Abstract
Collaborative efforts between public and private entities such as academic institutions, governments, and pharmaceutical companies form an integral part of scientific research, and notable instances of such initiatives have been created within the life science community. Several examples of alliances exist with the broad goal of collaborating toward scientific advancement and improved public welfare. Such collaborations can be essential in catalyzing breaking areas of science within high-risk or global public health strategies that may have otherwise not progressed. A common term used to describe these alliances is
Keywords
Introduction
Over the last few decades, a multitude of initiatives have emerged that build on precompetitive collaborations and sharing of assets. Within the life science sector, such initiatives involve companies, academic institutions, governments, and nonprofit and nongovernment organizations, in combinations that range from two-party agreements to large, privately and publicly funded consortia. Sharing, through open access as well as regulated collaborations, can be very valuable for both research areas currently in focus and those currently outside the spotlight. It may open up opportunities for new discoveries and consequently improve public health. Although many models of asset sharing can be found within the pharmaceutical sector, the public-private partnership (PPP) model has gained popularity as an effective way to boost innovation at different stages of drug discovery and development. In a common definition, a PPP involves at least one public (nonprofit, academic, or government) unit and at least one private for-profit partner. 1 However, PPPs can take many forms, and within the biomedical/pharmaceutical sector, these range from bilateral agreements between academic institutions and industrial partners to large, multistakeholder collaborative models.2,3 Figure 1 provides an overview of the main PPP participants in drug development, lists some of their core strengths, and highlights a few key benefits to the overall drug discovery pipeline. This review describes different types of PPPs, assets shared, and applications in the pharmaceutical sector obtained from various public and peer-reviewed sources. Although many topics and examples are discussed here, the premise of PPPs in the pharmaceutical sector is very broad, and this review is by no means exhaustive of existing PPPs relevant for the industry.
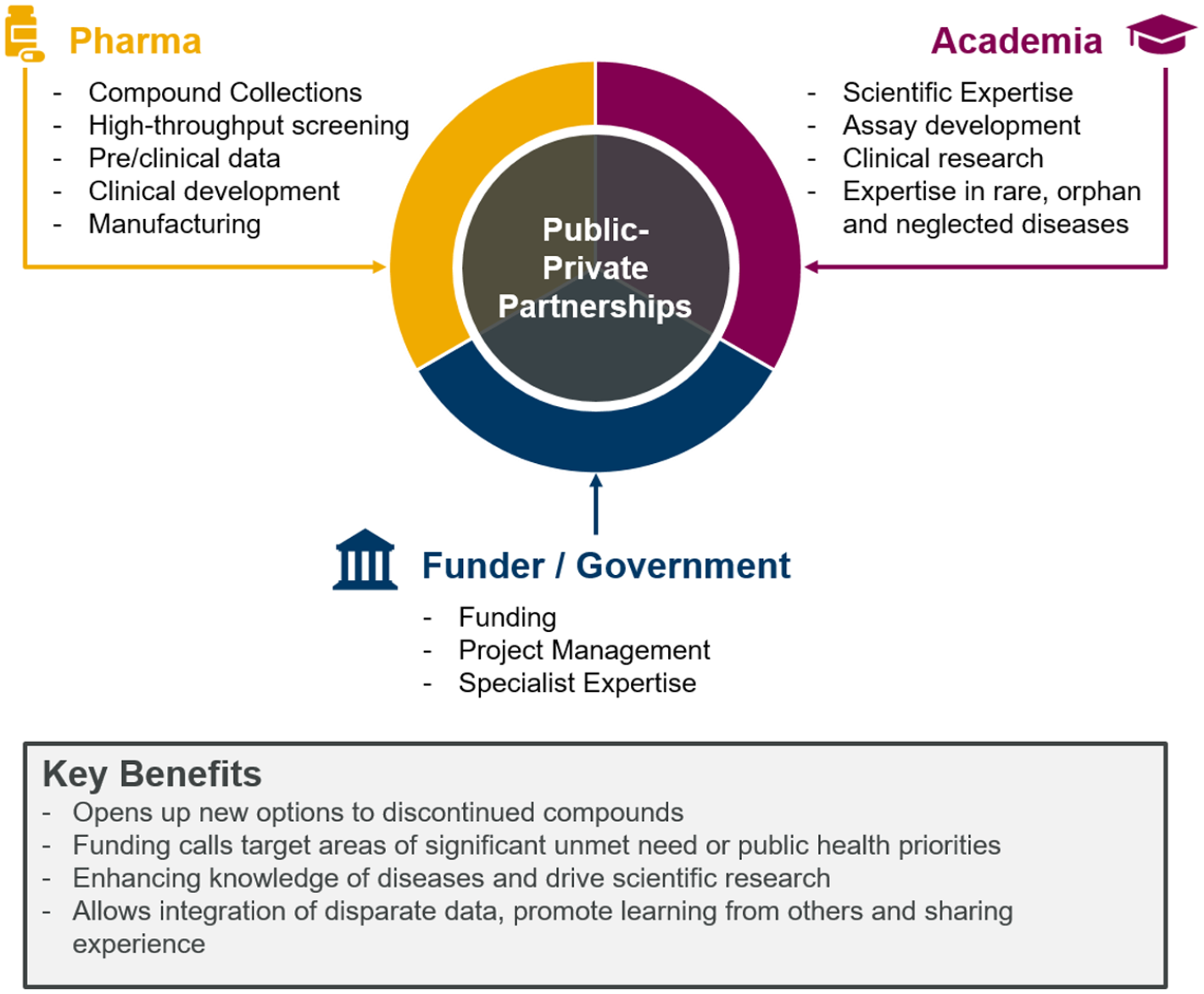
Main participants in a PPP—core strengths and key benefits in drug discovery/development. The figure provides an overview of the main participants in a PPP, highlights their core strengths, and lists important benefits across the various stages of the drug discovery pipeline.
