronal Medium base (StemCell Technologies, Vancouver, Canada) in 384-well plates coated with poly-L-ornithine (PLO) and laminin (Aurora Microplates, Whitefish, MT). i3Neurons were plated at either 7500 or 15,000 cells per well in 40 µL of cortical neuron media. CAG>mtTagGFP2 lentivirus (packaged by Vigene Biosciences, Rockville, MD) was mixed with the cell suspension prior to plating at the following MOIs: 1, 3, 5, 7, and 10. On day 5, 20 µL of media with lentivirus was removed, and 60 µL of fresh cortical neuron media was added. On cell day 7, the i3Neuron 384-well plates were imaged on the IN Cell Analyzer 6000 confocal microscope (GE Healthcare Life Sciences, Marlborough, MA), capturing the TagGFP2 signal in three slices ΔΖ = 0.7 µm; aperture = 1.0 AU; optimum excitation (λexc) = 488 nm; and optimum emission (λem) = 515 to 535 nm, using a 60× objective as described in Varkuti et al. 18 Four field images were collected per well. The imaging data were analyzed as described below to determine the optimal cell density and MOI to use for further experiments for each batch of i3Neurons prepared.
I3Neuron Immunocytochemical (ICC) Studies
iPSCs were dissociated into single cells and plated in 96-well plates at a cell density of 24,000 cells per well. On the second day after plating, cells were fixed for 15 min with a final concentration of 4% paraformaldehyde in each well, permeabilized with 0.5% Triton X-100 in 1× DPBS for 15 min, and blocked with 3% bovine serum albumin (BSA) in 1× DPBS for 1 h at room temperature. Wells were washed twice with 1× DPBS between all steps of the protocol. The following primary antibodies were added at a 1:500 dilution in blocking buffer (3% BSA in 1× DPBS) overnight at 4 °C: polyclonal rabbit Oct4 (cat. no. ab18976, Abcam, Cambridge, MA), monoclonal mouse Sox2 (cat. no. ab79351, Abcam), polyclonal rabbit MAP2 (cat. no. ab32454, Abcam), and monoclonal mouse Tuj1 (cat. no. ab78078, Abcam). The following secondary antibodies were added at a 1:1000 dilution in blocking buffer to the appropriate wells and incubated for 1 h in the dark at room temperature: Alexa Fluor 568 goat anti-rabbit (cat. no. A11011, Thermo Fisher) and Alexa Fluor 488 goat anti-mouse (cat. no. A11029, Thermo Fisher). Wells were washed twice with 1× DPBS, and in the final wash, NucBlue Live Stain ReadyProbes reagent was added at a concentration of 2 drops per 1 mL of 1× DPBS. Wells were imaged using a 20× objective in 2D after a 5 min incubation using the IN Cell Analyzer 6000 confocal microscope. i3Neuron ICC studies were performed similarly with exceptions described below. iPSCs were dissociated into single cells and plated on 384-well plates coated with Matrigel at a cell density of 3000 cells per well. The cells were then exposed to doxycycline in induction media to induce differentiation for 3 days, and ICC studies were commenced at this time. Day 3 i3Neuron ICC images were taken with a 60× objective. I3Neuron day 7 and day 10 images were obtained by reviving and plating previously cryopreserved cells at a density of 7500 cells per well on 384-well PLO and laminin dual-coated plates in cortical neuron media. Once plated, i3Neurons were aged until the appropriate time point, and the ICC procedure outlined above was commenced. i3Neuron day 28 images were obtained by co-culturing human astrocytes (iCell Astrocytes, Cellular Dynamics, Madison, WI) on PLO and laminin dual-coated plates in cortical neuron media. Human astrocytes were plated at a cell density of 5000 cells per well 1 day prior to i3Neuron plating (cell density = 10,000 cells per well). Cells were then aged to the appropriate time point, and ICC was commenced with images taken using a 60× objective.
i3Neuron High-Throughput Mitochondrial Assay
The i3Neurons were revived and immediately plated with CAG>mtTagGFP2 lentivirus at a MOI of 5 in 384-well plates coated with PLO and laminin at a cell density of 15,000 i3Neurons in 40 µL of cortical neuron media/well. On cell day 5, 20 µL of media with lentivirus was removed per well, and 60 µL of fresh cortical neuron media was added. On cell day 6, 20 µL of media was removed per well, and compounds solubilized in DMSO were added at the appropriate concentrations so that DMSO concentration remained constant in all wells of the plate at 0.125%. On cell day 7 (24 h after exposure to the compounds), i3Neuron plates were imaged as described above. The i3Neuron plates were also imaged on cell day 8 to obtain 48 h exposure data. For experiments using 4-hydroxychalcone (Indofine Chemical Co., Hillsborough, NJ) and 2,4-dihydroxychalcone (Millipore Sigma), two independent assay plates derived from separate cryovials of i3Neurons were plated on the same day and measured in parallel ( Fig. 3 and Suppl. Fig. 3 ).
Astrocyte Extended Culture Assay
i3Neurons to be used in extended culture assays were prepared as described above, except that CAG>mtTagGFP2 lentivirus was added to the differentiating i3Neurons on day 2 of differentiation rather than cell day 4 after being revived from cryopreservation. 700,000 single-cell-passaged iPSCs were added to each well of a Matrigel-coated six-well dish on day 0. On day 2 (during doxycycline induction), the differentiating neurons in one well were dissociated to determine the cell count to be used for MOI calculation. CAG>mtTagGFP2 lentivirus was added at a MOI of 5 to the media of each i3Neuron well to be transduced. On day 3, the transduced i3Neurons were dissociated as single cells and cryopreserved. One day prior to plating i3Neurons, human astrocytes (Cellular Dynamics) were plated at a cell density of 5000 astrocytes per well in 40 µL of cortical neuron media/well. After 24 h, 10,000 i3Neurons per well were then plated with the human astrocytes in an additional 40 µL of cortical neuron media/well (bringing the total volume to 80 µL of cortical neuron media/well). Three identical plates of i3Neurons and human astrocytes were plated and maintained in culture. 75% media changes were performed with cortical neuron media one time each week as the experiments progressed. 4-hydroxychalcone and 2,4-dihydroxychalcone were added to the plates as described above at cell days 13, 20, and 27, and imaged 24 h later at cell days 14, 21, and 28.
