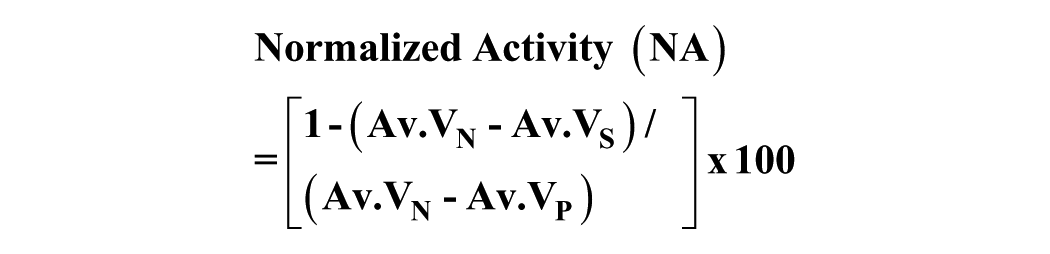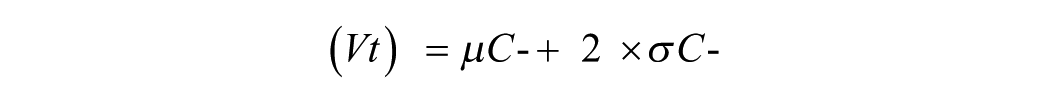Abstract
The current methodologies used to identify promising new anthelmintic compounds rely on subjective microscopic examination of worm motility or involve genetic modified organisms. We describe a new methodology to detect worm viability that takes advantage of the differential incorporation of the fluorescent molecular marker propidium iodide and the 2,1,3-benzothiadiazole core, which has been widely applied in light technology. The new assay developed could be validated using the “Pathogen Box” library. By use of this bioassay, it was possible to identify three molecules with activity against Caenorhabditis elegans that were previously described as effective in in vitro assays against other pathogens, such as Schistosoma mansoni, Mycobacterium tuberculosis, and Plasmodium falciparum, accelerating the identification of molecules with anthelmintic potential. The current fluorescence-based bioassay may be used for assessing C. elegans viability. The described methodology replaces the subjectivity of previous assays and provides an enabling technology that is useful for rapid in vitro screens of both natural and synthetic compound libraries. It is expected that the results obtained from these robust in vitro screens would select the most effective compounds for follow-up in vivo experimentation with pathogenic helminths.
Introduction
Helminthiasis is a serious global health problem. Diseases caused by soil helminths are among the most common infections worldwide, affecting primarily marginalized, low-income, and resource-poor populations. It is estimated that more than 1 billion people, mostly in developing countries in Africa, Asia, and America, are infected with one or more species of helminths. These infections were responsible for a burden of around 5.18 million disability-adjusted life years in 2010. 1 Besides the high rate of human infection, livestock farming also suffers from extensive economic losses due to helminth infections of cattle. The production of meat, fiber, and milk is impacted due to these parasite infections throughout the world. 2
There are currently few treatment options for helminth infections. 3 In areas where helminthiases are highly endemic, the World Health Organization recommends that the control strategy is the preventive therapy of populations at risk, mainly school-aged children, with albendazole and mebendazole. 4 Despite the drugs available, the emergence of resistance to many of the currently available anthelmintic drugs is a major concern for animal and human health. 5 Veterinary medicine is already facing this resistance problem in all the major groups of anthelmintics, mainly with sheep and goats, 6 and resistance to human treatment has already been reported in some endemic regions. 7
Despite the dreary perspectives, human helminth infections have not yet benefited from drug discovery efforts. 8 Most of the currently used anthelmintics have evolved from veterinary medicine and have been discovered through in vivo animal model testing. 3 Apart from the low investments directed to these neglected diseases, the difficulties in maintaining parasitic nematodes in the laboratory, by virtue of their complex life cycle, and the inability to adapt some helminthic parasites in laboratory conditions make the screening of compound libraries against living worms extremely challenging, thus remarkably decreasing the chances of discovery of novel chemical entities for antiparasitic chemotherapy.
Considering these limitations, in vitro whole-organism screening has gained relevance over the last years. In particular, the soil nematode Caenorhabditis elegans has been proposed as a surrogate model enabling anthelmintic drug discovery.9,10 Some of the advantages of the C. elegans model for drug discovery is that it is an easy and inexpensive culture, reproduces rapidly and robustly in vitro, is amenable to large-scale culture, has a small size that is compatible with screening miniaturization, and is a readily available genetic tool for target deconvolution. 11 The majority of marketed anthelmintics are active against C. elegans, 12 and this animal model has been useful in progressing the understanding of the mechanism of action of anthelmintics such as levamisole, benzimidazole, and amino-acetonitrile derivatives. 13
Among the available methods using C. elegans for drug discovery, most of them rely on the direct observation of movement, and the criteria to select compounds are based on the paralysis of worms. Some of them are based on observation and manual counting, and others use automated devices coupled with software to measure the nematode movement.14,15 Although automated, movement evaluation is not a suitable criterion to select nematicide compounds as the worms may be paralyzed but not dead. Another method is the use of the automated worm-handling technology COPAS Biosort (Union Biometrica, Somerville, MA), which is a C. elegans sorter similar to fluorescence-activated cell sorting for cells and sorts individual worms based on fluorescence. 16 These methods, besides being labor-intensive, are subjective and not applicable to high-throughput screening. In order to accelerate the process of drug discovery for new anthelmintics, there is a need for a reliable and robust screening method.
Here we report the development and validation of an improved methodology to objectively detect worm viability based on the differential incorporation of two fluorescent markers: DB-1, a 2,1,3-benzothiadiazole (BTD) fluorescent derivative, and propidium iodide (PI), a DNA intercalating agent. BTD is a heterocyclic widely applied in light technology. 17 The high electron affinity of the BTD core allows for an efficient design of π-extended fluorescent derivatives with push–pull-like molecular architecture, 18 rendering these structures as highly stable fluorogenic dyes. 19 One of the advantages of this class of fluorescent markers is that they typically have large Stokes shifts and well-defined emission wavelengths, 20 avoiding in this context overlaps of the excitations and emissions during bioimaging experiments. 21 We have been developing novel BTD derivatives for more than a decade with success, 17 and we wish to disclose herein the use of one of these fluorescent markers as a new strategy for anthelmintic drug discovery using C. elegans. This assay was validated using the Pathogen Box library, which contains 400 diverse drug-like molecules. Using this bioassay, it was possible to identify new molecules with activity against C. elegans, accelerating the search for new anthelmintic lead molecules.