PPPs in Drug Discovery
Several initiatives have been launched involving government units, leading pharmaceutical companies, and academic institutions to increase productivity, leverage collective expertise, and decrease attrition rate during the drug discovery/development process. 4 The Innovative Medicines Initiative (IMI), launched in 2008 to drive the development of new medicines, is currently the largest and most notable example of a PPP within the pharmaceutical sector. 5 It was formed between the European Federation of Pharmaceutical Industries and Associations (EFPIA) and the European Union (EU). IMI projects operate in four key areas, where there are limited incentives for private sector investments alone: tackling complex diseases where precompetitive consortia are necessary, accelerating disease understanding to a point where product development could be initiated, providing technology platforms where private and public organizations can pool resources to improve drug development, and addressing gaps in the drug discovery/development ecosystem. 6 Since its launch, IMI has funded 144 projects with 3127 participants and a budget of €5.3 billion. IMI projects cover a wide area of topics ranging from drug discovery and development of new medicines based on small molecules to vaccines, drug safety testing, clinical trial design, animal models, standards, biomarkers, standard operating protocols, screening platforms and their utility, tools, and new methodologies.
Several other precompetitive PPPs have also been formed to focus on optimizing technology platforms and research tools either for target selection or to advance the knowledge of diseases.1,4 The U.S. Food and Drug Administration (FDA) launched the Critical Path Initiative (CPI) 7 program in 2004 to drive innovation in the scientific processes for the development, evaluation, and manufacture of medical products. This led to the creation of a PPP—the Critical Path Institute (C-Path) 8 —in 2005 to develop precompetitive standards and approaches, which is achieved through an innovative, collaborative approach to sharing data and expertise. In 2012, the National Center for Advancing Translational Sciences (NCATS) 9 at the U.S. National Institute of Health (NIH) was established to transform the translational research pipeline and accelerate the delivery of new treatments. NCATS divisions have been organized to span the entire spectrum of translational science, from preclinical to clinical stages, across a wide range of human diseases.
PPPs have also been launched to advance research in specific disease and therapeutic areas or to achieve specific objectives like target identification, target validation, biomarker development, and drug efficacy and safety evaluation. A few examples of disease and therapeutic area-specific PPPs are the Coalition Against Major Diseases (CAMD), 10 Accelerating Medicines Partnership (AMP), 11 Alzheimer’s Disease Neuroimaging Initiative (ADNI), 12 Dundee Kinase Consortium, 13 National Cancer Institute (NCI) Community Cancer Centers Program (NCCCP), 14 Drugs for Neglected Disease Initiatives (DNDi), 15 and European Rare Diseases Therapeutic Initiative (ERDITI). 16 CAMD was launched by C-Path to accelerate new treatments for neurodegenerative diseases leading to dementia like Alzheimer’s (AD) and Parkinson’s (PD) diseases. On its 10th anniversary in 2018, CAMD was rebranded as Critical Path for Alzheimer’s Disease (CPAD) to primarily focus on AD and has achieved quite a few milestones, including the first-ever qualification opinion by the European Medicines Agency (EMA) to use low hippocampal volume for patient enrichment in predementia clinical trials and the first-ever drug–disease trial model, as well as clinical trial simulation tool, to meet FDA and EMA standards. 17 AMP was launched in 2014 as a PPP between the NIH, the FDA, several non-profit organizations, and biopharmaceutical/life science companies to focus on three disease areas: AD, type 2 diabetes, and autoimmune disorders like rheumatoid arthritis (RA) and systemic lupus erythematosus (lupus). The main objective of this initiative has been the identification and validation of promising new biological targets of these diseases for the development of novel diagnostics and therapeutics. 18 An AMP project for PD was launched in January 2018 and more than US$350 million has been provided as the current 5-year funding commitment. 11 The four ongoing AMP projects have already begun to deliver results, and an offshoot—Partnership for Accelerating Cancer Therapies (PACT)—has also been launched in 2017 by the NIH and 12 drug companies to validate biomarkers for cancer immunotherapies. 19 ADNI was launched in 2003 to accelerate drug development by validating imaging and blood/cerebrospinal fluid biomarkers for AD clinical treatment trials. 