Data Analysis
Each 60× image stack was merged into a projection image using a custom macro supplied with the GE Developer Toolbox software (version 1.9.2, build 2415, GE Healthcare Life Sciences). These final images were segmented into individual mitochondria, as described in Varkuti et al., 18 masking the somatic mitochondria to yield images containing only axonal and dendritic mitochondria, which were analyzed together. Supplemental Figure 1 contains images that illustrate how the projection images were segmented. The following data were generated for each image collected: cell body count, mitochondrial count, the sum of the total mitochondria area [cumulative area (CA)], the median circularity of the mitochondria per field, and the median length of the mitochondria per field. Since four images were taken per well, the parameter values for each image within a well were averaged together and reported as a well value. These well values were then analyzed along with the corresponding values from other wells. If an experiment consisted of multiple plates assayed in parallel, robust z-scores were calculated from the well data using the DMSO well data on the same assay plate.
Results
Human iPSC Differentiation into Homogeneous Excitatory Neurons
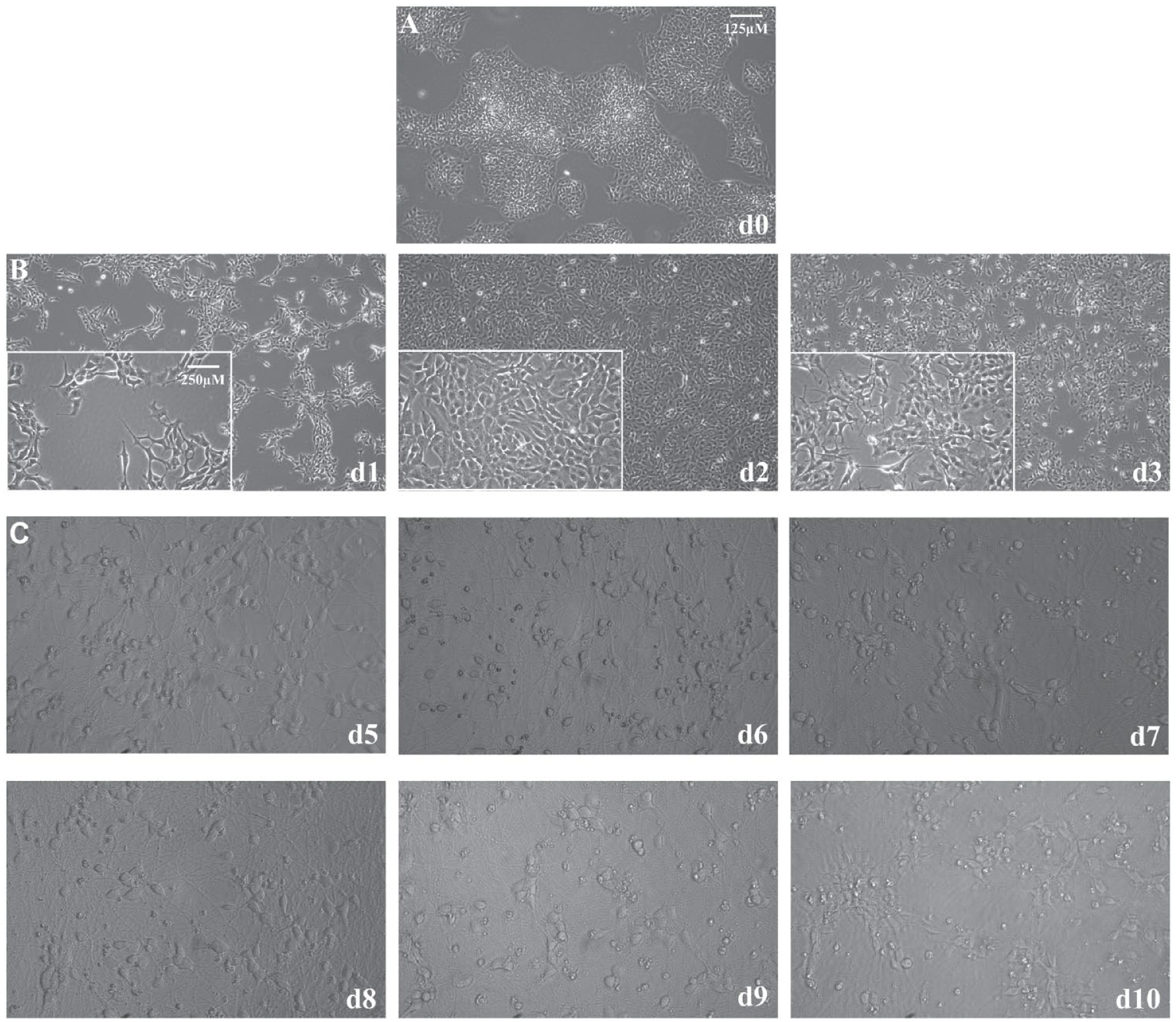
iPSCs with a stably integrated Ngn2 gene in the genome under the regulation of a TetOn promoter can produce large quantities of high-quality human excitatory neurons on the addition of doxycycline to induce expression of the transcription factor. 16 After only 3 days of Ngn2 expression, the human iPSCs visibly shift from compact colonies of cells with large nuclei and little cytoplasm ( Fig. 1A ) to young excitatory neurons with distinct somas and neuritic extensions ( Fig. 1B ). At this time point, the i3Neurons exhibit expression of the neuronal markers MAP2 and Tuj1 ( Suppl. Fig. 2 ). At day 3 of differentiation, i3Neurons must be re-plated in the desired assay format or cryopreserved because the neurites become so extensive that passaging them produces damage. The i3Neurons survive on PLO laminin-coated plates for approximately 2 weeks ( Fig. 1C ); however, after cell day 10, the cells begin to clump, and their overall health begins to deteriorate.
