Abstract
The European Lead Factory (ELF) consortium provides European academics and small and medium enterprises access to ~0.5 million unique compounds, a state-of-the-art ultra-high-throughput screening (u-HTS) platform, and industrial early drug discovery (DD) expertise with the aim of delivering innovative DD starting points. From 2013 to 2018, 154 proposals for eight target classes in seven therapeutic areas were submitted to the ELF consortium, 88 of which were accepted by the selection committee. During this period, 76 primary assays based on seven different readout technologies were optimized and mainly miniaturized to 1536-well plates. In total, 72 u-HTS campaigns were carried out, and follow-up work including hit triage through orthogonal, deselection, selectivity, and biophysical assays were finalized. This ambitious project showed that besides the quality of the compound library and the primary assay, the success of centralized u-HTS of large compound libraries across many target classes, various assay types, and different readout technologies is also largely dependent on the capacity and flexibility of the automation on one hand and the hit-triaging phase on the other, particularly because of undesired compound-assay interference. Thus far, the delivered hit lists from the ELF consortium have resulted in spinoffs, patents, in vivo proof of concepts, preclinical development programs, peer-reviewed publications, PhD theses, and much more, demonstrating early success indications.
Keywords
Introduction
While success rates in discovery and development of new medicines remain low and unmet medical needs are still a huge burden on global health care,1,2 many academic discoveries with great potential for pharmaceutical exploitation have remained underutilized.3,4 The underlying reason for this is that, on one hand, academic labs typically lack the resources, infrastructure, and expertise to progress early-stage biomedical discoveries toward development and, on the other hand, pharmaceutical companies and venture capitals are mainly attracted to investing in the later-stage de-risked projects. 5 Over the past decade, the awareness about this so-called innovation gap has resulted in notable changes to the traditional pharmaceutical research and development (R&D) model. A shift from closed innovation to open innovation 6 which is based on complementarity of assets, collaborative efforts, and shared risk within academia, biotechnology companies, large pharmaceutical companies, and contract research organizations (CROs). 7 Such public-private partnerships hold the promise of not only bridging the innovation gap between academia and industry but also integrating the fragmented efforts in the drug discovery (DD) landscape through open innovation.
The European Lead Factory (ELF) consortium is a collaborative public-private partnership with more than 30 partners (2013–2018) funded by the EU commission through the Innovative Medicines Initiative (IMI) and the European Federation of Pharmaceutical Industries and Associations (EFPIA). In its first phase (IMI-1 funding period 2013–2018), this consortium provided researchers in Europe with a unique early DD platform to translate innovative biology and chemistry concepts into high-quality starting points for DD.3,8–10 The ELF consortium accomplished this by capitalizing on two pillars consisting of the first Joint European Compound Library (JECL) and the first European Screening Centre (ESC; Fig. 1A ).
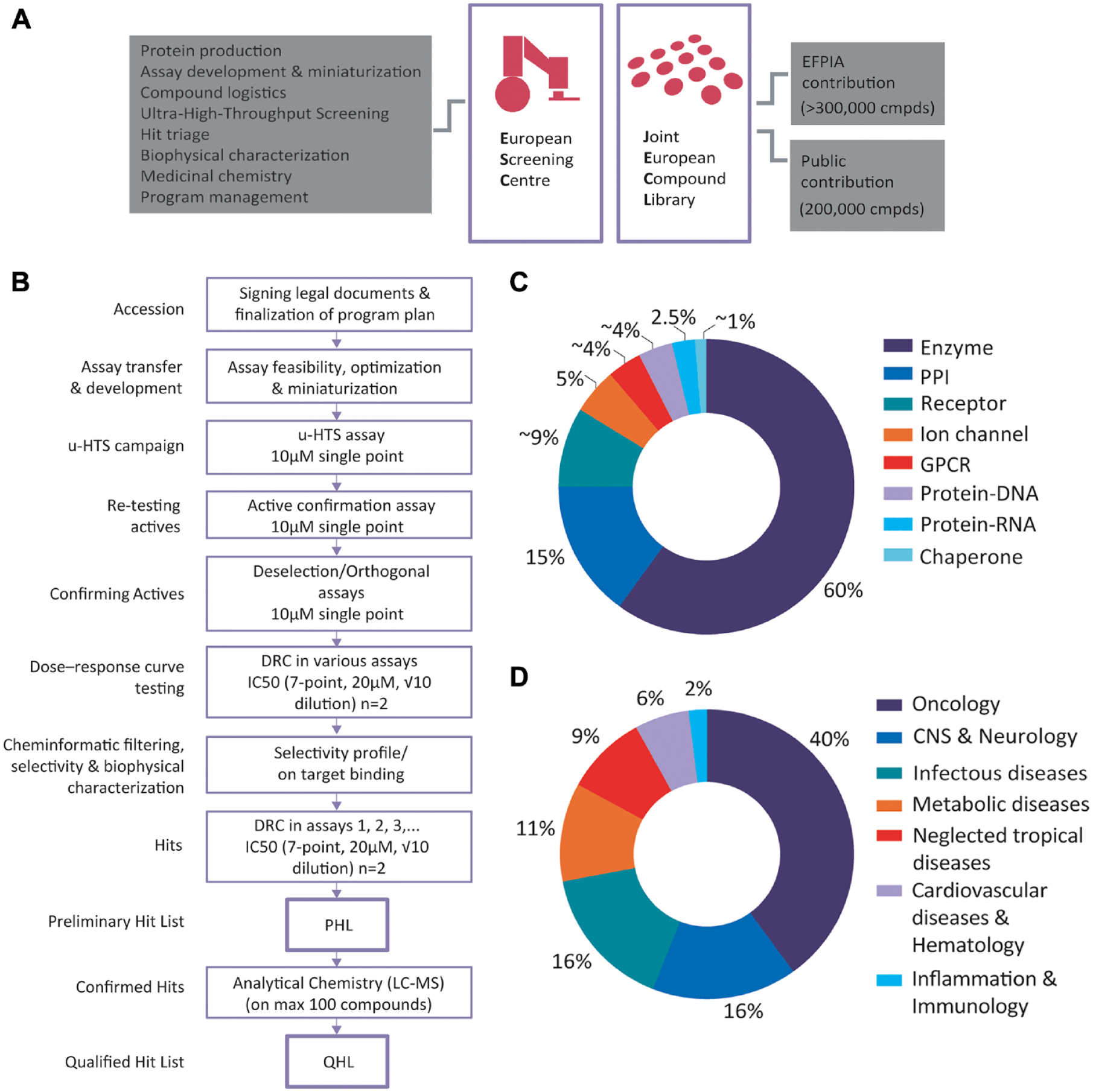
The European Lead Factory (ELF) consortium and the associated screening cascade for public target programs in different target classes and disease areas. (
The JECL is a diverse pool of ~0.5 million drug-like compounds, partly contributed by the EFPIA companies and partly generated by chemistry CROs and academic labs (the latter called the public compound collection). Two studies on the JECL showed that the library is high quality with respect to drug-likeness and physiochemical profile, chemically diverse with each subset driven from a distinct region of chemical space and predicted to have activities on a wide spectrum of biological targets, based on Bayesian models built using data from the ChEMBL database.11,12 Another study on the public compound collection described the approaches and challenges for successful generation of a large screening collection by academic labs and chemistry CROs while populating a biologically relevant chemical space that is distinct from the EFPIA subsets. 13 These overall attributes formed the foundation of a unique and attractive screening collection for small-molecule lead discovery. The ESC, consisting of five different partners, collectively assured the identification of the most promising chemical starting points for further development through their unique and complimentary assets. Participating partners in ESC were the Pivot Park Screening Centre (PPSC) (for assay development, ultra-high-throughput screening [u-HTS], data analysis, and part of hit triage); University of Oxford (for protein production and structural biology); University of Dundee, Newhouse (for biophysical hit characterization and medicinal chemistry); BioAscent (for compound logistics); and Lygature (for program management). Even though the ultimate goal of the ELF consortium is to facilitate and accelerate the discovery of new medicines, it also resulted in unique opportunities such as access to external innovation, knowledge and resource exchange, an extended network, and independent partnerships (https://www.europeanleadfactory.eu/elf-2013-2018/results-2013-2018).