20 The first 5-year phase (ADNI1) began in 2004 with a budget of US$67 million to develop biomarkers as outcome measures for clinical trials; it was completed in 2010.12,20 ADNI entered its second phase (ADNI2) in 2011 with an additional $67 million in funding, and ADNI3 began in 2016 with an expanded goal of determining the relationships between the clinical, cognitive, imaging, genetic, and biochemical biomarker characteristics across the entire spectrum of AD. 12 ADNI has developed standardized biomarkers for use in clinical trial subject selection and as surrogate outcome measures and generated data used in more than 600 publications, leading to the identification of novel AD risk alleles and an understanding of the relationship between biomarkers and AD progression.21,22 The University of Dundee’s Division of Signal Transduction Therapy (DSTT), also known as the Dundee Kinase Consortium, was founded in 1998 to explore new approaches to treat diabetes, cancer, and arthritis.4,13 Almost £60 million has been invested in this long-standing collaboration between academia and industry, with another £7.2 million to be provided by 3 major pharmaceutical companies to support fundamental research in multiple therapeutic areas until 2020. The collaboration has led to the development and clinical approval of more than 40 drugs targeting kinases, mostly as therapeutics for cancer. 23 The NCI launched NCCCP as a PPP with community hospitals in 2007 to advance cancer care and research, and the pilot phase ended in 2010. 24 Since 2014, NCI has replaced the NCCCP by integrating aspects of it into the newly formed NCI Community Oncology Research Program (NCORP).25,26 DNDi, founded in 2003, constitutes an example of a product development PPP to address all aspects of drug discovery and development for neglected tropical diseases (NTDs) like malaria, leishmaniasis, African trypanosomiasis, lymphatic filariasis, and Chagas disease. It is structured like a virtual biotechnology company for hit identification and lead optimization. 27 DNDi has developed eight treatments for five deadly diseases 15 since 2003 and is currently working on over 40 projects, including more than 20 novel chemical entities and conducting over 20 clinical trials. 28 ERDITI is a PPP established to advance the development of drug treatments for rare diseases. It is sponsored by the European Science Foundation and coordinated by the French Institute of Rare Diseases Research with four major pharmaceutical partners. 29 The main objectives of ERDITI are to provide access to compounds developed by pharmaceutical companies, facilitate efficient collaboration between public and private partners, and support research efforts from the preclinical phase until drug development and marketing.16,29 The NIH and FDA have also worked extensively in order to expand collaborative opportunities and establish PPP initiatives to meet the therapeutic needs of the rare diseases community. 30
Other examples of PPPs established to support collaborative research with specific objectives or to address specific challenges during the drug discovery process include the Biomarkers Consortium, 31 SNP Consortium (TSC), 32 RNAi Consortium (TRC), 33 Center for Therapeutic Target Validation (CCTV), 34 Predictive Safety Testing Consortium (PSTC) 35 and IMI eTOX 36 project. The Biomarkers Consortium, managed by the Foundation for the National Institutes of Health (FNIH), brings together resources and expertise to rapidly discover, develop, and qualify biomarkers to support drug development, medical diagnostics, and clinical care and improve regulatory decision-making.4,31 TSC was launched in 1999 by 10 leading pharmaceutical companies and five leading scientific laboratories to map single-nucleotide polymorphisms (SNPs) in the human genome. 37 Although the initial goal was to map 300,000 SNPs in 2 years, 1.4 million SNPs had been released to the public domain by 2001. 38 TRC was launched in 2007 to create specific RNA interference (RNAi) inhibitors for targeting human and mouse genes and successfully built a 160,000 short-hairpin RNA (shRNA) library targeting 15,000 mouse and 15,000 human genes during the first phase.39,40 TRC is now part of the Genetic Perturbation Platform (GPP) 41 and available via the GPP web portal. 42 CTTV was formed by GSK, the Wellcome Trust Sanger Institute, and the European Bioinformatics Institute (EBI) to combine resources for the discovery of potential drug targets and target validation. 34 It has been renamed the Open Targets Consortium43,44 and serves as a platform to collect, organize, and exploit information in the scientific literature to identify novel drug targets. 45 The Open Targets Consortium hosts the Open Targets Platform, 46 which integrates data from the public domain to allow target identification and prioritization, and the Open Targets Genetics Portal, 47 which identifies targets based on genome-wide association studies (GWAS) and functional genomics. PSTC was launched in 2006 to bring together pharmaceutical companies to share and validate innovative safety testing methods under the advisement of three regulatory agencies: the FDA; its European counterpart, the EMA; and the Japanese Pharmaceutical and Medical Devices Agency (PMDA). 