Abstract
Protein kinases are attractive targets for both biological research and drug development. Several assay kits, especially for the detection of adenosine diphosphate (ADP), which is universally produced by kinases, are commercially available for high-throughput screening (HTS) of kinase inhibitors, but their cost is quite high for large-scale screening. Here, we report a new enzyme-coupled fluorescence assay for ADP detection, which uses just 10 inexpensive, commercially available components. The assay protocol is very simple, requiring only the mixing of test solutions with ADP detection solution and reading the fluorescence intensity of resorufin produced by coupling reaction. To validate the assay, we focused on CDC2-like kinase 1 (CLK1), a dual-specificity kinase that plays an important role in alternative splicing, and we used the optimized assay to screen an in-house chemical library of about 215,000 compounds for CLK1 inhibitors. We identified and validated 12 potent inhibitors of CLK1, including a novel inhibitory scaffold. The results demonstrate that this assay platform is not only simple and cost-effective, but also sufficiently robust, showing good reproducibility and giving similar results to those obtained with the widely used ADP-Glo bioluminescent assay.
Introduction
Protein kinases are one of the largest families in eukaryotes, catalyzing protein phosphorylation and thereby mediating signal transduction pathways in many cellular processes, including proliferation, differentiation, metabolism, and cell cycle regulation. Hyperactivation or deficiency of various protein kinases is associated with a number of diseases, and so they are considered attractive targets for both biological research and drug development. Fasudil was the first kinase inhibitor to be approved for clinical use (for cerebral vasospasm) in 1995, followed by more than 30 other kinase inhibitor drugs. 1 Most of them are anticancer drugs, such as imatinib, a Bcr-Abl kinase inhibitor, and gefitinib, an EGFR tyrosine kinase inhibitor, but there are others, such as tofacitinib, a Janus kinase (JAK) inhibitor, which was recently approved for treatment of rheumatoid arthritis.2,3 More than 130 kinase inhibitors are currently in clinical trials for a range of diseases. Nevertheless, it is noteworthy that currently approved drugs or kinase inhibitors under advanced development cover only about 10%–15% of the whole kinome, suggesting that many other kinases remain to be explored as potential therapeutic targets.4,5
CDC2-like kinase 1 (CLK1) is a dual-specificity kinase involved in the regulation of alternative splicing,6–9 that is, combinatorial rearrangement of exons and/or parts of introns into mature RNA, which occurs during the expression of more than 95% of the genes in the human genome and influences many biological cell processes, such as apoptosis, angiogenesis, and neuromuscular degeneration. 10 Therefore, CLK1 inhibitors are considered to be candidate drugs for the treatment of various disorders, including Alzheimer’s disease. 11 The first selective kinase inhibitor of CLK1, TG003, was discovered by Hagiwara and coworkers in 2004, 12 and since then other CLK inhibitors, including KH-CB19, 13 Leucettine L41, 14 and other compounds,15–18 have been reported by many groups, although none of these inhibitors are yet in clinical use. Here, we aimed to find new kinase inhibitors with potential as drug leads or research tools.
Many assay technologies have been developed for high-throughput screening (HTS) to find kinase inhibitors, including 32 P-labeled adenosine triphosphate (ATP) incorporation assays, such as the scintillation proximity assay (SPA; PerkinElmer, Waltham, MA), and many peptide-based kinase assays, such as the HTRF KinEASE homogeneous time-resolved fluorescence immunoassay (Cisbio Bioassays, Inc., Codolet, France) using biotin-tagged substrates and IMAP assays (Molecular Devices, Sunnyvale, CA) measuring the fluorescence polarization signal of fluorescent substrates.19–24 Among them, the adenosine diphosphate (ADP) quantitative assay may be the most commonly used platform for HTS nowadays, because ADP is the universal product of all kinases, in contrast to the phosphorylated products, which vary widely and are sometimes difficult to detect. These ADP assays include the ADP-Glo assay (Promega, Madison, WI), in which ADP is converted to ATP and the ATP is measured by means of luciferase/luciferin reaction;25–27 the Transcreener fluorescence polarization assay (Bellbrook Labs, Madison, WI) using anti-ADP antibody and far-red fluorescence-labeled ADP;28–30 and the ADP-Hunter fluorescence assay (DiscoverX, Fremont, CA), which generates H2O2 from ADP by enzyme coupling reaction followed by production of resorufin.31,32 These generic ADP detection assay platforms do not employ radiolabeling and are homogeneous, so they are suitable for HTS, but their cost is quite high for large-scale screening, and this may be an issue especially in academia.
Here we report a new one-step, enzyme-coupled assay for ADP detection that uses 10 inexpensive, commercially available components and requires only mixing of test solutions with ADP- detection solution before reading the fluorescence intensity of resorufin produced by the coupling reaction. To validate this fluorescence assay, we optimized it to measure CLK1 activity and conducted HTS of an in-house chemical library of about 215,000 compounds. Our results confirm that this simple, inexpensive fluorescence assay is highly sensitive and robust. The results are essentially equivalent to those of the ADP-Glo assay, which is used widely for kinase screening.
Materials and Methods
Materials
Recombinant human CLK1, CK1δ, DYRK1A, and GSK3α were purchased from Carna Biosciences (Kobe, Japan). The substrate peptide Dyrktide was purchased from AnaSpec (San Jose, CA). Nucleoside diphosphate kinase (NDPK; 1.9 kU/mg protein) from baker’s yeast, phosphatase inhibitor cocktail 3, the LOPAC1280 compound library, and ATP (purity ⩾ 99.0%) were obtained from Sigma-Aldrich (St. Louis, MO). ADP-hexokinase (52.6 U/mg solid) from Thermococcus litoralis was purchased from Asahi Kasei Pharma (Tokyo, Japan). Diaphorase I (1.8 kU/mg protein) from Bacillus stearothermophilus was from Nipro (Osaka, Japan). Recombinant glucose-6-phosphate dehydrogenase (G6PDH; 754 U/mg protein) from Leuconostoc sp. and NADP+ were from Oriental Yeast Co. (Tokyo, Japan). Triton X-100 and Tween 20 were purchased from Alfa Aesar (Lancashire, UK) and Tokyo Chemical Industry (Tokyo, Japan), respectively. ADP, dithiothreitol (DTT), N-ethylmaleimide (NEM), bovine serum albumin (BSA), resazurin, and other chemicals were from Fujifilm Wako Pure Chemical (Osaka, Japan). Small-volume, nonbinding, 384-well black microplates and 384-well polypropylene microplates were purchased from Greiner Bio-One (Frickenhausen, Germany) for kinase assay and for electrophoretic mobility shift assay, respectively. FL-Peptide 34 (5-FAM-RRRFRPASPLRGPPK-OH) was purchased from PerkinElmer and myelin basic protein (MBP) was from Fermeus (Turku, Finland).
