Abstract
High-throughput screening (HTS) often yields a list of compounds that requires prioritization before further work is performed. Prioritization criteria typically include activity, selectivity, physicochemical properties, and other absolute or calculated measurements of compound “value.” One critical method of compound prioritization is often not discussed in published accounts of HTS. We have referred to this oft-overlooked metric as “compound natural history.” These natural histories are observational evaluations of how a compound has been reported in the historical literature or compound databases. The purpose of this work was to develop a useful natural history visualization (NHV) that could form a standard, important part of hit reporting and evaluation. In this case report, we propose an efficient and effective NHV that will assist in the prioritization of active compounds and demonstrate its utility using a retrospective analysis of reported hits. We propose that this method of compound natural history evaluation be adopted in HTS triage and become an integral component of published reports of HTS outcomes.
Introduction
The goal of HTS (high-throughput screening) is to find compounds that can be optimized into efficient and selective modulators of a chosen target or phenotype. The major challenge to achieving this goal is the winnowing down of thousands to millions of compounds to the best 10–100 compounds via a carefully designed screening tree. This process is complicated by the fact that a large number of compounds poured into the screening funnel can interfere with the technology of the screening methods, the reagents, proteins present in the assay, or all of the above. This interference can confound the effective selection of good compounds from the majority of compounds that will be ineffective.
In this article, we will be referring to the properties and analysis of “hit” compounds. We define a hit compound as a compound identified from a primary HTS, whether virtual, biochemical, or phenotypic, which shows a favorable signal in comparison to the bulk of the screening signal. Best practices for this determination vary depending on the assay, but excellent discussion can be found in the National Institutes of Health’s (NIH) Assay Guidance Manual. 1 A hit compound is not a lead compound for further optimization, because it has not yet been validated as active at the target or phenotype under investigation. A rigorous screening tree (vide infra) is necessary to validate beyond hit status to further development.
Many biochemical and biophysical methods can assist in the hit prioritization and selection process. These should be standard operating procedures. Perhaps, however, the most straightforward method that can influence the prioritization of screening hits is neither biophysical nor biochemical, but data-driven. That is, an oft-overlooked tool in hit characterization and prioritization is the straightforward determination of hit “natural histories.” These natural histories are readily available using cheminformatic evaluation of the hits and searching of the literature for relevant references. Natural histories are the metadata that have been collected for compounds throughout the years in large, public compilations of screening data (e.g., PubChem) and in medicinal chemistry knowledge distilled into simple characterization models (e.g., Lipinski’s rule). 2 We propose herein a convenient natural history visualization (NHV) of compounds that will prove useful for hit evaluation and prioritization. This visualization conveys information related to the purity, promiscuity, quality, and literature reports of a compound. We recommend that researchers present NHVs of all hit compounds published in HTS reports in which no follow-up medicinal chemistry compound synthesis has been performed to validate the hits. The ready availability of the NHVs will enable editors and readers to immediately judge the potential of the hits that have been reported.
Many large pharmaceutical companies and mature academic HTS groups will have versions of these data that they already use in the evaluation of their screening campaigns; however, this information is not readily communicated in a simple format in literature-screening reports. With the expansion of HTS methodology to smaller start-ups and early-stage academic HTS groups, we propose this NHV model as an accessible way for groups at all levels of expertise to evaluate the quality of HTS hits.
Why “natural history”? Natural history “is a domain of inquiry involving organisms, including animals, fungi, and plants, in their natural environment, leaning more towards observational than experimental methods of study” (https://en.wikipedia.org/wiki/Natural_history). We modify this definition here to read that the natural history of a hit is the domain of inquiry involving “hits” in the laboratory environment, leaning not on experimental methods (as in the useful and important screening tree), but on what has been observed and recorded (metadata) with respect to the hit (compound) in its laboratory environment.
Methods
The Natural History Visualization
Our proposed visualization paradigm for natural histories is shown in Figure 1 . The natural histories components we chose to include in hit characterization include evaluations that are based on scientific evidence of value in the prioritization process: (Element 1) robustness of hit chemical purity and identity characterization; (Elements 2–3) potential for promiscuity that is extrapolated from publicly available screening data; (Element 4) a useful structural and physicochemical property flagging method based on high-quality, pharmaceutical company data; and (Element 5) an evaluation of previous references to the compound in the literature ( Fig. 1A ). These are ordered from left to right in order of their importance to judging hit quality. We chose the tools to generate the element values ( Fig. 1B ) using the following reasoning: (1) The tools should be readily accessible without specialized software and should be based on current, best triage science; (2) the tools report on orthogonal aspects of hit quality; and (3) the tools do not require excessive time or effort to generate the value for each element. Element 1 should be known (or reported, if building NHVs retrospectively). All other elements can be generated starting with the SMILES (simplified molecular-input line-entry system) string of the hit. Figure 1C shows the icons we chose to indicate the hit quality as gauged by each element tool. We chose to use black-and-white versions of icons (https://openmoji.org) to simplify the hit-quality presentation ( Fig. 1C ). An example of a completed NHV (natural history visualization) is shown in Figure 1D .