35 PSTC’s current emphasis is on developing and obtaining regulatory qualification of improved clinical safety biomarkers that would be used in drug development. PSTC serves as a third party to assess drug safety tests, facilitate the exchange of resources and expertise, and promote collaborative interactions to collect and summarize data.10,35 The eTOX project was launched in 2010 as an IMI consortium between 13 pharmaceutical companies, 11 academic institutions, and 6 small- and medium-sized enterprises (SMEs). 36 A major objective of the project was to develop innovative strategies and novel software tools to predict the in vivo toxicology of new chemical entities using information already available in the early drug development phases. 48 The consortium implemented the largest database of preclinical safety data, created ontologies to support chemistry- and biology-based searches, and developed 90 in silico models for safety prediction. 49 The project launched the eTOXsys 50 platform to access the generated data and models and was successfully concluded in 2018. 36
Access to biological materials like cells, tissues, or bodily fluids for investigations and research is a prerequisite for the preclinical phase of the drug development process. Numerous initiatives have emerged in recent years to facilitate structured access to quality-controlled patient or donor samples in a way that is ethically appropriate as well as privacy compliant. Large repositories of well-annotated biological specimens are often referred to as “biobanks” and can be heterogenous in terms of their design and uses.51–53 Although several bottlenecks exist for establishing PPPs in biobanking,53,54 their beneficial impact on the sustainability of biobanks has been considered.53,55 Currently, there are three leading international entities governing biobanking: the International Society for Biological and Environmental Repositories (ISBER), the Biobanking and BioMolecular Resources Research Infrastructure–European Research Infrastructure Consortium (BBMRI-ERIC), and the European, Middle Eastern and African Society for Biobanking (ESBB). 51 The pan-European BBMRI-ERIC constitutes a research infrastructure for biobanking and brings together all the main players in this field. 56 BBMRI-ERIC has also developed the concept of Expert Center as a PPP to serve as a key intermediary between public and private sectors engaged in the analysis of biological samples under internationally standardized conditions. 57 A recent study has been performed to examine the progress of PPP development involving academic biobanks in the European biobank field, focusing on the community established by BBMRI-ERIC. 58
Table 1 provides a summary of the various PPP examples in the order these are discussed in this review, along with their main objective/area of focus and web links.
A Summary of PPP Examples and Their Main Focus/Objectives and Web Links.

Compound-Sharing Partnerships
The sharing of physical compound samples as well as compound-associated data is a common form of contribution from pharmaceutical companies to precompetitive collaborations. Such shared assets range across therapeutic areas and phases of research and development.
Preclinical Compounds
Initiatives where screening compounds are shared or swapped between two pharmaceutical companies provide an opportunity for each partner to access chemical space beyond their own compound collection. The two principal ways of accessing compounds through a swap have been by either sending physical samples of compounds to the partner or transferring the assay for screening at the compound owner’s facility. Physical swapping of samples was the basis of an AstraZeneca-Sanofi collaboration where 210,000 compounds were exchanged between the companies. 59 When the two pharmaceutical companies compared their compound collections, it was found that the structural overlap was small and that the collections complemented each other. 60 An example of the latter was the AstraZeneca-Bayer Boomerang collaboration. 61 Assays were transferred from the target owner to the compound owner, and screening data had to be analyzed by the compound owner. The structures of the hit compounds were shared with the target owner after successful completion of the workup. Such a setup requires that one company (i.e., the target owner) relies on the other company to perform their data analysis for them to the best of their abilities. Since the compound owners can apply internally available computational workflows in the triage, data sets can be handled with the same scrutiny and level of effort as is applicable for an internal project. Although the aforementioned examples are pharma–pharma partnerships and thus outside the PPP framework, these have been included to illustrate the types of assets that could be shared within a collaboration.