In this perspective, we mainly focus on PPSC as an independent CRO with an emphasis on activities and services, the impact of the ELF consortium on PPSC, and the contributions of PPSC to the ELF consortium. Furthermore, we summarize the overall hit-finding efforts in the ELF consortium public target screening programs, crowd sourced from biomedical researchers at European academic organizations and small biotech companies from 2013 to 2018, and describe a chemical biology approach to systematically evaluate assays before the screening phase and how such an approach could help with tailoring screening assay conditions and guiding postscreen follow-up activities. Moreover, we illustrate the application of cheminformatics tools to enrich blinded primary screen results and reflect on the lessons learned during assay development and u-HTS operations.
PPSC (Partner in ESC)
PPSC is a quickly growing CRO that specializes in early DD services, including cellular and biochemical assay development, u-HTS, hit-to-lead biological profiling, and laboratory automation support, with customers ranging from big pharma to small and medium biotech companies, nonprofit/nongovernmental organizations, and universities in Europe, the United States, and Asia. Upon the takeover of Organon/Schering-Plough by Merck Sharp & Dohme and the decision to close the lead discovery research site in Oss, the Netherlands, PPSC was established as a spinoff in 2012. PPSC currently operates with ~30 employees, including members of the management team, assay development and robot operator groups, as well as additional supporting administrative, finance, and human resource staff.
PPSC is a project-driven organization and as such is organized in a matrix structure, in which project teams are formed with internal members (including a project leader, one to two assay developers, one to two robot operators, and an account manager) and external outsourcing partner members. After initiation of projects based on an agreed work package with clear milestones, deliverables, acceptance criteria, and timelines, outsourced projects typically start with a kickoff meeting and scheduling biweekly or monthly progress report teleconferences, followed by transfer of an existing assay cascade from customers or development of novel screening assay cascades from scratch and subsequent optimization and miniaturization to a u-HTS-ready format. PPSC receives and screens customer libraries ranging from thousands to millions of compounds mainly in 384- or 1536-well formats of both echo-compatible and noncompatible plates or alternatively screens its own ~0.3 million diverse compound library, followed by fully automated data analysis. Identified active compounds are typically retested and then redelivered in REMP tubes and profiled in primary and follow-up assays for hit triaging and subsequent dose-response curve (DRC) testing and biophysical characterization for confirmed hit characterization. An assay development project typically takes 1 to 6 months, and the duration is dependent on the complexity of the assays, availability of commercial reagents, and the need for in-house reagent development. u-HTS projects including assay optimization, miniaturization, and follow-up triaging usually take 2 to 4 months. Hit-to-lead biological profiling services often include delivery of 10s to 100s of compounds to PPSC on a weekly to monthly basis, followed by reporting of structure-activity relationship (SAR) biological data to guide external chemical synthesis cycles. The current automation infrastructure at PPSC includes a state-of-the-art u-HTS lab for large-scale compound screening and an open-access lab with multiple automation islands and liquid-handling stations for medium-scale HTS, hit-to-lead biological profiling support, plate preparations, biophysical testing, and high-content screening. Other facilities at PPSC include assay development labs, tissue culture labs, and a molecular biology lab for wet lab work, as well as storage rooms for compound libraries, chemicals, reagents, and consumables.
With the existing state-of-the-art screening infrastructure, more than two decades of experience in screening, and industrial expertise in lead discovery, PPSC held a competitive position to become a partner for the ELF consortium. To this end, PPSC carried out all the assay development and miniaturization, u-HTS campaigns, and part of hit triaging for the public target screening programs. The overall public target screening programs and the interactions within ESC and the whole ELF consortium contributed to experience and knowledge advancement in novel biology, exposure to a wide range of target classes, therapeutic areas, assay readout technologies, hit-triaging cascades design, and the expansion of PPSC’s network within academia, chemistry CRO space, and pharmaceutical companies. PPSC aims to sustain its growth trajectory by maintaining an up-to-date u-HTS platform through investing in the integration of state-of-the-art screening technologies and novel complex biological assays to accelerate the early lead discovery efforts of its global clients.
