Abstract
During the past decade, mass spectrometry imaging (MSI) has become a robust and versatile methodology to support modern pharmaceutical research and development. The technologies provide data on the biodistribution, metabolism, and delivery of drugs in tissues, while also providing molecular maps of endogenous metabolites, lipids, and proteins. This allows researchers to make both pharmacokinetic and pharmacodynamic measurements at cellular resolution in tissue sections or clinical biopsies. Despite drug imaging within samples now playing a vital role within research and development (R&D) in leading pharmaceutical companies, however, the challenges in turning compounds into medicines continue to evolve as rapidly as the technologies used to discover them. The increasing cost of development of new and emerging therapeutic modalities, along with the associated risks of late-stage program attrition, means there is still an unmet need in our ability to address an increasing array of challenging bioanalytical questions within drug discovery. We require new capabilities and strategies of integrated imaging to provide context for fundamental disease-related biological questions that can also offer insights into specific project challenges. Integrated molecular imaging and advanced image analysis have the opportunity to provide a world-class capability that can be deployed on projects in which we cannot answer the question with our battery of established assays. Therefore, here we will provide an updated concise review of the use of MSI for drug discovery; we will also critically consider what is required to embed MSI into a wider evolving R&D landscape and allow long-lasting impact in the industry.
Introduction
The essence of pharmaceutical research and development (R&D) is the selection of an effective drug molecule that can reach the right target receptors in the right tissue at a sufficiently high concentration to deliver required efficacy, while simultaneously avoiding any induced toxicity. 1 This is achieved through deployment of a battery of techniques and technologies that measure the abundance, metabolism, and distribution of the active pharmaceutical ingredient (API) and relate to measurements of any induced cellular response or unwanted toxicity, typically measured in tissue samples. In drug discovery, plasma concentration measurements have traditionally been used as a surrogate for the concentration of drug in tissues, and pathophysiology has been used to image cellular changes. This can provide an inaccurate or incomplete picture of the molecular events happening in the sample.
Mass spectrometry is a powerful analytical technique, and when operated as an imaging modality it allows molecular maps to be visualized in 2D and 3D and at the nano- to the macroscale. Mass spectrometry imaging (MSI) is a label-free technique capable of simultaneous visualization of the molecular distribution of endogenous compounds, 2 drugs, 3 lipids, 4 proteins, 5 peptides, 6 and drug delivery systems 7 in biological tissues. The process involves ionization of the sample from a known location, moving the tissue, and repeating the ionization process. A mass spectrum is acquired at every ionization location or “pixel,” and linking the abundance of detected ions to a spatially resolved pixel map of the surface of the sample allows images for every detected molecule to be reconstructed.
The array of technologies applied to perform MSI has continued to grow during the past decade. Instruments now routinely deliver the required acquisition speed, sensitivity, and specificity researchers require to support many aspects of drug discovery and development. While performance has significantly improved during the past decade, as will be discussed later in more detail, there remain many challenges and opportunities within the expanding scope of MSI. Platforms vary but typically either are capable of broad applicability or are optimized to deliver a specific task at the expense of wider applicability. The selection of MSI platform is therefore best determined by the application. There are, however, common themes around sample collection, processing, and storage that are agnostic of MSI platform. There are also common challenges concerning scope to scale MSI study size, data analysis, and integration with other imaging modalities and wider analytical data streams. We will therefore start by briefly considering the key technologies and review the major components in sample preparation and analysis pipeline. We will then outline the main applications of MSI in pharmaceutical research before spending some time considering the challenges for MSI in delivering sustained impact, at scale, in pharmaceutical research.
Key MSI Technologies
MSI systems consist of two main parts, a sampling source that ionizes molecules and a mass analyzer that detects them. The performance of mass analyzer varies considerably and balances specificity (the ability to distinguish molecules of similar molecular weight) with accuracy and speed of analysis. The ionization method paired with any mass analyzer will significantly influence the sensitivity, spatial resolution, speed, and range of detected molecules. These have previously been reviewed in detail;8,9 however, brief consideration will be given here to their utility in pharmaceutical R&D. Matrix-assisted laser desorption ionization (MALDI) allows spatially resolved detection of small and large intact molecules from surfaces and thin tissue sections. The technique requires the application of an ultraviolet (UV)-absorbing chemical matrix to the sample surface. 10 MALDI is the most widely used MSI technology in industry and academia, and it can provide high spatial resolution analysis and high acquisition speeds. The range of detected endogenous metabolites or drugs is far from exhaustive, however, even when using high-mass-resolving mass analyzers. Therefore, alternative ionization methods are a powerful addition to any pharma lab conducting MSI. The most complementary method is desorption electrospray ionization mass spectrometry imaging (DESI). 11 This ionization method overlaps with many of the molecules detected by MALDI MSI, allowing cross-validation of results, but has been proven to be alternatively sensitive for a range of different endogenous and exogenous targets. 12 In contrast to most commercially available MALDI platforms, DESI operates under atmospheric conditions and requires no sample processing other than standard fresh-frozen tissue sectioning. Currently, the only major limitation in DESI MSI is the spatial resolution achievable, which is typically not greater than 30 µm. This limits its application to projects requiring cellular resolution imaging in fine structures like intestinal villi 13 or lung tissues. 14 It has been extensively demonstrated, however, that what it lacks in spatial resolution it makes up for in detected therapeutics and overall sensitivity.3,15,16
For those researchers seeking the highest-resolution MSI, secondary ion mass spectrometry (SIMS) can be used to perform molecular imaging at subcellular lateral resolution, taking MSI beyond tissue-wide imaging to an organelle-length scale. There are three main variations of SIMS to consider. Most widely used is time-of-flight secondary ion mass spectrometry (ToF-SIMS), which uses a high-energy focused ion beam to desorb biomolecules from the surface samples. While ToF-SIMS imaging is comparable to MALDI imaging in sensitivity for some target species, it can undertake 3D imaging by using the ion beams to remove surface material sequentially from imaging acquisitions. This allows the compilation of 3D MSI maps of the tissue or sample. ToF-SIMS is not able to match MALDI imaging in the range of intact molecules detected or the speed of acquisition.