Pharmaceutical companies have also generated publicly available screening libraries, like the Tres Cantos Antimalarial Set (TCAMS), which was compiled by GlaxoSmithKline (GSK). This collection contains more than 14,000 antimalarial compounds that were identified after screening 1.9 million compounds from the GSK corporate compound collection. TCAMS has been widely used across the malaria community with good success.62–69
For example, a TCAMS screen identified 14 primary sulfonamides that inhibited the in vitro growth of
Since
Novartis has also made available a large collection of malaria screening data (5600 compounds tested in dose–response and confirmation assays) known as the Novartis-GNF Malaria box. 72 However, these compounds were not always available for follow-up screening. Therefore, in 2011 Medicines for Malaria Ventures (MMV) made a compilation, named the Open Access Malaria Box, from Novartis, St. Jude Children’s Hospital, Memphis, and the GSK compound collections that focused on commercial availability. 73 The set represented 400 different chemotypes, 200 of which were picked for their druglike properties and the remaining were selected as chemical probes. In addition to malaria, the Malaria Box was also a useful source of leads for other neglected diseases like giardiasis, lymphatic filariasis, toxoplasmosis, and animal piroplasmosis.74–76
Building on the success of the Malaria Box, the Pathogen Box was created, covering hits for tuberculosis, malaria, kinetoplastids, helminths, cryptosporidiosis, toxoplasmosis, dengue, and other reference compounds. As an example, the Pathogen Box, together with FDA-approved compounds, was screened against
The Malaria and Pathogen Boxes are no longer physically available since the samples of constituent compounds and funding for preparation and dispatch have been consumed. However, they have been widely successful, resulting in more than 90 publications so far. 78 In 2019, MMV and DNDi made available the Pandemic Box—400 diverse druglike molecules active against bacteria (201 compounds), viruses (153 compounds), or fungi (46 compounds). The boxes have been made freely available given that any results would be made publicly available in an open-access publication within 2 years.
The IMI consortium-driven European Lead Factory (ELF) 79 project provides a unique example of creating a joint screening compound collection through PPP. It consists of well-known pharmaceutical partners, academic labs, the Joint European Compound Library (JECL), and the European Screening Centre. 80 The compound library is available to partners for screening without initial disclosure of molecular structures and is open to any EU researcher from academia as well as SMEs that have a potential drug target or screening assay available, with all costs covered by IMI.
During its first iteration, launched in 2013, the JECL consisted of two parts. The first set was a library of 321,000 compounds contributed by the seven participating pharmaceutical companies. 81 The second part of the compound collection was composed of 200,000 novel compounds designed and synthesized by ELF public partners, as an effort toward increased coverage of chemical space that was more complex and natural product-like. 82 Many academic groups got the opportunity to screen the JECL without any cost via a target nomination and selection process. The identified hits were also further developed within the ELF project. The participating pharmaceutical companies had the opportunity to later license the compounds from the academic projects. The screening activities have resulted in spin-off companies like Keapstone Therapeutics and licensing agreements as exemplified by Servier and Scandicure.83,84
Technical and operational solutions can facilitate compliance to agreed processes, as well as streamline data flows. Several cloud-based commercial computational and data storage platforms suitable for collaboration are also available, such as the BIOVIA ScienceCloud and CDDVault.85,86 An example can be provided from the ELF, where access to data throughout the life cycle of screening programs, program management and audit, and communication between ELF partners are mitigated through BIOVIA’s Honest Data Broker (HDB) system. 87 The data analysis, also enabled by functionalities in the HDB environment, is done by an ELF program team member who does not have access to view chemical structures but relies on screening results and metadata. An experienced medicinal chemist engaged by each program team works as an independent consultant and is allowed to view a limited number of compound structures without disclosing them to the program team, and ultimately requests the final list to be disclosed. After the request for structure release has been submitted, the compound owner is notified through the HDB and has a time window available to perform any necessary internal intellectual property (IP) reviews. In the last step, structures are released and/or rejected through the HDB and shown to the program team members. Upon release, the name of the compound owner is also disclosed, using the HDB functionalities.
The Structural Genomics Consortium (SGC) 88 is a nonprofit PPP created to determine protein structures that represent potential targets for drug discovery endeavors. It is supported by several pharmaceutical companies, including AbbVie, Bayer, Boehringer Ingelheim (BI), Janssen, Merck Sharp and Dohme (MSD), Pfizer, and Takeda. The SGC also provides the Chemical Probes 89 set, which is a collection of potent, selective, and well-characterized small-molecule modulators, well suited for phenotypic screens. It also contains compounds donated by pharmaceutical partners. 90 This set includes associated data, inactive structural analogs, and co-crystal structures. 91 Each probe molecule meets stringent quality criteria, has been characterized in a panel of more than 500 assays, and is available as an open-source reagent that can be requested for biomedical research. As a recent example, a probe molecule for PRDM-9 was discovered in a collaboration between the SGC and MSD. 92
An open PPP was also proposed to identify chemical probes functionally annotated and to broaden the biological evaluation as well as pharmacological understanding of the untargeted kinome. 93 This led to the creation of an annotated set of 367 small-molecule ATP-competitive kinase inhibitors called the GSK Published Kinase Inhibitor Set (PKIS). 94 The PKIS compounds have been extensively utilized by the scientific community and profiled in activity assays with 224 recombinant kinases and 24 G protein-coupled receptors, leading to the identification of chemical starting points for hitherto untargeted kinases (such as LOK, BRSK1/2, MKNK1, and AAK1) with modest affinity values and selectivity profiles. 95 SGC and GSK have also expanded PKIS to create PKIS2 for the identification of novel kinase drug targets. 96
Several other examples of smaller compound-sharing collaborations have taken place, and also for more advanced preclinical as well as clinical-stage small and large molecules, for the testing and validation of scientific hypotheses. For example, BI has made available its selective protein tyrosine phosphatase nonreceptor type 5 (PTPN5; STEP) allosteric inhibitor tools for use by the scientific community. 97 Dysregulation of STEP has been linked to neurodegenerative and neuropsychiatric diseases, highlighting this enzyme as an attractive therapeutic target for drug discovery.