(
Development of a High-Content, Mitochondrial Screening Platform for Human Neurons
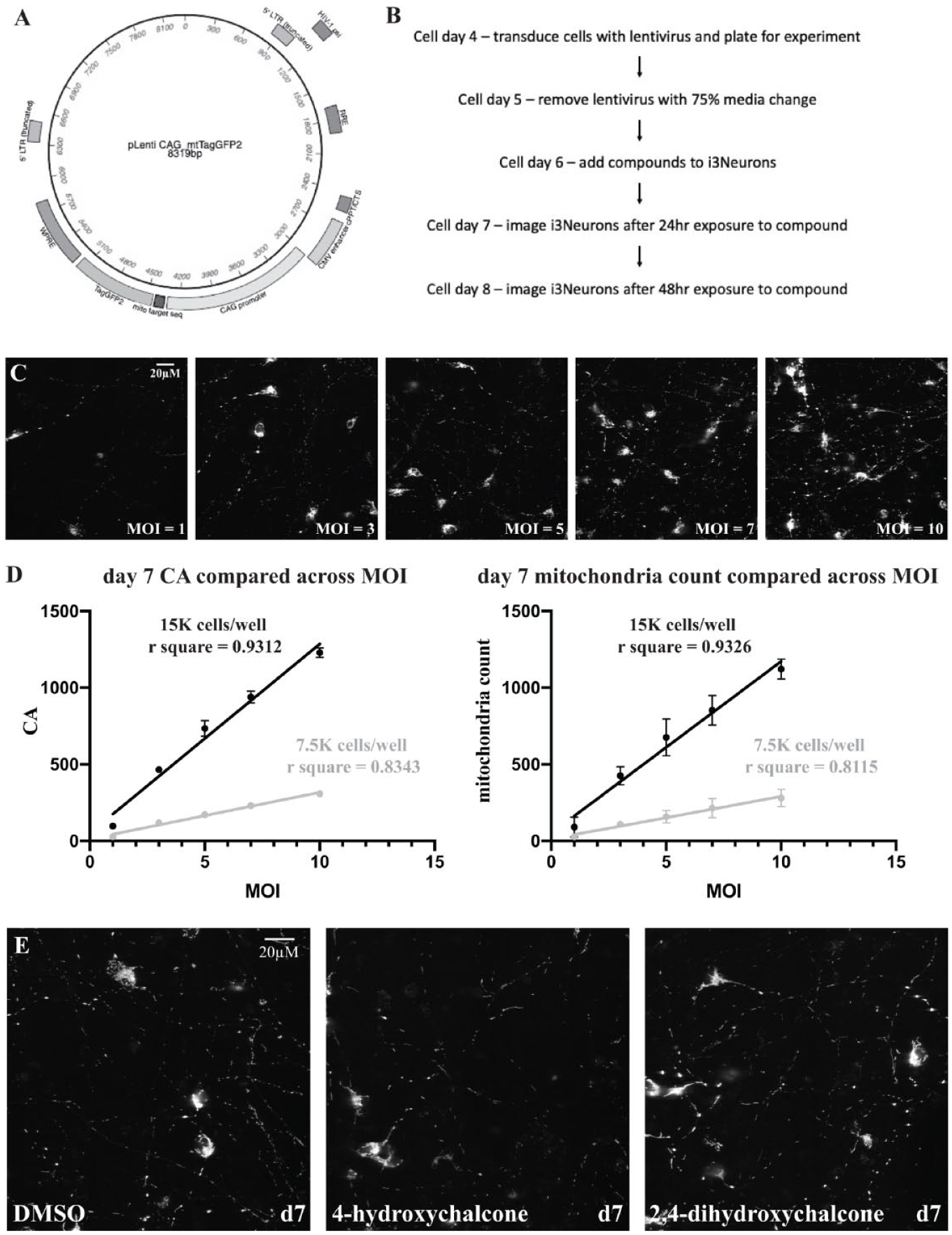
Varkuti et al. describe a high-content assay and screen to search for compounds that increase mitochondrial content, health, and/or function using mouse cortical neurons. 18 This assay used a mitochondrial-targeted version of the TagGFP2 reporter engineered into the Rosa26 locus behind a conditional promoter to sparsely label mitochondria in the cultured neurons. Expression of the reporter is dependent on treating the cultures with adeno-associated virus (AAV) expressing Cre recombinase. A similar scheme was used to develop an assay for human neurons differentiated from iPSCs. First, a lentivirus ( Fig. 2A ) was created to express mitochondrial-targeted TagGFP2 in the human neurons under the control of the strong, ubiquitous CAG promoter that contains the cytomegalovirus (CMV) early enhancer element, a minimal promoter, the first exon and intron of the chicken beta-actin gene, and the splice acceptor of the rabbit beta-globin gene.19,20 iPSCs and iPSC-derived cells have been found to silence gene expression from certain promoters via methylation and histone modification.21,22 Therefore, it was critical to use a robust promoter that was not susceptible to these silencing processes.
(
We next turned our attention to finding the optimum cell density per well and MOI that would offer the sparse but robust signals from mitochondria required for accurate segmentation. We found from experience that images obtained from random fields containing too few objects tend to increase the variability of the assay. Too many objects within a field, in contrast, lead to overlapping objects and inaccuracy in the assay. We analyzed a single plate with either 7500 or 15,000 i3Neurons per well with varying MOIs of CAG>mtTagGFP2 lentivirus (1, 3, 5, 7, and 10 on cell day 4), and we imaged the neurons at cell day 7 and cell day 8.
i3Neuron CAG>mtTagGFP2 Lentivirus MOI Titration: Day 7 Results for Cell Density of 15,000 Cells per Well.