The highest spatial resolution of any MSI technologies is provided by NanoSIMS, a nanoscale imaging that achieves unparalleled lateral resolution through use of a targeted labeling strategy. This requires the detection of specific elements or stable isotopes, but allows highly successful hypothesis-led molecular biological studies. NanoSIMS has many diverse applications, including microbiology, cell biology, pharmacology, geology and space sciences, material research, nanotechnology, plant sciences, and beyond. Unlike other MSI techniques that seek to preserve and detect whole molecules, NanoSIMS uses energetic reactive primary ions to break the molecule in the sample. Since the different isotopes of one element can be mapped, it is possible to measure the local isotopic enrichment of a molecule labeled with an isotope at high resolution. This successfully allows NanoSIMS to map the distribution of labeled molecules in tissues or cells at 50 nm resolution.
A recent addition to the SIMS technology stable is OrbiSIMS. This new instrument combines a ToF mass analyzer for high-speed analysis and the Orbitrap (Fourier transform mass spectrometer). This ultra-high-resolution hybrid analyzer is ideally suited for imaging complex biological samples. 17
A range of other MSI ionization methods are used to support the MSI mainstays of MALDI, DESI, and SIMS; their main aim is to aid pharma MSI by providing further complementary data. To date, none of these platforms are used in isolation for meaningful stand-alone R&D but are powerful bolt-on capabilities. For example, liquid extraction surface analysis (LESA) has been demonstrated to add extra sensitivity at the expense of spatial resolution18–20 when applied to metabolic profiling, 21 drug imaging, 3 or challenging intact protein targets.22,23 Combining similar surface droplet microjunction techniques with pre-ionization chromatography increases the specificity and range of detected targets, but compromises on overall throughput. 24
There are extensive other ionization methods that readers could continue to consider for future roles in MSI in R&D, including variations on the themes just discussed [such as laser ablation electrospray ionization 25 (LAESI), laser desorption ionization 26 (LDI), or NanoDESI 27 ] or more tangential techniques such as pixel imaging mass spectrometry. 28 Implementation within the pharmaceutical industry for many of these technologies is currently challenging, however, usually due to high costs, restricted project demand, and applicability, or due to limited commercial availability.
While MSI is a powerful label-free technology, it is worth noting that an exciting emerging field of research is providing deep metabolic tissue imaging with deep cellular phenotyping. This is achieved by using mass spectrometry to detect specially labeled multiplex immunohistochemistry (IHC) alongside MSI. The principle has long been published, 29 and the use in standalone multiplex histology has gained high-impact recognition.30,31 While to date there has been little published pharma-generated literature, 32 there are multiple pharma companies actively deploying this technique within their business alongside MSI platforms. Using a complementary imaging technique in a study exemplifies many of the new challenges for those working in the drug discovery MSI community, specifically data analysis, multidimensionality integration, and overall interpretation. While some challenges are overcome by new working groups, for example greater project pathologist alignment, many are still to be solved and will be considered later.
Sample Preparation and Processing Pipeline
Standard sample preparation and processing protocols for MSI were laid down many years ago, and we have previously reviewed them in detail. 33 Throughout the years, however, groups have explored extensive optimization and modifications of the sample collection, preparation, and processing prior to MSI analysis that have primarily targeted increased reproducibility and target analyte sensitivity. For example, pH-controlled sample washing 34 or buffer wash and a cyclohexane pre-extraction 35 have been demonstrated to increase MALDI sensitivity for pharmaceutical compounds, such as cimetidine, imipramine, and cobimetinib. DESI protein imaging has been shown to be improved by tissue washing. 36 MALDI matrix modification 37 or tissue derivatization methods for MALDI MSI38,39 have proven effective in increasing target sensitivity. Such approaches aid the wider applicability of MSI within pharma and beyond.
Two key factors in sample preparation and processing that are of utmost importance to wider expansion of MSI for pharma R&D are (1) sample stability to enable scalability and (2) exogenous and endogenous target quantitation. The latter has become a routine requirement in most studies, 40 with researchers using internal standards to correct for ionization suppression, calibration spots for drug quantitation, and deuterated endogenous metabolite analogues for metabolic quantitation.41–46 The issue of sample stability is critical to allow accurate measurement of exogenous and endogenous compounds. MSI-detected abundance has been demonstrated to decrease in tissue sections allowed to warm for moments prior to sample desiccation 47 and is even worse for endogenous targets, as exemplified by extensive work by the Andrén lab studying neuropeptides by MALDI-MSI48–50 and more recently by Swales et al. studying tumor metabolism. 51 The simplest way to mitigate such sample instability is to keep samples frozen and minimize the time when tissue sections are in the cryostat. This means that those wishing to perform larger studies, increasingly required to power efficacy studies, need to process all sections at the same time. The most effective way is to embed all samples in MSI-compatible media, and then all samples are exposed to the same cryostat conditions and subsequent slide conditions. To increase stability and minimize analyte delocalization, resulting from surface condensation following freezer retrieval, it is highly recommended that samples are vacuum packed following desiccation in the cryostat. 51 Heat stabilization has been muted on and off throughout the years; however, it is no longer recommended for those studying drug or metabolic targets. Heat stabilization has proven beneficial, however, for protein profiling using LESA-MSI. 23
Applications in Pharmaceutical Research
Traditionally, drug distribution studies were performed at later stages of the development pipeline, using techniques such as quantitative whole-body autoradiography (qWBA).52,53 Tissue abundance was determined by using tissue homogenization followed by liquid chromatography–mass spectrometry (LC-MS).54,55 Label-based techniques still have relevance in the progression of a compound, but they are often time-consuming, costly, or misleading due to the nonspecific method of detection being used. Radiolabeled drugs have the potential to undergo metabolism, and often their primary metabolites can contain the radiolabel; this can misrepresent the distribution profile of the drug, rendering the data ambiguous. Homogenization of tissues results in the loss of all spatial information, making it unsuitable for the analysis of drugs that are localized. It is therefore advantageous to have a label-free method of direct tissue analysis to elucidate drug distribution at an early stage in a drug’s life cycle to confirm drug delivery at the active site or to highlight drug accumulation in an organ in which toxicity may have been encountered. Therefore, MSI has really brought forward the ability to assess biodistribution of drug projects. As previously alluded to, the applications of MSI go well beyond mere compound mapping and quantification, although, as summarized in Table 1 , this is now readily done. The rationale for MSI investment is the additional applications to which the technologies can be applied to add value and rapid return on investment. We will now touch on a number of the diverse applications to which MSI can be applied.
