Abstract
The Labcyte Echo acoustic liquid handler allows accurate droplet ejection at high speed from a source well plate to a destination plate. It has already been used in various miniaturized biological assays, such as quantitative PCR (q-PCR), quantitative real-time PCR (q-RT-PCR), protein crystallization, drug screening, cell dispensing, and siRNA transfection. However, no plasmid DNA transfection assay has been published so far using this dispensing technology. In this study, we evaluated the ability of the Echo 550 device to perform plasmid DNA transfection in 384-well plates. Due to the high throughput of this device, we simultaneously optimized the three main parameters of a transfection process: dilution of the transfection reagent, DNA amount, and starting DNA concentration. We defined a four-step protocol whose optimal settings allowed us to transfect HeLa cells with up to 90% efficiency and reach a co-expression of nearly 100% within transfected cells in co-transfection experiments. This fast, reliable, and automated protocol opens new ways to easily and rapidly identify optimal transfection settings for a given cell type. Furthermore, it permits easy software-based transfection control and multiplexing of plasmids distributed on wells of a source plate. This new development could lead to new array applications, such as human ORFeome protein expression or CRISPR-Cas9-based gene function validation in nonpooled screening strategies.
Introduction
Nanovolume liquid handlers have opened new ways to perform biological experiments, ranging from micro- to picoliter scale, enhancing the high throughput of various biological assays. Several noncontact dispensing methods have been described and are mostly based on inkjet printing technologies. 1 Out of these, acoustic droplet ejection (ADE) uses focused acoustic energy to control ejection of a specific volume of liquid. 2 The Echo (Labcyte) is an accurate, precise, and efficient device that uses this technology and produces highly reproducible, fully automated, noncontact, and tipless dispensing of liquid at high throughput. 3 Moreover, it is fitted with an integrated dynamic fluid Analysis technology (Labcyte) for the determination of the fluid composition, the liquid height in the source plate, and the power needed to eject 2.5 nL droplets into the destination microplate. 4 Volumes up to 500 nL can be handled by transferring several hundreds of droplets per second. Initially, this device was only able to deliver solutions containing at least 70% DMSO, but new settings have circumvented this limitation to deliver more aqueous solutions. 4 To date, ADE has been successfully used to miniaturize biological applications such as quantitative PCR (q-PCR) and quantitative real-time PCR (q-RT-PCR), drug and RNA interference screening, 5 and protein crystallization 6 and is also expected to be applied soon to high-throughput sequencing 7 and cell dispensing. 8 Thanks to the Cherry Pick software (Labcyte), the user can elaborate a study plan to control the volume to be dispensed and the source well and destination well, within the source and destination plates, respectively. A customized picklist file can be generated in spreadsheet software to enable multiplexing of several fluids dispensing from different source plates in the expected wells of different destination plates.
Given that transfection is the starting point of many cell-based experiments, having an easy and reproducible way to perform it in a high-throughput manner would be an extremely valuable tool, even for nonspecialists. High-throughput transfection allows for rapid optimizations of the parameters, which ensures efficient development of the miniaturized cellular experiments. However, high-throughput transfection is a challenging issue, and whereas several attempts have been successful for siRNA screening, 9 very few examples of DNA plasmid transfection have been reported so far. A jetPEI-based method has been described using the Biomek liquid handling system in 96-well plate format. 10 A microfluidic chip has been developed for the independent transfections of 280 chambers, but it requires specialized skills in this field. 11 Despite all the advantages of the ADE technology, to date no plasmid transfection assay has been developed using this technology. Plasmid DNA transfection is, however, time-consuming and error-prone if various multiplexed conditions and several DNA dilutions are required. An easy, efficient, and high-throughput cell transfection protocol using the ADE technology would therefore be a valuable tool for several reasons: (1) the ability to miniaturize, leading to the use of a smaller amount of the prepared DNA; (2) the high throughput and reproducibility over the manual protocol (even for nonexperts); and (3) the device-controlled dispensing from a predefined picklist, allowing the DNA amount to dispense from a single diluted DNA stock solution, and easy multiplexing of several plasmids’ DNA in the same destination well. Beyond calcium phosphate precipitation-based DNA delivery systems, efforts have been made to enhance DNA cell delivery, such as encapsulating nucleic acids in liposome-like complexes using lipids or cationic polymers or, more recently, using nonliposomal polymeric chemistry systems. 12 However, transfection can be challenging as every cell type has a different set of requirements for optimal introduction of foreign DNA. The transfection methods relying on DNA encapsulation are relatively simple and straightforward and generally provide high transfection efficiency in many cell lines. 13 Reverse transfection protocols have been developed to enhance the throughput and quickly test and combine several plasmid transfections. 14 These reverse protocols seem suitable for use of the Echo liquid nanodispenser, as nucleic acid complexes with the transfection reagent are formed before seeding the cells.