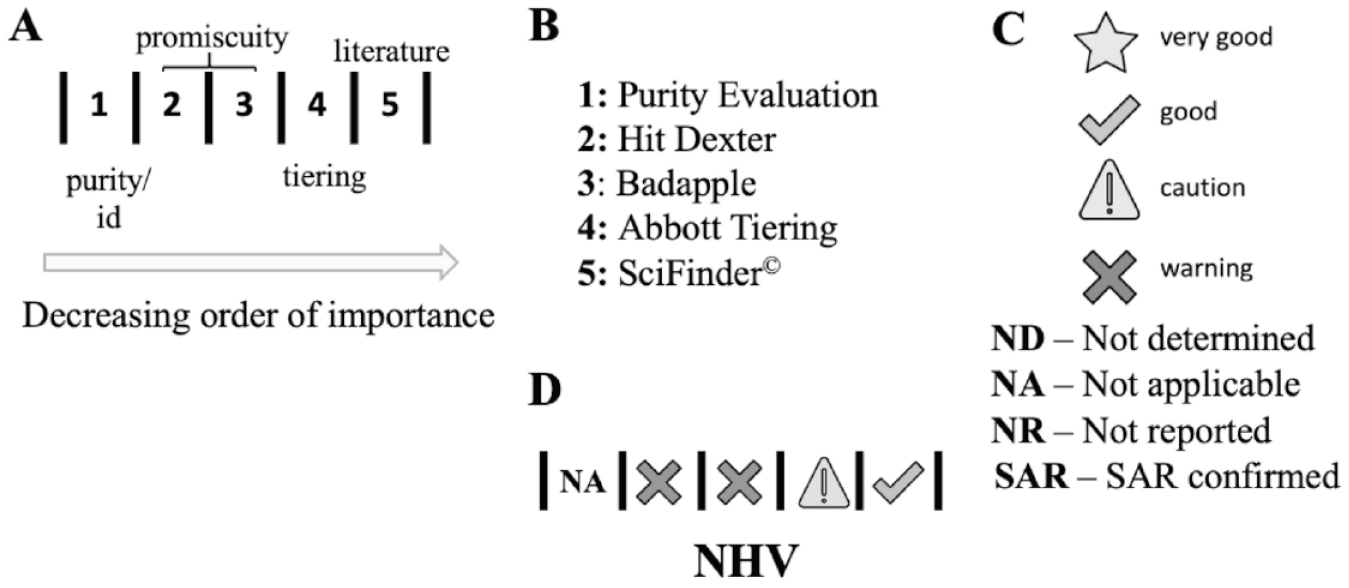
The development of an NHV (natural history visualization) of a hit compound.
How We Generated the Values for Each Element in the Visualization
Purity Evaluation (Element 1)
The reported purity and identification (purity/id) of hit compounds is of paramount importance when considering their activity. This is because some targets and biochemical systems are extremely sensitive to the reagents that are often remaining from compound synthesis, even in commercial compound samples. We have previously outlined different levels of hit compound characterization, but have simplified this process as shown in Table 1 . 3 These values can be assigned by the researchers who report the hits, or in this case, retrospectively by researchers who want to validate a set of hits that were reported.
Rules for Scoring for Each of the Elements in the NHV.