Summary of Different qMSI Methods Used to Quantify Drugs in Tissues.
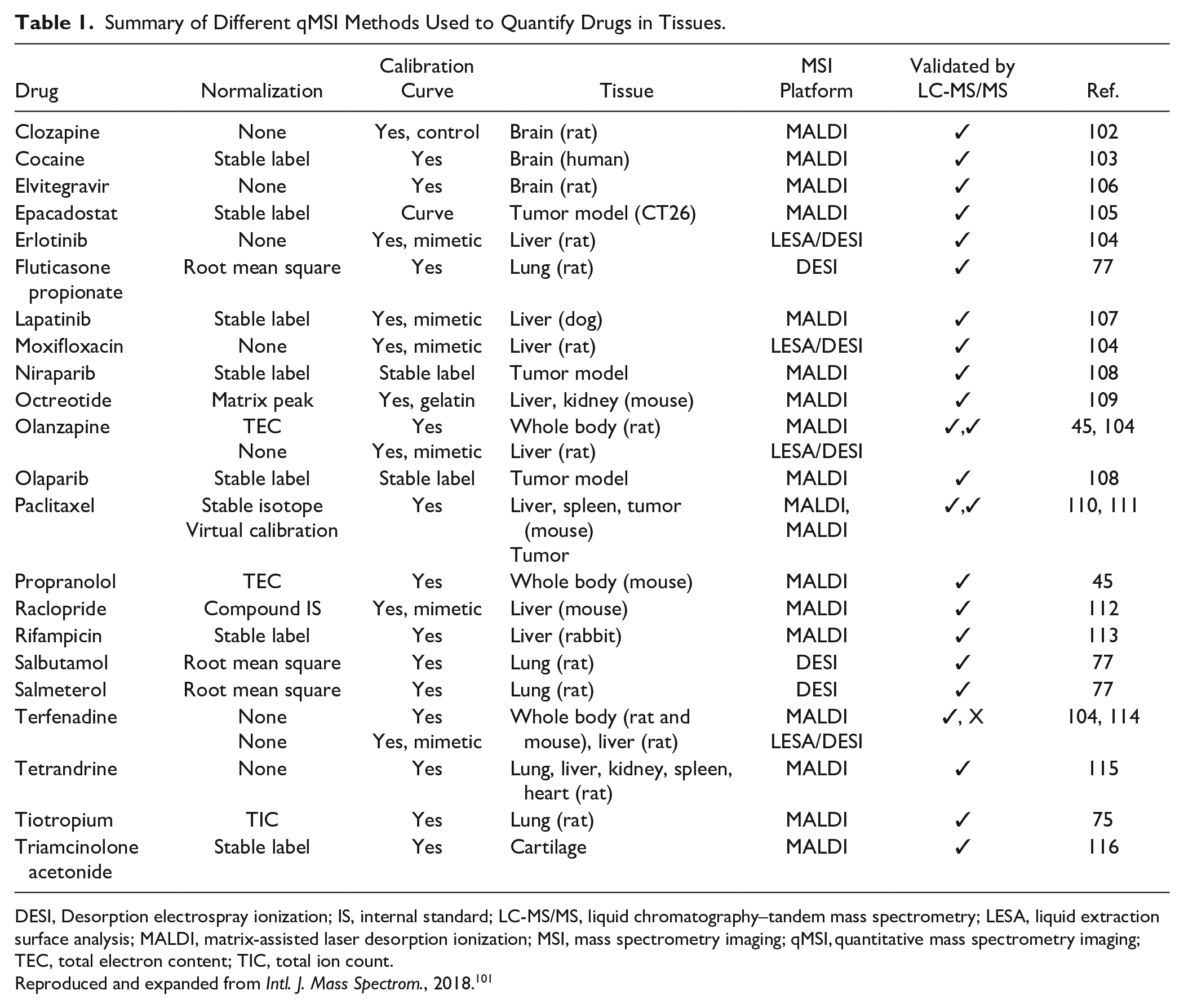
DESI, Desorption electrospray ionization; IS, internal standard; LC-MS/MS, liquid chromatography–tandem mass spectrometry; LESA, liquid extraction surface analysis; MALDI, matrix-assisted laser desorption ionization; MSI, mass spectrometry imaging; qMSI, quantitative mass spectrometry imaging; TEC, total electron content; TIC, total ion count.
