Abstract
DNA-encoded library (DEL) technology has been used as an ultra-high-throughput screening approach for hit identification of drug targets. This process is an affinity-based selection and requires incubation of DEL molecules with the target. Currently, in most reported cases, the input (i.e., the copy number) of individual DEL molecules varies from 105 to 107. With the ever-increasing DEL size and screening cost, lowering the input of DEL molecules while maintaining an appropriate signal-to-noise ratio in a selection is of paramount importance. In this article, we varied the input of DEL ranging from 103 to 105 in selections with two different protein targets to explore the lower limit of DEL molecule input. The results could facilitate the optimization of the DEL selection process and reduce costs related to library consumption.
Introduction
DNA-encoded library (DEL) technology, inspired by and originated from the phage display library, is considered a powerful chemical tool, especially for its potential in small-molecule drug discovery. Typical DELs consist of mixtures of small molecules conjugated to unique DNA oligo tags, and structural information of each small molecule is encoded into its corresponding DNA sequence.1–3 When DELs are applied against proteins of pharmaceutical interest in a selection, the library members with higher affinity are enriched after multiple rounds of wash, and their structural identities are revealed by subsequent DNA sequencing.4–6 The DEL selection process possesses great advantages in terms of speed, cost, and scale when compared with traditional high-throughput screening. To date, multiple pharmaceutical companies such as GSK, Roche, Amgen, and so forth are actively involved in DEL construction, selection service, or drug discovery activities based on the DEL platform. Compounds discovered by DEL technology have already progressed to the clinical phase, such as receptor-interacting serine/threonine-protein kinase 1 (RIPK1) inhibitor GSK2982772 7 and soluble epoxide hydrolase inhibitor GSK2256294. 8
As DEL technology has gradually matured over the past decades, the size of individual DEL(s) has been constantly increasing, reaching trillions as claimed in literature 9 or by the DEL service provider. 10 To deal with this large number of DNA-encoded small molecules, appropriate parameters of the chosen analytical methodology (usually based on DNA sequencing) are essential for successful DEL selection as well as accurate data analysis. It was observed that the enrichment signal from the DEL selection decreased when the input of individual DEL molecules decreased, 11 yet the threshold where the enrichment signal could be indistinguishable with background noise was not rigorously examined. Previously, it was empirically suggested that each DEL member molecule should have at least 105 copies in the library for one selection.12,13 In Table 1 , we summarize several DEL selection practices from the literature in terms of their library input.
