Abstract
Epigenetics continues to emerge as an important target class for drug discovery and cancer research. As programs scale to evaluate many new targets related to epigenetic expression, new tools and techniques are required to enable efficient and reproducible high-throughput epigenetic screening. Assay miniaturization increases screening throughput and reduces operating costs. Echo liquid handlers can transfer compounds, samples, reagents, and beads in submicroliter volumes to high-density assay formats using only acoustic energy—no contact or tips required. This eliminates tip costs and reduces the risk of reagent carryover. In this study, we demonstrate the miniaturization of a methyltransferase assay using Echo liquid handlers and two different assay technologies: AlphaLISA from PerkinElmer and EPIgeneous HTRF from Cisbio.
Introduction
Epigenetics is generally defined as the study of changes in the regulation of gene expression that do not involve changes in the sequences of DNA.1,2 Epigenetic changes are thought to play a role in many diseases, including cancer, 2 cardiovascular disease, neuropathologies, inflammation, and diabetes.1,3 A class of enzymes known as methyltransferases play a role in histone modification, which affects the regulation of gene transcription. 1 These enzymes are therefore intriguing targets for drug discovery.4–6 In particular, the histone H3 methyltransferase G9a epigenetically activates the serine-glycine synthesis pathway to sustain cancer cell survival, proliferation, and ultimately tumorigenesis.7,8 SET7/9 methylates histone H3 and is able to modulate p53 activity in a human cancer cell line, implying that it may also play a significant role in human tumorigenesis. 9