Reproduced and expanded from Intl. J. Mass Spectrom., 2018. 101
Building on biodistribution of drug and drug metabolites is using MSI to inform our understanding of pharmacokinetic-pharmacodynamic (PK/PD) relationships. MSI experiments can relate the distribution of the drug to the distribution of changing endogenous metabolites and those providing molecular maps of the target tissue or tumor architecture. 56 This is further exploited by using the sensitivity and spatial resolution of analysis, exemplified in Figure 1 , to explore the differential absorption profiles of drugs along the crypt–villus axis. 13 The MSI-based study consisted of propranolol and metoprolol (highly permeable compounds) and atenolol (low- to moderate-permeability compound) orally co-administered to rats. MSI experiments were conducted to identify the site of drug absorption via high-spatial-resolution MALDI-MSI. Imaging data integrated with quantitative measurement of drug concentration in tissue homogenates. MALDI-MSI experiments were carried out in positive reflectron ionization mode over a mass range of m/z 120–1020 using two MALDI-TOF/TOF instruments.
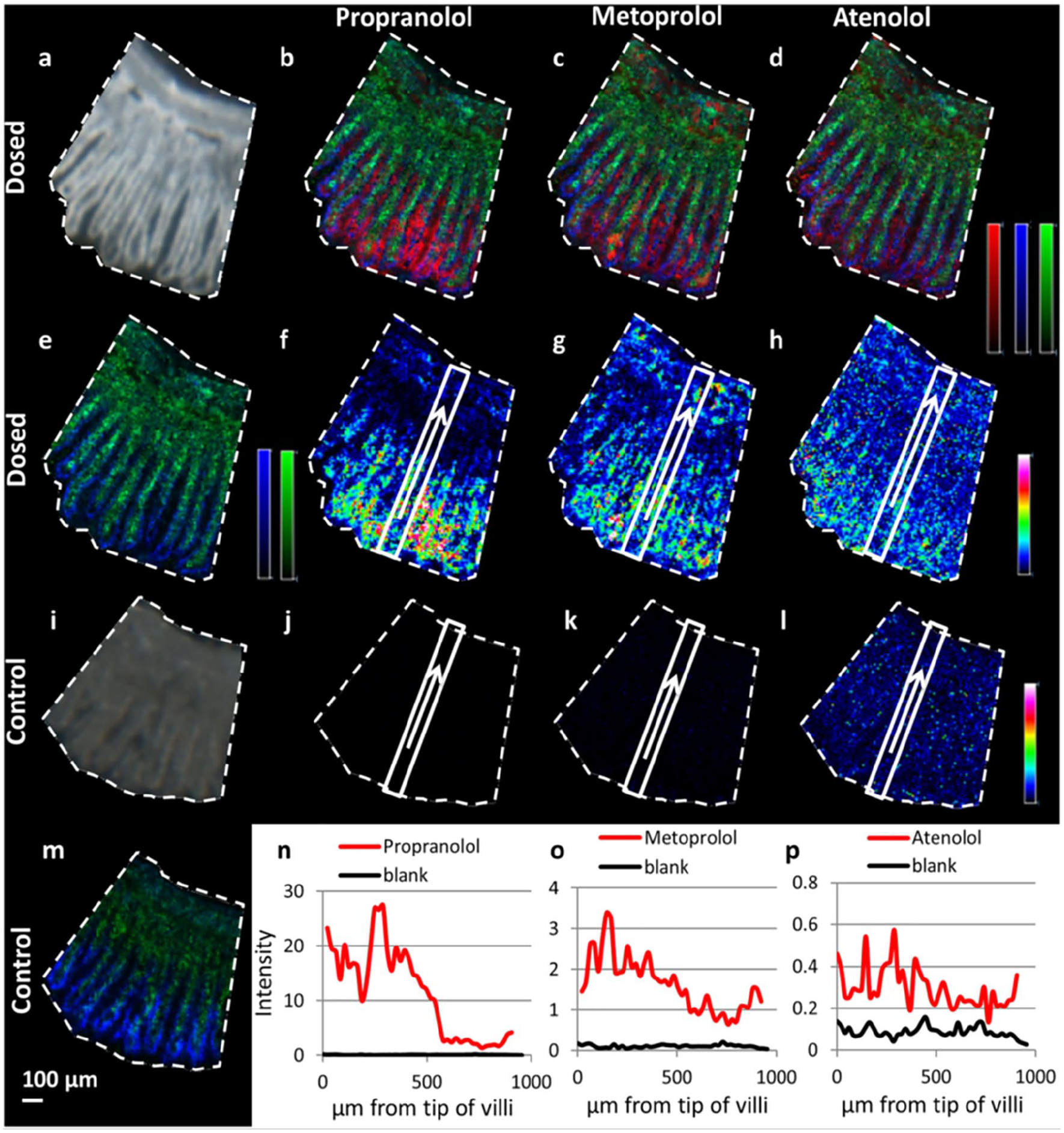
Distribution profiles of the three selected drugs (propranolol, metoprolol, and atenolol) in rat small intestine following oral dosing. (
As previously stated, there are methods to detect previously poorly detected classes of endogenous targets. One important class of poorly detected targets in drug discovery is steroids. Imaging experiments can be accomplished, however, using on-tissue derivatization with Girard T reagent. This modifies the targets to generate increased ionization, so quantitation of substrate and product (11-dehydrocorticosterone and corticosterone) of the glucocorticoid-amplifying enzyme 11β-HSD1 could be made. 57 MSI analysis of metabolomic profiles in samples, both at baseline and following therapeutic interventions, adds significance to both understanding and characterizing the tumor microenvironment in the preclinical and clinical setting. MSI can be widely used for investigatory toxicity and safety assessment.