Kinase Assay of CLK1
CLK1 assay was performed in small-volume, nonbinding, 384-well black plates at a final volume of 5 µL/well. Test compounds (1 mM solution in DMSO; 50 nL/well) were dispensed into wells of columns 3–22 of the plates using an Echo 555 acoustic dispenser (Labcyte, Sunnyvale, CA), and 50 nL of DMSO was dispensed into the remaining wells. Then, a Multidrop Combi Reagent Dispenser (Thermo Fisher Scientific, Waltham, MA) was used to add a solution of CLK1 (7.2 nM, 2.5 µL/well) in buffer A (20 mM HEPES [pH 7.4], 0.01% Tween 20, 2 mM DTT, 5 mM MgCl2). The reaction was initiated by the addition of 2.5 µL of substrate solution (50 µM Dyrktide, 200 µM ATP in buffer A) into the wells. The plates were incubated at room temperature for 2 h in a closed container with well-wetted tissues. Phosphorylation reactions were stopped and quantified using an enzyme-coupled fluorescence ADP detection assay as described below.
ADP Detection Assay
Five microliters per well of kinase reaction solution or ADP solution (10 µM for reference assay) was mixed with 5 µL aliquots of ADP detection reagent (2 mM glucose, 200 µM NADP+, 100 µM resazurin, 20 mM NEM, 2 U/mL ADP-hexokinase, 2 U/mL G6PDH, and 2 U/mL diaphorase I in buffer B [10 mM MgCl2, 0.01% w/v Triton X-100, 0.01% w/v BSA, 100 mM Tris-HCl, pH 7.5]) in 384-well plates, and the plates were incubated at room temperature for 60 min in the dark. Fluorescence intensity due to the formation of resorufin was measured with a PHERAstar microplate reader (BMG Labtech, Offenburg, Germany) with excitation at 540 nm and emission at 590 nm. NEM, a thiol reagent, was included in the ADP detection reagent to terminate the target kinase reaction as well as to block the reactive thiol in DTT, which is often used in kinase assays as a kinase activator.
Kinase Selectivity Assay
Kinase selectivity assays for CLK1, CK1δ, DYRK1A, and GSK3α were performed in basically the same way as described for the CLK1 assay, except that MBP was used as the substrate at a concentration of 1 µM. Kinase concentrations were varied depending on the activity—3.6 nM for CLK1, 16 nM for CK1δ, 2.2 nM for DYRK1A, and 6.4 nM for GSK3α—and the incubation time was 2 h. Phosphorylation reactions of all kinases were evaluated using an enzyme-coupled fluorescence ADP detection assay.
Electrophoretic Mobility Shift Assay
Twenty microliters of 5.2 nM CLK1 solution, 20 μL of 4.5 μM FL-peptide 34 and 300 µM ATP, and 20 μL of test compound solution (5% DMSO) were added per well and the plates were incubated for 2.5 h at room temperature in buffer A; then the reaction was stopped by addition of 3 μL of 500 mM EDTA. Phosphorylated and unphosphorylated peptides were separated and detected with a Lab-Chip EZ Reader II (PerkinElmer).
Data Processing and Analysis
The percentage inhibition was calculated based on the DMSO control as 0% inhibition, and the background (no-kinase) control as 100% inhibition. The calculated percent inhibition values were analyzed by TIBCO Spotfire (Somerville, MA) and used for the calculation of IC50 values by fitting the dose–response curves to a four-parameter logistic model in GraphPad Prism (GraphPad Software, La Jolla, CA).
Results and Discussion
Optimization of Enzyme-Coupled Fluorescence Assay for CLK1 Screening
We have previously reported on a universal enzyme-coupled fluorescence assay for glycosyltransferases. 33 It employs a nucleotide kinase reaction to generate ADP from the nucleotides produced in the glycosyl transfer reaction, followed by a cascade of coupled enzyme reactions. Here, we used a part of this enzyme-coupled reaction cascade for the detection of ADP produced by kinase activity and we optimized the assay for kinase HTS, as shown in Figure 1A . The assay protocol is simple, requiring only the mixing of test solutions with ADP detection solution, followed by reading the fluorescence intensity of resorufin. The reaction cascade requires the use of only 10 inexpensive, commercially available components (dotted line in Fig. 1A ): three enzymes (ADP-hexokinase, G6PDH, and diaphorase), their substrates (glucose, NADP+, and resazurin), and additives (BSA, NEM, MgCl2, and Triton X-100) in Tris buffer.