Materials and Methods
C. elegans Culture
The wild-type C. elegans Bristol N2 strain, kindly provided by Professor Ricardo Fujiwara (UFMG, Brazil), was grown on nematode growth medium (NGM) with Escherichia coli strain NA22 as the food source according to the methodology previously described. 22 After 4 days of culture in a BOD incubator (CE-120, Cienlab, Campinas, Brazil) at 22 °C, the plates were washed with M9 medium 23 and a synchronous worm population was obtained by filtration through a 20 μm pore sieve (43-50020-03, Pluriselect, Leipzig, Germany). Filtered L1 larvae were collected and washed by centrifugation at 200g for 3 min, followed by two washes with M9 medium.
Viability Evaluation
In order to evaluate the viability of C. elegans through the automated differentiation of live and dead larvae, a method based on the selective incorporation of two fluorescent cell markers, DB-1 and PI (P1304MP, Molecular Probes, Eugene, OR), was developed. To assess whether PI would be a good indicator of viability, worms were incubated for 72 h with M9, 2% DMSO (D4818, Sigma-Aldrich, St. Louis, MO), and 5 and 20 mM levamisole hydrochloride (31742, Sigma-Aldrich) in a black polystyrene 384-well plate (781091, Greiner Bio-One, São Paulo, Brazil). After that, 5 μL of PI (final concentration of 20 μM) and 15 μL of DB-1 (final concentration of 1.56 μM) were added to the plates, followed by a further 24 h of incubation. At the endpoint, treated wells were observed under microscope to check the pharynx pumping rate, and the plate was fixed with 15 μL of 12% paraformaldehyde (158127, Sigma-Aldrich) and immediately processed for image acquisition and analysis.
To assess whether the dye concentration used causes toxicity and thereby influences the viability, worms were incubated with different concentrations of PI and DB-1. The DMSO tolerance was evaluated in the same conditions with DMSO concentrations ranging from 0% to 20%. In order to validate the developed methodology of viability evaluation, 1 M of the reference compound levamisole was serially diluted by a factor of 2 and tested in dose–response with different incubation times. The interexperiment variability was also evaluated by performing the test on two different days.
Assay Setup
The first step in the assay setup was the addition of controls in the 384-well plate. The positive control, which consisted of 20 mM levamisole hydrochloride, was added in columns 1 and 2, followed by the negative control, 2% DMSO, in columns 23 and 24, and then all samples to be tested, which were in a 100% DMSO stock solution. Each sample was diluted and added to a single well at the final concentration of 50 μM (resulting in a concentration of 2% DMSO). After addition of the controls and samples on the plate, the previously synchronized L1 larvae suspended in M9 medium, containing approximately 6000 worms/mL, were added to all wells of the plate with the aid of a multichannel pipette so that each well held a final volume of 50 μL. After 72 h of incubation at 22 °C, plates were stained with 20 μM PI and 1.56 μM DB-1, followed by another 24 h of incubation; after that the worms were immobilized with 2% paraformaldehyde for 30 min and subjected to image acquisition and analysis.
Imaging Acquisition and Processing
Nine fields per well were imaged using a 10× objective with two different filters, Texas red (excitation filter, 560–580; emission filter, 590–640) and Alexa 488 (excitation filter, 460–490; emission filter, 500–550), using the Operetta high-content automated imaging system (PerkinElmer, Waltham, MA). After the acquisition, images were subjected to high-content analysis using Harmony software (Perkin Elmer). The software is capable of recognizing and individualizing each worm by means of the emitted signal of the green fluorescence provided by the incorporation of the DB-1 dye by the helminths. After identifying the population based on this green signal, the software analyzes the image acquired in the red channel and measures the average intensity of the signal provided by the incorporation of PI by each worm. This intensity was calculated in order to classify if the worm was alive or dead based on a threshold defined according to the different intensities of PI in the positive and negative controls. As output information the software provides the total number of worms in the field, the average intensity of PI, and, in a second step of the analysis, the number of dead worms and the viability of the population analyzed.
Data Analysis
The raw data regarding the viability values were normalized as a function of the negative and positive controls, to determine the normalized anthelmintic activity, according to the following equation:
where Av.VN is the average viability of negative control wells, Av.VP is the average viability of positive control wells, and Av.VS is the average viability of the sample well.
The coefficient of determination (R2) of the two screening runs was calculated using Prism 6 software (GraphPad Software Inc., La Jolla, CA).
Data were processed using nonlinear regression and sigmoidal dose–response to obtain EC50 values (compound concentration corresponding to 50% normalized activity). The Z′ factor was used to evaluate the performance of the screening test. Data generated in the assay validation and primary screening of compounds were processed and analyzed by Excel (Microsoft, Redmond, WA) and Spotfire (TIBCO Software Inc.).