In this study, we have evaluated the ability of an acoustic-based dispensing system to perform plasmid DNA transfection in 384 wells. We simultaneously co-optimized several parameters involved in the transfection process: the total amount of DNA, the nature of the transfection reagent, and the starting DNA concentration. We then improved throughput, evaluated the stability of preloaded plates, and finally implemented optimized conditions to perform co-transfection experiments.
Materials and Methods
Plasmid DNA Vectors
The tdTomato-N1 plasmid expressing the tandem dimer Tomato was a gift from Michael Davidson and Roger Tsien (Addgene plasmid 54562). The humanized mVenus coding sequence was amplified by PCR and cloned in the same vector backbone instead of the tdTomato.
Plasmid DNA Extraction
Plasmid DNA was isolated using miniprep NucleoBond extraction columns (Macherey Nagel) according to the manufacturer’s protocol.
Transfection Reagents
jetPEI (Polyplus) or TransIT-X2 Dynamic Delivery System (Mirus) transfection reagents were used with their respective diluents: 150 mM NaCl and Opti-MEM medium (Life Technologies) according to the supplier’s instructions.
Echo 550 Reverse Transfection
The Echo 550 device was set to 384LDV_AQ_B2 mode. Three picklists were generated from a Microsoft Excel macro file to design the position of each source well, the expected transferred volume, and the destination wells (step 1: diluent dispensing; step 2: DNA transfer; step 3: transfection reagent mix dispensing). Generated picklists were imported in the Echo Cherry Pick 1.5.3 software for transfers. A 384LDV Microplate (Labcyte) and a 384-well Microplate µClear Black (Greiner Bio-one) were used as the source plate and destination plate, respectively. In the optimized protocol, diluent was dispensed using the MicroFlo dispenser (BioTek, 1 µL cassette).
Cell Culture
HeLa cells were grown in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% v/v fetal bovine serum, 4.5 g/L glucose, 100 U/mL penicillin, and 0.1 mg/mL streptomycin (Life Technologies). To reach a cell density suitable for proper image analysis after 48 h of growth, 1500 cells were seeded per well using the MicroFlo dispenser (BioTek, 10 µL cassette) after the Echo 550 dispensing and incubated at 37 °C, 5% CO2.
Cell Fixation
Transfected cells were fixed for 15 min with 10% formalin (Sigma-Aldrich). Nuclei were stained for 20 min with 1 µg/mL Hoechst (Invitrogen) diluted in 1× phosphate-buffered saline (PBS; Life Technologies), and washed three times using 1× PBS.
Image Acquisition and Analysis
Images of two or three fluorescent channels (Hoechst, tdTomato, mVenus) were sequentially acquired (three fields) using the INCell Analyzer 6000 (GE Healthcare, Life Sciences) with 10× objectives and a proper emission filter set (DAPI, dsRed, and FITC, respectively). Transfection efficiencies were analyzed based on nuclei staining using Columbus software (PerkinElmer) and expressed as the percent of transfected cells.
Data Analysis
Statistical analyses were performed using GraphPad Prism version 7.00 software using one-way ANOVA analysis and Dunnett posttests, or two-way ANOVA and Bonferroni posttests.
Results and Discussion
Plasmid DNA Transfection Feasibility Using Echo Acoustic Dispenser
To test the ADE technology for automated plasmid transfection upon a reverse transfection protocol, we used HeLa cells and two different transfection reagents: jetPEI, a lipopolyplex, and TransIT-X2, a nonliposomal system. To easily monitor transfection efficiency by automated confocal imaging, a tdTomato red fluorescent protein expressing plasmid was chosen due to its exceptional brightness. 15 Four main parameters can affect DNA transfection efficiency: (1) DNA amount, (2) ratio of transfection reagent to DNA, (3) cell density at the time of the transfection mixture addition, and (4) incubation time of the reagent–DNA complex. We focused our effort on the first three parameters since DNA solution and transfection reagent incubating time before cell addition cannot easily be controlled in the 384-well format. Using three-step dispensing, we studied the impact of these parameters by changing the diluent volumes from 15 to 4000 nL, the amount of DNA from 5 to 100 ng, and the concentration of the final dispense of 500 nL containing the DNA transfection reagent (1× and 2× concentrations). In our preliminary experiments, we noticed that DNA solution dispensed could evaporate before addition of the transfection reagent mixture (data not shown), so a diluent was dispensed before DNA to circumvent this problem. The 500 nL transfection mixture volume per well was the same for both reagents, as calculated according to the manufacturer’s instructions, and was not optimized in this study. As this mixture requires vortexing of a fresh preparation before addition to diluted DNA solutions, we filled the source plate just after vortexing.