MSI is used in a targeted fashion, such as identifying the accumulation of histopathological observed drug metabolites, in situ and with sufficient spatial resolution to map the deposition of the crystals.58–60 It is also widely used to measure CNS exposure, as exemplified in the disposition and metabolism analysis of fosdevirine, a non-nucleoside reverse transcriptase inhibitor, in the CNS. 61 MSI can be used to understand advanced drug delivery, such as tracking the distribution and release from an encapsulation form of the drug from delivery particles that help target specific tissues or protect it from early degradation or metabolism. 62 Further uses include blood–brain barrier penetration,18,63 imaging microphysiological systems and 3D cell cultures64–67 ( Fig. 2 ), assessing tumor hypoxia, 56 and clinical translation and application.68–71 MSI was able to provide novel insights to the 3D cell culture study presented in Figure 2 , because the localized distributions of metabolites within the microenvironments of such systems are hard to accurately assess. The study used high-spectral-resolution Fourier transform ion cyclotron resonance (FT-ICR) MALDI MSI to image the distribution of endogenous metabolites in breast cancer MCF7 spheroids. The wider study was able to show that known specific metabolite markers (adenosine phosphates and glutathione) indicated that the central region of these cell culture models experiences increased hypoxic and oxidative stress. The study also described the sample collection and processing needed to handle these challenging MSI samples.
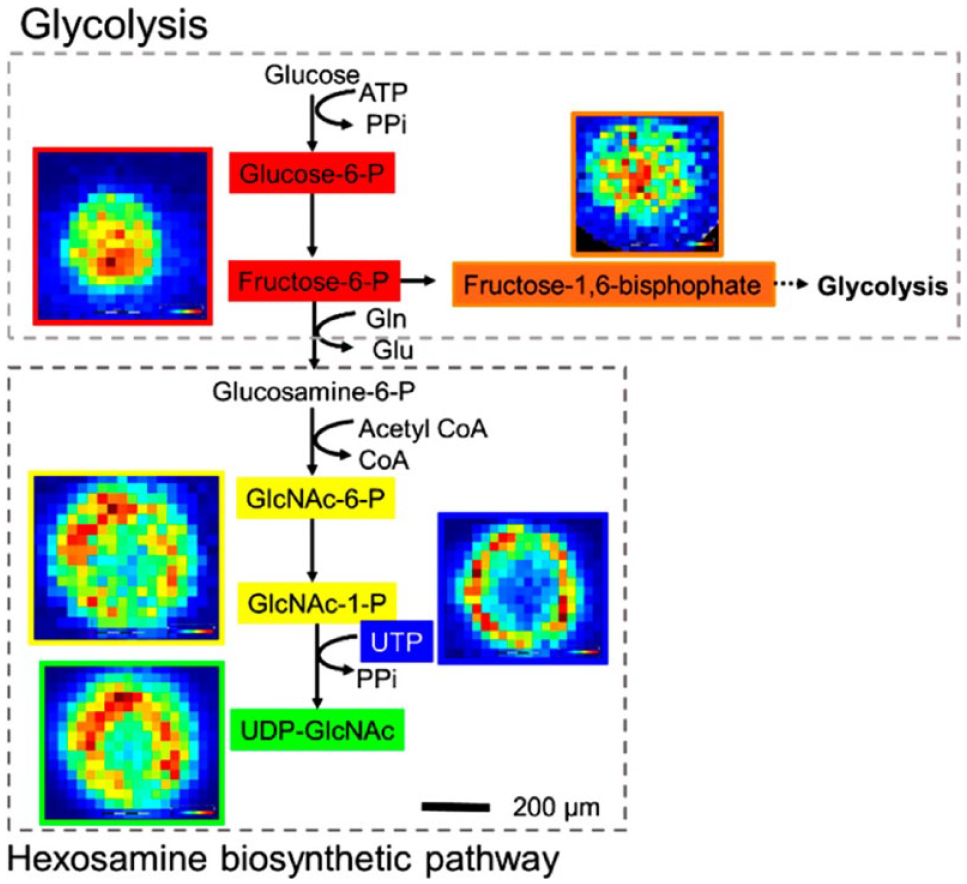
Elemental formula of the metabolites assigned can be mapped onto the hexosamine biosynthetic pathway (HBP). Note how the HBP is upregulated in the outer region, whereas glycolysis intermediates are upregulated in the center region. Glucosamine 6-phosphate (glucosamine 6P) was not observed. Reproduced from Anal. Chem., 2019. 67
To date, most MSI studies within pharmaceutical R&D have revolved around questions of drug efficacy as well as measurement of endogenous metabolites as biomarkers for efficacy, or in support of DMPK (drug metabolism and pharmacokinetics) and tissue exposure studies. MSI is, however, a powerful tool in investigatory toxicology studies answering pathology-driven hypotheses; for example, when pathological findings have suggested a toxicological response in tissue, MSI can be used to highlight a co-localization of drug or drug metabolite in the affected area. 72 More refined application uses MSI assessment of tissue to offer mechanistic insights relating to toxicity, offering the ability to measure and monitor molecular changes prior to gross morphological changes. For example, the nephrotoxicity of polymyxin antibiotics has been examined by MSI. The toxicity of different polymyxin analogues and compound abundance was correlated to the renal cortex and medulla in rats. 73
Expanding the applicability of MSI in toxicity studies has recently been illustrated by researchers exploring preclinical drug-induced lipidosis in tissues and cells. The research described the detected distributions of the administered drug, drug metabolites, lipid molecules, and a putative marker of lipidosis, di-docosahexaenoyl (22:6)-bis(monoacylglycerol) phosphate (di-22:6-BMP). 74 Figure 3 shows that combining positive-mode parent molecule MS1 images of amiodarone and metabolite M11 and negative-mode images of di-22:6-BMP/di-22:6-PG (phosphatidylglycerol) shows the spatially localized increase in this lipid alongside the presence of drug and metabolite. The data were acquired at either 50 µm for the imaging MS and tandem MS (MS/MS) repeat analysis of the single-dosed tissues or 100 µm for the imaging of all dosing conditions in positive- and negative-ion modes.