Comparison of Molecule Copy Numbers in DNA-Encoded Library (DEL) Selection Practices.
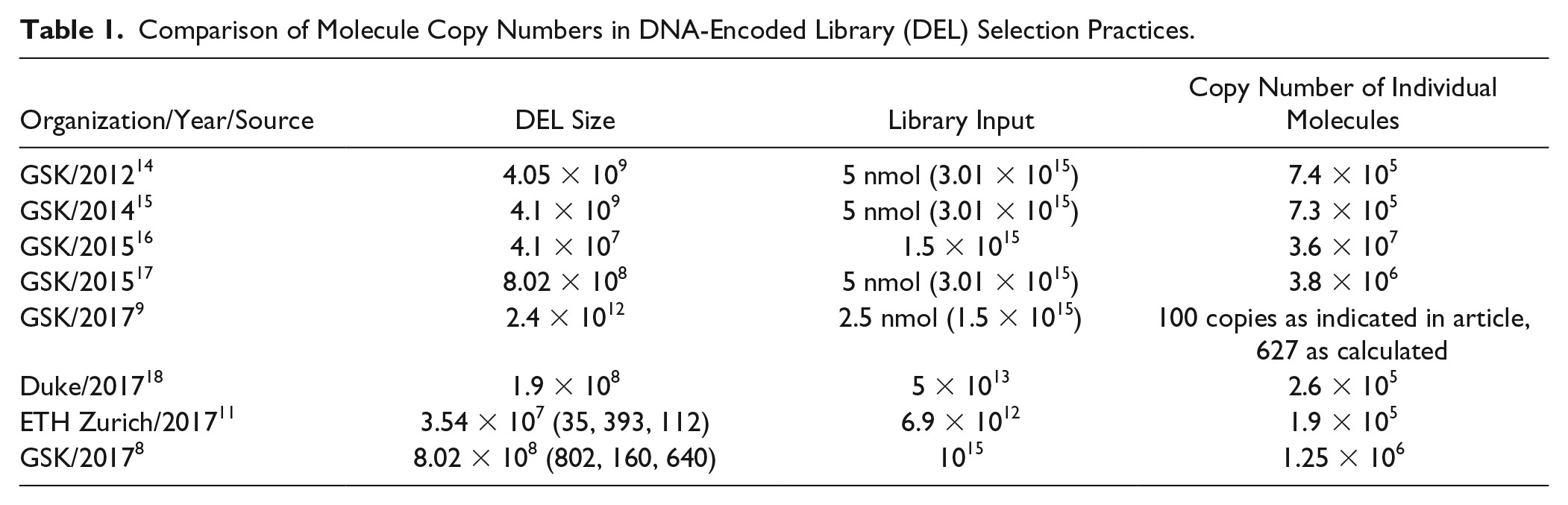
These practices, however, apply certain restrictions to the DEL selection process. For instance, to ensure this level of library input, one would have to increase the amount of DEL used in one selection as the number of library members increases. At a certain level, an increase in the amount of DELs will face technical and economic problems, including solubility, viscosity, or cost issues. Also, considering the sophisticated and expensive preparation process required for DELs, it would be economically significant if one could lower the input of a DEL for individual selection without affecting its performance (i.e., the ability to determine the enrichment signal in a statistically significant manner). Therefore, determination of the minimally required molecule copy number in individual DEL selection under given conditions would allow researchers to have significantly higher efficiency when using DEL to discover new chemical entities, especially those with pharmaceutical interests. It could also have a positive impact on DEL selection practice and guide the direction of optimization for future DEL screening experiments.
In this article, we describe the selection of two different soluble protein targets with various inputs of individual DEL molecules ranging from 103 to 105 to explore the lower limit of individual DEL molecules in selection and thus to facilitate the optimization of DEL selection.
Materials and Methods
Materials
Rho-associated protein kinase 2 (ROCK2, 19-417) was expressed in insect cells. Constructs were subcloned into pFastBacHTA vector, and the bac-to-bac system was used to produce baculoviruses to infect sf9 cells. It was purified by Ni-NTA affinity column chromatography (GE Healthcare, Chicago, IL), followed by size exclusion chromatography (HiLoad 16/600 Superdex 200 pg, GE Healthcare). The final purified protein was formulated in 20 mM Tris-HCl, pH 8.0, 150 mM NaCl, 20% glycerol, and 1 mM DTT and then stored at −80 °C. SAE is a heterodimer of SAE1 (SUMO-activating enzyme subunit 1) and SAE2 (SUMO-activating enzyme subunit 2); SAE1 (1-346) was cloned into pET-22b to encode a native polypeptide, and SAE2 (1-640) was cloned into pET-28a to encode an N-hexahistidine fusion protein. The constructs were coexpressed in Escherichia coli Rosetta (DE3) and purified by Ni-NTA affinity chromatography (GE Healthcare) followed by gel filtration (HiLoad 16/600 Superdex 200 pg, GE Healthcare). The protein was concentrated to 10 to 15 mg/mL in 50 mM Tris-HCl, pH 7.5, 350 mM NaCl, 1 mM DTT before freezing in liquid nitrogen and storage at −80 °C. DEL (with 167 billion compounds) was from HitGen. Ni-charged MagBeads were purchased from GenScript (Piscataway, NJ; L00295), salmon sperm DNA was from Sigma (St. Louis, MO; 31149), and all other reagents were from commercial sources.