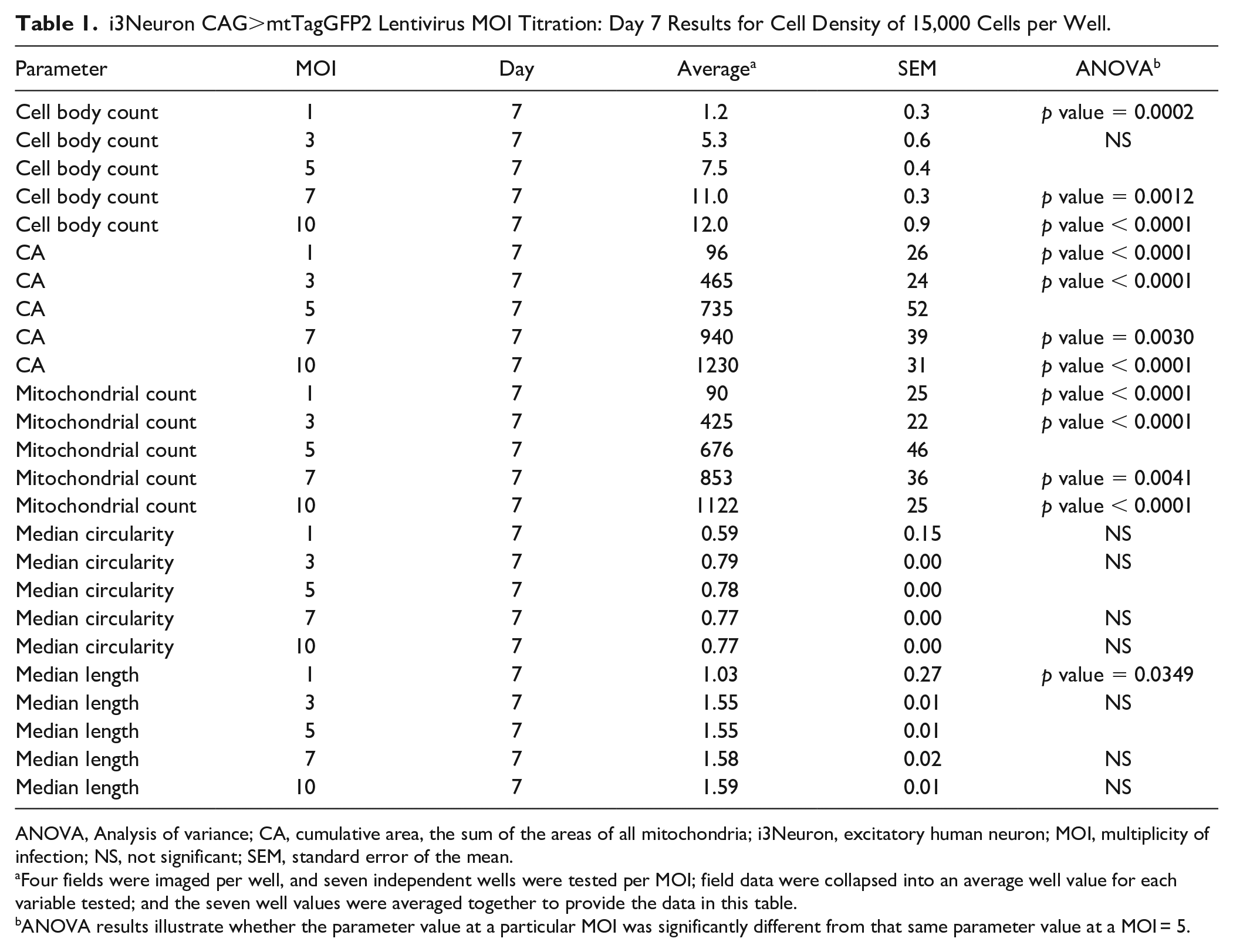
ANOVA, Analysis of variance; CA, cumulative area, the sum of the areas of all mitochondria; i3Neuron, excitatory human neuron; MOI, multiplicity of infection; NS, not significant; SEM, standard error of the mean.
Four fields were imaged per well, and seven independent wells were tested per MOI; field data were collapsed into an average well value for each variable tested; and the seven well values were averaged together to provide the data in this table.
ANOVA results illustrate whether the parameter value at a particular MOI was significantly different from that same parameter value at a MOI = 5.
i3Neuron CAG>mtTagGFP2 Lentivirus MOI Titration: Day 8 Results for Cell Density of 15,000 Cells per Well.