HCS Assay Validation and Screening of the Pathogen Box
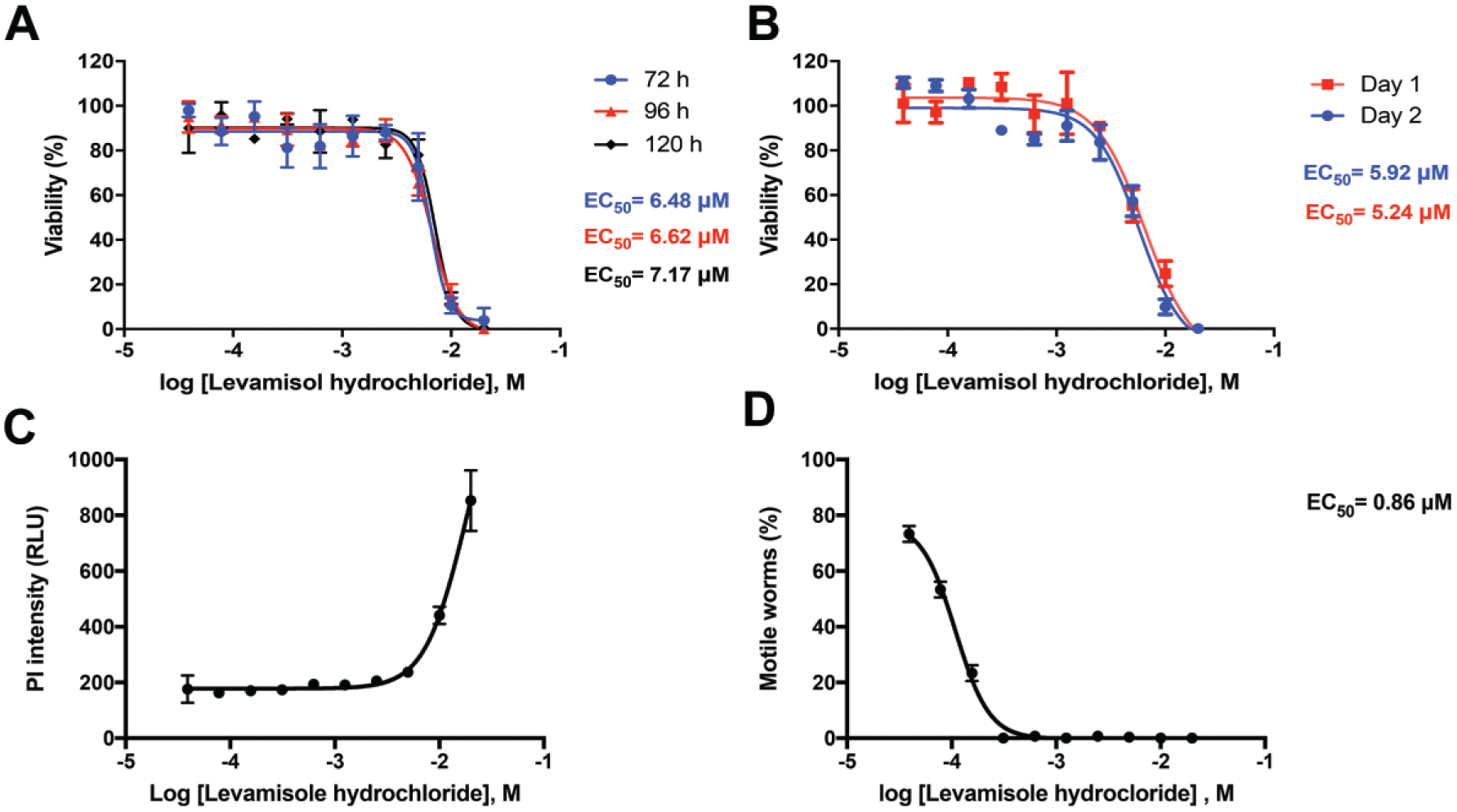
In order to validate the use of levamisole as a positive control of the assays, it was tested in dose–response and incubated for 72, 96, and 120 h following the assay predefined conditions. After establishing the adequate incubation time, the experiment was performed on 2 different days, to evaluate the variability of the EC50 among the experiments.
The Pathogen Box library (www.pathogenbox.org) was reformatted to 384-well plates in 10 mM DMSO. The plates were received in 96-well plates (10 µL/well) at a stock concentration of 10 mM. Samples were manually transferred onto 384-well polypropylene stock plates (784201, Greiner Bio-One, São Paulo, Brazil). Stock plates were kept frozen at −20 °C, protected from light, and submitted for a maximum of three freeze–thaw cycles. The screening was performed according to the assay setup described previously, and in order to validate the methodology and check the reproducibility, the assay was performed twice, on 2 different days.
Hit Analysis
Samples that showed normalized activity greater than 50% were visually inspected and false positives were removed (primarily wells in which the compound had precipitated, interfering with the worm recognition by the software). After the visual confirmation the approved samples were selected as potential hits for further analysis.
Results and Discussion
We initially tested the ability of fluorescent BTD derivatives to stain C. elegans; these derivatives, in general, were very efficient to stain cancer cells, as we have already demonstrated. 24 BTD derivatives were selected because of their outstanding photophysical properties, including high photostabilities, and no notable blink or fade-off (naked eye) during biological experiments using cells 24 or complex models such as C. elegans. 25
From all BTD derivatives tested that efficiently stained the C. elegans, DB-1, whose structure is shown in Figure 1A , afforded a clean and intense green fluorescent signal ( Fig. 1B ). The software was capable of identifying most of the worms in the field. Some of them were not identified because there was an overlap of the worms, and the software could not separate each one. Considering the use of the dye to stain all the worms, it was pretty satisfactory ( Fig. 1C ). Four fields of the well were analyzed by the software and also visually counted using brightfield microscopy, and a comparison of the number of worms counted was performed. There was no statistically significant difference observed between the two methods used (p < 0.05), demonstrating that the DB-1 staining allows the software to recognize the worms effectively ( Fig. 1D ). In this way, DB-1 dye was selected to further application in the current work. Based on the viability analysis of the worm using two fluorescent markers that were incorporated differentially in live and dead worms—PI, which is a DNA intercalator used as a marker for cell viability tests, 26 and DB-1—a high-content screening (HCS) assay was developed to evaluate the activity of compounds against the model organism C. elegans. It has been shown that PI can penetrate only dead cells, whose membranes are compromised, and has also been previously used to determine the viability of C. elegans and other helminths such as Schistosoma mansoni. 8 This selectivity was confirmed through the observation in lightfield microscopy of the pharynx pumping rate and measuring the intensity of PI emitted by the worms; the ones treated with 5 and 20 mM levamisole did not show any pharynx pumping and the ones treated with M9 and 2% DMSO showed an average pumping rate of 12 contractions/10 s ( Suppl. Fig. S1 ).