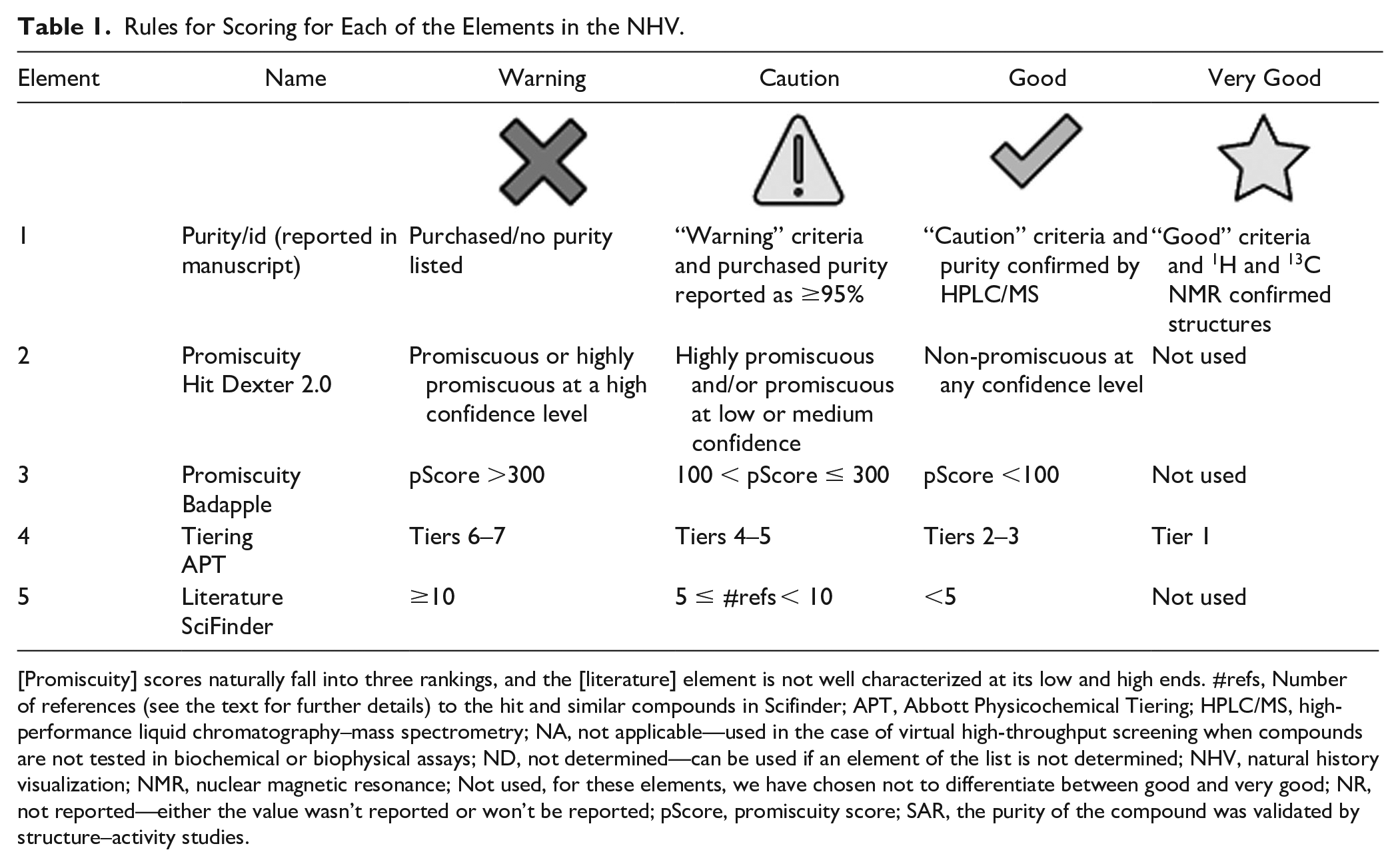
[Promiscuity] scores naturally fall into three rankings, and the [literature] element is not well characterized at its low and high ends. #refs, Number of references (see the text for further details) to the hit and similar compounds in Scifinder; APT, Abbott Physicochemical Tiering; HPLC/MS, high-performance liquid chromatography–mass spectrometry; NA, not applicable—used in the case of virtual high-throughput screening when compounds are not tested in biochemical or biophysical assays; ND, not determined—can be used if an element of the list is not determined; NHV, natural history visualization; NMR, nuclear magnetic resonance; Not used, for these elements, we have chosen not to differentiate between good and very good; NR, not reported—either the value wasn’t reported or won’t be reported; pScore, promiscuity score; SAR, the purity of the compound was validated by structure–activity studies.
Hit Dexter (Element 2)
We assigned a value to the [promiscuity] or potential polypharmacology of hits based on their evaluation in the online tools Hit Dexter 2.04,5 and Badapple. 6 Hit Dexter is a machine learning approach to estimate how likely a small molecule is to trigger a positive response in biochemical assays. The models were derived from a dataset of approximately 300,000 compounds [garnered from the PubChem Bioassay database of ~1 million compounds measured in ~1000 PSAs (primary screening assays)] with experimentally determined activity for at least 50 different protein groups. Hit Dexter uses machine learning models (binary classifiers) to identify compounds that have the potential to be either non-promiscuous, promiscuous, or highly promiscuous in PSA. Hit Dexter scoring is based on complete molecular structures. This sets it apart from the tool used to generate Element 3, Badapple, which uses scaffold-based promiscuity scoring. We use the “Assessment” offered by Hit Dexter to assign the value to this element of the NHV (https://nerdd.zbh.uni-hamburg.de/hitdexter/).
Badapple (Element 3)
Badapple (bioactivity data associative promiscuous pattern learning engine) calculates the promiscuity score (pScore) of a compound according to a formula based on each scaffold of the molecule. This model was built on approximately 430,000 compounds measured in a total of more than 800 different assays gathered from the NIH Roadmap Molecular Libraries Program (MLP) screening centers. The pScore cutoffs recommended by Badapple are used to assign the value to this element (https://datascience.unm.edu/tomcat/badapple/badapple).