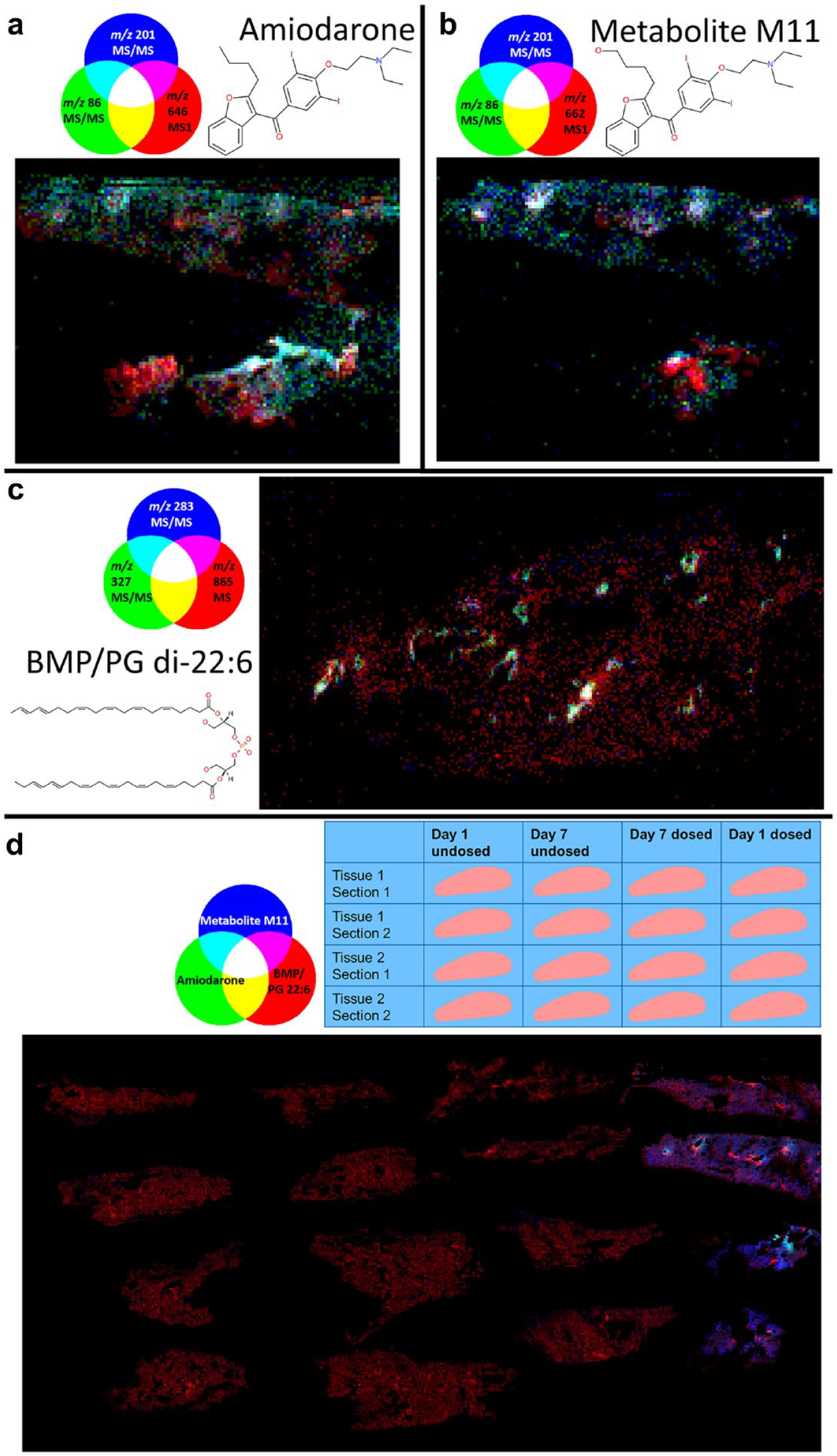
Registered red-green-blue (RGB) overlay images from repeat-analysis experiments, including the two biological replicates of the day 1 dosed tissue showing parent molecule MS1 (red) and major tandem mass spectrometry (MS/MS) fragments (blue and green) of (
Another application of MSI that is distinctively applicable to drug discovery is in the development of drug delivery systems, and therefore it is worth calling out in this review. The ability to simultaneously image and differentiate drug delivery systems, released or delivery system–bound pro-drugs or APIs, and drug metabolites offers nuanced insights to researchers trying to develop systems to target or prolong tissue exposure or smooth out Cmax and mitigating potential toxicity relating to high peak exposure levels. 7 Such a holistic analysis overview is hard to obtain from bulk tissue homogenization or bioanalysis.
MSI has been extensively used to monitor compounds in lung tissue, with early reports quantifying drugs such as tiotropium 75 and more recent reports measuring differential routes of pulmonary drug administration of the anticholinergic drug tiotropium bromide. 76 The latter study demonstrated that the distribution of the drug in the lungs of guinea pigs varied considerably when administered by nebulized inhalation compared to intratracheal instillation at doses that induced significant anti-bronchoconstrictive activity. A more recent refinement of lung retention and localization has been reported in which compounds were co-administered. For example, Figure 4 shows imaging of inhaled salmeterol, salbutamol, and fluticasone propionate, so that lung targeting and retention can be discerned in the same tissue using DESI-MSI. 77 Within the study, rat samples of lung tissue were obtained, high-resolution DESI-MSI analyses were performed at spatial resolutions between 30 and 70 µm in negative-ion mode, and mass spectra were collected in the mass range of 200–900 Da. An even more elegant refinement for determining tissue targeting has used MSI to uncover the regional localization of inhaled salmeterol retention in the lung. Regional and histological localization of salmeterol retention in the lungs after inhalation was compared to systemic administration. This was achieved via a novel dual-isotope study, in which salmeterol was delivered via inhalation and d3-salmeterol delivered intravenously to the same rat. 14 The use of co-administration or formulation of deuterated analogues and subsequent differential biodistribution mapping is a powerful new MSI strategy for drug discovery.