Scientific Review and Proposal Selection
All of the eligible public target proposals underwent a rigorous scientific review, while the proposals were kept confidential. This was carried out by the screening selection committee consisting of members within EFPIA, ESC, and external members who evaluated the proposals based on scientific quality, target validation, novelty of target and/or approach, clear need for hits, availability of high-quality assays and supporting data, compatibility of assays with u-HTS and available readouts, chemical tractability, level of innovation, differentiation compared with existing treatments, and exploitation potential. Together with assessment of elements such as patient benefit and risk diversity of the portfolio, proposals were selected that contributed to a scientifically sound, diverse, risk balanced, and economically justifiable portfolio of screening programs. Interestingly, the overall distributions of disease areas and target classes across 154 submitted proposals were largely maintained in the 88 selected proposals. Upon approval of the target program proposals and signing of the legal documents, a program team was formed including the program owner (PO) and experts from the ESC, who wrote a comprehensive program plan defining the strategy to retrieve the best <50 hits (i.e., the qualified hit list [QHL]). 4
Assay Development and Miniaturization
There are many approaches to small-molecule lead discovery. One of the most prominent ones is random high-throughput screening (HTS). Over the past decades, unbiased HTS has remained one of the core approaches of many pharmaceutical and biotechnology companies to identify novel starting points for drug development. In fact, a recent analysis of clinical candidates suggests that ~30% of novel leads are generated through random HTS. 14 The quality of the outcomes from the HTS campaigns is dependent on several factors such as robustness and quality of the primary assay, buffer composition, readout technology, kinetics including incubation times and temperatures, composition of the compound library, quality of automation, as well as the postscreen hit-triaging strategy. A major challenge in evaluating the hits from HTS campaigns is to select hit compounds that are worth pursuing as a chemical lead in a DD program. Therefore, having appropriate follow-up assay cascades in place is crucial for any successful hit-finding effort. 15
Upon ELF consortium onset, u-HTS assay development procedures required rapid evolvement into a pharma-grade early DD operation with smooth assay feasibility testing, validation, optimization, and miniaturization pipeline. To standardize the assay transfer from POs, several documents were created to support every program. Among others, the assay input requirement (AIR) document (describing detailed assay reagents and protocols), program plan (describing the outline of the screening cascade), and general guideline regarding ELF consortium (explaining the workflow and terminology) were used for all programs. The screening cascade outline included in the final program plan described the general public target program screening cascade ( Fig. 1B ). Upon signing the legal documents, AIR documents were transferred, and the program plan was agreed upon. PPSC tested the feasibility of original conditions and reagents in the assays as described in the AIR document and proposed alternative conditions, setups, or readouts if, for example, the PO results were not reproducible, assay parameters were suboptimal when miniaturized, the original assay setup was biased toward a high hit rate or picking up assay-interfering compounds despite optimization efforts, and so forth. In addition, based on the identified liabilities in the screening cascades, alternative or tailored assay conditions were included for optimal hit triaging. u-HTS campaigns on the JECL were carried out in single point (10 µM), and depending on the primary active rate, active compounds were repicked from the same source plates and retested in the primary assay. Confirmed actives (plus a selection of nonactive compounds chosen based on near neighbor analysis of actives and Bayesian model activity predictions) were redelivered from BioAscent, and depending on the project, orthogonal assays (e.g., same assay setup with different readout, alternative enzymatic product with same or different readout, etc.) and deselection assays (counterscreen assays such as same assay readout without the target and/or liability-based assays such as resonance energy transfer quenching, sensitivity to non–drug-like mode of actions such as metal ion chelation, aggregation, etc., as well as viability assays) were employed to confirm activity and deselect false-positives due to, for example, technology interference and uninteresting compounds (e.g., due to toxicities, chelation, aggregation, etc.). The remaining compounds were redelivered in DRCs and tested in the primary and appropriate follow-up assays in duplicates (n = 2) at seven different concentrations (7-point) with a maximal dose of 20 µM and 1/2 log (√10) dilution series. Duplicate data points seemed to be sufficient due to having seven dose points per compound and infrequent outliers. A combination of these activities at PPSC; biophysical characterization of hits at the University of Dundee, Newhouse; and cheminformatics filtering formed the basis for selecting the preliminary hit list (PHL), followed by evaluation of compound structures by medicinal chemists at the ESC, selection of up to 100 most promising compounds for liquid chromatography–mass spectrometry (LC-MS) analysis, and delivery of QHL. On average, the transfer of assays from POs to ESC took about 1 month. On few occasions, POs visited PPSC to accommodate assay transfers and/or learn about screening operations. After full assay transfer and arrival of reagents, ~2 to 3 months was spent on assay panel feasibility testing, optimization, and miniaturization; ~1 to 2 weeks for u-HTS and active confirmation; ~2 to 3 months for hit validation, DRC testing, biophysical characterization, and PHL registration; ~1 to 2 months for LC-MS analysis; and ~1 to 2 months for QHL registration.
The general public target program screening cascade as outlined in the program plan may seem routine to pharmaceutical R&D community and academic screening groups. However, a great deal of discussion between ESC members and POs was vital to justify the basic criteria for u-HTS and acceptable parameters for u-HTS–ready assays; especially terms such as assay robustness, reproducibility, window, and quality, which were interpreted in many different ways. Other challenges ranging from technical to biological aspects were encountered during the course of the ELF consortium operations. As an example, the interaction with more than 40 potential POs prior to the submission of the application was essential to raise the ELF consortium awareness, guide assay development experiments, and inculcate the concepts relevant to the early DD process. Furthermore, 25 proposals passed the selection process, despite not meeting all requirements for the ELF consortium and were as such “conditionally” accepted. For instance, those programs were based on assays with kinetic readout, large volumes, suboptimal Z-prime, inclusion of washing steps, no compatible readout (e.g., radioactivity), or lack of stable cell lines. All 25 conditionally approved programs except one were further developed at PPSC to meet u-HTS standards. Although the total number of assay development activities needed for these programs to reach u-HTS–ready stage was higher compared with the other programs, accepting these programs had an added value to the ELF consortium public target portfolio in the form of innovativeness, novelty, and attractive biology and knowledge expansion of POs. In general, the number of experiments per program prior to the transfer to screening mainly depended on the familiarity of the POs with HTS. The overall ELF consortium public target portfolio in the first phase (IMI-1 funding period 2013–2018) belonged to eight different target classes including enzymes (60%), protein-protein interactions (PPIs; 15%), receptors (~9%), ion channels (5%), G protein–coupled receptors (GPCRs; ~4%), protein-DNA interactions (~4%), protein-RNA interactions (2.5%), and chaperone (~1%; Fig. 1C ). These targets related to seven different disease areas including oncology (40%), central nervous system and neurology (16%), infectious diseases (16%), metabolic diseases (11%), neglected tropical diseases (9%), cardiovascular diseases and hematology (6%), and inflammation and immunology (2%; Fig. 1D ).