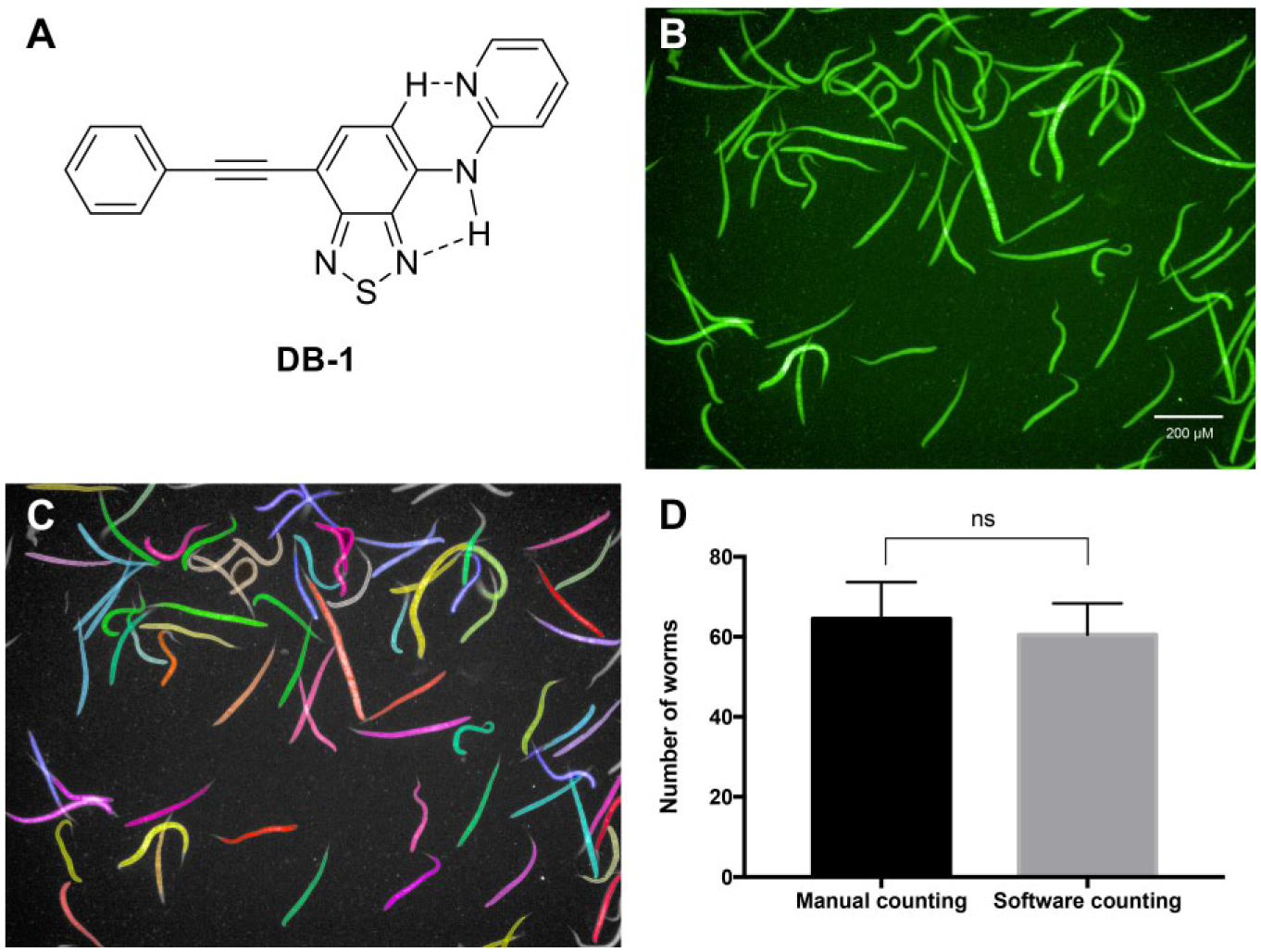
Applications of DB-1 dye in C. elegans staining and analysis. (
The measured average intensity of PI in each case is showed in Figure 2A . It is observed that the worms, incubated with 2% DMSO and M9, present a lower intensity of PI, around 150 RLU, while the ones that are dead, and did not show any pharynx pumping, presented higher values, ranging from 250 up to 800 RLU, which is significantly different from the live ones. This differential incorporation of PI in viable and nonviable worms was explored by software analysis in order to determine the viability of the population evaluated in the high-content assay.
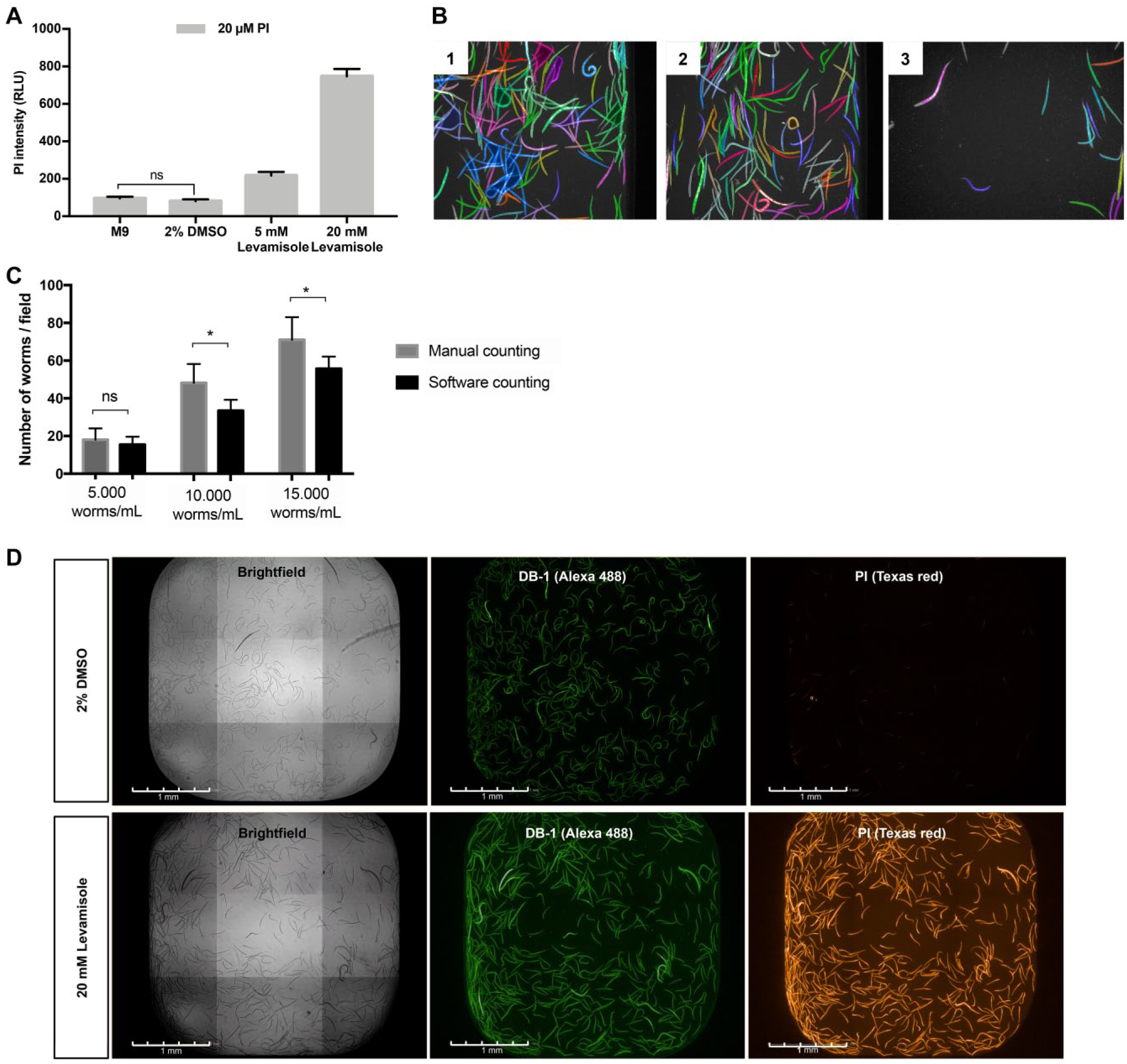
Standardization of live–dead staining protocol using the molecular markers DB-1 and PI. (
In order to ensure that the concentration of the dyes used in the assay was not toxic to the larvae, contributing to a decrease in the viability of the population, different concentrations were tested; Supplemental Figure S2A shows that concentrations up to 50 µM PI and 100 µM DB-1 did not lead to decreased viability of worms incubated previously with 2% DMSO for 72 h, and an additional 24 h with the dyes. Thus, 20 µM PI and 1.56 µM DB-1 were established as final dye concentrations to be used. DB-1, the developed BTD derivative, could be used, as noted, at a considerably lower concentration due to its intense fluorescent signal.
Considering most of the samples tested were stored in DMSO, worm tolerability from 0.5% up to 20% was investigated, and it was shown that concentrations lower than 5% did not decrease the viability of the larvae during the 96 h incubation ( Suppl. Fig. S2B ). It was previously demonstrated that up to 2% was tolerable,27,28 so the concentration of 2% was chosen for the assay setup.
In order to determine the optimal worm number per well that the software is capable of identifying and counting, three different worm suspensions were evaluated. In Figure 2B one field of the well is represented for each worm suspension evaluated; the colored worms represent the identification of the worms by the software, and each individual worm selected is indicated by a different color. The graph in Figure 2C shows the manual counting of the whole well imaged (nine fields), in each condition previously imaged versus software counting. A significant difference was not observed in the number of worms counted manually and with the software when considering the suspension of 5000 worms/mL, but with the higher concentrations of worms evaluated, this difference was significant (p < 0.001). With a higher density of worms in the well, many of them overlap and the software cannot identify a single worm; in this way, the proportion of 5000 worms/mL was chosen to be used in the assay.