Abbott Physicochemical Tiering (APT) (Element 4)
APT 7 is a hit prioritization paradigm that places hits into tiers (from 1 = most favorable to 7 = least favorable) based on a consideration of their calculated physicochemical properties and structural features. This method includes the “consideration of parameters—lipophilicity (ClogP), molecular weight, number of hydrogen bond donors and acceptors, total polar surface area, number of aromatic rings (NAR), and the fraction of sp3 carbons (Fsp3). These parameters describe the overall greasiness (ClogP), size (MW), polarity (tPSA, HBD, HBA), aromaticity (NAR), and three-dimensionality, or level of saturation (Fsp3) of a molecule, and as such should adequately describe the overall physical behavior of a molecule.” 7 This prioritization is included to point out the potential strengths or weaknesses of compounds that may not be represented in the datasets used in building the Hit Dexter 2.0 and Badapple models. We used a custom protocol in Pipeline Pilot to generate the values for this element, but the values could also be generated using values generated in SwissADME (http://www.swissadme.ch/index.php).
SciFinder (Element 5)
We have previously noted that HTS publications often do not cover literature precedence with enough detail to reveal data associated with hits that have already been reported. 8 Therefore, we chose to include a simple-to-determine value of literature relevance in our NHV. This value is determined by a structural search in Scifinder (or equivalent) that returns analogs ≥85% similar to the hit compound, with the criteria that analogs be single substances, be commercially available, and fall into the “Studies: Biological” class. This is then further refined by only retrieving all of the “journal” references associated with the aforementioned analogs that mention “screening” as a research topic. The final count of references remaining after this filtering process determines the value of the [literature] element ( Table 1 ).
Results
To demonstrate the utility of these visualizations, we retrospectively applied these principles of NHV to three specific cases selected from the recent literature, and then individual, illustrative hits. We use [name of element] to indicate the elements of each visualization.
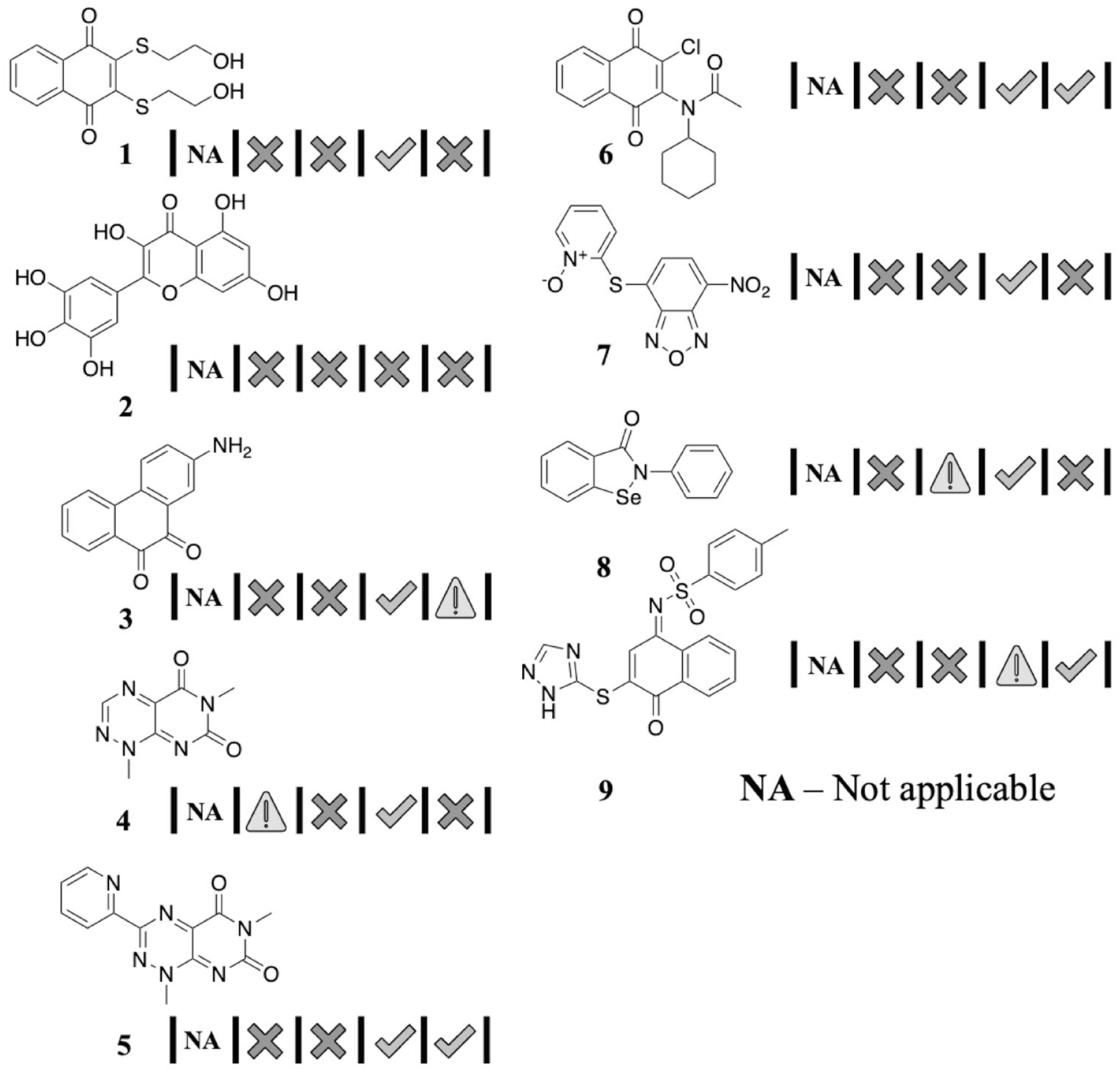
Natural history visualizations (NHVs) of highly promiscuous compounds.