Upon transfer of all materials to PPSC, for every single project, assay feasibilities were tested using the detailed protocol and reagents provided by the PO. If meeting all requirements, the program progressed to assay optimization and miniaturization. During assay development and optimization, several experiments were performed for each project to check for appropriate assay component concentrations, stability of reagents and their minimal required concentrations or volumes, kinetics (such as incubation time, temperature, Km/Kd determination), assay linearity, EC50/IC50 of reference compounds, inter-/intraplate variation, DMSO tolerance, optimal final reagent concentrations and volume per well required, plate effects as well as robustness of the assay using reagents, chemicals, and microtiter plates from different vendors. Furthermore, the awareness for the need of orthogonal, deselection, selectivity, and other assays within academia was rather low. PPSC together with the University of Dundee, Newhouse, team (currently largely employed at BioAscent), developed most of these assays from scratch, reaching >200 overall assays ( Fig. 2A ). For this, several factors beyond and above assay robustness and window were taken into consideration, such as the choice of appropriate protein (state) and biochemically sound assay composition including concentrations of assay components, types of salts, pH, reducing agents, detergents, primary assay pitfalls, and so forth.
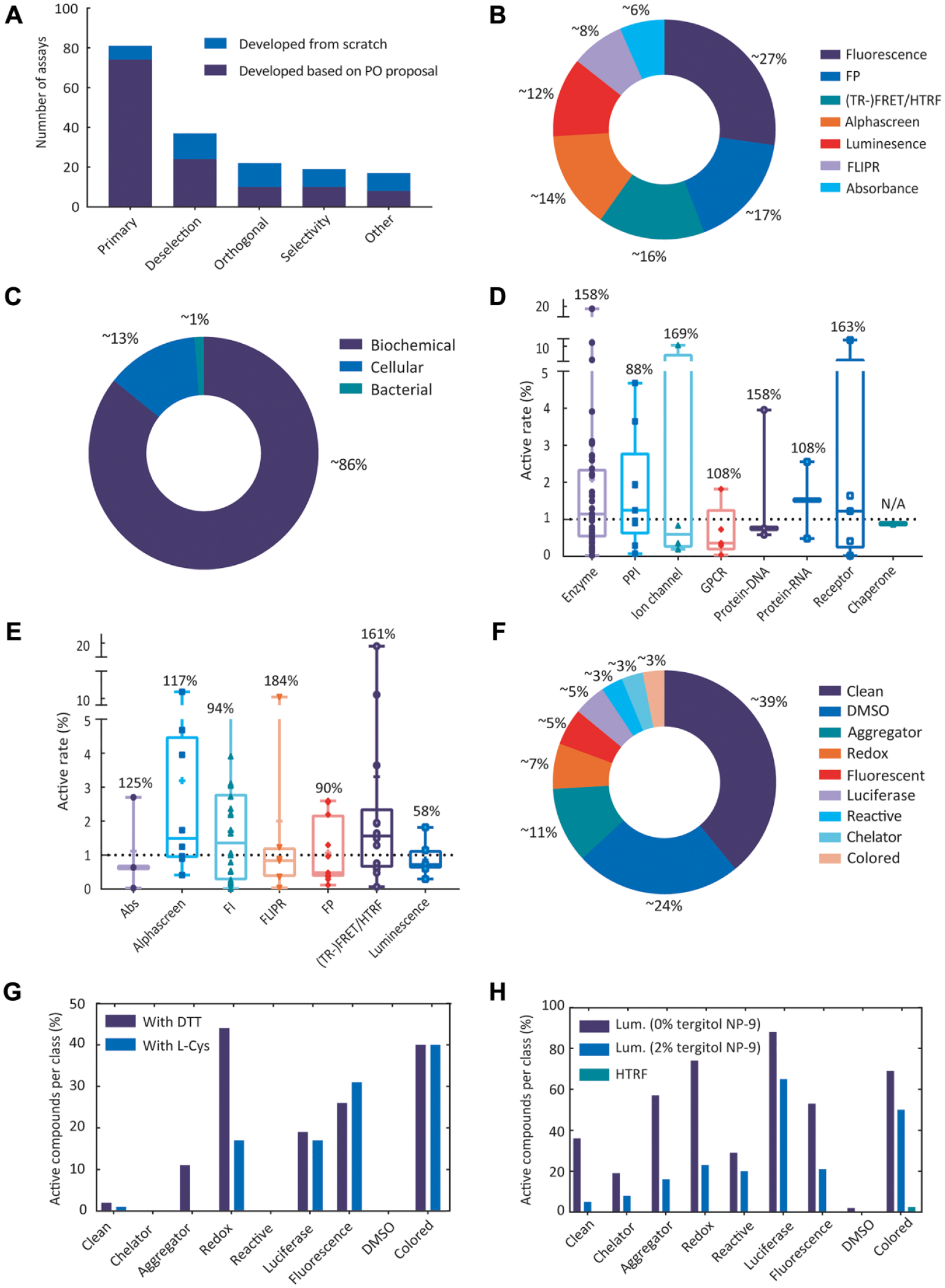
Assay development and miniaturization at the European Screening Centre (ESC). (
The assays employed by PPSC covered a wide range of readout technologies including fluorescence polarization (FP), fluorescence intensity (FI), fluorescence resonance energy transfer (FRET), homogeneous time-resolved fluorescence (HTRF), time-resolved fluorescence resonance energy transfer (TR-FRET), fluorescence imaging plate reader (FLIPR), AlphaScreen (AS), luminescence (Lum), and absorbance (Abs; Fig. 2B ). These readouts were based on biochemical (~85%), cellular (~13%), and bacterial (~1%) assays ( Fig. 2C ). To reduce reagent consumption and processing time on the u-HTS equipment to cover as many programs as possible, all assays, except three, were successfully miniaturized to 1536-well format with a final volume per well ranging from 2 to 8 µL. From 82 transferred protocols, 2 were not feasible because of a lacking active protein within a reasonable price range, and 2 others were stopped because of time constrains for further optimization within the first ELF consortium funding period. In summary, 80 primary assays were shown to be feasible, of which 8 could not reach the screening phase, partly because of the end of project running period. Of these eight, four POs received a u-HTS–ready protocol (meaning fully automated and miniaturized assay protocol) ready to be applied to screening in 1536-well format. The remaining 72 programs progressed to the u-HTS phase. The data from primary screens of these programs showed that the average primary active rate (based on Z-score >4 or <−4) is dependent on neither the target class ( Fig. 2D ) nor readout technology ( Fig. 2E ) and large variation in primary active rates within each target class (CV = 88%-169%) and readout technologies (CV = 58%-184%) suggested that in HTS campaigns, biological targets from the same class do not necessarily behave the same. This observation was not entirely unexpected, as each target class contained variety of subclasses with different target protein structural domains, catalytic activities, and assay components. In addition, the data provided counterevidence to general assumptions with respect to superiority of certain readout technologies and confirms that the primary active rate is a poor predictor of the “real” hit rate, especially for challenging targets such as protein interactions with typically large interaction surfaces. Nevertheless, the collected data from these campaigns showed that the median of primary active rates for ion channels, GPCRs, and protein-DNA interactions as well as Abs, FLIPR, FP, and luminescence readouts are less than 1%, and luminescence primary assays had the least variation (CV = 58%) compared with other readouts. It is of importance to note that a Z-score >4 or <−4 was used here to make an unbiased comparison across readouts/targets, and even though the same criteria were used to identify primary actives in many programs, depending on the data spread, the primary active selection for some programs was based on the %Effect alone or in combination with Z-score ranging from >10 to 4 or <−10 to −4.