Clinical-Stage Compounds
Sharing of clinical-stage compounds across the pharmaceutical sector is heavily skewed toward investigator-sponsored studies involving marketed drugs, or those that are actively pursued in pharmaceutical development. Such studies are often supported by project teams that seek out external collaborations to broaden knowledge and data packages around the asset and to explore life cycle management opportunities. While such studies are led by the academic collaborator, project teams from the industry partner are available not only to provide access to the preclinical/clinical supply needed to support the study, but also to aid compound information, regulatory data packages, safety reporting, and other study-related requirements. Many of these studies are funded by the industry, although strategic alliances with funding bodies have also been established to encourage wider availability of clinical assets for research into areas of unmet need. 98
The government funding body–pharmaceutical company partnership model was pioneered in 2011 when the UK Medical Research Council (MRC), in partnership with AstraZeneca, launched the “Mechanisms of Disease” initiative, 99 with the aim of enhancing the understanding of human disease through experimental medicine studies. This was followed in 2012 by the “Discovering New Therapeutic Uses for Existing Molecules” repurposing initiative, started by the NIH. 100 The Pharmaceutical Assets Portal project initiated by the Clinical and Translational Science Award (CTSA) consortium has also worked toward creating a multistakeholder PPP to facilitate drug repositioning efforts. 101 Most recently, a similar scheme—the “Clinical Compound Bank”—was piloted through the IMI across the EU.102,103 These schemes operate according to standardized industry–academic collaboration agreements. Preexisting IP stays with the company or collaborating institution, while any new IP is owned by the institution, with the company having first right of refusal to negotiate a license. Several industrial partners have also shared discontinued clinical assets through such schemes for both preclinical and clinical studies. The assets companies make available are often supported by previous clinical data, thus offering a potential fast route to market to benefit patients in alternative indications. Unfortunately, a lack of dedicated industry resource to support studies with discontinued compounds has prevented some companies from sharing their assets for clinical studies. Funders are now offering schemes that cover supportive studies such as toxicological assessment, as well as the costs associated with drug manufacture and supply.100,104 These include the highly competitive MRC Development Pathway Funding Scheme (DPFS) and the NIH New Therapeutic Uses program. These schemes provide academics with real opportunity to translate academic discoveries toward the clinic, as well as aim to reduce some of the challenges faced by pharmaceutical companies when sharing assets for studies that are driven by scientific need rather than alignment with company strategic interest. Some examples of clinical research that has been driven forward using the government funding body–pharmaceutical company partnership model include 11β-HSD1 inhibition in interstitial intercranial hypertension and Src kinase inhibition in AD and cancer-induced bone pain.105–107
Efforts have also been made by SGC toward a pharma-backed PPP known as ArchPOCM with the goal of improving efficiency and lowering the drug development expenditure by generating a portfolio of small molecules that hit new therapeutic targets as well as carry out early clinical work up to phase II trials. 108 The core tenet of this project was to create an IP-free open-sharing environment and extend precompetitive cooperation to establish phase II clinical proof of mechanism for novel targets. Despite lack of success in launching ArchPOCM due to insufficient support from the industry, a smaller-scale project without any pharmaceutical backing was launched to identify a candidate compound effective against KDM4B, which is implicated in people with breast cancer. 109
PPP to Develop ML Models
Computational modeling collaborations, especially targeted toward machine learning (ML) and deep learning (DL), have gained popularity in the last few years. 110 For instance, such collaborations have been formed around data access, method development, and model building. Both high-quality and large volumes of data are needed to build accurate ML models. Since data are considered to be one of the most valued assets within a company, direct data sharing to build models is not a viable option in many cases. To overcome this challenge, a technique called privacy-preserving ML has been developed, where transfer learning is used to improve ML models without sharing underlying data. For example, it can be used to improve the accuracy of models for predicting biological activity and physicochemical properties of small molecules. In 2019, an IMI2 project, Machine Learning Ledger Orchestration for Drug Discovery (MELLODDY), 111 was launched to achieve this objective. The consortium consists of 10 pharmaceutical partners and 7 public partners and will be running for 3 years. There will be yearly executions of the ML workflow build to assess how much the models will be improved for a pharmaceutical partner when trained with privacy-preserving access to data from 9 other companies.