Natural history visualizations (NHVs) of hits reported from a high-throughput screening (HTS) targeting main protease (Mpro) from severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2).
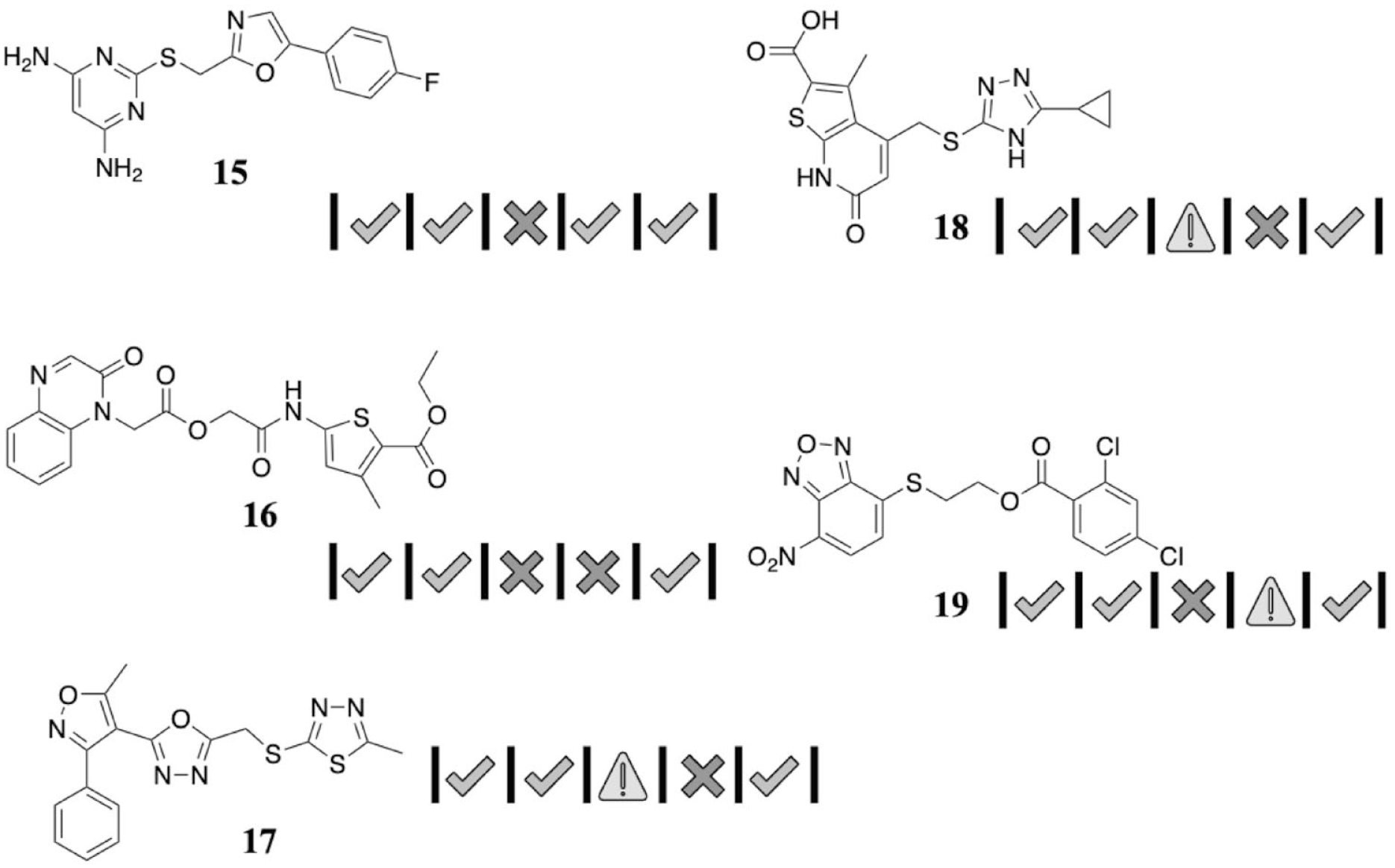
Natural history visualizations (NHVs) of hits from a virtual high-throughput screening (vHTS; followed by assays) targeting druglike small-molecule inhibitors of aspartate N-acetyltransferase (ANAT).