Robustness Set Compound Collection
It is widely accepted that besides the quality of the library and the primary assay, the success of any u-HTS campaign is largely dependent on the postscreen hit-triaging phase, particularly because of undesired compound-assay interference leading to false-positive data. The undesired false-positive effects of these compounds disturb the hit selection process, thus requiring orthogonal assay readouts and adjustments in the primary assay conditions. For organizations running many u-HTS assays with the same compound library, such interfering molecules in the long run become recognized as “frequent hitters.” The main underlying causes for such interferences are aggregation, metal ion chelation, redox activity, autofluorescence, absorbance, luciferase inhibition, and chemical reactivity.16–18 To dissect such liabilities early in the assay development phase, the “robustness set” compound collection was established together with the University of Dundee, Newhouse. The robustness set comprises selected molecules from the literature with well-defined assay interfering properties and so-called clean compounds for which we did not expect assay interference. These were supplemented with many DMSO controls, all in a ready-to-employ single 1536-well plate ( Fig. 2F ).
The application of the robustness set within the ELF consortium target programs not only provided a framework to identify assay liabilities and tailor u-HTS assay conditions such that minimal interferences occur but also helped to design suitable follow-up assay cascades for successful hit triaging. Based on the lessons learned from early ELF consortium target screening programs, the established framework prior to calling an assay u-HTS ready was to perturb the assay with different classes of molecules with known assay interfering properties and a small set of chemically diverse leadlike compounds without obvious assay interfering properties. Active compounds within the robustness set were then identified by quantifying responses to these perturbations (%Effect relative to DMSO controls). Depending of the assay type and responses to reference compounds, typically a compound in the robustness set was called “active” when it resulted in a %Effect of >25 to 50%. Once active compounds were identified, a postanalysis was performed to calculate the percentage of active compounds per each class that showed a %Effect >25 to 50%. Although the percentages of actives in the clean compound class provided an indication of the expected hit rate, the percentages of actives in other compound classes informed on assay liabilities and biases toward picking up undesired compounds. Depending on the nature of identified liabilities, appropriate adjustments were made to assay conditions.
As an example, we profiled the performance of an FP assay from a PO using the robustness set. The original assay protocol indicated dithiothreitol (DTT) as a reducing agent in the assay buffer. Testing robustness set compounds resulted in the identification of assay liabilities, mainly toward redox-cycling compounds (~45% of compounds in this class were active), autofluorescent compounds (~25% of compounds in this class were active), and colored compounds (~40% of compounds in this class were active). DTT and oxygen present in HTS samples may generate H2O2, which can oxidize essential functional groups in proteins (e.g., cysteines). Such processes are catalyzed by compounds capable of redox cycling, thus resulting in false-positives. 19 To tailor the assay buffer such that it becomes less biased toward redox compounds, a weaker reducing agent (L-cysteine) was used, resulting in picking up fewer redox compounds (~17% of compounds in this class were active; Fig. 2G ). Even though using a weaker reducing agent is not expected to have an impact on liabilities toward autofluorescent and colored compounds, the results from the robustness set test guided the decision on the type of orthogonal and deselection assays to be used in the hit-triaging phase.