After this final use of acoustic dispensing, cells were seeded using BioTek MicroFlo at a density of 1500 cells/well, as this ensures proper final automated image analysis 48 h posttransfection.
The TransIT-X2 reagent showed significantly higher transfection efficiency ( Fig. 1 ) than jetPEI at both concentrations used (up to 89.3 ± 1.15% and 43.75 ± 9.2% positive cells, respectively; p < 0.05). The higher concentration of jetPEI showed generally higher transfection efficiency than 1× for 5–30 ng of transfected DNA ( Fig. 1B ) and 500–2000 nL diluent volumes ( Fig. 1A ). The poor efficiencies obtained using jetPEI could be explained by the suppression of the vortex step normally required to form optimal DNA–jetPEI complexes, which can hardly be set up with a 384-well plate during the dispensing time. Better efficiencies could then be expected for transfection reagents avoiding vortex steps, such as the nonliposomal system TransIT-X2 in our case. However, more studies would be necessary to determine if non-liposomal-based transfection leads to higher efficiencies than liposome-based transfection in our high-throughput protocol. The best transfection efficiencies for TransIT-X2 were obtained with 5–30 ng of transfected DNA and diluent volumes ranging from 250 to 2000 nL. In these conditions, both concentrations of reagent led to the same results ( Fig. 1A , B ): significantly higher transfection efficiencies were obtained for a quantity of transfected DNA ranging from 5 to 30 ng with 1000 nL of diluent ( Fig. 1B ). The Echo dispenser was developed to dispense 2.5 nL droplets from source wells filled with 3–12 µL of aqueous liquid using an LDV plate, with a 9 µL working volume. Using 100 ng/µL DNA solution allows a total dispense of 900 ng, which represents 45 different destination wells that can be filled with 20 