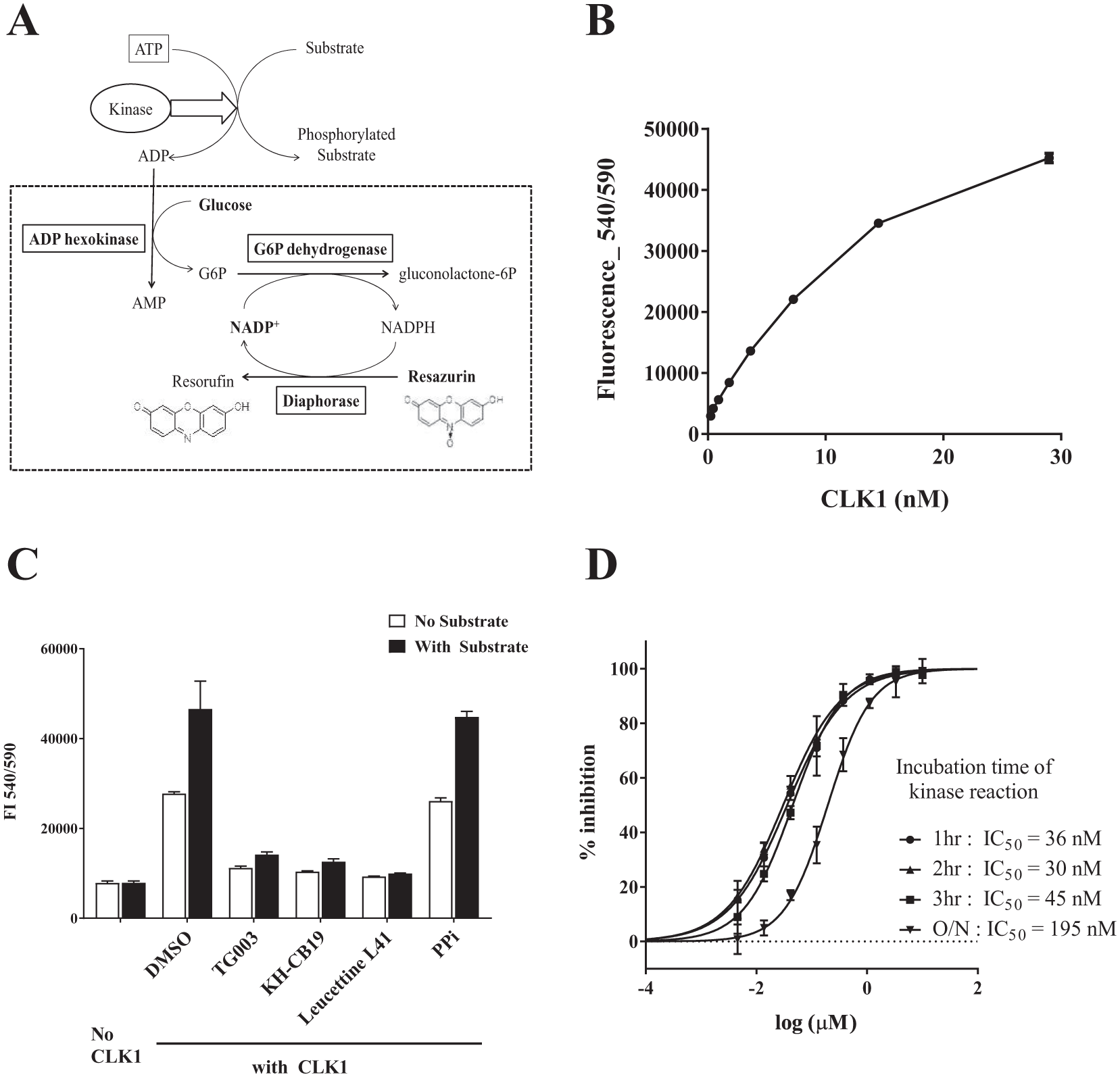
Development of the enzyme-coupled ADP detection assay and optimization for CLK1 inhibitor screening. (
To validate this ADP detection assay for kinase screening, we optimized it for HTS of CLK1 inhibitors. We first confirmed the linearity of the ADP standard curve from 1 to 40 µM ADP, and established that 1.2 µM ADP gave a sufficient signal-to-background (S/B) ratio of more than 2 (the signal of 1.2 µM ADP was 2554 vs the background signal of 1014). In addition, this fluorescence signal was stable, being unchanged after overnight incubation at room temperature ( Suppl. Fig. S1A ). We previously showed that additives such as 200 mM NaCl, 10 mM MnCl2, 5 mM EDTA, and 1 mM ATP did not affect the fluorescence intensity, and 2 mM DTT in the kinase solution also had no effect. 33 Next, we examined the dose dependency of CLK1 and chose a final CLK1 concentration of 3.6 nM from the viewpoints of reaction linearity, good S/B ratio (larger than 2), and economy of kinase samples ( Fig. 1B ). We also checked the time course of the CLK1 reaction and found that the fluorescence intensity increased linearly up to 3 h, so the incubation time of the kinase reaction was set at 2 h ( Suppl. Fig. S1B ). We confirmed that DMSO concentrations up to 5% were well tolerated ( Suppl. Fig. S1C ).
Because of the principle of ADP detection in the kinase assay, contamination of samples with ATP-consuming enzymes could be a serious problem; for example, if phosphatase were present in the kinase solution, ATP would decompose to ADP, providing a false-positive result. To avoid this issue, we tested ADP production in the absence of substrate, and we also examined the effect of phosphatase inhibitors. In the absence of substrate, the reaction of CLK1 amounted to almost half of that with the substrate, but known CLK1-specific inhibitors suppressed the reaction nearly to baseline, while phosphatase inhibitors had no effect ( Fig. 1C ). This indicates that ADP production in the absence of substrate is likely derived from CLK1-specific autophosphorylation 11 and intrinsic hydrolysis of ATP, 34 and that the decrease of ADP production in this assay can be used for the evaluation of kinase inhibitors.
Under the optimized assay conditions, we examined the IC50 values of known CLK1 inhibitors TG003, Leucettine L41, and KH-CB19; the values obtained were 102, 34, and 12 nM, respectively, which are reasonably close to the reported values of 49, 15, and 20 nM ( Suppl. Table S1 ).13,14 We also observed that the IC50 values of Leucettine L41 measured at intervals over a 3 h incubation period were 30–40 nM, which indicates that the stability of this assay is well maintained at least to 3 h under these conditions ( Fig. 1D ). We also confirmed the stability of the fluorescence readout and IC50 value after addition of detection reagent. Again, the measured IC50 value remained essentially constant for at least 3 h ( Suppl. Fig. S1D ).
Based on the above results, we fixed the CLK1 assay conditions for HTS as follows: CLK1 concentration 3.6 nM, Dyrktyde substrate concentration 25 µM, ATP concentration 100 µM, and incubation time 2 h at room temperature. The Km value of CLK1 for ATP is reported to be 11 µM, 35 but we used 100 µM ATP for HTS to obtain sufficient fluorescence signals.