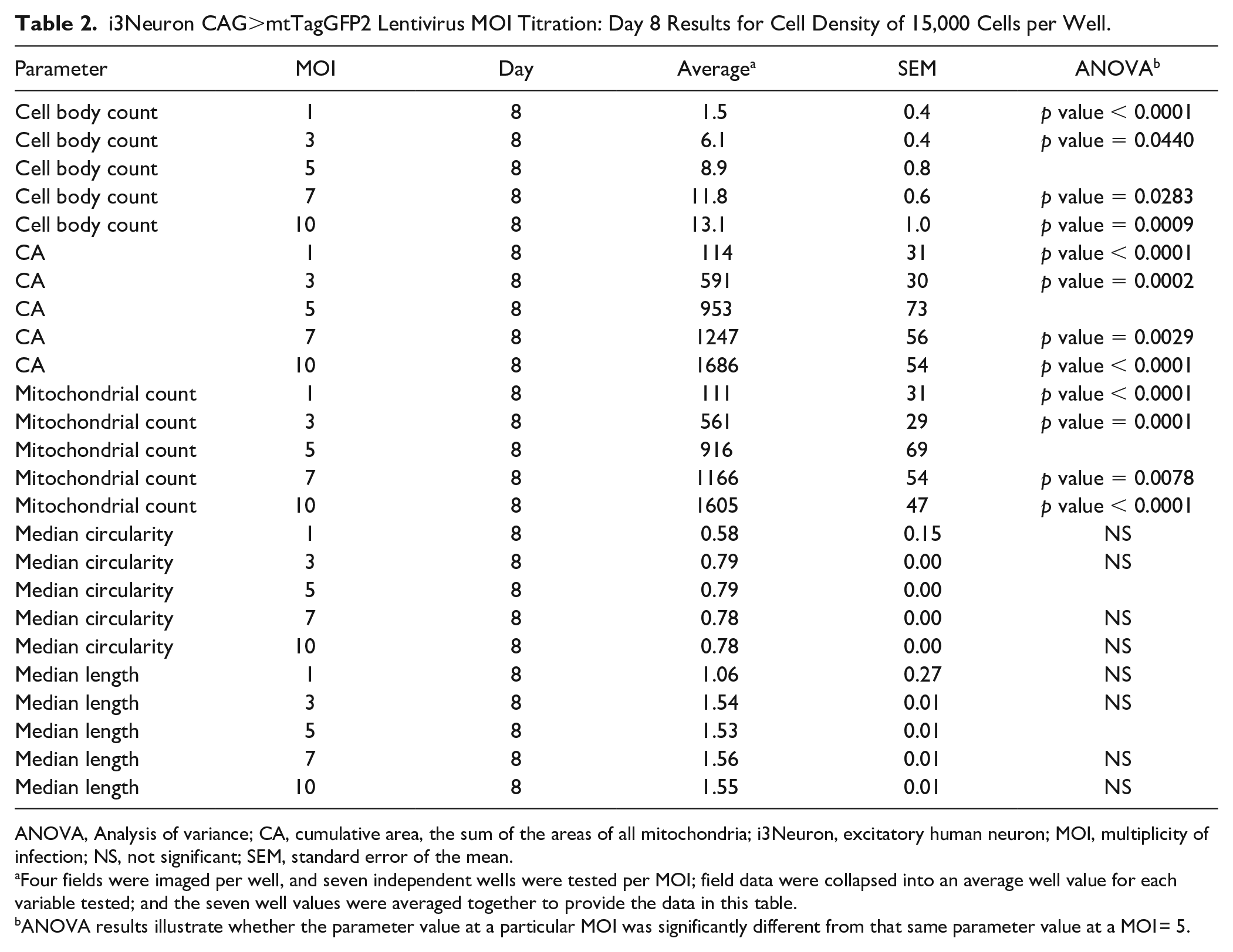
ANOVA, Analysis of variance; CA, cumulative area, the sum of the areas of all mitochondria; i3Neuron, excitatory human neuron; MOI, multiplicity of infection; NS, not significant; SEM, standard error of the mean.
Four fields were imaged per well, and seven independent wells were tested per MOI; field data were collapsed into an average well value for each variable tested; and the seven well values were averaged together to provide the data in this table.
ANOVA results illustrate whether the parameter value at a particular MOI was significantly different from that same parameter value at a MOI = 5.
As anticipated, increasing the MOIs led to significantly increased cell body count, CA, and mitochondrial count compared to a MOI = 5 for the data obtained using 15,000 cells per well (
Fig. 2C
, and
i3Neurons Reproduce Mitochondrial Phenotypes Observed in Mouse Cortical Neurons
Our prior study using mouse neurons identified multiple compounds named MnMs (modulators of neuronal mitostasis) that influence mitochondrial content, elongation, and circularity. We chose to use two of the compounds, 4-hydroxychalcone and 2,4-dihydroxychalcone, to develop the high-content assay using i3Neurons. These compounds were found to reduce median circularity and dramatically increase median mitochondria length. 18 The i3Neurons were plated on cell day 4, followed by compound addition on cell day 6 with imaging on cell day 7 (24 h after compound exposure) and cell day 8 (48 h after compound exposure) ( Fig. 2B ). Each compound was solubilized in DMSO and assayed over a concentration range of 1 to 10 µM, with each concentration being assayed in four wells. DMSO alone was also placed in 20 wells of the plate as a control at a final concentration of 0.125%. Two independent plates were assayed in parallel, and robust z-scores were calculated using the DMSO wells on each plate as the control population for other wells on that plate. Figure 2E shows representative images for the DMSO control and both chalcone compounds at a concentration of 3 µM, illustrating that these compounds dramatically lengthen mitochondria to the point of being visually apparent. At this concentration, there was no significant difference in mitochondrial content as calculated by CA (data not shown) between the chalcones and DMSO.
Dose–response curves were constructed for each compound for the median circularity and median length parameters ( Fig. 3 ). The EC50 values after 24 h of exposure to 4-hydroxychalcone were 1.9 µM for both mitochondrial median circularity and median length. These values remained fairly constant after 48 h of exposure to the compound, increasing slightly to 2.6 µM and 2.4 µM, respectively. The EC50 values after 24 h of exposure to 2,4-dihydroxychalcone were similar to those for 4-hydroxchalcone at 2.0 µM for median circularity and 2.1 µM for median length, increasing slightly with 48 h of exposure to 3.1 µM and 3.2 µM, respectively. The experiment illustrated in Figure 3 was repeated ( Suppl. Fig. 3 ) exactly as stated in the Materials and Methods section, except that a different batch of i3Neurons was used. The response to the compounds remains consistent even when using different batches of i3Neurons.