DEL Selection
Automated selection was carried out using a KingFisher Duo Prime Purification System (ThermoFisher, Waltham, MA) in a 96-well plate. 19 After Ni-charged MagBeads were equilibrated with selection buffer (40 mM Tris, 20 mM MgCl2, 50 µM DTT, 0.1% Tween-20, 0.3 mg/mL salmon sperm DNA, 10 mM imidazole, pH 7.5), beads were transferred to a new row and incubated with 3 µM ROCK2 at room temperature (RT) for 30 min in selection buffer. Then, the immobilized protein along with the beads was incubated with DEL sample in selection buffer at RT for 1 h followed by a 1 min wash at RT in 100 µL selection buffer for nine times. Retained DEL members were recovered by heat elution in elution buffer (40 mM Tris, 20 mM MgCl2, pH 7.5) at 75 °C for 15 min. After the first round of selection, the second round was repeated with the heat-eluted portion of the previous round used as the input to the successive round with fresh protein. After each round, the output was quantified by quantitative PCR. After two rounds, the selection was done, and the output was amplified by PCR and further purified by QIAquick PCR Purification Kit (QIAGEN, Hilden, Germany; 28006) according to the manufacturer’s instructions. Sequencing was then performed for PCR-amplified samples on an Illumina HiSeq 2500. SAE selection was performed in a similar condition in solution mode, in which 2.5 µM free protein was incubated with DEL and then captured by Ni-charged MagBeads in selection buffer (50 mM Tris, 5 mM MgCl2, 0.2 mM DTT, 0.01% Tween-20, 0.3 mg/mL ssDNA, 10 mM imidazole, pH 7.5).
Data Analysis
Samples were decoded, and the result was presented in DataWarrior (developed by Openmolecules) cubes, with each dimension representing one cycle of DEL construction. The enrichment of certain cycle(s) of DEL construction was defined as “features,” and the difference between the groups was compared by feature intensity. Feature intensity is the enrichment score calculated as the “sum of sequence count” of the feature divided by the average of the “sum of sequence count” of all possible parallel features in the library. Background subtraction was performed by removing signals that appeared in no target control in the selection performed previously, enriched features were further examined by structures, and imidazole-like binders were excluded; promiscuous features were also highlighted and cross-checked by comparing with other selections in HitGen database.
Results and Discussion
The experiment design in this study is outlined in Table 2 . The HitGen library, containing 167 billion DEL compounds, was used for the selection of two different soluble protein targets, ROCK2 and SAE. ROCK2 is a serine/threonine-protein kinase involved in the regulation of the cytoskeleton and cell polarity. 20 SAE functions as E1 in the SUMOylation cascade and mediates the adenosine triphosphate–dependent SUMO ligation. 21 The input of each DEL molecules varied among 105, 3.3 × 104, 104, and 103. After two rounds of selection, the DEL molecules recovered were PCR amplified and sent for sequencing. After decoding and data analysis, the DNA sequences enriched in selection were translated into chemical structures and visualized by DataWarrior files. Each axis of the cube represented one cycle of the DEL synthesis; for four-cycle DELs, one of the cycles was prefixed in the cube. We use “feature” to represent the clusters enriched after selection, and the “line feature” indicated that two cycles of the DEL were fixed and 1 was variable, whereas the “plane feature” indicated that one cycle was fixed and the other two were variable.
Experiment Design in DNA-Encoded Library Selection.