A 10× objective was used for imaging acquisition of the conditions tested above, allowing the visualization of an average of 200 larvae in the nine fields acquired, providing a representative number of worms for viability analysis. The image of whole wells containing worms previously treated for 72 h with either 20 mM levamisole or 2% DMSO and stained with 20 μM PI and 1.56 μM DB-1 for an additional 24 h is shown in Figure 2D . Both conditions were able to incorporate the DB-1 dye; however, only a minimum presence of the PI was observed in the worms that were treated with 2% DMSO, likely due to ingestion of the marker and the presence inside the digestive tract. In contrast, the larvae of the levamisole-treated population are intensely stained with PI, confirming the dye selectivity.
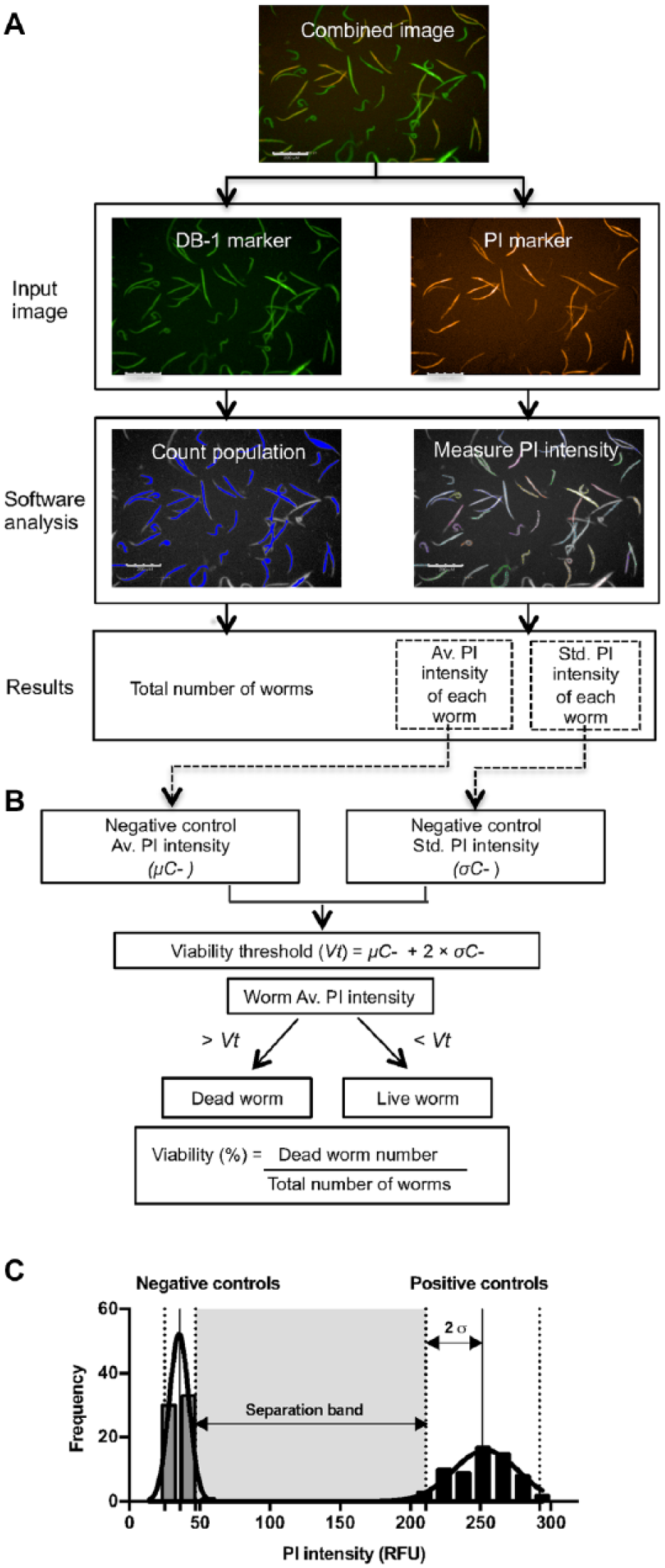
For the high-content analysis of C. elegans viability, the processing system was adjusted to provide the following information: number of total larvae, mean intensity of PI in each of the larvae, number of dead larvae, and percentage of viability. In Figure 3A , the first image called “combined image” represents the superimposition of the two individual images obtained with each of the fluorescent markers used (DB-1 and PI). Each input image was analyzed separately by the software. In the first step, the worm population was identified and counted based on the fluorescence of the DB-1 marker (green fluorescence) emitted by all the worms in the population. Based on the worm area identified, the software analyzed the second image provided by PI marker emission (red fluorescence) and calculated the average as well as the standard deviation of PI intensity in the population selected. As shown in Figure 3C , the PI intensity in the control wells follows a normal distribution; thus, the viability threshold to be used in the analysis to distinguish live from dead worms is then calculated by
where Vt is the viability threshold, μC- is the mean PI intensity of the negative control, and σC- is the standard deviation of PI intensity in the negative control.
High-content analysis of live–dead C. elegans. (
The software analyzes the average fluorescence intensity of the PI in each worm of the well and, based on the established viability threshold, classifies the worms into dead (mean intensity of PI > Vt) and live (mean intensity of PI < Vt). The percentage viability of the population analyzed is the ratio of the number of dead worms to the total number of worms.