A pharmaceutical company might also want to leverage the expertise within the community for building ML models on their data. The Kaggle platform was introduced to fill this need, and competitions involving data from participating companies have been launched continuously. 112 The Merck challenge has been the most widely known Kaggle competition, which used proprietary bioactivity data from a pharmaceutical company. 113 The outcome was published in a scientific article, and a DL architecture gave the best results. 114 As ML algorithms are costly to develop and test, several consortia have been founded with the mission to precompetitively develop these for drug discovery. The three most well known are the MLDPS 115 consortia based at MIT, RETHINK 116 at ETH, and the ATOM 117 consortium. Of these, MLDPS has the largest number of pharmaceutical companies involved as partners. It has recruited PhD students and postdoctorates to develop novel methods in compound synthesis and property prediction. The participating companies get early access to the developed software, which is later open sourced to benefit the whole scientific community. ATOM consortium has developed the ATOM Modeling PipeLine (AMPL)—an end-to-end modular and extensible software pipeline to build and share ML models for a wide array of molecular properties that are required for in silico drug discovery. 118
Data and Intellectual Property Sharing
Data integration is important for PPP since all pharmaceutical companies are integrating external public data with their in-house data assets. Therefore, there is a need for both common data standards and precompetitive efforts to make data easily interoperable. An early project that addressed this need was the IMI OpenPhacts project. 119 Building on this, and the need for machine-readable data that has emerged with the enablement of high-throughput data science, the FAIR (Findable, Accessible, Interoperable, Reusable) principles were published in 2016. 120 Many precompetitive initiatives have arisen that promote FAIR data, some of which are run within PPPs, and others are facilitated by nonprofit organizations or other organizational bodies. As an example of the former, the IMI has launched the IMI2 FAIRplus project, which aims to apply the FAIR principles to data generated within IMI. 121 Several examples of the latter are run within the Pistoia Alliance, 122 which is a pharma-funded nonprofit organization that enables precompetitive collaborations. The FAIR Implementation Project, which has developed a publicly available toolkit, is one such example. 123 Others are the Ontologies Mapping Project, which is implemented in collaboration with EMBL-EBI, and the DataFAIRy project, which in its first phase aims at increasing the body of publicly available FAIR metadata for published bioassays.124,125 Pistoia Alliance has also worked with Imperial College London (ICL) and the University of Michigan (U-M) to bring together resources to form the tranSMART Foundation 126 —an open-source PPP community to develop a comprehensive informatics-based data-sharing and analysis cloud platform for clinical and translational research. 127 The first iteration of tranSMART was developed by Johnson & Johnson (JnJ) in 2010 to serve as an internal research data warehouse, and Janssen R&D, Inc., a JnJ company, made it open source in 2012.128,129 The Allotrope Foundation is another non-PPP collaborative platform that brings together players from industry and aims at developing ontologies, data models, and a universal format for storing experimental data. 130 Two additional non-PPP foundations relevant to the pharmaceutical sector are Lhasa Limited which focuses on data sharing mainly related to safety, and the Cambridge Crystallographic Data Centre (CCDC), which specializes in crystallographic data and related software development.131,132
In addition to compound sharing, a PPP also provides excellent collaborative instances to support the sharing of additional data. An example is the Shared Platform for Antibiotic Research and Knowledge (SPARK) initiative, which is run by the Pew Trust. 133 The platform contains curated published data as well as data sets from pharmaceutical companies. Novartis 134 and Achaogen 135 have both contributed to SPARK by providing data sets stemming from their own discontinued antibacterial programs.
Leading pharmaceutical companies, product development partnerships, and public organizations have joined with World Intellectual Property Organization (WIPO) to establish the WIPO Re:Search consortium. The main objective has been to offer a mechanism for sharing IP that has the potential to be used in the discovery and development of new, more effective products to treat neglected diseases.136–138 WIPO Re:Search is sponsored by WIPO, which is a United Nations (UN) agency, and is administered by the nonprofit BIO Ventures for Global Health (BVGH). It consists of three components: an IP database available for licensing from a provider hosted by WIPO; a Partnership Hub managed by BVGH to provide information about the consortium’s licensing, research collaboration, funding, and networking possibilities; and services to support and facilitate the negotiation of licensing agreements. WIPO Re:Search has started more than 140 collaborations so far, and over two-thirds of these are R&D projects.