SET7/9 and G9a are enzymes belonging to the family of histone lysine methyltransferases. Both are SET domain–containing proteins. SET7/9 catalyzes the monomethylation of lysine 4 on histone H3,
10
while G9a methylates lysines 9 and 27 on histone H3.
1
Both enzymes use S-adenosyl-
There are currently a number of assay technologies available for studying the activity of these methyltransferases. Here we examine the miniaturization of two of these assay platforms: HTRF (Homogenous Time Resolved Fluorescence) and AlphaLISA. Both assay technologies are sensitive and robust and can readily be used for high-throughput screening.12,13 The costs associated with these assays, however, can be quite high. Because these assays are homogenous, mix-and-read, no-wash assays, they can be quite amenable to miniaturization, 14 resulting in savings on kit reagent costs and thereby significantly reducing the costs of running high-throughput screens while not compromising data quality.
Miniaturizing to significantly lower assay volumes requires the ability to accurately and precisely dispense submicroliter volumes of kit reagents, enzyme, substrate, and compound. Acoustic dispensing with the Echo liquid handler15,16 enables dispensing of volumes down to 2.5 nL while minimizing dead volumes and enabling assembly of these assays at very low volumes.
In our study, for both assay platforms and enzymes, we used biotinylated peptides: histone H3 (residues 1–21) for the SET7/9 and G9a enzymes. The EPIgeneous HTRF Kit is a universal assay for methyltransferases that measures the conversion of SAH from SAM during the methylation of the peptide ( Fig. 1 ). 17 We used this assay to study both the G9a and SET7/9 enzymes. We also investigated the miniaturization of an AlphaLISA assay that directly measures the methylation of lysine 4 residue on histone H3 to study SET7/9 activity ( Fig. 2 ). Alpha beads are small (250–350 nm) and thus not prone to settling, which makes them ideal for acoustic transfer. 18
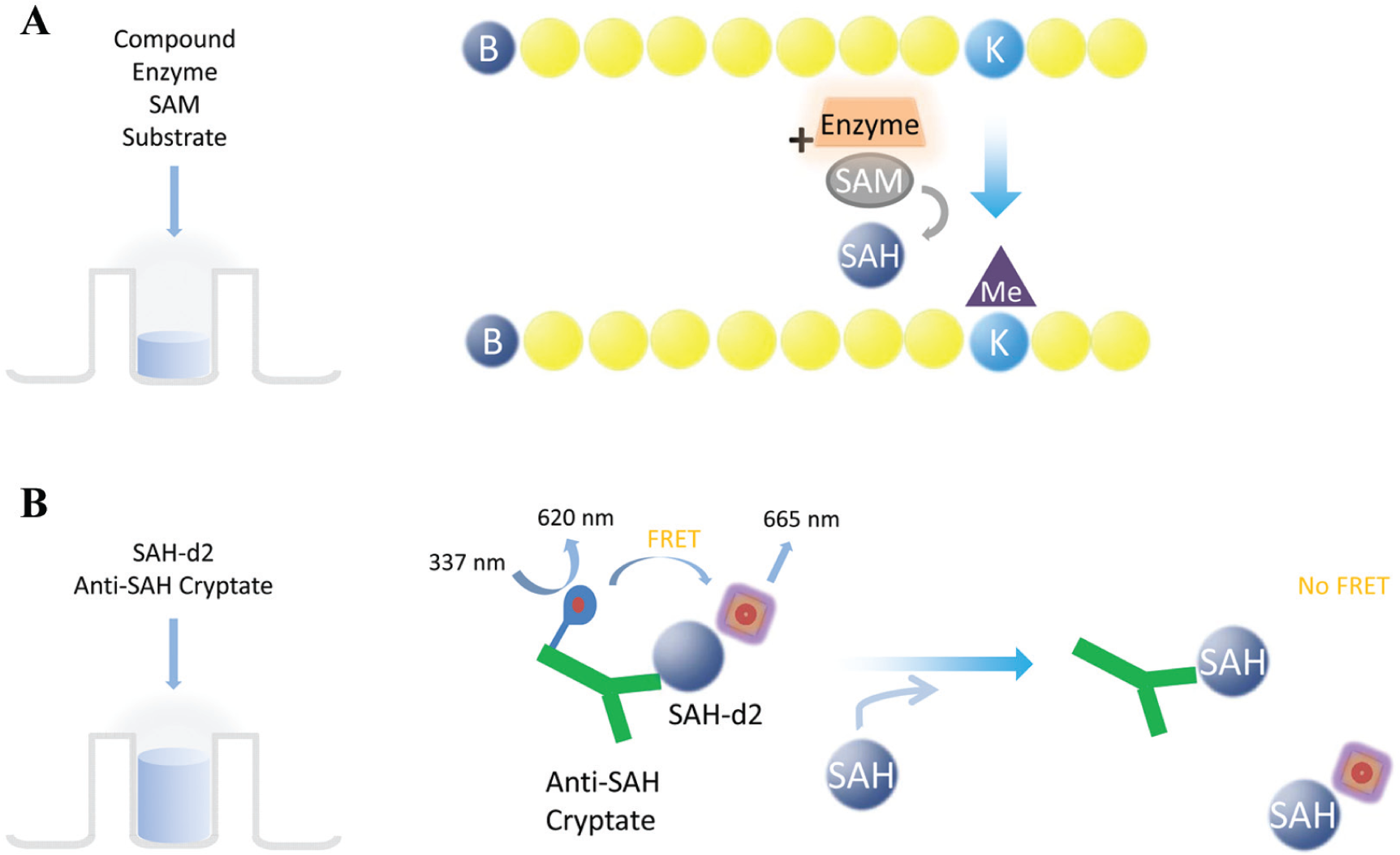
The EPIgeneous HTRF Methyltransferase Assay from Cisbio is a universal kit for methyltransferases and their substrates in a two-step, single-well reaction. (
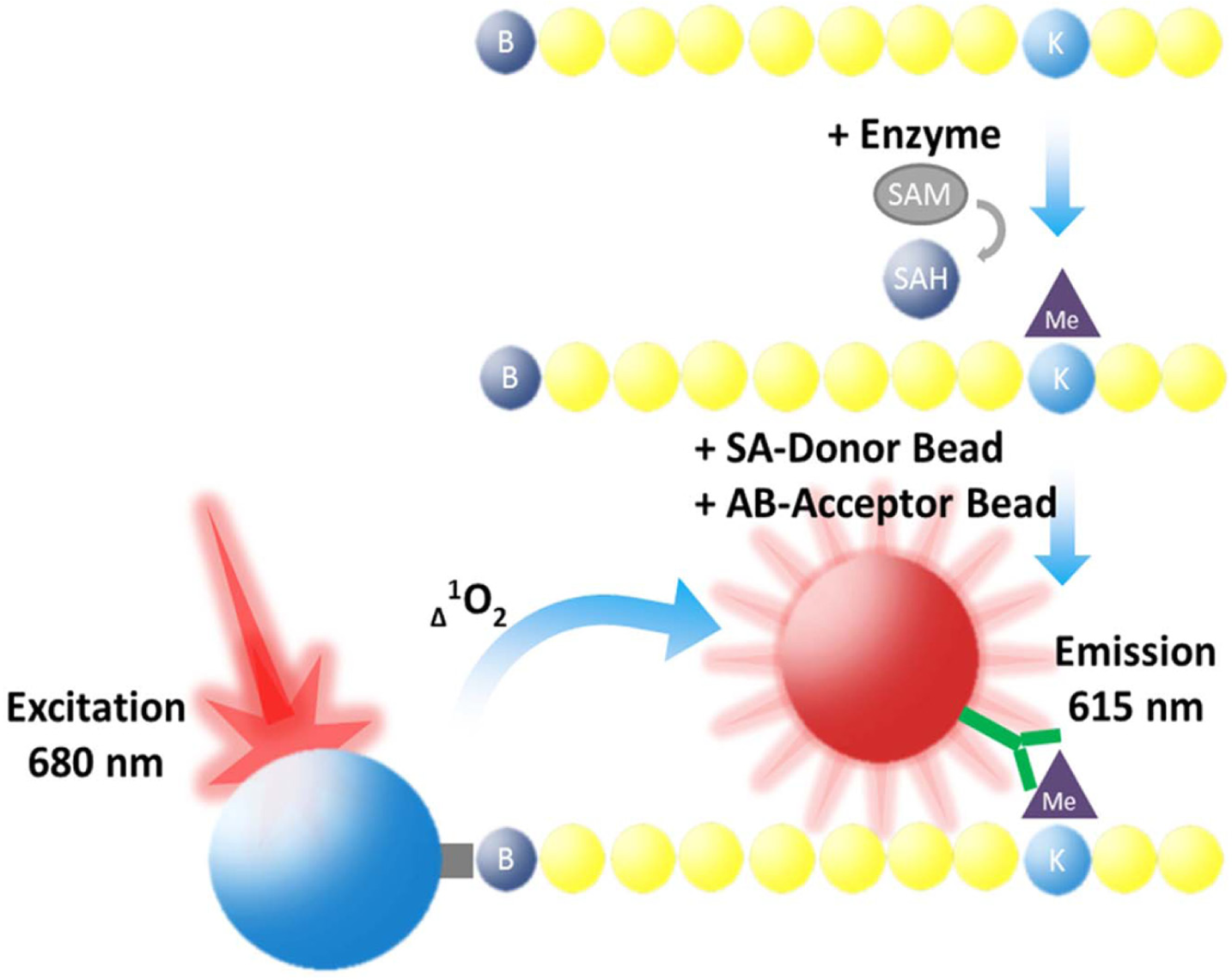
AlphaLISA assay. Acceptor beads bind specifically to the modified histone H3 peptide. The streptavidin donor beads then bind to the biotinylated peptide. The bound beads are then in close enough proximity to each other that when the donor bead is excited by laser light, a transfer of energy to the acceptor bead occurs via a singlet oxygen, resulting in 615 nm emission.
Materials
An EPIgeneous Methyltransferase Assay Kit (cat. 62AHPEB), which included SAH-d2, anti-SAH lumi-Tb cryptate, and detection buffers 1 and 2, was purchased from Cisbio US (Bedford, MA). For the EPIgeneous assays, SET7/9 (cat. 51010) and G9a (cat. 51000) enzymes were purchased from BPS Bioscience (San Diego, CA).