With the assay set up, the reference compound levamisole, which acts as a nicotinic acetylcholine receptor agonist in nematode muscle, inducing spastic paralysis, 29 was tested in dose–response with different incubation times. After the incubation, the treated wells were subjected to high-content analysis, and the viability was scored according to the procedure described before. The increase in the incubation time did not interfere in the EC50 value; all the time points evaluated presented an average of 6.76 ± 0.30 mM (mean ± standard deviation) ( Fig. 4A ). The total time of incubation used in the setup of the assay was 96 h. With use of this established condition, the experiment was also performed on 2 different days in order to check the variability between the experiments and showed an average EC50 value of 6.4 mM; no significant difference was observed between the days (Student t test, p < 0.05) ( Fig. 4B ). The EC50 values reported in the literature regarding the activity of levamisole in C. elegans are particularly variable, since the viability is measured with different methodologies, with incubation time and drug concentrations differing from one study to another. In this way, it is difficult to establish a comparison parameter of EC50; its values may vary from 0.5 mM with 72 h of incubation 30 to 0.05 mM with 2 h of incubation. 31 Compound solubility, stability in DMSO solution over time, plate storage conditions, and dilution protocols can all contribute to variances in activities between screening platforms and between laboratories.
Optimization of reference compound incubation time. (
Through screening of the Pathogen Box library, a collection of 400 compounds with known anti-infective activity, the C. elegans HCS assay was validated. Figure 5A shows the assay scheme with all the steps involved in the screening process. The library was screened in two independent runs, at 50 µM. The Z′ factor obtained at run 1 was 0.80 ± 0.08 (mean ± standard deviation), and 0.75 ± 0.05 at run 2, demonstrating that the assay is robust. Figure 5B shows the average dispersion of the compounds and controls used according to their normalized activity in the two assays performed. A linear correlation between the runs was observed (R2 = 0.88) ( Fig. 5C ), demonstrating that the results are reproducible. As part of the validation process, the images from wells containing compounds that presented a percentage of normalized activity greater than 50% were visually inspected to confirm the results. Some of these compounds had formed precipitates that interfered with image analysis, generating false positives. After this visual inspection, three compounds were selected as hits: MMV007920, MMV688938, and MMV688766.