Pilot Screening Using a Known Bioactive Library
To assess the accuracy and usability of this fluorescent kinase assay for HTS, we performed a pilot screen of 1279 LOPAC compounds (one restricted compound was excluded from the LOPAC library) at 10 µM. The overall quality of four assay plates was high, with average values of S/B = 3.96, Z′ = 0.86, coefficient of variation (CV) of high control (HC) = 2.7%, and CV of low control (LC) = 3.4%.
We also conducted a reference assay in the absence of CLK1 but in the presence of 10 µM ADP to detect false positives due to inhibition of the coupling enzyme activities. The quality of this reference screening was also high (S/B = 6.32, Z′ = 0.90, CV of HC = 2.5%, and CV of LC = 2.2%), and only one compound, Ro 90-7501, showed more than 50% inhibition; in fact, Ro 90-7501 showed nearly 100% inhibition in both CLK1 and the reference assay, suggesting that it strongly inhibits at least one of the enzymes catalyzing the coupling reactions (
Suppl. Table S3
). Scatterplots of the pilot screening results are shown in
Figure 2A
(CLK1 assay) and
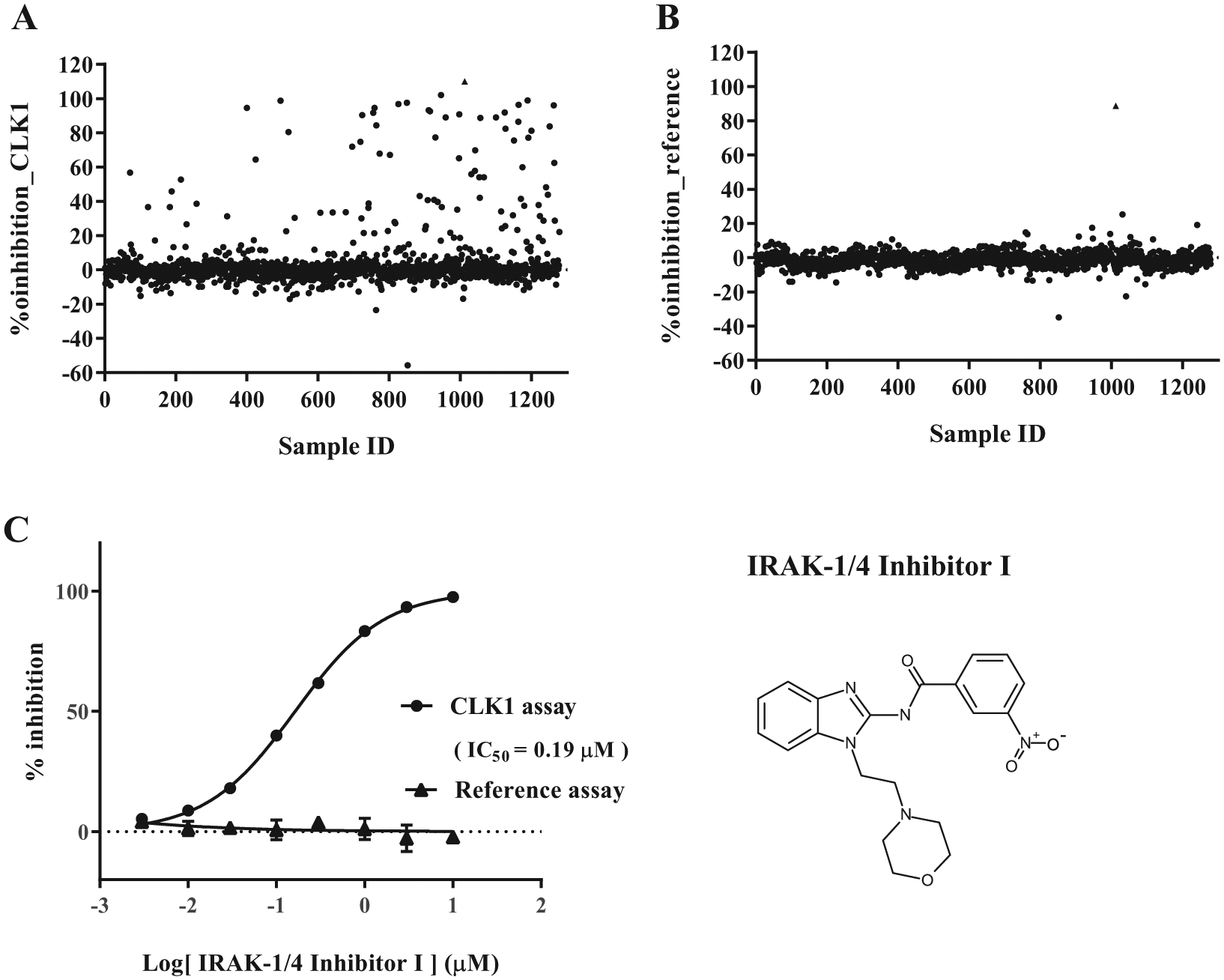
Results of pilot screening. Scatterplots of percent inhibition values of the CLK1 assay (
The number of compounds showing >50% inhibition at 10 μM in the primary screening was 40. Only one compound, IRAK-1/4 inhibitor I, 36 showed more than 50% inhibition at 0.2 μM, and it inhibited CLK1 activity with the IC50 value of 0.19 μM, without showing inhibitory activity in the reference assay ( Fig. 2C ). We also performed electrophoretic mobility shift assay, which detects the phosphorylated substrate directly, and confirmed that this hit compound actually inhibits the phosphorylation of the substrate peptide, with the IC50 value of 0.72 μM. The difference between the IC50 values of two methods could be due at least in part to the difference of the substrates used in the assays.
Thus, our enzyme-coupled fluorescence assay identified a potent inhibitor of CLK1 phosphorylation, which indicates that this ADP detection assay is effective for finding kinase inhibitors.