ng of DNA from the same source well. In order to increase this number, we examined the effect of the starting DNA solution concentration on the transfection efficiency. HeLa cells were transfected using 1× concentrated TransIT-X2, with 5–100 ng of DNA, but using a serial dilution of 400, 200, and 100 ng/µL of the dispensed DNA solution in three separate wells of the source plate. For all the diluent volumes tested, a starting DNA concentration of 100 ng/µL leads to significantly higher transfection efficiencies than 200 and 400 ng/µL for 5–30 ng of DNA dispensed ( Fig. 1C ). This could be due to variations in liquid surface tension, a crucial parameter for droplet ejection. 16 DNA greatly increases the viscosity of water and might then impair the dispensing process in acoustic liquid delivery. However, to confirm this point, we would have to test each cell line used.
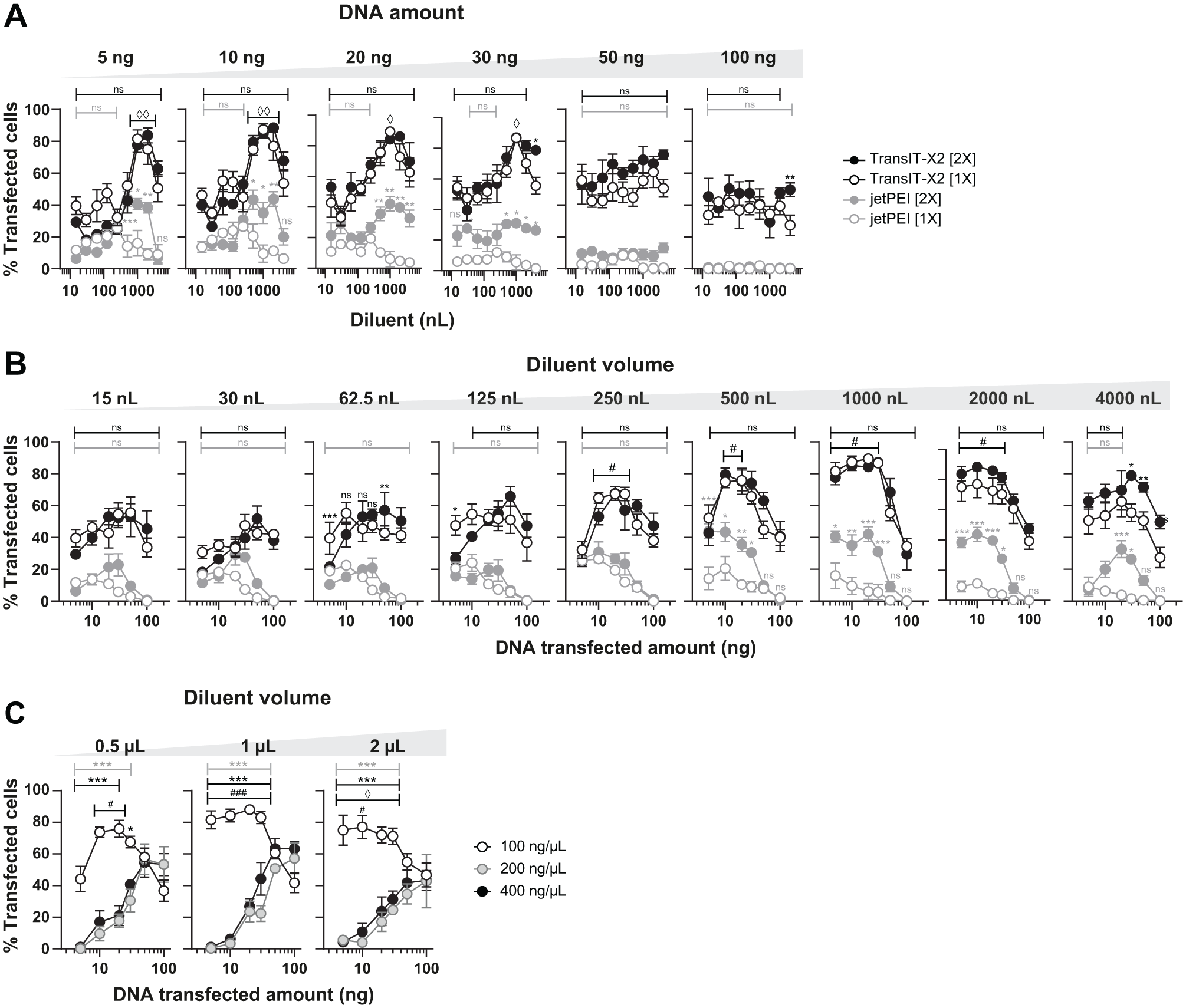
Transfection feasibility using an Echo 550 acoustic nanovolume dispenser. HeLa cells were reverse transfected using the Echo 550 device and TransIT-X2 (black) or jetPEI (grey) transfection reagents. The 1× concentration recommended by the manufacturer or a 2× concentration was used (unfilled and filled circles, respectively). Volumes from 15 to 4000 nL of the recommended diluent (Opti-MEM medium for TransIT-X2; 150 mM NaCl for jetPEI) were used with increasing amounts of DNA (5–100 ng). Transfection efficiencies were determined 48 h later using image-based analysis software. (
These first results showed that our three-step Echo 550-based reverse transfection DNA-dispensing protocol is efficient and accurate. For further optimization, we used the best parameters obtained for HeLa cells: TransIT-X2, 1 µL of Opti-MEM diluent, a DNA source concentration of 100 ng/µL, and a total DNA dispensed amount of 10–30 ng.