Alpha streptavidin donor beads (cat. 6760002), anti-methyl-Histone H3 Lysine 4 (H3K4me1-2) AlphaLISA acceptor beads (cat. AL116), and AlphaLISA 5× Epigenetics Buffer (cat. AL008) were purchased from PerkinElmer (Waltham, MA). For the AlphaLISA assay, SET7/9 enzyme (cat. ALX-201-178-C100) was purchased from Enzo Life Sciences (Farmingdale, NY). Biotinylated Histone H3 (1–21) (cat. 61702) peptide was purchased from AnaSpec, Inc. (Fremont, CA). (R)-PFI 2 hydrochloride ((1R)-1-[[3-(trifluoromethyl)phenyl]methyl]-2-oxo-2-(1-pyrrolidinyl)ethyl 1,2,3,4-tetrahydro-6-isoquinolinesulfonamide hydrochloride, cat. 4892) and UNC 0646 (N-(1-cyclohexyl-4-piperidinyl)-2-[hexahydro-4-(1-methylethyl)-1H-1,4-diazepin-1-yl]-6-methoxy-7-[3-(1-piperidinyl)propoxy]-4-quinazolinamine, cat. 4342) methyltransferase inhibitors were purchased from Tocris Biosciences/R&D Systems (Minneapolis, MN). Sinefungin (cat. ALX-380-070-M005) was purchased from Enzo Life Sciences. SAM (cat. A7007) and SAH (cat. A9384) were purchased from Sigma-Aldrich (St. Louis, MO).
Assays were performed in 384-well white, low-volume, flat-bottom assay plates (cat. 784075) from Greiner Bio-One (Monroe, NC) and 1536-well white, flat-bottom assay plates (cat. 3729) from Corning (Corning, NY).
Methods
Kit reagents, beads, enzymes, substrates, and compounds were all transferred with the Echo 555 and 525 liquid handlers from 384-well Echo-qualified polypropylene source plates (cat. P-05525) from Labcyte (Sunnyvale, CA). The Echo 555 liquid handler transfers aqueous and DMSO solutions in 2.5 nL increments, while the Echo 525 liquid handler transfers aqueous solutions in 25 nL increments. The Echo 525 was utilized for larger-volume transfers for increased throughput.
Plates were read on a PHERAstar FS plate reader (BMG LABTECH, Cary, NC) equipped with AlphaLISA and HTRF optic modules, as well as AlphaScreen and HTRF lasers for excitation. The HTRF optic module used 330/80 nm excitation and 620/10 and 665/80 nm emission filters, along with a 650 nm cutoff dichroic mirror with matched dual photomultiplier tube (PMT) technology. The AlphaLISA module used 680 nm excitation and 615 nm emission. Data were analyzed using MARS Data Analysis software (BMG LABTECH), and HTRF ratio values are reported as the ratio of acceptor fluorophore intensity/donor fluorophore intensity (665/620 nm)*10,000. The Z′ factor determination is used to describe the robustness of an assay with a theoretical upper limit of 1.0 and calculated as described by Zhang et al. 19 with Microsoft Excel (Microsoft, Redmond, WA). Dose–response analysis and titrations were fit to a four-parameter logistic equation using GraphPad Prism (GraphPad Software, La Jolla, CA).
Reagent Preparation
A stock solution of SAM was prepared at 30 mM in 5 mM H2SO4/10% ethanol (v/v) in H2O. Assay buffer for the EPIgeneous HTRF assays consisted of 50 mM Tris-HCl (pH 8.8), 10 mM NaCl, 1 mM dithiothreitol (DTT), and 0.01% Tween-20. Standard curves and working solutions of enzymes, peptides, and SAM were prepared in assay buffer. Detection buffer 1 was ready to use, requiring thawing at room temperature (RT), due to the presence of DMSO, prior to use. SAH-d2 was prepared in detection buffer 2. The concentration of SAH-d2 was prepared in accordance with the manufacturer’s recommendations dependent on the SAM concentration used for each methyltransferase reaction. Anti-SAH lumi-Tb cryptate was prepared in accordance with the manufacturer’s recommendations in detection buffer 2. The volume of detection reagents was added in accordance with the manufacturer’s recommendation to reach a final assay volume of 2× the enzymatic reaction volume.