HTS of DDI Library
Next, we performed HTS of 214,106 compounds from our in-house library (Drug Discovery Initiative [DDI] library) at 10 µM using this ADP detection method to discover potent CLK1 inhibitors. In the HTS campaign, we screened 60 plates per batch and ran one or two batches a day (about 20,000–40,000 compounds per day), which enabled screening of all 214,106 compounds in a week. The cost of detection reagents for the whole primary HTS was only US$2500, which was less than the cost of the plates. Such low costs should greatly extend the availability of HTS in academia.
To ensure robustness of the assay, especially that the sensitivity remained the same throughout the CLK1 HTS campaign, we included positive control plates containing serial dilutions of Leucettine L41 first in every assay batch, and checked the IC50 value as a parameter to monitor the quality of the screening performance. The IC50 values of Leucettine L41 were highly reproducible through the 13 batches of kinase assay (the average value was 48 ± 5.7 nM), suggesting that the sensitivity of this assay did not change during the HTS campaign ( Suppl. Fig. S2 ).
Across the entire 675-plate run of the primary HTS, we obtained very good Z′ (0.90) and S/B (4.5) values, as in the pilot screening ( Fig. 3A ). The percent inhibition values of all 214,106 compounds are plotted in Figure 3B . The compounds showing significantly higher inhibitory activity than 3 SD from the DMSO control (about 30%) can be considered positive hits, but as this resulted in a large number of primary hits (4760, hit rate 2.3%), we selected the 2307 compounds with more than 50% CLK1 inhibition as primary hits (hit rate 1.1%) in the present case.
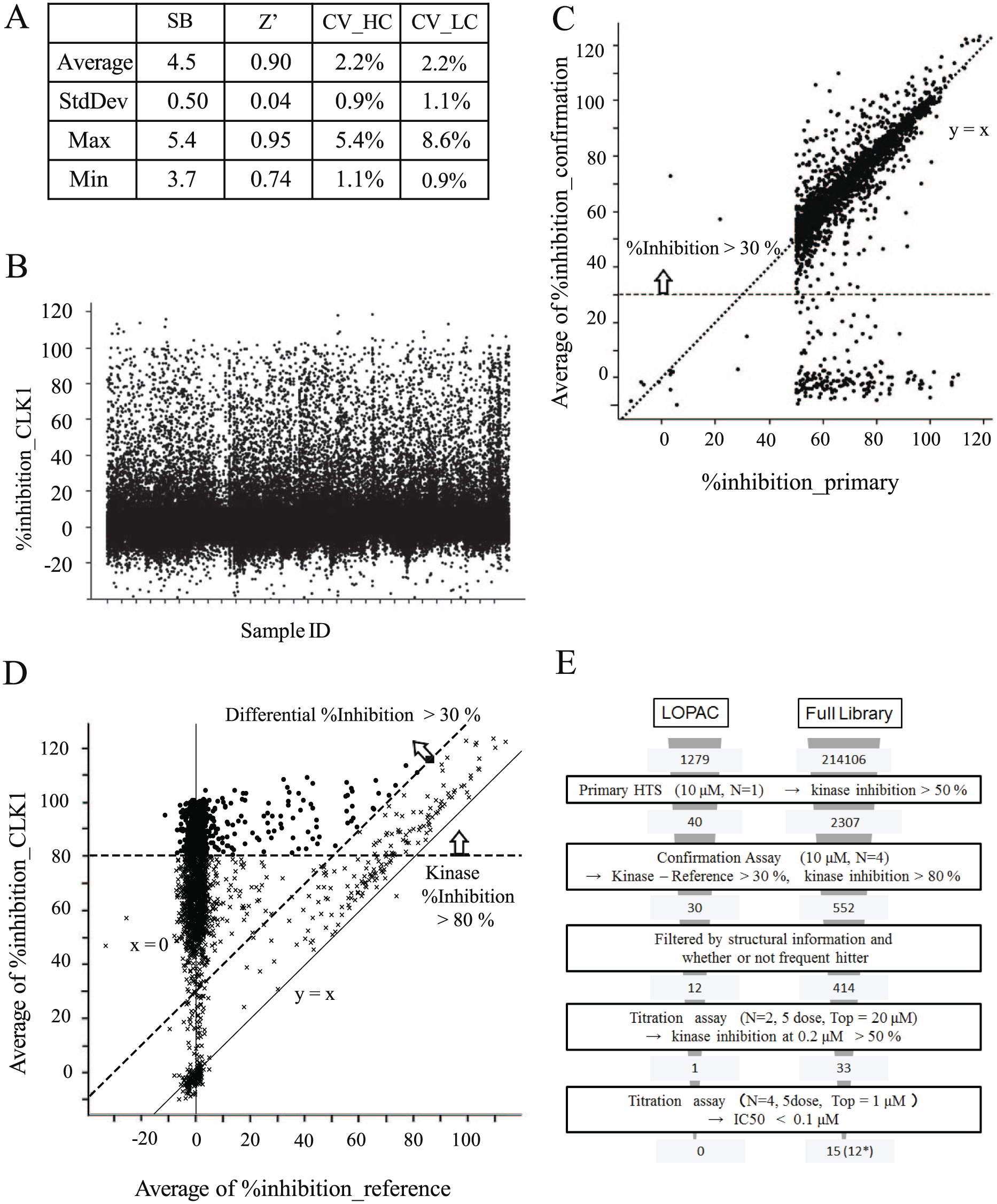
Summary of HTS of the DDI library. (
In order to evaluate the false-positive rate due to compound effects on the coupling reaction of this new assay, we also conducted a reference assay against the same 214,106 compounds; the Z′ (0.88) and S/B (5.7) values obtained were as good as those of the original CLK1 assay ( Suppl. Fig. S3A ). A scatterplot of the whole reference assay is shown in Supplemental Figure S3B , and the correlation plot between the CLK1 and reference assays is shown in Supplemental Figure S3C . We found that 124 compounds showed more than 50% inhibition in the reference assay, so that the false-positive rate (124/214,106 compounds) was 0.06%, which corresponds to 5.4% of the 2307 hit compounds. The rate of false positives as a percent of the set of hit compounds remained almost the same when other criteria, that is, 30% and 40% inhibition, were employed ( Suppl. Table S2 ). False-positive hits included known compounds, such as Ro 90-7501, staurosporine, and dynasore ( Suppl. Table S3 ), which were not excluded from the primary hits, so that the quality and reproducibility of the primary results could be properly evaluated.