Improvement of the Protocol
We thought that our protocol could be further improved in terms of throughput. We estimated the throughput of our protocol by measuring the time required for each step: 25 or 50 min to dispense 1 or 2 µL of diluent, respectively; 5 min to deliver 20 ng of DNA (starting from a 100 ng/µL concentration); 15–20 min for the 500 nL transfection mixture; and only 45 s for dispensing cells using MicroFlo. Our initial settings for the overall dispensing process then required nearly 1 h to transfect a 384-well plate, including the manual plate handlings between the different steps. The handling of both the diluent and transfection reaction mixture appears to be the throughput-limiting step. Acoustic dispensing is particularly well suited to prepare the low volumes of transfection mixture required for this kind of experiment, saving this usually expensive reagent. We therefore decided to keep this technology for this step. However, MicroFlo is able to dispense 1 and 2 µL in 8.5 and 10.5 s, respectively. As the largest volume to be dispensed is the diluent, we compared Hela cell transfection using Echo- or MicroFlo-based dispensing for the diluent step. Both dispensing methods led to similar results ( Fig. 2A ), and no significant differences in transfection efficiencies were noticed. MicroFlo lowered the overall process time to less than 30 min, enhancing the throughput of the protocol by twofold, up to 16 plates per day. At this rate, another expected limiting factor would be the cell preparation, as an entire workday would be necessary to fill 16 plates, assuming a rate of one plate every 30 min. We then tested whether plates could be preloaded with DNA in order to add the transfecting reagent and cells extemporaneously. Whereas more than 4 years of DNA conservation has previously been reported using different optimized media, 17 such as Biomatrica or polyvinyl alcohol, we avoided their use in order to keep our initial working protocol (DNA congelation or drying in Opti-MEM medium). One microliter of diluent was then dispensed using acoustic dispensing or MicroFlo, and DNA was dispensed using acoustic dispensing. The prepared plates were either (1) used immediately to dispense the transfection reagent and cells (initial protocol, day 0 control), (2) frozen to −20 °C, or (3) dried in an oven and used the same day once thawed or rehydrated, respectively (day 0), or 2 or 7 days later (days 2 and 7, respectively). Both conservation methods led to the same results as no significant differences were seen between the initial full Echo or MicroFlo/Echo-enhanced protocol and the preloaded plate used at day 0, 2, or 7 ( Fig. 2B ). This indicates that DNA preloaded plates can be stored and used to transfect cells at higher rates. Indeed, as the transfection reagent dispensing only requires 15–20 min, we are able to transfect nearly 24 plates during a workday at a rate of 3 plates per hour. Up to six plates can therefore be prepared using the same cell batch for 2 h. At this stage, we have proven that our full Echo or MicroFlo/Echo transfection protocol can be easily used to set up a transfection protocol for a given cell line.
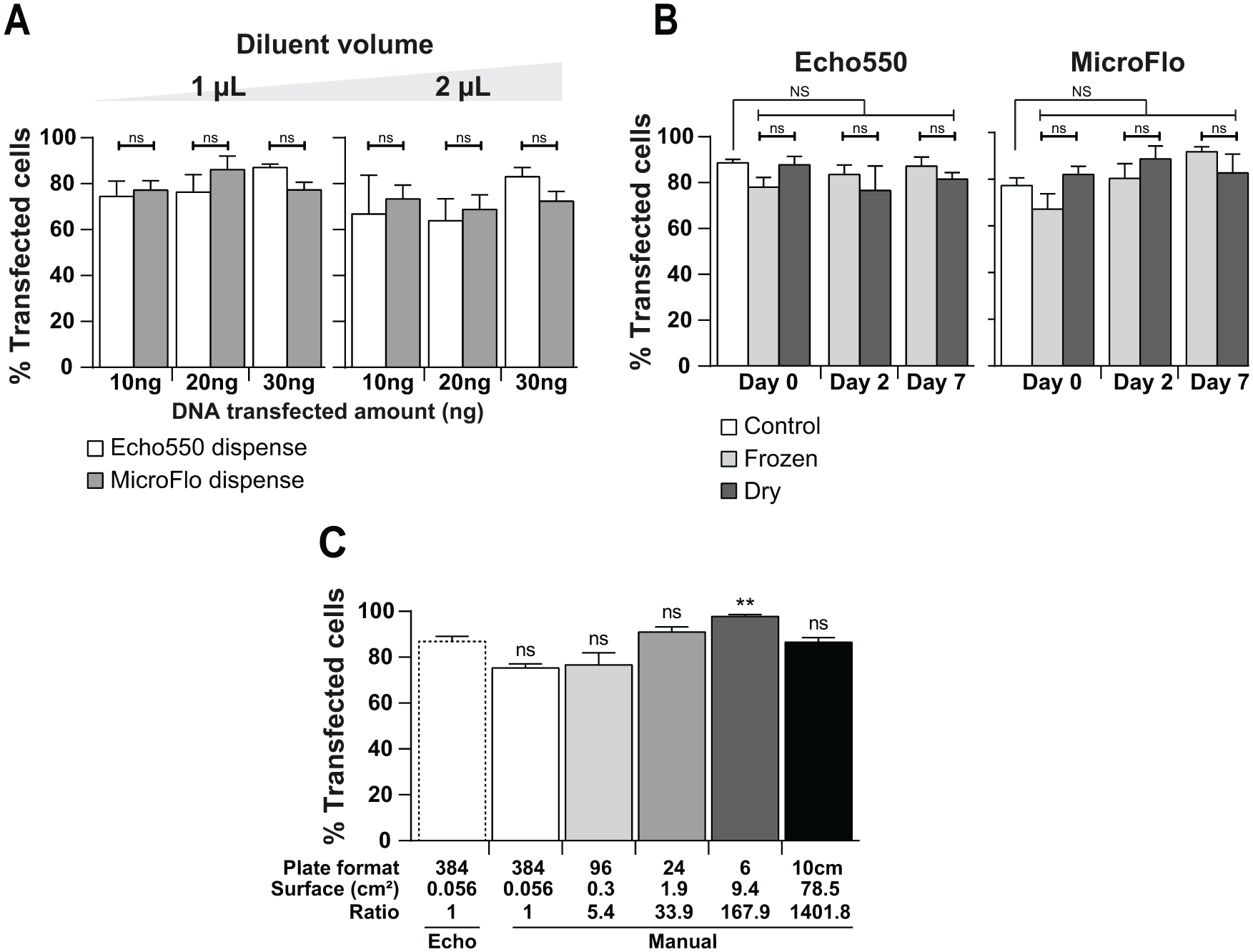
Improvement of the protocol. (