Assay buffer for the AlphaLISA assay consisted of consisted of 50 mM Tris-HCl (pH 8.8), 5 mM MgCl2, 1 mM DTT, and 0.01% Tween-20. Anti-methyl-Histone H3 Lysine 4 (H3K4me1-21) AlphaLISA acceptor beads were diluted in 1× Epigenetics Buffer 1 at a concentration of 100 µg/mL, and then added to a 384-well Echo-qualified source plate prior to transfer to the assay plate. Streptavidin donor beads were diluted in 1× Epigenetics Buffer 1 at a concentration of 50 µg/mL and added to a 384-well Echo-qualified source plate prior to transfer to the assay plate. Because the source plates with the beads could not be centrifuged due to risk of settling the beads, the beads were manually reverse-pipetted into the source plate wells in order to avoid generating bubbles.
All compounds and reagents were added to 384-well Echo-qualified source plates, and then transferred to the assay plates with either an Echo 555 or 525 liquid handler ( Fig. 3 ).
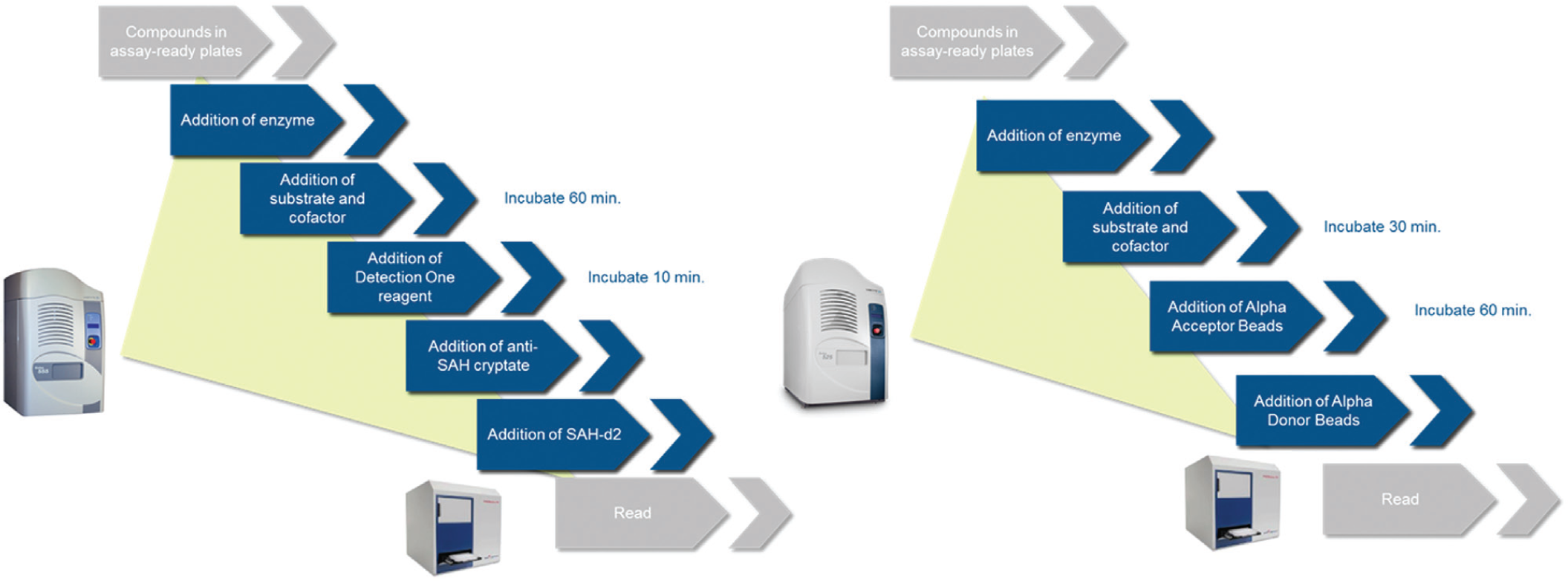
The Echo 555 and 525 liquid handlers were used to transfer kit reagents, beads, enzymes, and substrates from 384-well Echo-qualified polypropylene source plates. The Echo 555 was used to transfer compounds to assay plates.