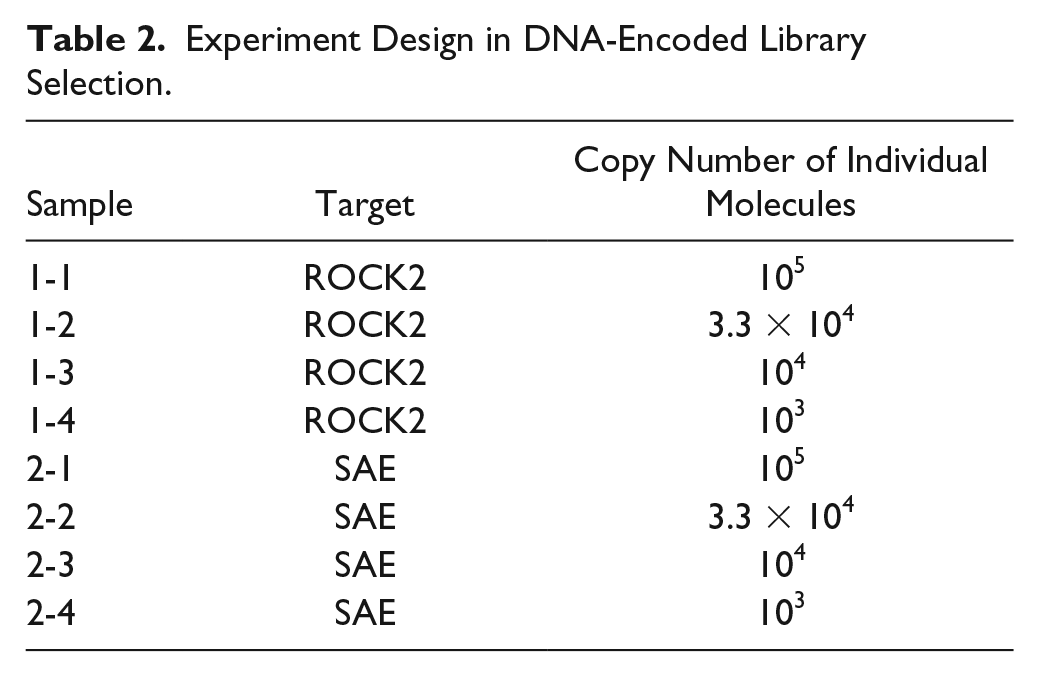
As the selection of these two targets has been done previously at HitGen and validated active hits have been found from the HitGen library, our data analysis mainly focused on the identification of features representing the validated active hits.
For ROCK2, the feature intensity of one strong feature ( Fig. 1A ), in which a couple of active hits with IC50 ranging from 50 nM to 860 nM in enzymatic assays were identified, decreased with the decrease of input, as indicated by enrichment scores of 658.55, 548.49, 255.2, and 7.31, respectively. This feature was also reproducible in the lowest input 103 in our design. For one weak feature ( Fig. 1B ) in which the active hits with an IC50 of about 7 µM were identified, we were able to identify this feature with 104 input but failed to observe the feature with 103 input; the feature intensity was 546.74, 79.56, 71.38, and 4.17 for the four groups, respectively. Further analysis of these features revealed their corresponding molecules as typical kinase hinge binders. Direct comparison of feature intensity supported the conclusion that strong features were reproducible in the 103 input whereas weaker features were not. All validated features that showed up in the 105 input group were reproducible in the 3.3 × 104 and 104 groups, indicating that the input could be decreased to 104 in selection without losing significant enrichment signals.
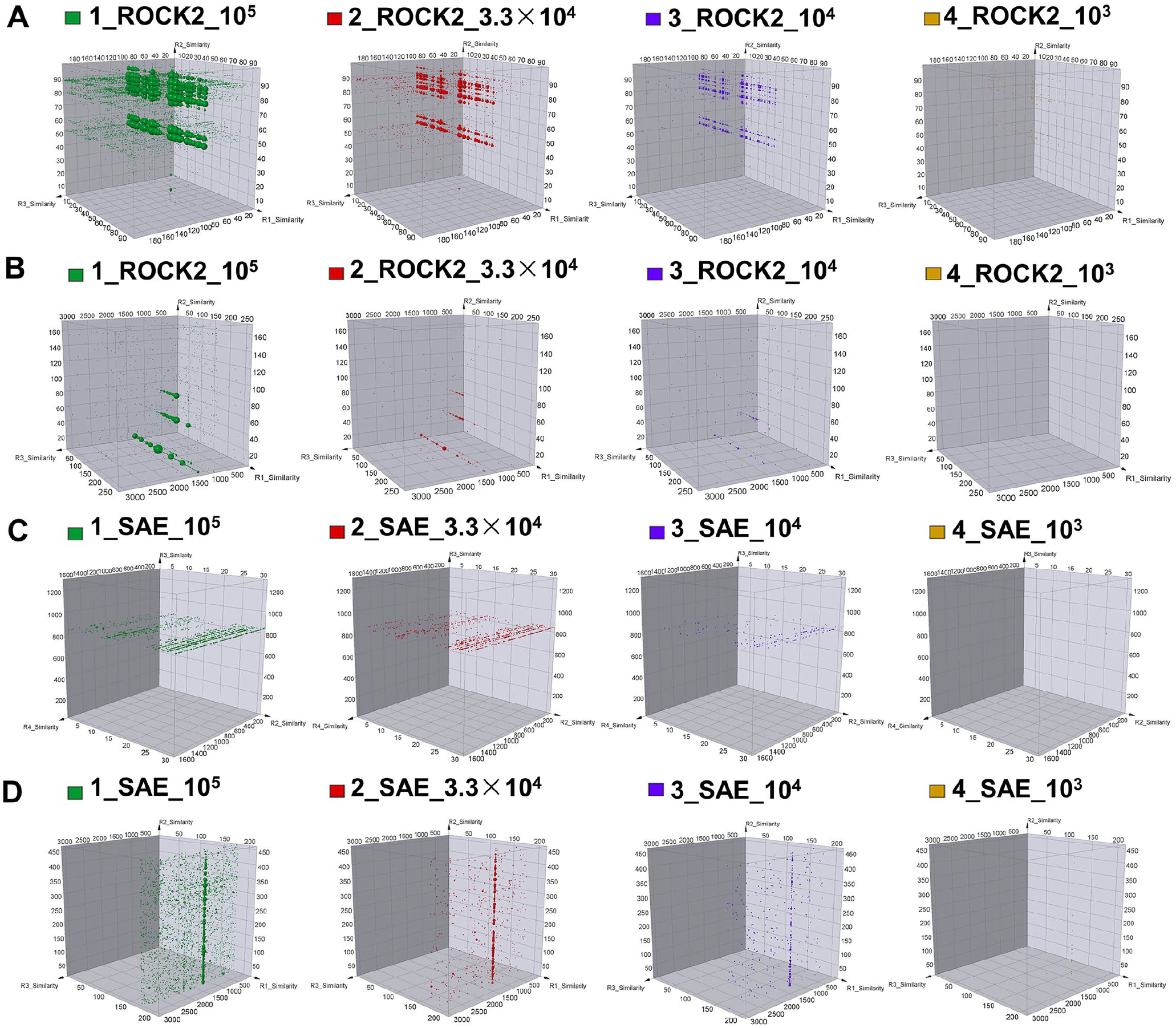