Scaling up after screening, we then examined whether the identified optimal protocol could be implemented at a higher scale. The DNA amount (20 ng), volumes (1 µL of diluent, 0.2 µL of DNA, and 0.5 µL of reagent), and cell number (1500 cells/well) used in the 384-well MicroFlo/Echo-based transfection protocol were used as a reference to calculate the parameters for larger-scale transfections. Ratios were calculated from a 384-well culture surface to other culture well or dish surfaces (bottom of Fig. 2C ). HeLa cell transfection in 384-, 96-, 24-, and 6-well plates and a 10 cm culture dish was then manually performed using the indicated ratios and starting with the above-mentioned 384-well condition references. Under these conditions, all plate formats were efficiently transfected and no significant differences were seen compared with the MicroFlo/Echo-enhanced protocol, except for the six-well plate, which marginally showed higher transfection efficiency ( Fig. 2C ). This means that our high-throughput protocol could be used for the identification of optimal transfection settings for larger-scale manual transfections. Our MicroFlo/Echo protocol was thus not only as efficient as the manual protocol but also much faster. Furthermore, in this protocol, DNA dispensing is performed automatically using a customized picklist, avoiding errors associated with the cumbersome repetitive manual dispensing (i.e., more than 1 h for the manual protocol vs three plates per hour for the MicroFlo/Echo protocol). We still tried to improve the manual protocol to reduce the overall time process, focusing on the transfection reagent dispensing step. Using an automated pipette and a reservoir, 34 s were required (still more than the MicroFlo device), and on top of that, a larger quantity of reagent was needed, leading to waste at least 500 µL of dead volume of transfection mixture. Thus, although feasible, this way does not seem to be advantageous.
Our Echo acoustic dispense-based high-throughput transfection protocol appears to be a valuable tool to transfect cells with different plasmids and screen conditions applicable to larger-scale transfections. However, we thought that it would be a huge advantage if it could also be used for co-transfection experiments.
Plasmid DNA Co-Transfection
Two fluorescent plasmids were used to evaluate the multiplexing potential of acoustic dispensing. The tdTomato-expressing plasmid was modified to express mVenus, a bright yellow fluorescent protein, and 1.7 ratio of mVenus over tdTomato was used due to differences in their brightness and quantum yield. 18 The DNA dispense picklist was modified in order to dispense 20 ng of total DNA, including both plasmids mixed in the same source well (control) or in distinct source wells, into a single destination well. Both plasmids were efficiently transfected when mixed in either the same source well or separate source wells and multiplexed by the Echo picklist-driven dispense ( Fig. 3A ). The transfection efficiencies were similar for both the mixed and sequential DNA plasmid dispenses ( Fig. 3B ), and no significant differences were observed. This implies that the hard and time-consuming step of multiplexing DNA for co-transfection even in a dose-ranging manner would be easily software driven by the Echo 550 customized picklist. The co-transfection efficiencies seemed lower than previously with the use of 20 ng of DNA. This can be explained by the ratio used, as mentioned above, which lowers the transfected amount of tdTomato and mVenus. However, whereas not all the cells were transfected, the results expressed as the percent of co-transfected cells within the mVenus- or tdTomato-positive populations showed no significant differences, as both almost reached 100% ( Fig. 3C ). Taken together, these results indicate that ADE-driven DNA multiplexing can be efficiently used for co-transfection and that transfected cells can be co-transfected by both plasmids using our experimental settings.