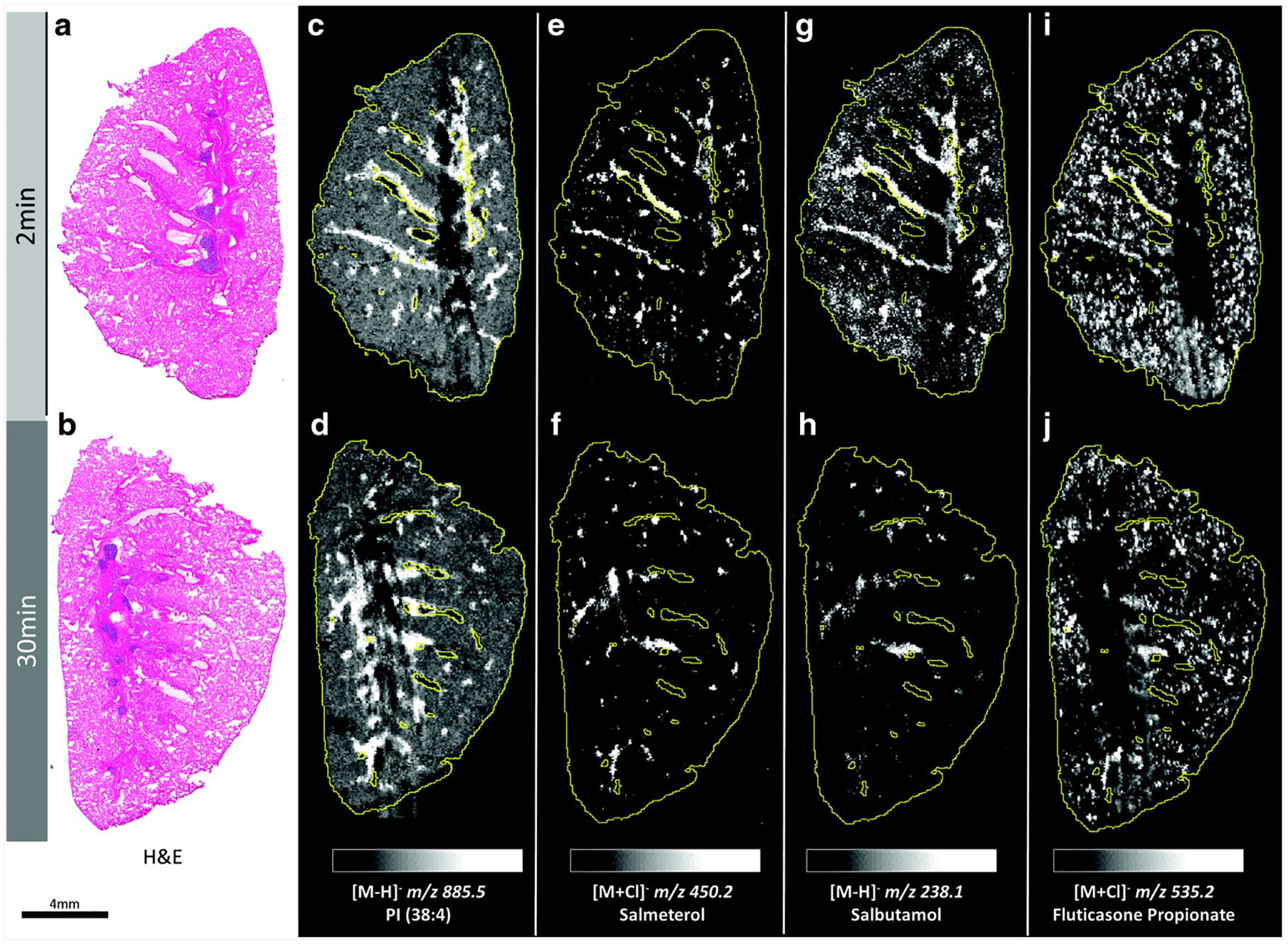
Whole-lung imaging by desorption electrospray ionization–mass spectrometry imaging (DESI-MSI) in negative detection mode at 70 µm spatial resolution. Hematoxylin and eosin (H&E) stain images from (
Data Challenges for MSI at Scale in Support of Drug Discovery
One significant challenge for expanded use of MSI is that high-performance platforms generate gigabytes of complex high-dimensional data for each experiment. Efficiently and effectively exploiting this ocean of data requires as much investment in time and resources as that applied to the acquisition of the data. Analysis of MSI data is separated into pre-processing (correction for variations in the data, experimental artifacts, and noise; results in peak picked data) and post processing (statistical analysis performed on the peak picked data to identify trends within the data). Protocols for pre-processing are often instrument dependent or vendor software determined. Failure to adequately understand any automated or user-defined pre-processing will affect the eventual results. Situation-dependent pre-processing can include dead-time correction, smoothing, baseline correction, normalization,78,79 peak detection, 80 peak alignment, 81 charge state deconvolution, and deisotoping. 82 Postprocessing and holistic data mining remain major challenges in exploiting remaining hidden value from MSI data. Depending on the question at hand, one or more univariate analysis, 83 multivariate analysis, 84 clustering, 85 or classification 86 algorithms can be applied. 83 Only a few are implemented in available software, however, and even fewer are available in software capable of handling data from multiple platforms,87,88 requiring the development of bespoke software routines. Processing of datasets that are larger than the available memory in the processing computer can also be problematic or—in some cases without memory-efficient routines—impossible. Random projection has been shown as an extremely computationally efficient means of reducing the dimensionality (and thus size) of a given dataset while retaining compatibility with subsequent analyses, such as principal component analysis (PCA), non-negative matrix factorization (NMF), and image segmentation (or clustering) routines. For lossy compression, randomized approximation methods can be used for significantly improved compression ratios. 89 A further benefit of these methods is that multivariate methods such as PCA and k-means clustering can be performed directly on the compressed data, resulting in faster execution and reduced data size limitations. Other lossy, dimensionality reduction techniques include PCA 90 and wavelets. The use of any of these methods, however, would require the development of custom software because currently no software has integrated these techniques.