DataWarrior cube view of active features for ROCK2 and SAE target. The feature represented the binding of DNA-encoded library (DEL) compound with tens to hundreds nanomolar (
For SAE, after data analysis, one plane feature in which active hits with an IC50 of about 1 to 9 µM on enzymatic activity was identified, was readily reproducible, and feature intensity decreased with the decrease of input. When the input was lowered to 104, the feature became very weak, and we failed to observe this particular feature at 103 (
Fig. 1C
). The feature intensity of each group was 58.13, 20.12, 1.93, and 0.00. Another strong feature (
Further analysis of all other features from these 2 selections described above indicated that the top 100 features in 105 sample were reproducible and well correlated in 3.3 × 104 and 104 samples for ROCK2. However, there were seven features that did not show up in the 103 sample and 49 features with enrichment score below 4, which was very weak and gave little information regarding chemical diversity ( Fig. 2A ). With fewer and weaker features identified for SAE, the top 50 features were analyzed, and good correlation was identified in 105 and 3.3 × 104 samples with five features’ intensity below 4 in the 3.3 × 104 sample. In the 104 sample, 5 of the 50 features exhibited a feature intensity greater than 4, and 23 features were not identified. In the 103 input sample, no correlation was found, and most features disappeared in this sample ( Fig. 2B ).