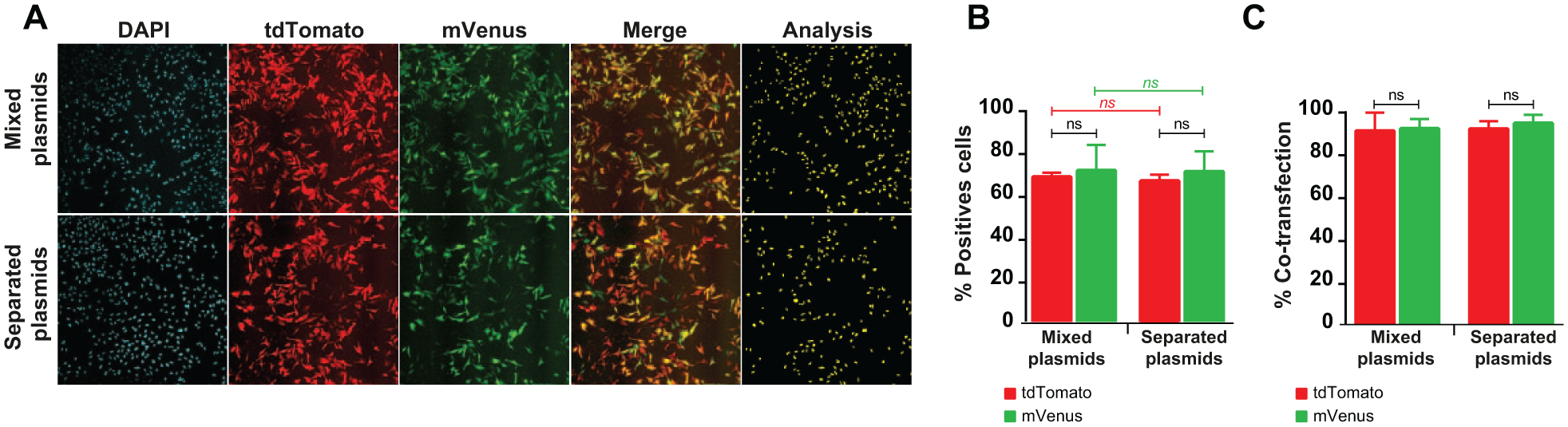
Plasmid DNA co-transfection ability. HeLa cells were transfected using the MicroFlo/Echo-enhanced protocol: the MicroFlo device was used for 1 µL of diluent dispense and the Echo 550 device was used for the DNA and TransIT-X2 reagent (1× concentration) dispenses. Both plasmids (total of 20 ng), in a mixed solution (20 ng using a 1.5 ratio of mVenus over tdTomato) loaded in a unique well or in two separate source well transfection efficiencies, were compared 48 h posttransfection using image-based analysis software. Representative field of 10× objective fluorescence microscopy acquisition (
In conclusion, we developed an easy, accurate, and high-throughput transfection method using automated acoustic dispensing ( Fig. 4 ).
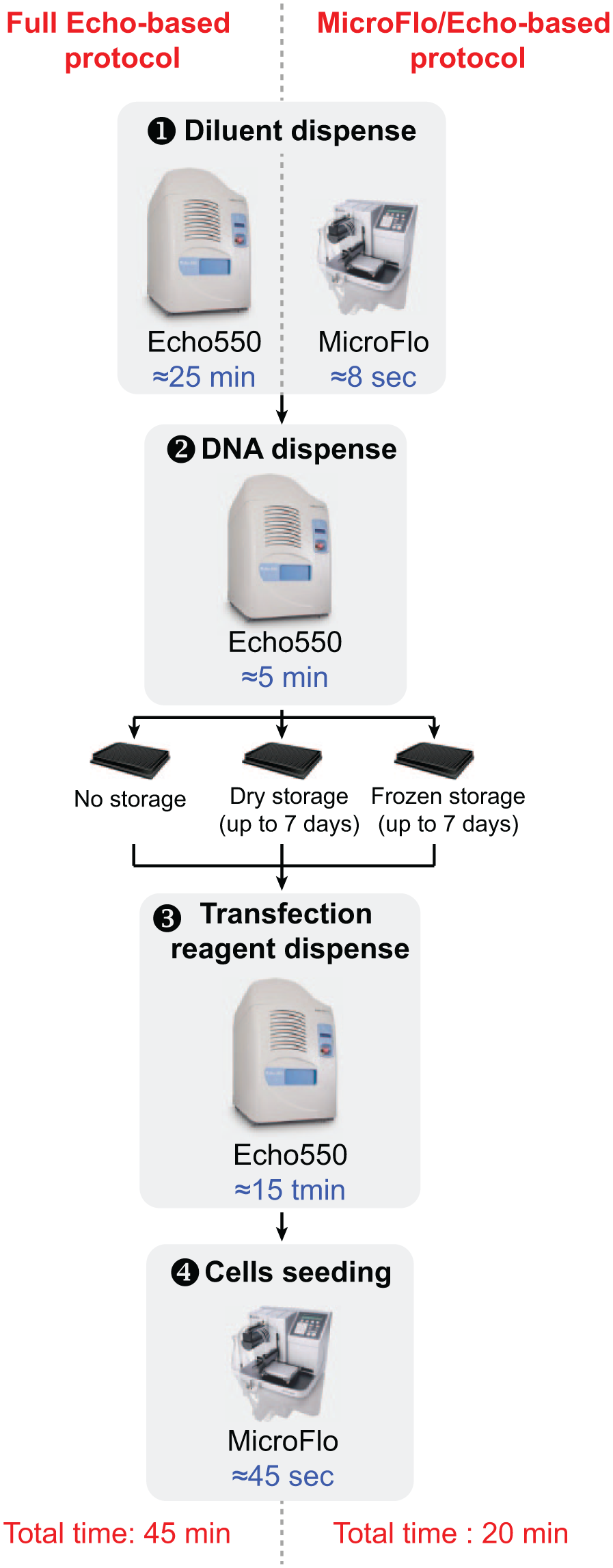
Experimental workflow. Schematic representation of our automated high-throughput reverse transfection protocol for the different conditions tested. The time needed for each step is indicated in blue, depending on the device used.
Using fluorescent proteins expressing plasmids, and high-throughput image-based software analysis, we demonstrated the feasibility of ADE-based liquid dispensing to transfect cells in 384-well plates using three dispensing steps (diluent, DNA, and transfection reagent), before cell seeding using MicroFlo device. This method enables simultaneous evaluation and comparison of different parameters in order to optimize transfection parameters.
On top of its high throughput, this method has several other advantages: (1) it reduces the reagent costs using lower volumes of transfection reagents; (2) it avoids waste and saves DNA preparation labor, using amounts of only a few nanograms, and is also efficient, using cheap mini-column-based plasmid DNA minipreparation, which is compatible with plate-based high-throughput plasmid purification protocols; and (3) it ensures successful transfection, even nonexperimented biologists, as few manual steps are required: diluting the source DNA to 100 ng/µL, dispensing it on a source plate, and preparing the cell suspension before seeding. The nanodispenser executes the time-consuming and error-prone steps according to a customized picklist: dose delivery and multiplexing of plasmids.