Confirmation Assay
Next, we performed CLK1 confirmation assay of all 2307 primary hits in quadruplicate, and in order to validate the assay reproducibility, percent inhibition values in the primary HTS were plotted against the average of those in the confirmation assay. As a result, we found that 2030 compounds (88%) showed inhibitory activity of more than 30%, while the other 277 compounds (12%) did not reproduce their activity in the primary screening activity ( Fig. 3C ). These nonreproducible hits would have arisen during the kinase reaction step, because a similar plot for the reference assay showed high reproducibility and no false positives ( Suppl. Fig. S3D ).
The correlation between the results of the confirmation assay and those of the reference assay is shown in Figure 3D . Compounds that lie near the y = x line are thought to be inhibitors of ADP detection steps, while compounds near x = 0 are likely to be acting on CLK1, and compounds between the y = x line and the y axis are likely to be inhibitors of both CLK1 and coupled enzymes. We selected 552 compounds that showed both more than 80% CLK1 inhibition and more than 30% inhibition after deduction of the reference inhibition from CLK1 in the confirmation assay, and filtered them down to 414 compounds by removing frequent hitters and pan-assay interference compounds (PAINS). 37
Titration Assay
The 414 confirmation hits were tested by five-dose titration assay twice; first from 0.2 to 20 μM and second from 0.001 to 1 μM. All compounds showed IC50 values of less than 10 μM, and finally we identified 15 compounds with very potent activity (IC50 less than 0.1 μM). As final hits, we selected 12 compounds, after excluding 3 with low purity. The flowchart of the whole HTS campaign is shown in Figure 3E .
Final HTS Hit Compounds
The chemical structures and IC50 values of the 12 hits are shown in Table 1 , and the dose–response curves are shown in Supplemental Figure S4 . The hits showed little or no effect in the reference assay, suggesting that hit selection in the confirmation assay had worked correctly. We also performed electrophoretic mobility shift assay for some of them and confirmed that they inhibited the phosphorylation of substrates with IC50 values similar to those determined by the ADP detection assay ( Suppl. Table S4 ).
Hit Compounds in CLK1 HTS.
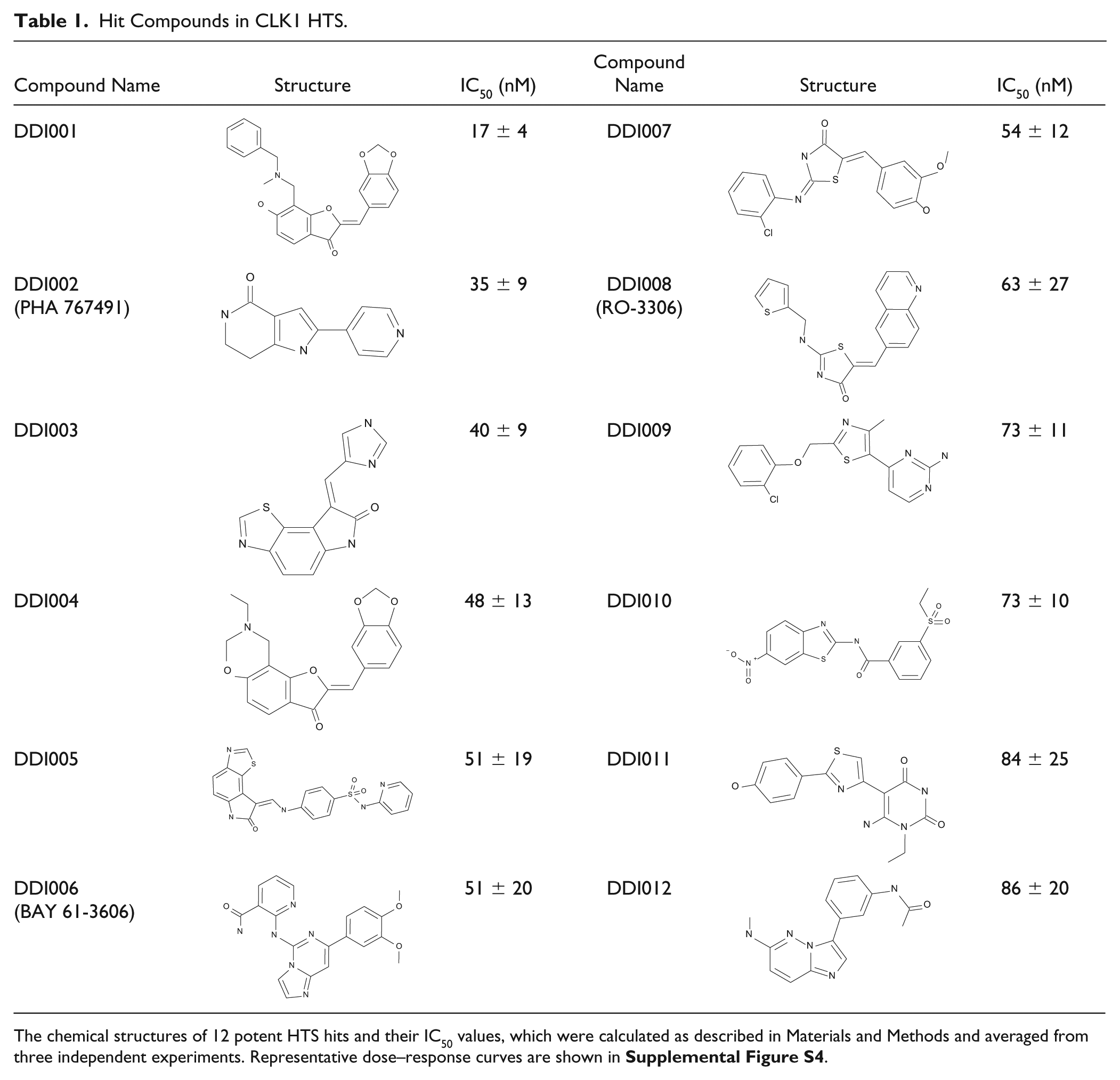
The chemical structures of 12 potent HTS hits and their IC50 values, which were calculated as described in Materials and Methods and averaged from three independent experiments. Representative dose–response curves are shown in Supplemental Figure S4 .