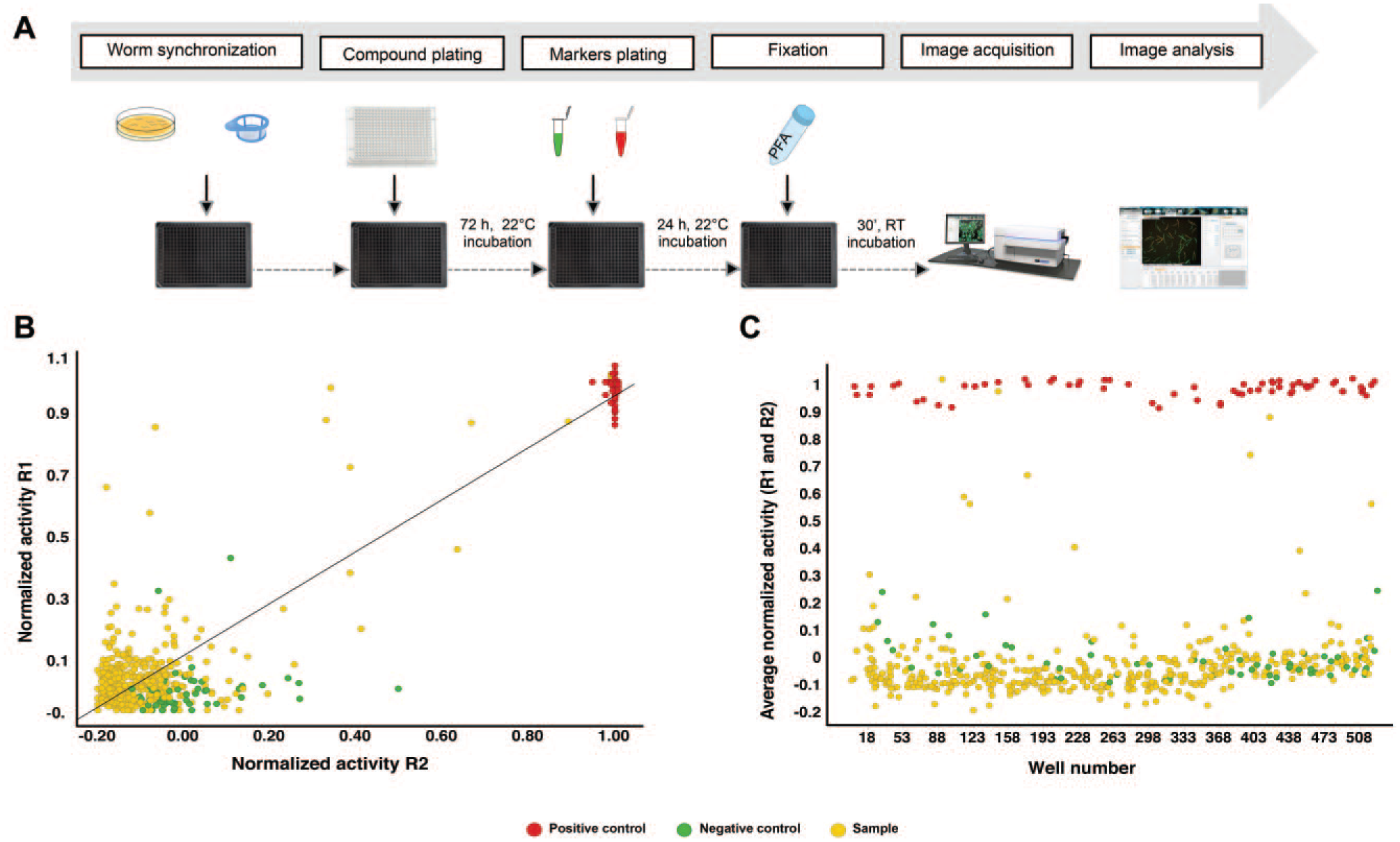
Overview of the Pathogen Box screening in the C. elegans HCS assay. (
The new HCS strategy described here using the model nematode C. elegans is based on image acquisition and analysis after incubating worms with two different cell markers. One of the dyes was DB-1, a fluorescent BTD derivative, which is a green emitter with large Stokes shifts (111–156 nm), as we demonstrated elsewhere. 24 It has in its structure a planar core, that is, the BTD coupled with the aminopyridine moiety on one side bearing a strong electron-withdrawing ability. On the other side, there is a phenylacetylene group capable of donating electrons—thus a classical push–pull molecular architecture of an efficient emitter. As noted in the C. elegans staining assays, DB-1 returned a clean and intense signal and therefore was the dye selected to further the application in the current work. The use of two different fluorescent dyes allows the software to recognize all the worms in the field (which is not possible using only brightfield and PI), allowing for the screening of hundreds of compounds in the same well plate. The greatest differential of this assay is the fact that it is not based on the evaluation of worm motility, as some of the recently developed assay works.32,33 The evaluation of motility can be subjective, and in many cases the worms have to be touched with fine needles to check if they are really immotile. Moreover, there are some drugs that could paralyze the worms for some time, which does not mean that they are dead. Previously reported assays that also use PI intensity in order to evaluate the C. elegans viability rely only on the fluorescence measurement of the whole well. 34 Our image-based assay is capable of identifying each of the worms treated in the well and evaluating their individual viability; moreover, image acquisition allows the evaluation of many other parameters, such as the number of worms and the average PI intensity of each worm, a configuration of a high-content assay.
The measurement of the incorporation of live and dead markers has already been used in cell toxicity assays 35 and also in some anthelminthic assays.8,16,34 Considering that many fluorophores are used in viability measurements, PI and DB-1 were shown to be suitable for the differential staining of live and dead worms, contributing to the development of a reliable and simple image-based assay that has the advantage of identifying dead worms, not only immotile ones. The suitability of the assay as a mechanism to identify compounds with activity against C. elegans was also demonstrated by assessing the Pathogen Box library for potential hits. The high Z′ factor values obtained in all the assays (>0.5) demonstrate that the assay is of excellent quality and is a promising tool for drug discovery. 36
The screening of new compounds using libraries such as the Pathogen Box leads to the possibility of repurposing compounds previously tested in other pathogenic models and finding new applications for known molecules. This approach significantly reduces the time, cost, and effort of the process, because much information, such as the pharmacology and toxicology of the existing drug, is already known.