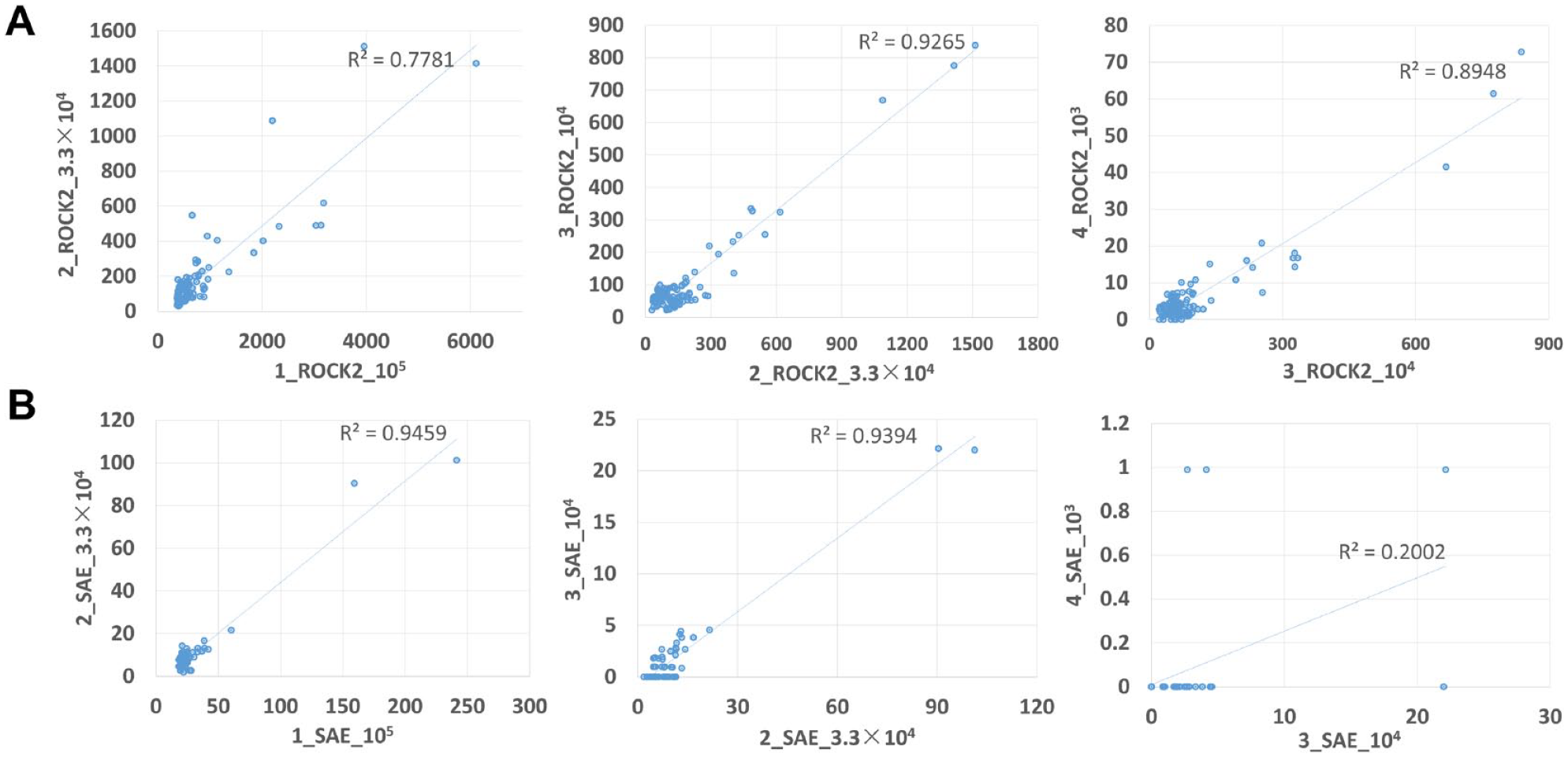
Feature intensity correlation. The top 100 features in ROCK2 (
In our design, SAE was chosen as it represented the targets with relatively weak features, whereas ROCK2 represented the targets with relatively strong features from past DEL screening practices at HitGen. The selection of two targets was performed in different modes typically used in DEL selection: immobilization mode and in-solution mode. For ROCK2, it was done in immobilization mode; that is, the target protein was immobilized on Ni-charged MagBeads before incubation with the DEL molecules. For SAE, free protein was incubated with the DEL molecules, then the protein along with the DEL binders was immobilized by the Ni-charged MagBeads. The two targets represented two different selection modes with strong or weak features in selection, to draw a more generalized conclusion on DEL selection.