In another example, a luminescence-based primary assay for the activity of an epigenetic enzyme was miniaturized to 1536-well plate format based on the adaption and optimization of the PO protocol. Results from testing the robustness set showed that the assay is responsive to luciferase inhibitors as expected (90%) and predicted a high hit rate based on the percentage of actives in the clean class (~38%). Furthermore, the assay setup was prone to picking up different classes of assay interfering molecules such as autofluorescent, colored, redox, and aggregators. Even though the addition of 2% tergitol NP-9 to the assay buffer reduced these liabilities, switching to HTRF readout technology significantly improved the primary assay quality ( Fig. 2H ) and minimized efforts needed in the hit-triaging phase.
u-HTS
In addition to the assay development activities, all u-HTS campaigns for the ELF consortium public programs were performed at PPSC. This facilitated the use of screening equipment in assay development. PPSC started operations in 2013 with a newly installed, state-of-the-art u-HTS platform developed together with HighRes Biosolutions. The system was designed to run biochemical and cellular assays in a fully automated manner using 1536-well plates, running >20 full-deck screens per year on a 2.5 million compound library. It had built-in flexibility to support different types of assays and projects and featured redundant instruments to allow for continuing operations in case of instrument failures. The u-HTS system consisted of three pods with Staubli robotic arms surrounded by >50 instruments and connected by a conveyer belt system. The three pods operated as a single system for large and complex projects or independently to perform different (smaller) tasks. Most instruments were positioned on carts, so that the configuration of the system could be easily adapted to the needs of individual programs and future technology upgrades. The system was accompanied by a unique and custom-designed safety system consisting of enclosures and hurricane doors. The enclosures shielded the users from the robot arms, and the hurricane doors were closed when needed to separate individual instruments from the robot. This allowed for trouble shooting, reagent top-ups, and even assay development activities for new programs in parallel, without interrupting the robot activities. For example, one of the ELF consortium screening programs required supplementing freshly prepared enzyme solution every hour because of enzyme instability, which was smoothly accommodated by the platform. The compound preparation pod featured a 12-sided pod including four incubators for storage of compounds in controlled conditions.
The JECL compounds, prepared in 1536-well format by BioAscent in Scotland, were stored online for short-term use in sealed plates within the environmentally controlled incubators (~10% humidity with constant N2 flushes at room temperature) that were directly accessible by the robot for screening. To limit and control compound oxidation, the compound source plates were refreshed every 3 months or used in <10 operations to ensure that compound integrity is maintained. u-HTS–ready assays were tested for reproducibility, stability, and robustness during prescreen runs (methodology phase). Protocols were optimized to eliminate and, if not possible, minimize plate incubation and dispenser effects. Quality during screening was monitored by including DMSO dummy plates and quality control plates containing serial dilutions of available reference compounds. During primary screens, samples were transferred into assay plates (mostly 1536-well format, but the system also supported 384-well format) using ECHO 555 acoustic dispensers. Source plates were resealed and returned to the online store while the assay plates were transported to the other pods using the conveyer belt system. The two assay pods on the system were nine-sided and held all instruments that were required to perform various biological assays. Different types of dispensers were available to enable users to select optimal conditions for a wide range of reagent types and volumes, and seven available Cytomat incubators allowed different incubation steps at temperatures ranging from 4 to 37 °C. Assay readouts were performed in pod 3, which covered a wide range of readout modalities. To keep the u-HTS platform up to date and accommodate a broader range of biological assays and readouts, several novel instruments and technologies were implemented into the u-HTS system. Among these were reliable, flexible, and fast dispensers to increase accuracy, reduce dead volumes, and reduce clogging (ECHO 555 and Certus); plate washers suitable for robust washing steps in 1536-well format and medium replacement (BlueWasher, BlueCatBio, Concord, MA); FLIPR-Tetra suitable for ion channels and fast kinetics readouts such as calcium fluxes and membrane potential measurement; next-generation multimode readers (PHERAstar, BMG Labtech, Cary, NC); and, recently, RapifleX (Bruker, Billerica, MA) for label-free MALDI-TOF mass spectrometry readout capable of u-HTS to minimize the use of costly detection reagents, limit technology interference, and shorten the hit triage.
Data processing was performed using a combination of in-house and commercial software. The custom-built SAMTRACK software continuously tracked samples and monitored operations on the robotic system. Logfiles produced by dispensers such as the ECHO were processed for every sample across the screening campaign. These plate maps were then automatically fed into ActivityBase software platform (from IDBS) together with the data files from the readers without manual intervention, and users easily evaluated the data remotely in ActivityBase and performed data quality control. When necessary and if applicable, empirical orthogonal function in IDBS software was used to correct for reproducible plate effects. Z-factor (Z′) >0.6 and signal-to-background (S/B) >3 (>2 in HTRF) were used as quality parameters for approving data from plates. For a few cellular screening programs that were not robust, Z′ >0.5 was also accepted for data approval. After data approval, the final data were loaded into ScienceCloud in a batch process that ran automatically every night. The batch process was included later to increase efficiency. The ScienceCloud package was used for secure storage and analysis of all screening and triage data. To maintain confidential information (e.g., about the JECL structures), Honest Data Broker (HDB) 20 was specially integrated into ScienceCloud and used to analyze blinded u-HTS campaign results. To use the HDB as a tool throughout all ELF consortium processes, the consortium agreed on standardized metadata (e.g., for target classes, technology readout, and mode of action) as well as abbreviations for different triage phases (for instance, P for primary screen, C for confirmation, O for orthogonal, etc.). This was the basis to create a dedicated database that can be used for cross-program analysis to determine, for example, frequent hitters, technology interfering compounds, unique compounds, target class-related compounds, and so forth. For identification of primary actives, depending on the respective mode of action, a Z-score of >4 or <−4 was used as minimum criteria for initial active selection.
If the number of primary actives was above the selection criteria, “active confirmation” was fulfilled via repicking compounds from the full library. In general, the active rate was typically lower in the active confirmation than primary screen. This could be partly explained by statistical false-positives during the primary screen, resulting in a lack of activity within repicked compounds. In addition, the process of repicking the primary active compounds that are randomly distributed across the compound source plates is time-consuming and results in a 1536-well assay destination plate long unsealed time and risk of DMSO evaporation and compound precipitation. To avoid DMSO evaporation in assay destination plates during compound repicking, compounds were transferred to ≤10 columns of 1536-well destination assay plates. This reduced the destination plate unsealed time, resulting in better correlations with compound effects observed in primary screen. Furthermore, depending on the active rate, sometimes the selection criteria needed to become more stringent to stick within the compound limit of 1% to be reordered for testing as serial dilutions to generate DRC data. This cutoff made the follow-up work more economical.