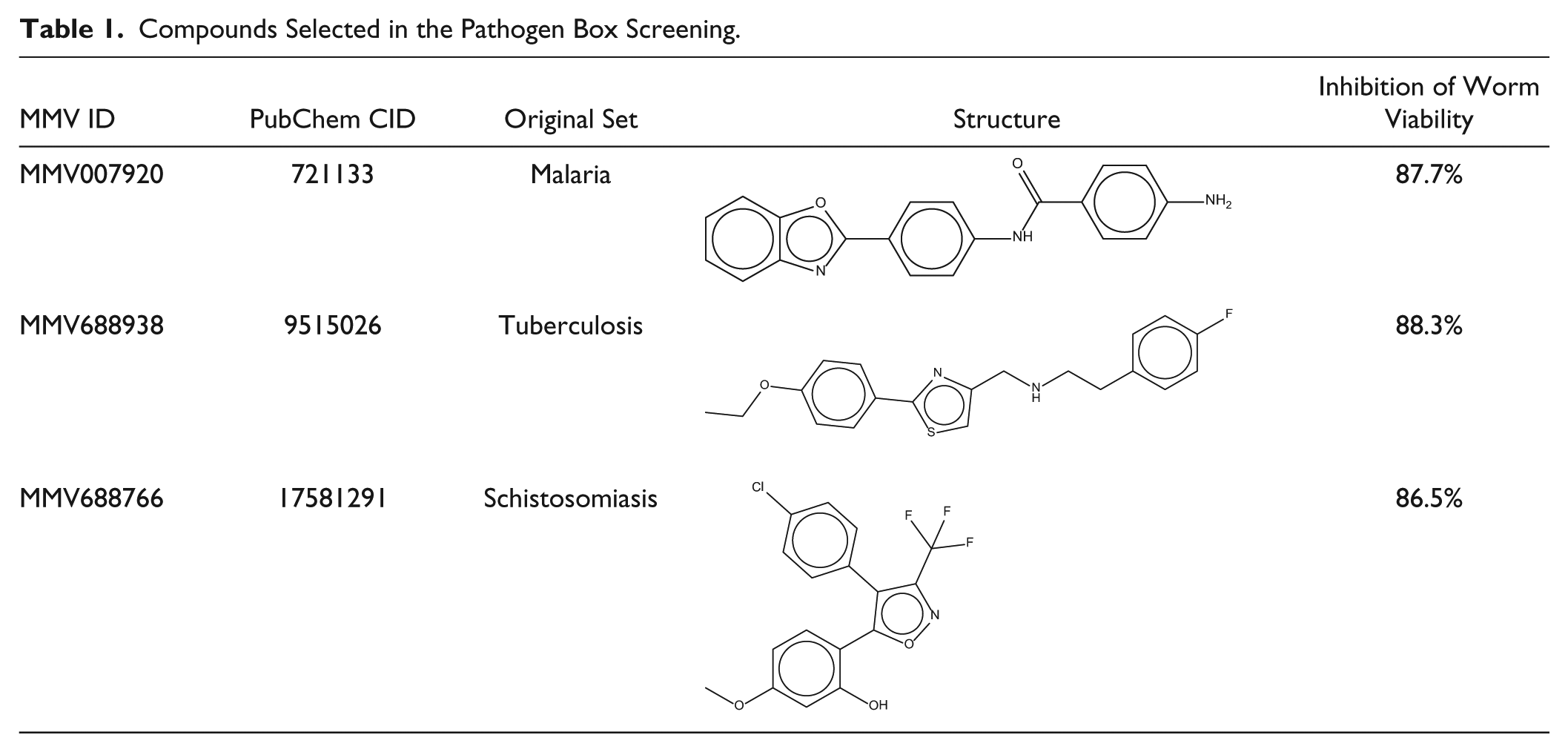
The active compounds identified in this screening have been previously evaluated in trials for some specific pathogens, and their structures are shown in Table 1 . The compound MMV007920 is a 4-amino-N-[4-(1,3-benzoxazol-2-yl)phenyl]benzamide, previously identified in an antimalarial screen and recently reported as an inhibitor of C. elegans growth. 37 It is a heterocyclic compound, derivative of benzoxazole, a compound that is reported to be a biologically potent molecule, showing anthelmintic and antioxidant properties. 38 The compound MMV688766, a trisubstituted isoxazole derivative present in the set originally tested for schistosomiasis, had also demonstrated antiplasmodial activity. 39 MMV688938 was previously described as having potential to inhibit Candida albicans SC5314 biofilm formation 40 and is present in the tuberculosis set. This is the first time that these three molecules are described as nematicidal against a C. elegans model. Concerning the cytotoxicity, the compounds of the library have been previously tested in some cell lines and showed adequate selectivity against the pathogens evaluated, being considered adequate for initial drug discovery. The compounds MMV007920 and MMV688938 were tested in HEP-G2 cells and presented CC50 values of >10 and 28.5 μM, respectively. MMV688766 showed a CC50 value of 8.31 μM in MRC-5 cells (data available in https://www.pathogenbox.org/about-pathogen-box/supporting-information). Thus, these results show that the viability of the developed HCS assay is robust for the screening of compound libraries and the discovery of novel compounds for anthelminthic drug discovery.
Compounds Selected in the Pathogen Box Screening.
The current study has therefore validated a new high-content assay using a C. elegans model as a surrogate model for anthelmintic drug discovery and identified three new molecules with activity against this model nematode that may warrant further investigation and add value to the current drug discovery efforts. This was possible through use of a fluorescent BTD derivative (DB-1) because of the high fluorescence intensity and stability. In this context, the toolbox of biological applications of fluorescent BTD derivatives has just been expanded.
Supplemental Material
Supplemental_material_DISC851130 – Supplemental material for Expanding the Biological Application of Fluorescent Benzothiadiazole Derivatives: A Phenotypic Screening Strategy for Anthelmintic Drug Discovery Using Caenorhabditis elegans
Supplemental material, Supplemental_material_DISC851130 for Expanding the Biological Application of Fluorescent Benzothiadiazole Derivatives: A Phenotypic Screening Strategy for Anthelmintic Drug Discovery Using Caenorhabditis elegans by Giovana A. S. Cintra, Brenno A. D. Neto, Pedro H. P. R. Carvalho, Carolina B. Moraes and Lucio H. Freitas-Junior in SLAS Discovery
Footnotes
Acknowledgements
We thank Prof. Ricardo Fujiwara at Universidade Federal de Minas Gerais for provision of the Bristol N2 C. elegans strain, Prof. Carlos Eduardo Winter at Universidade de São Paulo for provision of the laboratory structure to complete the project, Dr. Bruno dos Santos Pascoalino for technical help, and the Medicines for Malaria Venture (MMV) for providing the Pathogen Box. B.A.D.N. also acknowledges CNPq, CAPES, FAPDF, FINATEC, and DPP-UnB.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project was financially supported by the Medicines for Malaria Venture (MMV) through the “Exploiting the Pathogen Box Challenge Grant” (process no. PO15/01083[7]), and the Fundação de Amparo à Pesquisa do Estado de Sao Paulo (FAPESP process no. 2015/206326-7) and CAPES (process 23038.006737/2012-56) through a PhD fellowship for G.A.S.C.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.