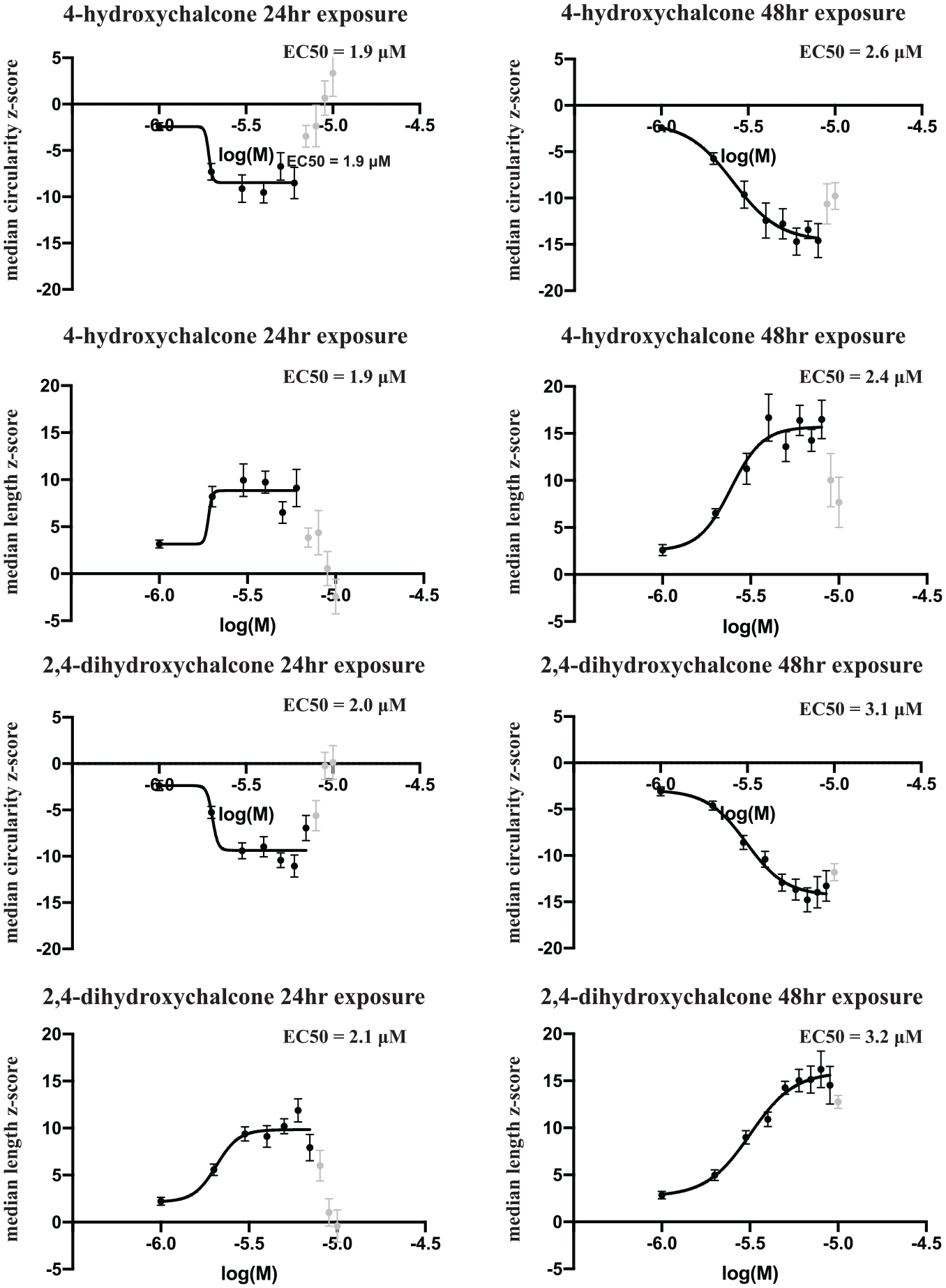
Dose–response curves for mitochondrial median circularity and length using 4-hydroxychalcone and 2,4-dihydroxychalcone. Each compound was assayed over a concentration range from 1 to 10 µM on two duplicate plates. Robust z-scores were calculated for mitochondrial median circularity and length values of the neuritic mitochondria for each well, and were plotted against the log10 of the molar concentrations of the compounds. Each point on the plot represents the robust z-score average ± SEM (standard error of the mean) of eight independent wells between the two independent plates. High concentrations of each compound caused toxicity in the wells, as illustrated by increased and/or decreased z-scores (light gray points) where appropriate. These values were excluded for calculating the EC50 values.
i3Neurons Respond to Compounds in Similar Ways under Extended Culture Conditions
The i3Neurons can be maintained in a healthy state in culture for more than 1 month by co-culturing the neurons with human astrocytes.