To enrich the primary active compound list while carrying out blinded u-HTS campaigns, the power of cheminformatic tools available on ScienceCloud was used to reduce the risk of early abandonment of false-negatives and potentially generate early SAR data using Bayesian models and near neighbor searches. The Bayesian model used molecular fingerprints of the compounds together with experimentally determined biological activities to derive features of active molecules that were associated with biological activity. Then, the model cross-referenced molecular fingerprints of every molecule in the full library against features associated with biological activity to find potential false-negative compounds. Typically, the top 250 to 500 molecules predicted to be active were included for follow-up retesting. Near neighbors were compounds showing similar structural scaffold as active compounds, yet they were not identified as actives in the primary screen. An acceptable chemical similarity score cutoff was set to find these molecules, which were later included in the primary active list for retesting. Eleven programs had QHLs containing at least one Bayesian model– or near neighbor–derived compound.
Even though during the project ramp-up phase fewer programs were recruited than planned, PPSC managed to complete >20 full-deck screens for ELF consortium programs in 2017 ( Fig. 3A ), reaching a u-HTS production of >12 million data points (in combination with non-ELF screening campaigns), an output typically expected from an established large pharmaceutical company. The throughput for every screen was optimized per assay based on required incubation times and stability of reagents. For screens carried out in 384-well formats, the maximum number of data points per day was up to 54,000 ( Fig. 3B ). Every u-HTS campaign started with a prescreen of 50 plates and continued if the assay performed within the acceptance criteria. By processing around 100 to 200 plates per day for screens based on 1536-well formats, the system routinely generated up to 200,000 data points per day and even ran an enzymatic assay on 350,000 compounds in less than 24 h ( Fig. 3B-C ). Although most screens were carried out in 3 to 4 days, some expanded to 5 days in order to retest failed plates, and a few campaigns based on 384-well format took up to 7 to 16 days, mainly because of assay instability. These were the same assays in which efforts to miniaturize them to 1536-well format failed.
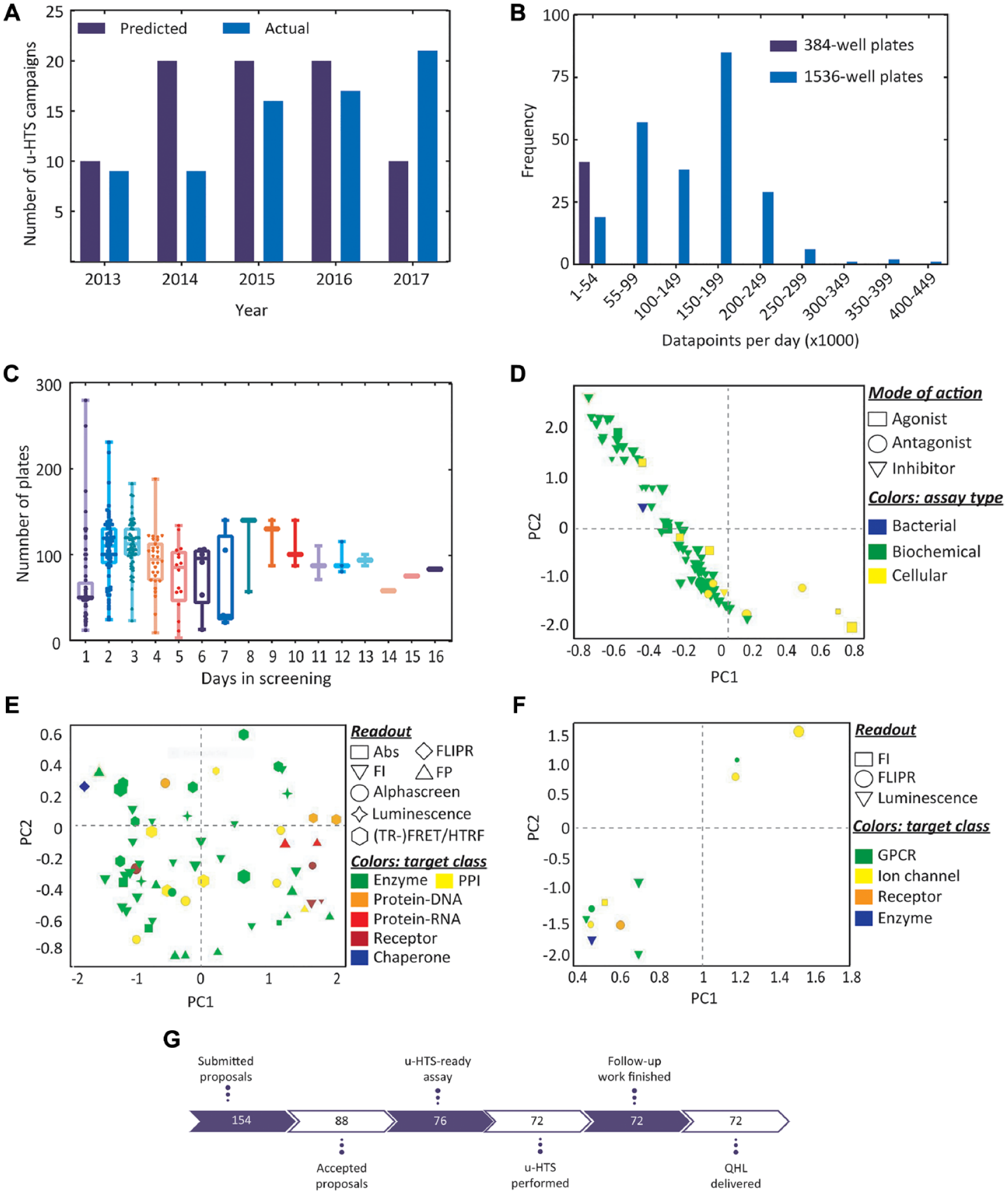
Ultra-high-throughput screening (u-HTS) output within the European Lead Factory (ELF) consortium. (
To identify generalizable trends in the data collected during the screening cascades for all programs, principal component analysis (PCA) was carried out on different variables such as excitation and emission wavelength (except luminescence and for absorption), plate format (number of wells in 384-well and 1536-well formats), volume per well, library size (the size of the library increased over the course of 5 years), number of screening days, assay robustness (Z′) and window (S/B), primary active rate, confirmed active rate, number of reordered compounds, PHL, number of LC-MS qualified compounds, QHL, and number of Bayesian model– and near neighbor–derived compounds in QHL ( Fig. 3D ). In a 2-dimensional PC space, PC1 mainly captured the variation in number of compounds registered in QHL, with higher numbers associated with some of the cellular assays, and PC2 was a function of screening days and well formats. This orientation in PC2 space was partly trivial, as the u-HTS campaigns based on some of the cellular assays that were carried out in 384-well format had larger volumes per well and took more days to finalize than others. Interestingly, these programs mainly focused on identification of agonists and antagonists but not inhibitors and had the highest number of possible compounds (50) registered in QHL. In contrast, programs focused on inhibitor finding mainly against protein-DNA, protein-RNA, receptors, and some PPIs, and enzymes had up to 10 compounds in QHL, in line with known challenges to identify inhibitors of protein interactions. A low number of compounds in the QHL list did not seem to be readout dependent. We carried out the same analysis on biochemical programs ( Fig. 3E ) and cellular programs (including a bacterial program; Fig. 3F ) separately to avoid biases toward assay type–specific trends. Whereas in the PCA of biochemical screening campaigns, besides relative short distances between protein-RNA, protein-DNA, and some receptors, no distinct clusters were identifiable in PC space, the PCA of cellular screening programs mainly captured differences in screening days and plate formats across similar targets and readout technologies, suggesting that the potential for miniaturization is not necessarily target class or readout technology dependent.