The most potent CLK1 inhibitor, DDI001, has the same scaffold as DDI004, and we found that six other compounds with the same scaffold were included in the 414 confirmation hits ( Table 2 ), suggesting that this could be a novel CLK1-inhibitory scaffold.
Derivatives of Potent CLK1 Inhibitor, DDI001, in the Confirmation Assay of Hit Compounds.
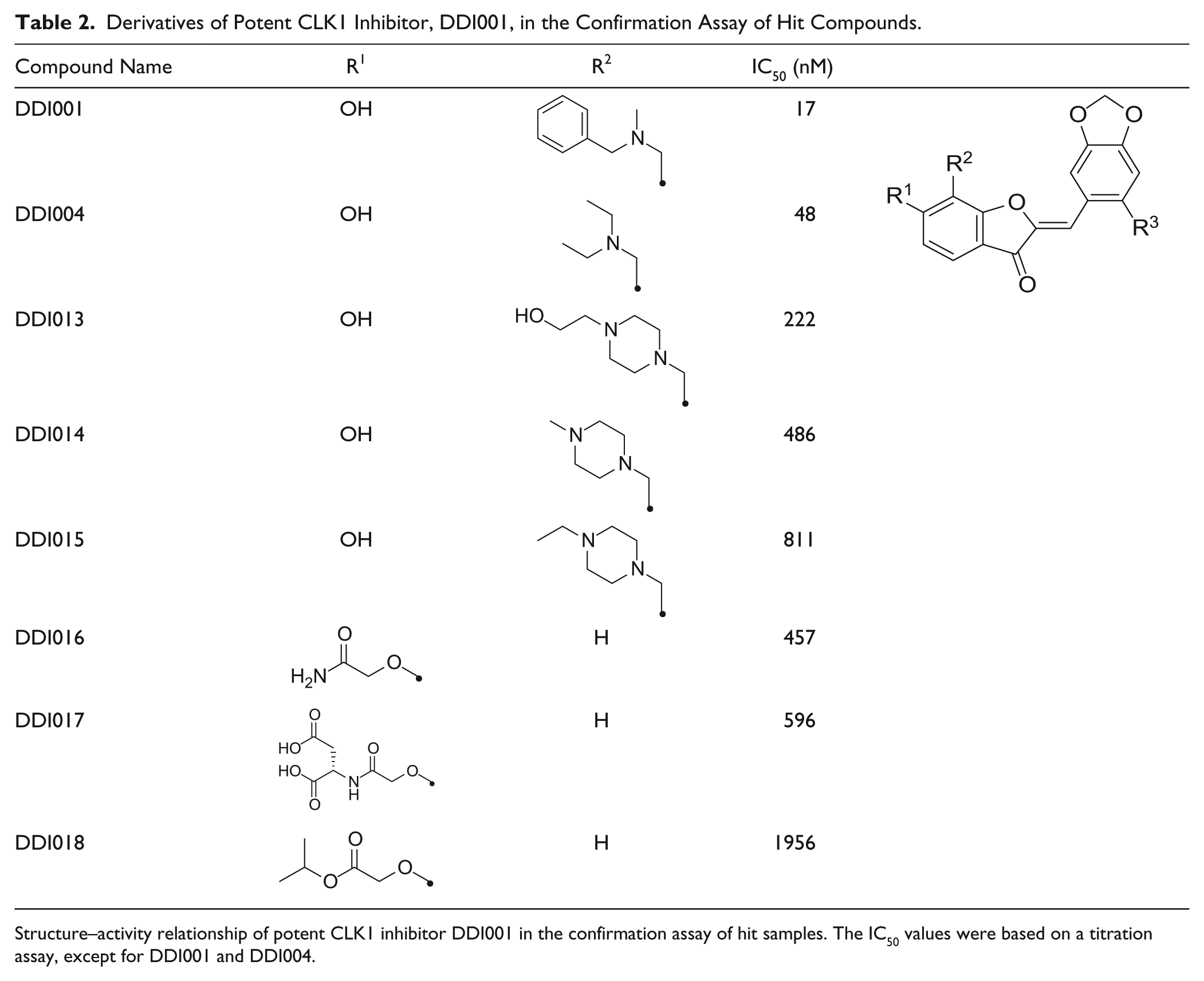
Structure–activity relationship of potent CLK1 inhibitor DDI001 in the confirmation assay of hit samples. The IC50 values were based on a titration assay, except for DDI001 and DDI004.
The kinase selectivity of the final 12 hit compounds was evaluated against a small panel of kinases, CK1δ, DYRK1A, and GSK3α, which are structurally related and reported to be involved in Alzheimer’s disease. 11 In the profiling assay, to minimize time and cost, we used MBP as a common substrate and employed ADP detection with our enzyme-coupled fluorescence assay. 38 As shown in Supplemental Table S5 and Supplemental Figure S5, DDI011 and DDI012 showed the most selective inhibitory activity. Two compounds with the same scaffold, DDI001 and DDI004, showed similar potent inhibitory activity toward CLK1 and GSK3α, less potent inhibitory activity toward DYRK1A, and weak activity against CK1δ.
Thus, our HTS campaign identified several potent inhibitors and a novel inhibitory scaffold for CLK1, although further studies will be needed to improve the selectivity. The present derivatives are thus promising leads for further development as potent pharmacological tools to study alternative splicing, as well as potential therapeutic lead compounds for diseases associated with abnormal alternative splicing.