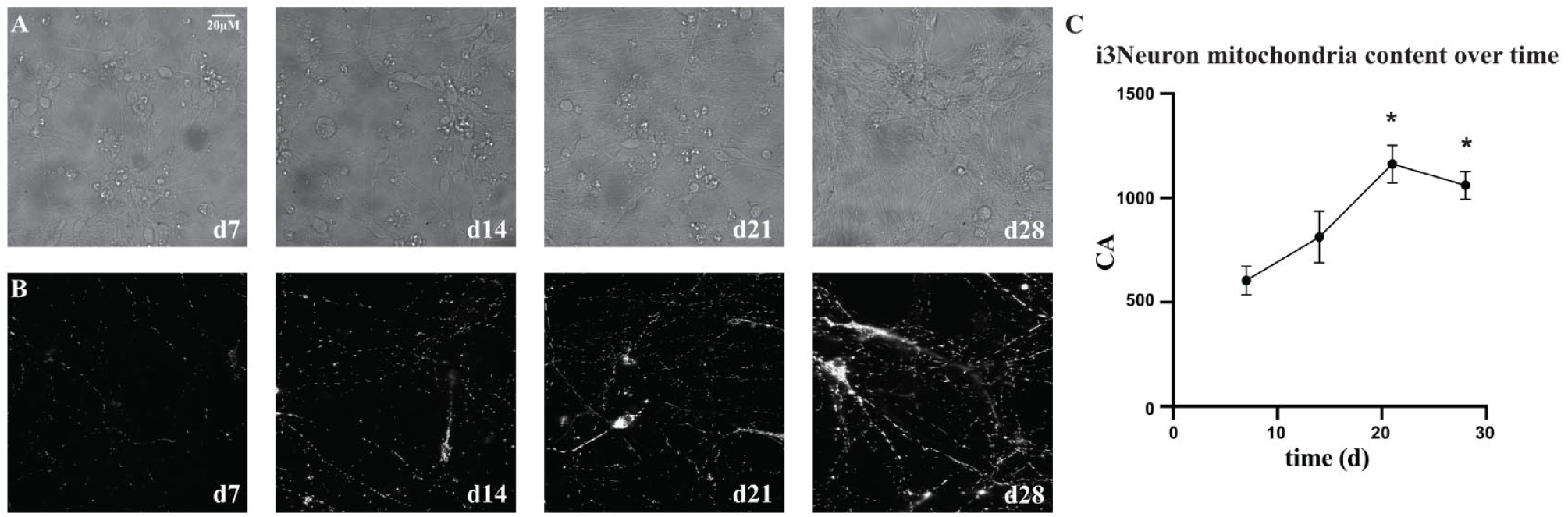
Excitatory human neuron (i3Neuron) health and viability extended with astrocyte co-culture. This figure contains cell-based screening data showing the accumulation of mitochondria [cumulative area (CA)] throughout days in culture. All of the data were obtained from four separate plates, each plated on the same day from four separate cryovials but imaged on different days. (
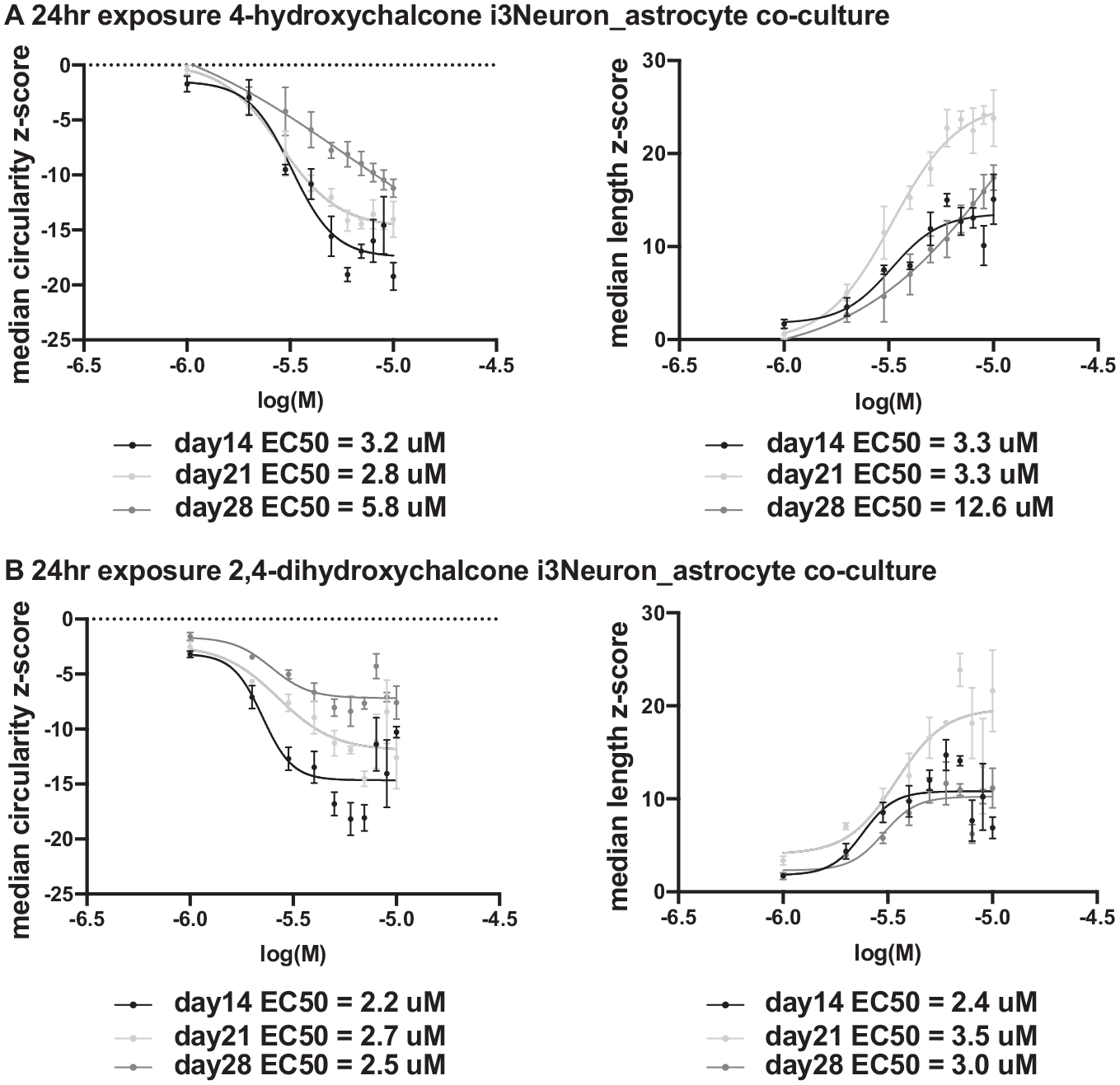
Dose–response curves for excitatory human neurons (i3Neurons) co-cultured with human astrocytes throughout a 3-week culture period. This figure contains cell-based screening data for three sets of dose–response data to two compounds, 4-hydroxychalcone and 2,4-dihydroxychalcone. All of the data were obtained from three separate plates, each plated on the same day from three separate cryovials but imaged on different days. (
Discussion
Mitochondrial dysfunction is a hallmark of aging for many different diseases, including neurodegenerative diseases and other neuropsychiatric disorders. For some genetic forms of PD, ALS, and Charcot–Marie–Tooth disease, an insult to mitochondria is the precipitating factor for the disease.23–25 For others, including AD, mitochondrial dysfunction remains as a very prominent aspect of the neuropathology. A hypothesis of increasing influence, however—the mitochondrial cascade hypothesis for sporadic AD—envisions mitochondrial dysfunction as an initiator or very near the top of the hierarchy of system failures involved in disease progression.26,27 The prominence of impairments in mitochondrial dysfunction among these diseases commands the development of new assays that may offer strategies for obtaining the needed therapeutics. To date, very few high-throughput assays have been developed that focus on mitochondrial phenotypic readouts to aid in the search for compound targets that improve mitochondria health and function.18,28–30 The assay presented here has great potential for mitotherapeutic discovery.