Conclusion
Discovery and development of new medicines is a risky and time-consuming business. Increasing costs and declining R&D productivity remain major challenges for large pharmaceutical companies and global health care. In response to this sustainability crisis, collaborative efforts and open innovation models have gained much attraction. The ELF consortium is an example of such collaborative efforts to accelerate the discovery of new medicines. This pan-European platform not only facilitates the translation of novel biology and chemistry into high-quality starting points for DD but also stimulates the cooperation of public institutes and the private sector.
Over the course of 5.5 years within the first phase of the ELF consortium, 154 target programs were submitted to the consortium. After review by the selection committee, 88 proposals were accepted, of which 6 were not progressed for legal reasons. The distribution of target classes and disease areas of accepted proposals were roughly similar to the distributions across all submitted proposals. For the remaining 82 programs, the feasibility of 78 primary assays was validated, and 76 assays were successfully optimized and miniaturized to 1536-/384-well format. u-HTS campaigns for 72 programs were completed, which followed the finalization of follow-up triaging and delivery of QHLs ( Fig. 3D ).
The results from these campaigns made it evident that, usually, HTS assays that are not adaptable to 1536-well format also perform suboptimally in lower plate formats, and the findings also reconfirmed the challenges in identifying small molecules against protein interactions with other proteins and nucleic acids. In addition, these results demonstrated that even within the same target class, outputs from different readout technologies are context dependent, and in-depth understanding of the molecular interactions between targets, compounds, assay components, additives, and buffer conditions is critical to design optimal u-HTS assays. Moreover, we showed that systematic perturbation of screening assays with well-characterized molecules is a reliable method to determine assay liabilities and design tailored follow-up cascades accordingly. This approach fits well with the existing infrastructure in screening facilities and provides a framework to execute successful screening programs rapidly, which is particularly important for project-based organizations with high project turnover. On the technical side, we exemplified the importance of miniaturization for maximized throughput and showed how flexibility of the automation could, on one hand, accommodate biologically and technically diverse screening programs and, on the other, facilitate the integration of various new instruments to keep a u-HTS platform up to date.
Together with its partners at ESC, PPSC actively contributes to the ELF consortium by disseminating knowledge and know-how to the public target portfolio, streamlining industry-standard operations from assay concepts all the way to confirmed hits, already delivering >200 high-quality assays and successful u-HTS campaigns and follow-up triaging results. In the meantime, PPSC has positioned itself as a well-known service provider in the international early DD market and academic space, thanks to the knowledge and experiences gained during the project as well as the network that accompanied the ELF consortium. On the EFPIA side of the ELF consortium, independent HTS campaigns on the JECL were carried out by partner companies (at their own in-house screening facilities), resulting in a combined industry target portfolio, which adds value to the internal DD programs of each partner through the expanded chemical space facilitated by the ELF consortium.
Access to an ever-expanding, biologically relevant chemical space, enabled through shared high-quality compound libraries, together with the widespread use of novel label-free u-HTS technologies such as MALDI-TOF mass spectrometry holds the promise of not only accelerating the discovery of de-risked new chemical entities for further development but also reducing the cost of large-scale screening and the duration of hit triage as well as hit-to-lead phases. We envision that in parallel with other lead-generation methods such as structure-based drug design, screening DNA-encoded libraries, fragment-based screening, and so forth, the method development efforts for label-free multiparametric/phenotypic readouts and maturation of physiologically relevant disease modeling in miniaturized assays based on organoid, induced pluripotent stem cell, and Crispr/Cas technologies have the potential to pave the way for the discovery of the next generation of high-quality starting points for drug development and ultimately improve pharmaceutical R&D productivity by reducing late-stage attrition rates.
It is commonly known that it takes 10 to 20 years on average to bring a single new medicine to the market. Therefore, it is still too early to truly evaluate the direct impact of initiatives such as the ELF consortium on health care. Nevertheless, early success stories and achievements from the ELF consortium now form the basis for continued operations of the ELF consortium concept through the ESC, a unique library for attractive biology (ESCulab; https://www.europeanleadfactory.eu/), an IMI-2 project.
Footnotes
Acknowledgements
We thank all of our colleagues in the assay development and operator groups at PPSC, members of the ESC, especially our collaborators at BioAscent (formerly at the University of Dundee, Newhouse), public target program owners, and all members of the ELF consortium for their feedback and helpful discussions. We would also like to show our gratitude to Jon de Vlieger (Lygature) for critical comments on the article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research leading to these results has received support from the Innovative Medicines Initiative Joint Undertaking under grant agreement no. 115489, resources of which are composed of financial contribution from the European Union’s Seventh Framework Programme (FP7/2007-2013) and EFPIA companies’ in-kind contribution.