Data analysis challenges are further exacerbated when performing studies among multiple facilities or collaborators that might not have access to the same analysis tools. Therefore, significant work in the development of open standards for MS has been made, resulting in the community open standard mzML. 91 The MSI community built on this, extending mzML to meet their needs and resulting in the open MSI community standard imzML. 92 A product of this is that the amount of software for visualizing MSI data has dramatically increased.87,88,93–95
Conclusions
This review has focused on the impact MSI has been having on pharmaceutical R&D; however, the topics covered are equally applicable to wider challenges that need to be continued to affect, perform at scale in, and integrate into the wider research landscape. Also, while most pharmaceutical MSI researchers and publications have focused on preclinical drug discovery, the approaches described here are applicable to a drug development and clinical translational setting. In this setting, a significant challenge is to maintain and standardize the collection of patient material and biopsies. Subsequent sample processing also requires further consideration due to any risk of infectious material. An exciting expansion of tissue-based MSI is in situ MS in surgery, which is providing molecular information directly from the cut tissue (REIMS and iKnife). 96
We have described the application of MSI to drug discovery; however, other current applications include diverse research fields such as disease state profiling, agrochemical investigation, 97 and even forensic analysis.98–100 Among all fields in which these technologies are applied, there remains a need to address the issue of sensitivity in MSI. There are a number of technical and operational procedures that can be used. As discussed earlier, sensitivity remains a challenge for some targets and classes of molecules. Approaches to compensate for inherent poor ionization efficiencies include operating as a probe-based spectroscopy, in which the larger the pixel area surveyed, the richer the spectrum in MSI methods, which provide higher ion intensities by virtue of sampling larger areas. In real terms, this is not the same as greater sensitivity per unit of tissue sampled, but lowering resolution does increase the coverage of compounds that can be surveyed. Higher-mass-resolving power instruments result in “narrower” peaks in the resulting spectra. When presenting images as a function of peak intensity, this does result in vastly improved ion intensities, an apparent measure of improved sensitivity. It is important, however, to also understand that there will remain situations in which MSI is unable to provide enough sensitivity at required spatial resolution for certain drugs. Therefore, we recommend MSI is deployed as a powerful capability alongside the suite of analytical tools in the drug discovery process, with integration to other quantitative and qualitative data.
Looking forward, a commonly claimed demand of researchers is for increased performance in lateral resolution and sensitivity to allow even increased cellular and subcellular imaging. This has always been the call from the MSI community, and it continues to be partially valid. Increased specifications are important, and there has been considerable progress in acquisition speeds during the past decade. It is, however, the context of the information obtained that is critical. For example, knowing the subcellular localization of a compound is interesting, but it becomes useful if it is known in context of the related efficacy or cellular toxicity and offers insights that can be used to modify the therapeutic compound’s development. Furthermore, there remains a significant unmet need in delivery of imaging systems that can fully deconvolute the biological complexity occurring in disease and healthy tissues or adequately resolve the toxicity and efficacy challenges in drug discovery. Industry researchers are increasingly demanding more from every sample, be it patient biopsy, advanced animal systems, or complex cellular models. There is the ambition to perform truly multi-omic and spatially resolved experiments. Greater efforts are needed to deliver integrated imaging solutions to concisely solve complex biology, through linking of histology with MSI for PK/PD imaging and deep cell phenotyping. The MSI community is embracing these challenges and being supported by significant investment by leaders in pharma as well as strategic funding bodies. This is exemplified by Cancer Research UK funding a $20 million project to use MSI and integrated imaging technologies to map tumors at the molecular and cellular levels. Key to such large-scale projects are the development of both advanced data analysis strategies and access to appropriate patient and disease models. Disseminating this research into active drug discovery programs is achieved by drug companies such as AstraZeneca embedding and supporting cutting-edge multidimensional science (https://www.cancerresearchuk.org/funding-for-researchers/how-we-deliver-research/grand-challenge-award/funded-teams-bunch).
The field of mass spectrometry imaging in support of drug discovery continues to evolve at pace. Current and emerging MSI technologies are providing new ways to map and deconvolute complex safety and efficacy biomarkers while also providing new insights into the underlying biological complexity of tissues. These technologies can be expected to be core capabilities in an increasing number of cutting-edge pharmaceutical companies.
Supplemental Material
SLAS_Goodwin_2020 – Supplemental material for A Critical and Concise Review of Mass Spectrometry Applied to Imaging in Drug Discovery
Supplemental material, SLAS_Goodwin_2020 for A Critical and Concise Review of Mass Spectrometry Applied to Imaging in Drug Discovery by Richard J. A. Goodwin, Zoltan Takats and Josephine Bunch in SLAS Discovery
Footnotes
Acknowledgements
The authors would like to thank Cancer Research UK Rosetta Grand Challenge consortium.
Declaration of Conflicting Interest
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Richard Goodwin is employed by and holds stock in AstraZeneca, and all authors’ research and authorship of this article were completed within the scope of their employment with AstraZeneca.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