This i3Neuron mitochondrial phenotypic assay offers many advantages: (1) It is performed using a homogeneous population of the target cells of interest. (2) It is based in human biology rather than that of a rodent. (3) It relies on live imaging of the mitochondria in culture, such that multiple imaging sessions can be performed throughout time. (4) The cells can be produced quickly and in large quantities. (5) The assay can be accomplished within 1 week. (6) The assay is adaptable to other types of cells in the CNS involved in disease pathology. The lentivirus MOI is batch dependent, however, and must be optimized for each new production of i3Neurons. Careful planning of cell-based screening assays will help ensure that the same assay optimizations can be used throughout the length of the screen.
Most assays used in the past were developed using cell systems that are easy to maintain, such as HEK293 cells. While these cells are derived from a human source, they may not offer the required environment for discovering effective therapeutics targeting neurons, given the unique anatomy and physiology of the neuron. Neuroscience-based drug discovery in the past has relied on rodent model systems. There have been many failed clinical trials for compounds passed through the rodent drug development pipelines, giving pause as to whether they can be considered reliable for mimicking human neuronal environments. 31 Substantial efforts in prior neurodegeneration research have relied on fixed, frozen tissue. Fixation procedures distort cellular morphology, while live-cell imaging circumvents these issues. High-throughput screens based on primary neuron cultures obtained from the mouse can be plagued from a lack of starting material at a critical time, given that cultures are inevitably started from embryonic or early postnatal tissues. The i3Neurons can be produced in large quantities and cryopreserved for all experiments for a screen. Finally, the response to tested compounds using the basal or extended culture conditions appears to remain stable; therefore, one can obtain reliable results after only 1 week of culture. Thus, the assay developed has many important advantages in screening for potential mitotherapeutics compared to previously developed assays using rodent neurons.
We validated the assay primarily using two potent positive compounds, 4-hydroxychalcone and 2,4-dihydroxychalcone, obtained from our prior screens using rodent assays. 18 In addition, several other compounds identified initially from our screens using rodent neurons have exhibited significant responses using this assay that are currently being studied. Not all of the compounds identified from the rodent screens, however, exhibit strong potency using the i3Neuron assay. To date, we have screened 26 compounds found to improve mitochondrial health and/or count using mouse primary neurons. Of these compounds, eight (including 4-hydroxychalcone and 2,4-dihydroxychalcone) have shown evidence of improving mitochondrial health. This observation is notable for three major reasons. First, identifying negative compounds during assay development is an important part of the validation of the assay. Second, the difference in efficacy may reveal authentic differences in the biology of rodent versus human neurons, commanding greater emphasis on the use of human neurons to discover new therapeutics. Finally, the disparity may be due to the simple differences in the two assays. For instance, the rodent assays use a heterogeneous set of neurons isolated from the forebrain, whereas the i3Neuron assay uses homogeneous glutamatergic neurons.
This i3Neuron mitochondrial phenotypic assay by itself can only suggest novel compounds for future preclinical development. Additional assays that help select the most promising compounds will complement these studies. Assays that mimic the insults to mitochondria that neurodegenerative diseases present include increasing oxidative stress, adding glutamate to mimic glutamate toxicity, or treating cultures with toxic peptides. 18 All three of these assay examples are amenable to the i3Neuron assay. In addition, there are options for pursuing studies to reveal mechanisms of action, including RNA-seq and proteomics approaches that may define the pathways in which selected compounds alter human mitochondrial dynamics. In sum, the phenotypic and high-throughput assay using iPSC-derived human neurons to select compounds altering mitochondrial dynamics has the potential to quickly identify key compounds for further research and development.
Supplemental Material
sj-pdf-1-jbx-10.1177_24725552211000671 – Supplemental material for High-Throughput Phenotypic Assay for Compounds That Influence Mitochondrial Health Using iPSC-Derived Human Neurons
Supplemental material, sj-pdf-1-jbx-10.1177_24725552211000671 for High-Throughput Phenotypic Assay for Compounds That Influence Mitochondrial Health Using iPSC-Derived Human Neurons by Courtney MacMullen and Ronald L. Davis in SLAS Discovery
Footnotes
Acknowledgements
We would like to thank Michael E. Ward of the National Institute of Neurological Disorders and Stroke, National Institutes of Health, for providing the Ngn2 iPSC line used to differentiate i3Neurons.
Supplemental material is available online with this article.
Abbreviations
AAV, Adeno-associated virus; AD, Alzheimer’s disease; ALS, amyotrophic lateral sclerosis; BDNF, brain-derived neurotrophic factor; DPBS, Dulbecco’s phosphate-buffered saline; EDTA, ethylenediaminetetraacetic acid; FBS, fetal bovine serum; HD, Huntington’s disease; ICC, immunocytochemistry; iPSC, induced pluripotent stem cell; MOI, multiplicity of infection; NEAA, non-essential amino acid; Ngn2, neurogenin-2; NT3, neurotrophin-3; PD, Parkinson’s disease; PDL, poly-D-lysine; PLO, poly-L-ornithine; Tuj1, beta-III tubulin.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by National Institutes of Health (NIH) grants R01MH109957 and 3R01MH109957-03S1 to RLD.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