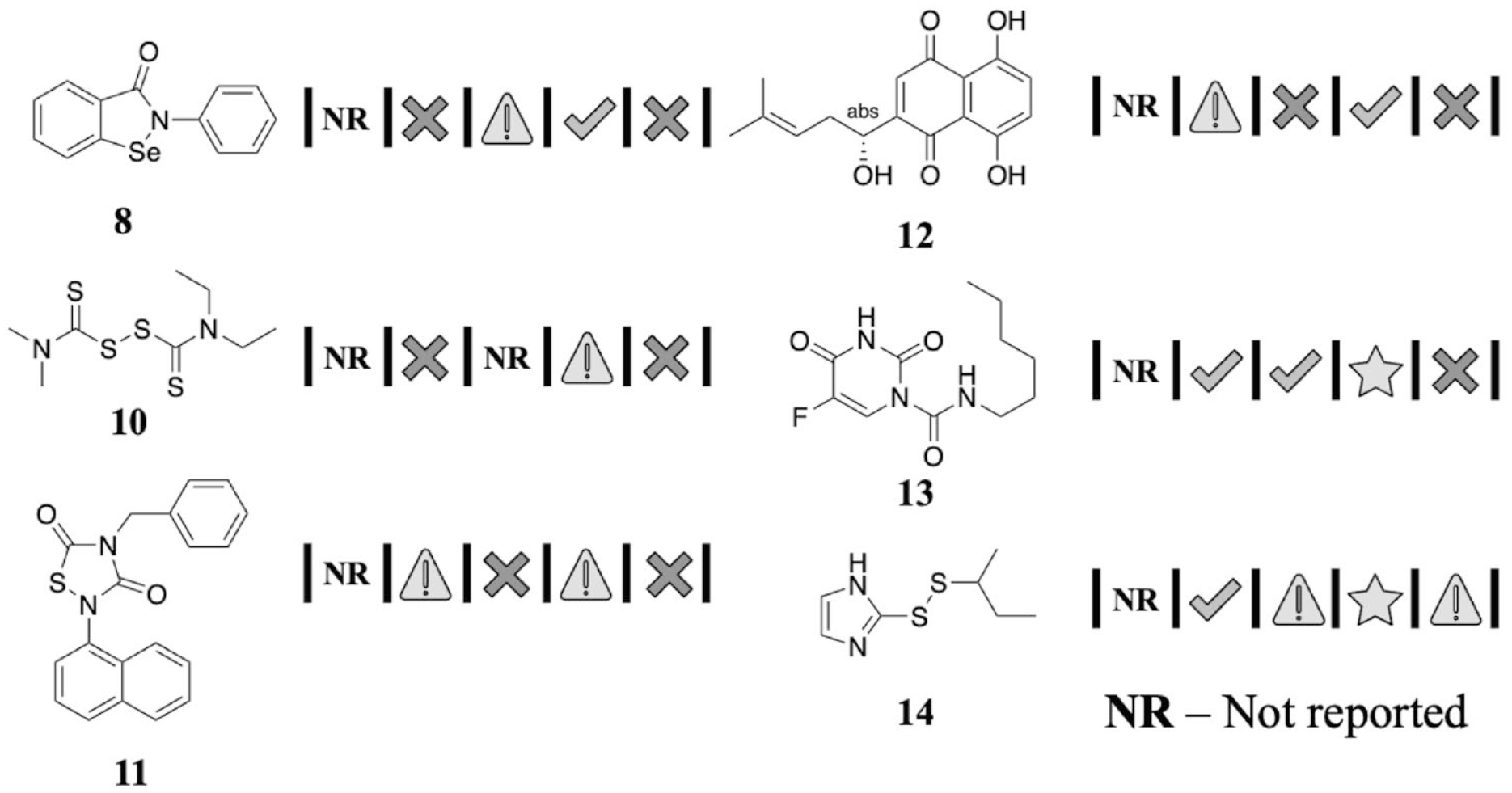
Two further illustrative examples of NHVs are shown in
Figure 5
. Compounds
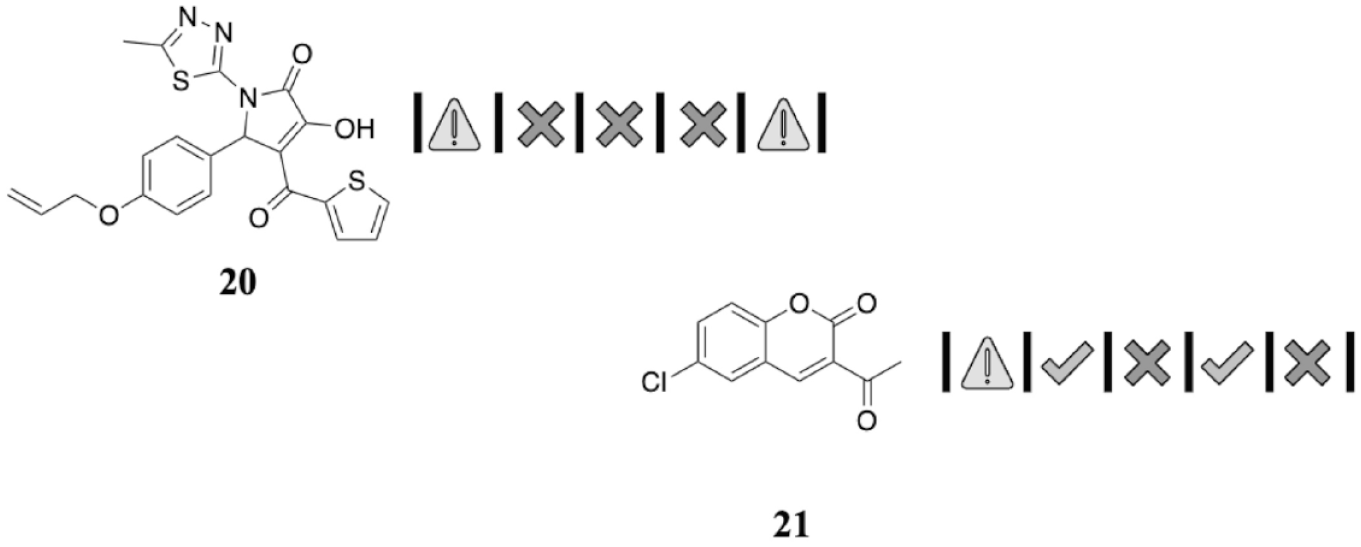
Further illustrative natural history visualizations (NHVs) from hits found in recent high-throughput screening (HTS).
Finally, we examined five articles in which the initial chemical matter was confirmed by SAR (structure–activity relationship) studies (
Fig. 6
). Other than
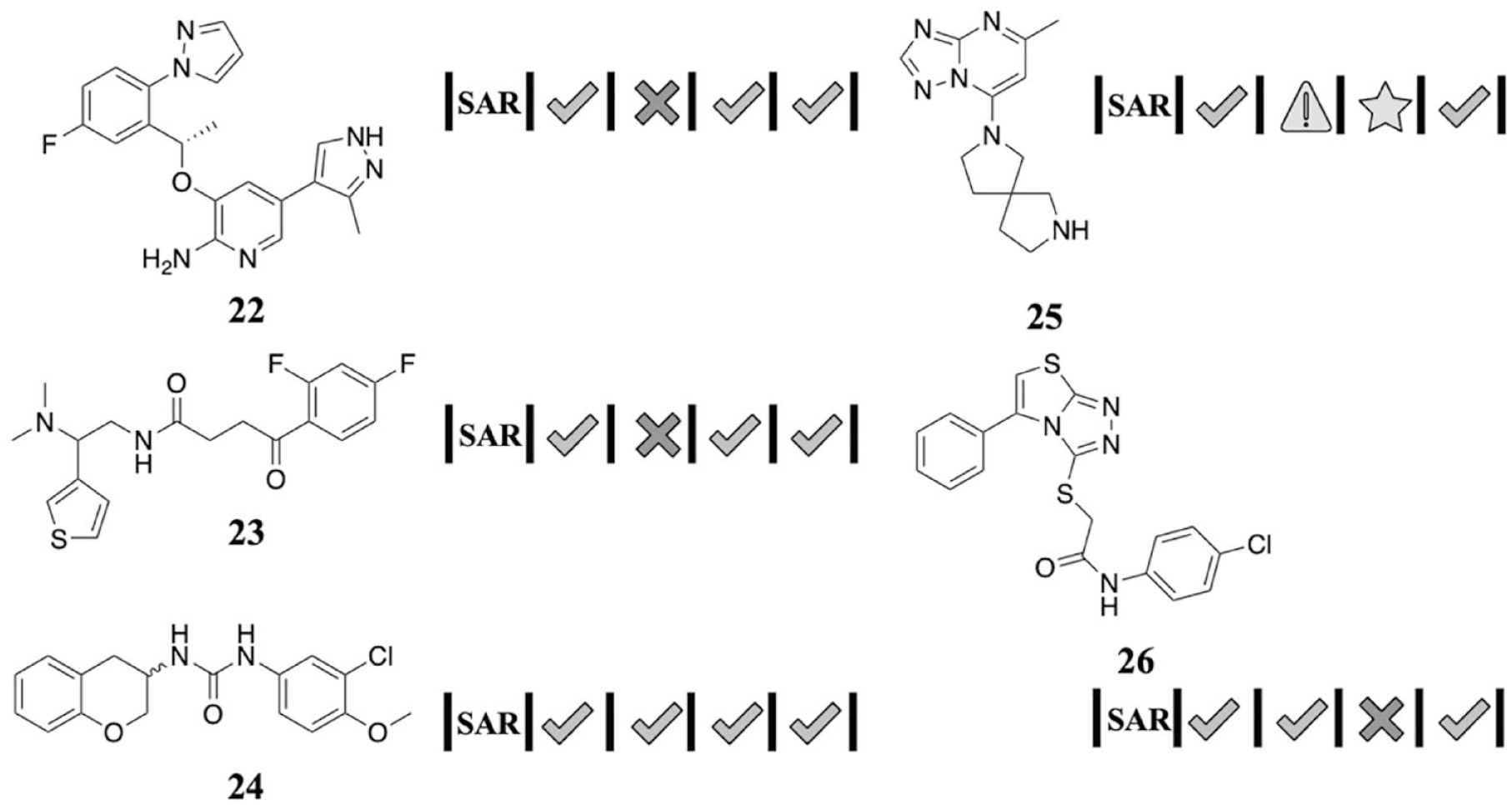
Natural history visualization (NHV) of hits that were confirmed in [purity/id] by the synthesis of analogs.
Discussion
We have developed a natural history visualization protocol that is based on a hit’s known or reported purity/id, its historical pharmacology, its calculated physicochemical and structural tiering, and references of the hit compound and close analogs in the literature. We developed this model because many HTS publications fail to follow best practices in considering these data. We expect that this model will allow researchers to easily and clearly characterize hits from their screens and allow reviewers and readers to immediately put the hits into the context of known compound metadata. Ultimately, NHVs will enable researchers to report and recognize the best screening science.
We chose the elements of this model to be easy to assign based on known data, free software, simple physicochemical property calculations, and readily available literature searches. Each of these elements should be easily accessible to any screening operation. With