The throughput of the full Echo protocol was further enhanced by replacing the diluent dispense-limiting step using ADE by an ultrafast MicroFlo dispense. This MicroFlo/Echo optimized process requires a total time of less than 30 min to transfect a 384-well plate with different plasmids. We further demonstrated that DNA preloaded plates could be stored (dry or frozen) and used later for the transfection mixture dispense and cell seeding steps only. Using such an approach improves the transfection rate to 3 plates per hour (i.e., 24 plates per day). This last optimized protocol could open up new ways to perform array-based experiments, such as expressing human protein-coding genes from the human ORFeome collection 19 or knocking down each of them using CRISPR-Cas9 library-based approaches. 20 Indeed, performing such array approaches would then require around 53 plates for the estimated 20,000 human genes and then take about 18 h on a dedicated automated platform, rather than 2–3 days of human work. Successful CRISPR-Cas9 high-throughput approaches have used pooled gRNA libraries and implied viral delivery and a strong discriminating phenotype to sort positive cells. 20 Due to its high efficiency and high-throughput performance, our protocol would achieve a nonpooled approach with the gRNA library/CRISPR-Cas9-expressing plasmids distributed in source plates. Such an approach would then allow screening applications using milder cellular phenotype changes, which are currently incompatible with the required cell-sorting step.
Finally, a further step would be to study the compatibility of long-lasting DNA storage mediums with our protocol when using DNA prefilled transfection-ready plates, awaiting the transfection reagent and cell dispense.
Footnotes
Acknowledgements
The authors wish to thank Dr. B. Villemagne, Dr. S. Moureu, Ms. M. Goasmat, and Ms. A. Guillome for critical review and corrections of the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project was supported by Inserm, Lille University, Lille Pasteur Institute, Conseil Régional Nord and PRIM (Pôle de Recherche Interdisciplinaire pour le Médicament).