EPIgeneous HTRF Methyltransferase Assays
Standard Curves
SAM/SAH standard curves were prepared in assay buffer. The standard curves mimic SAM-to-SAH conversion during the enzymatic reaction. A 16-point, 1:2 dilution standard curve including zero points for SAM and SAH calibrator was prepared in assay buffer with 1 μM SAM, with the exception of the zero SAM data point. Triplicates of each dilution were transferred to a 384-well assay plate and 1536-well assay plate at various volumes to mimic the enzymatic reaction volumes tested. The assay plate was prepared in accordance with the manufacturer’s recommendations. Detection reagents were added according to the manufacturer’s protocol. Following the 60 min incubation period with detection reagents, the plates were read on the PHERAstar FS and the HTRF ratio was plotted versus log SAH concentration.
Enzyme Titrations
Twelve- to 14-point, half-log dilution enzyme titration curves for G9a and SET7/9 were prepared in Echo-qualified, 384-well source plates. Enzyme titration curves were transferred to 384-well, low-volume assay plates in triplicate with the Echo 555 liquid handler. Assay buffer, peptide, and SAM were then added with the Echo liquid handler. The plates were then sealed and incubated RT for 60 min.
After incubation, detection buffer 1 was added to a 384-well Echo-qualified source plate. A volume of detection buffer 1 equivalent to 0.2× of the enzyme reaction volume was then transferred to the assay plate with the Echo liquid handler to stop the enzymatic reaction. The plate was resealed and incubated for 10 min. Anti-SAH lumi-Tb cryptate and SAH-d2 prepared in detection buffer 2 were each transferred with the Echo liquid handler in volumes each equivalent to 0.2× of the total assay volume. The plate was resealed and incubated for another 60 min. The plate was then read on the PHERAstar FS following the final incubation period and plotted as log enzyme concentration versus HTRF ratio.
Inhibitor Titration
A 12- to 14-point, half-log dose–response series, with a DMSO-only control point, was prepared for the G9a inhibitor UNC0646.8,20 Similarly, we generated a 12-point, half-log dose–response series of (R)-PFI 2 hydrochloride, a SET7/9 inhibitor, starting at the highest concentration of 100 µM in DMSO. Each point of the compound dose–response curves was first transferred in triplicate to 384-well or 1536-well assay plates with the Echo liquid handler. Assay buffer, enzyme, peptide, and SAM were added with the Echo liquid handler. For G9a inhibition, 40 nM enzyme, 1 µM peptide, and 1 µM SAM were used. For SET7/9 inhibition, 40 nM enzyme, 800 nM peptide, and 1 µM SAM were used. Enzyme reaction steps and detection reagent additions and incubations were performed as in the enzyme titration assay. The assay measurements were taken following the final incubation period and plotted as log inhibitor concentration versus HTRF ratio.
As a comparison, the SET7/9 inhibition assay was also performed in a 20 µL assay volume in a 384-well, low-volume assay plate. Fifty nanoliters of compound was added with the Echo 555 liquid handler; the remaining reagent additions were performed manually with a handheld repeating electronic pipettor. Reagent volumes were added in accordance to the manufacturer’s protocol; for the miniaturized assay volumes, reagent volumes were reduced in proportion to the total assay volume reduction ( Table 1 ).
Compound and Reagent Addition Volumes for the 20 µL (Manual Pipetting) and 2 µL (Echo Liquid Handler) Assay Volumes for the SET7/9 EPIgeneous HTRF Methyltransferase Assay.