Comparison with ADP-Glo Assay
The ADP-Glo assay, which is widely used for assaying kinases, is a bioluminescence assay with good sensitivity over a wide range of ATP/ADP concentrations. The ADP-Glo assay protocol consists of two steps after kinase reaction: in the first step, the remaining ATP in the reaction is depleted enzymatically, and subsequently, ADP is converted to ATP, which is measured by means of the luciferase/luciferin reaction. 39 This assay is homogeneous and highly sensitive, but quite expensive. To compare our fluorescence assay with the ADP-Glo assay, we performed a dose–response curve assay for hit compounds and known active compounds using the same kinase reaction conditions in the 384-well format. Figure 4 shows a good correlation of the IC50 values obtained in the two detection assays (the IC50 values are listed in Suppl. Table S6A ), and the scatterplot of percent inhibition values in each well showed good linearity with respect to the fluorescence assay and bioluminescence assay ( Suppl. Fig. S6 ). In addition, the quality control (QC) values indicated that the two assays are nearly equally robust ( Suppl. Table S6B ). Thus, our one-step fluorescence assay for ADP detection is stable and essentially as effective as the widely used ADP-Glo assay, while its cost is less than 1/10 of that of currently available assay kits. Furthermore, it is likely that false positives and false negatives obtained in screening by these two assay formats would be different, at least in part. Therefore, it could be useful to employ both assay formats to confirm the activity of compounds.
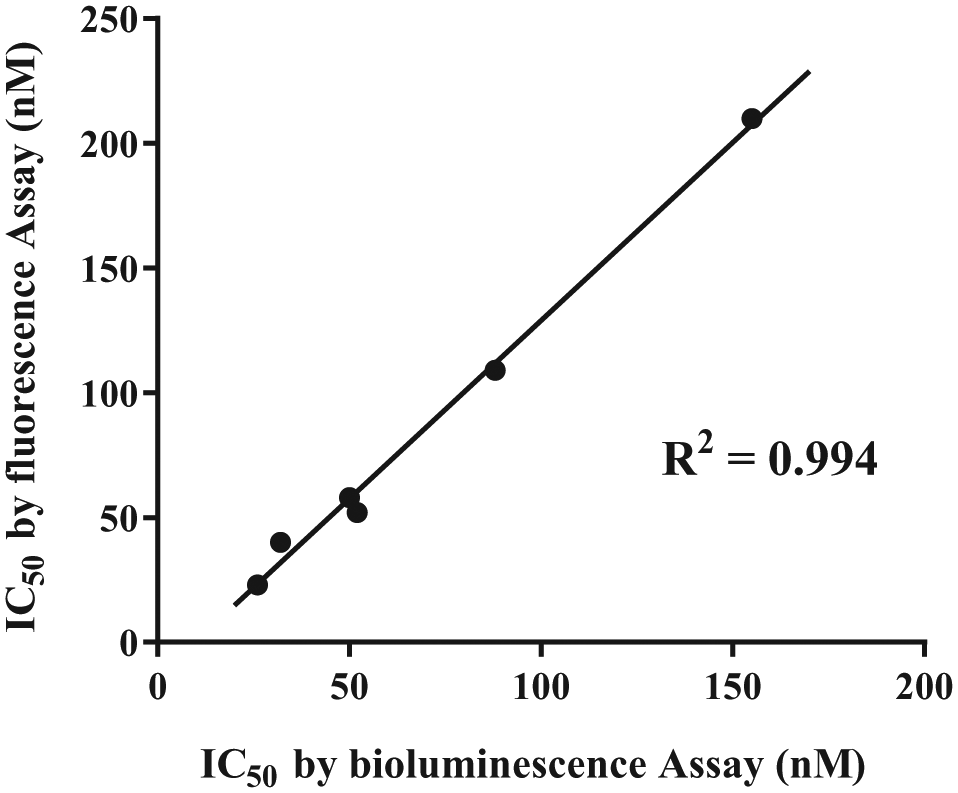
Comparison of our enzyme-coupled fluorescence assay with the ADP-Glo bioluminescence assay. Two parallel kinase assays were conducted in the 384-well format under the same conditions as used for the primary screening and detected by different two methods, one using enzyme-coupled fluorescence assay and the other using the bioluminescence ADP-Glo assay (conducted according to the manufacturer’s protocol). The IC50 values of four potent hit compounds and three known active compounds were calculated from the results of each assay, and the correlation was examined between the fluorescence assay and bioluminescence assay results. The line corresponds to the fitting of a simple linear equation to the data. All IC50 values are listed in Supplemental Table S6A .
In conclusion, we have developed a cost-effective enzyme-coupled fluorescence-based ADP detection assay employing a simple protocol: mixing a kinase reaction solution with the detection solution. In HTS, this assay showed good QC values and robustness, similar to those of the ADP-Glo assay. Application of the assay for HTS of CLK1 inhibitors yielded 12 potent hit compounds and a new CLK1-inhibitory scaffold, which seems suitable for further development. We believe that this assay platform enables simple, high-quality kinase screening at low cost, and it should be useful for the discovery and development of inhibitors for a variety of kinases that are considered candidate therapeutic targets.
Supplemental Material
Supple_Material_for_Inexpensive_HTS_of_Kinase_Inhibitors_by_Imamura_etal_rev_2 – Supplemental material for Inexpensive High-Throughput Screening of Kinase Inhibitors Using One-Step Enzyme-Coupled Fluorescence Assay for ADP Detection
Supplemental material, Supple_Material_for_Inexpensive_HTS_of_Kinase_Inhibitors_by_Imamura_etal_rev_2 for Inexpensive High-Throughput Screening of Kinase Inhibitors Using One-Step Enzyme-Coupled Fluorescence Assay for ADP Detection by Riyo Maruki Imamura, Kazuo Kumagai, Hirofumi Nakano, Takayoshi Okabe, Tetsuo Nagano and Hirotatsu Kojima in SLAS Discovery
Footnotes
Acknowledgements
We thank Shinichiro Egashira for his great help in data analysis by developing an Excel macro. We also thank Tomoko Futamata for her assistance with primary HTS, and Izumi Minegishi, Misato Hara, and other DDI staff for preparation of the plates containing the DDI library compounds.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Platform Project for Supporting Drug Discovery and Life Science Research from the Japan Agency for Medical Research and Development (AMED) under grant number JP18am0101086.
Supplemental material is available online with this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