our retrospective assignment of NHVs to 26 representative compounds, we have demonstrated that this protocol provides a simple, visual readout of hit quality that is an informative addition and complementary to the data produced by biochemical and biophysical methods.
Natural histories alone can’t qualify or disqualify a compound as a true modulator of a chosen target. This can be done only by a well-designed screening tree with the requisite counter-, interference, and orthogonal screens in place. NHVs can, however, provide guidance to a project team as to the risks that might be associated with following up on a particular compound series. These risks might be well-known in the primary literature. In addition, NHVs should be used for guidance, not as strict cutoffs. This is specifically why we didn’t propose a complex scoring system to generate an overall hit “rating.”
Our model alone does not completely address the issue of compounds that may have novel mechanisms of interference or are not similar enough to known database compounds to have an associated pharmacological history. For example, while
High-throughput screening remains the best method for finding chemical matter for novel targets. We propose that this, or similar methods, be used and reported to characterize the quality of compounds found in HTS campaigns and to prioritize compounds for further consideration. Adoption of these methods would also assist reviewers, assigned to judge the outcome of a particular screening campaign, to quickly assess the potential liabilities associated with each of the hits in the material under review. In the future, it would be helpful to have these or similar analysis tools under the purview of an organization such as NCATS (National Center for Advancing Translational Sciences), at least in the United States, to provide a curated and evergreen source of promiscuity and tier scoring. We are currently investigating artificial intelligence (AI)-based models to augment this process and will report these results in due course.
Supplemental Material
sj-pdf-2-jbx-10.1177_24725552211017518 – Supplemental material for The Communication of Hit Quality Using Natural History Visualizations (NHVs)
Supplemental material, sj-pdf-2-jbx-10.1177_24725552211017518 for The Communication of Hit Quality Using Natural History Visualizations (NHVs) by Kathryn M. Nelson and Michael A. Walters in SLAS Discovery
Research Data
sj-xlsx-1-jbx-10.1177_24725552211017518 – for The Communication of Hit Quality Using Natural History Visualizations (NHVs)
sj-xlsx-1-jbx-10.1177_24725552211017518 for The Communication of Hit Quality Using Natural History Visualizations (NHVs) by Kathryn M. Nelson and Michael A. Walters in SLAS Discovery
Footnotes
Acknowledgements
We would like to acknowledge seminal discussions with Daniel Erlanson (VP of Chemistry at Frontier Medicines) and Jonathan Baell (Professor of Medicinal Chemistry, Monash University) throughout the past 10 years regarding high-throughput screening triage.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