Reduction of input may have an impact on the discrimination of hits and promiscuous binders. We examined the promiscuous binders in our database and found their overall feature intensity was lower than the intensity of real features, the reduction of input resulted in a decrease of both signals; however, with the lower intensity of the promiscuous binders, the decrease was faster than those of the real features. For targets in which the promiscuous binders are stronger than real features, the trend may be different, but other cases are required to prove this.
In our test, we failed to identify some positive binders with 103 copy of DEL molecules. In the published literature, 9 hundreds of copy numbers for the DEL molecules were used as the input in selection, active signals were enriched, and binding of compounds was confirmed. The major reason for this difference is that the DEL was a peptide library, and most active hits exhibited more than 50% binding to the target protein. Peptides may have higher binding affinity than small molecules from the HitGen library; in addition, only compounds with very high binding potency can be identified in such a low-input design.
It has been reported that selection performance dramatically dropped when the input was below 105, and at least a 105 (and preferably 106) input should be used to ensure successful DEL screening. 13 In this article, the selection was performed with carbonic anhydrase IX as the target, and different ligands with known affinity were conjugated with DNA tags and screened along with a DEL of 360,000 compounds to serve as positive controls. The major difference between the conclusions in the previous study and our study is that the ligands were spiked in and identified as a single molecule in the previous article; however, in large-scale DEL selection, such as in our case (more than 100 billion compounds), the library is constructed by the “spilt and pool” strategy, in which the structures vary by the dimension of building blocks. In our study, the reproducibility of signals among different inputs was not identified as an individual compound but as features—the enrichment of a cluster of compounds with chemical similarities. For a single compound, a high sequence count is required for the compound to stand out and be seen among all background signals; however, for the enrichment of a certain dimension of building blocks, they stand out as lines or planes, which are much easier to be identified in cube-style figures. In our case, input less than 105 is also compatible with feature identification.
On the other hand, DEL structures and selection strategies are nonnegligible factors affecting the result. In our case, the DELs were constructed by the “split and pool” strategy, with small molecules attached to double-strand DNA tags, and the DNA was purely a barcode for compound identification. The binding process is determined by the reversible interaction of the small molecules with targets. In some studies, the DEL molecules were with covalent warheads, 22 cross-linking ligands, 23 or target proteins conjugated with complementary DNA tags 24 ; the selection strategies in these cases were very different, and our conclusion does not apply.
In most publications, inputs of 105 copies and greater were included in the selection. Theoretically, a higher input results in higher signals and more details for feature distribution. However, when the DEL is screened on a large scale with billions of compounds, the cost of library preparation, solubility issues, and viscosity issues are fundamental factors to be considered. Our study is the first report to use a DEL with a billion-scale size to explore the lower limit of individual DEL molecules required in selection. For targets with reported strong binders and well-known druggable pockets, usually a more stringent potency criterion is required. In our case, such as with the ROCK2 target, lowering the input to 103 to 104 is feasible and can filter the weak binders. For some targets with limited reported binder and flat or less druggable pockets, a higher input (no less than 3.3 × 104) should be used. In summary, the input can be adjusted along with targets and screening purposes. In general practice, we recommend using no less than 3.3 × 104 in selection, which allows for the identification of both a strong and weak binder, as well as retaining all the details and possible structure-activity relationship information in enriched features.
Footnotes
Acknowledgements
We thank Dr. Wei Cui for critical review of this article.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: All authors are employed by HitGen Inc., and their research and authorship of this article was completed within the scope of their employment with HitGen Inc.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