139
For example, researchers at the University of California validated
As a second example, in malaria research, academic researchers at Liverpool School of Tropical Medicine (LSTM) found that inhibition of the protease-activated receptor 1 (PAR1) pathway might reduce brain swelling in cerebral malaria, an often fatal complication of
Conclusions and Future Outlook
The emergence of diverse PPPs and consortia during the last decade has led to enhanced precompetitive research in various phases of the drug discovery and development process. 140 During this period, the sharing of assets, often through PPPs, has become an established tool for pharmaceutical innovation, which can deliver increased value throughout drug discovery and development. Here, we have illustrated these opportunities with several examples of asset sharing in the pharmaceutical industry and discussed PPPs that have successfully brought together the positive aspects of academia, industry, and government to drive innovation. Nevertheless, it is important to note that examples of PPPs in drug discovery that either have failed to launch (with the exception of ArchPOCM) or provide any return on investment (ROI) have not been reported in the public domain.
PPPs bring together the best of both academic and pharmaceutical worlds by advancing academic discoveries and biological insights, and translating them to working therapies for treating diseases via industry resources. These also enable the building of collaborative networks between academic partners. 141 Irrespective of their overall goals, PPPs often have to face some intrinsic challenges, such as, sustainability with respect to funding and delivering outcomes of high impact given the long timelines for drug development and the establishment of a clear framework.142,143 Performance measurement systems to evaluate success and long-term impact are a key challenge, and implementing a system endorsed by multiple stakeholders can be difficult.142–144 Recently, various frameworks and performance indicators to assess the value of PPPs in the pharmaceutical sector have been proposed.144–146 Although there are fundamental difficulties in quantifying the impact of biomedical PPPs, there is clear evidence that precompetitive PPPs like SGC, ADNI, and IMI have started to generate substantial outputs, in the form of, publications, patent applications, tools, open-source databases, and in silico and animal models.144,147 However, due to the intangible nature of many outcome indicators, the true measure of success for PPPs with multiple stakeholders would have to be measured by suitable proxies. 144 It must also be noted that research impact assessment is a relatively novel field in scientific endeavor and assessments are often based on mixed methodologies.144,148 Improvements in metrics to measure PPP performance and outcome could also help identify the various factors influencing the ROI and allow quantitative assessments to understand their true impact.
While it is difficult to assess the financial impact and enhanced external reputation gained by PPPs, an increase in funding from governments as well as pharmaceutical companies strongly indicates the high level at which such initiatives are considered valuable. It is expected that the impact of PPPs will grow in the future considering the increase in both data that need to be analyzed and continuous public investment in research. The current global public health crisis due to COVID-19 is an excellent case to demonstrate how PPPs can help provide a collaborative platform to accelerate the development of therapies to prevent or treat a serious pandemic. The response in initiating public and private enterprises to meet the public health crisis has been rapid and unprecedented in its scale during peacetime. Projects initiated across the globe range from clinical trials to repurposing marketed drugs or those in early- or late-stage development to vaccine development and new drug discovery activities. 149 For example, in drug repurposing, the UK’s 1800-patient ACCORD trial in hospitalized patients with COVID-19 is setting out to test up to six agents initially that could then be advanced into larger trials, including nebulized heparin, Ra Pharmaceuticals’ complement C5 inhibitor zilucoplan, BerGenBio’s AXL inhibitor bemcentinib, and AstraZeneca’s IL-33 mAb MEDI-3506 and BTK inhibitor acalabrutinib. In vaccine development, Accelerating COVID-19 Therapeutic Interventions and Vaccines (ACTIV) is a PPP led by the NIH. 150 The NIH has partnered with its sister agencies, such as the FDA, the Centers for Disease Control and Prevention (CDC), and other U.S. government departments; the EMA; representatives from academia and philanthropic organizations; and more than 15 biopharmaceutical companies. This forum allows for discussions and consensus on vaccine trial designs, rapid data sharing, and close collaborations between the public and private sectors to rapidly and efficiently conduct vaccine efficacy studies. The IMI2 call “Development of Therapeutics and Diagnostics Combatting Coronavirus Infections” yielded 144 project proposals, of which 8 were funded to a total of €72 million. 151 These include five diagnostics and three treatment projects, involving 94 organizations, including universities, research organizations, companies, and public bodies. SMEs are particularly well represented in the successful proposals, accounting for more than 20% of the participants and 17% of the budget. 152
Although it is difficult to estimate the monetary value of precompetitive collaboration, the accelerated collaborative response to COVID-19 is a clear piece of evidence that the effective use of combined resources can truly advance medicine and benefit public health on a global scale.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