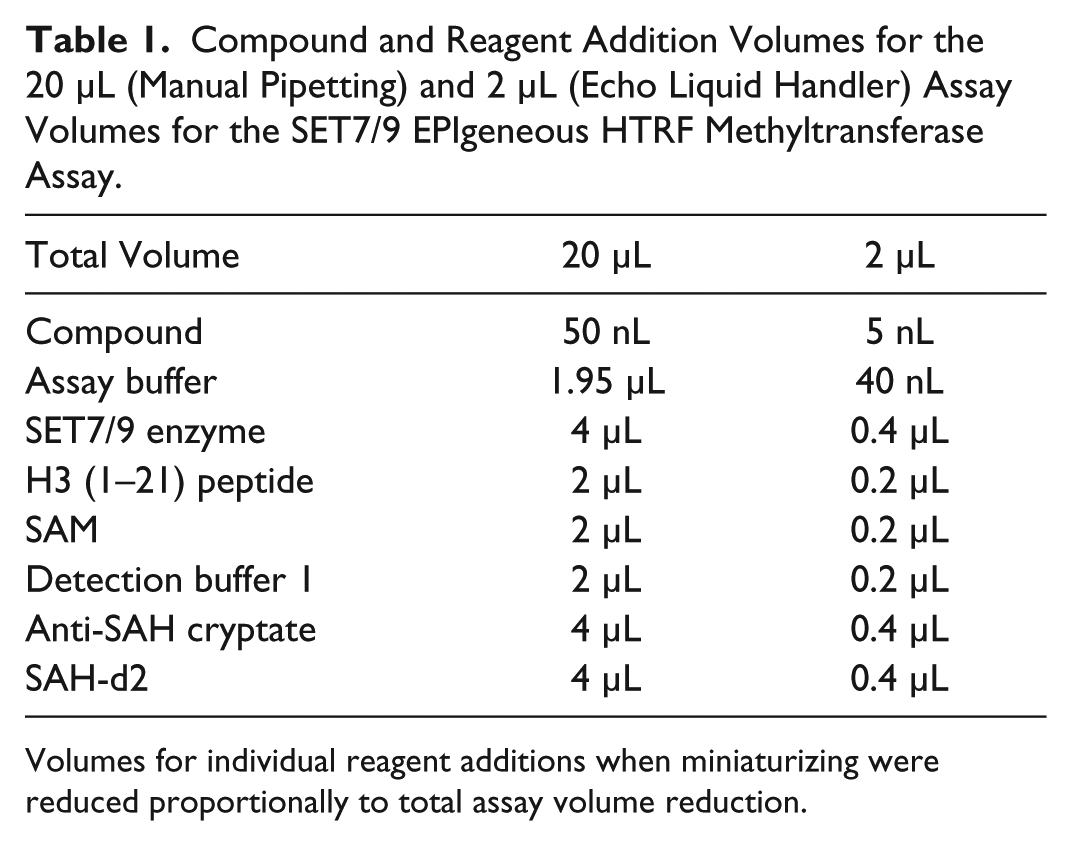
Volumes for individual reagent additions when miniaturizing were reduced proportionally to total assay volume reduction.
AlphaLISA Methyltransferase Assay
SET7/9 Inhibitor Titration
Dose–response curves of two known SET7/9 inhibitors, S-(5′-adenosyl)-
Anti-methyl-Histone H3 Lysine 4 (H3K4me1-21) acceptor beads were then transferred to the assay plates with the Echo 525 liquid handler to a final concentration of 20 µg/mL. Plates were resealed, briefly centrifuged at 1000 rpm, shaken for 15 s on a Teleshake 384 (Thermo Scientific, Waltham, MA) magnetic shaker, and then incubated for 60 min at RT in the dark.
Streptavidin donor beads were transferred to the assay plates with the Echo 525 liquid handler to a final concentration of 20 µg/mL. Plates were resealed, briefly centrifuged at 1000 rpm, shaken, and then incubated for 30 min at RT in the dark. Plates were then read on the PHERAstar FS plate reader and AlphaLISA signal plotted against log inhibitor concentrations. Volumes of reagent additions for each total assay volume are listed in Table 2 .
Compound and Reagent Addition Volumes for the 25 µL (Manual Pipetting) and the 5 and 10 µL (Echo Liquid Handler) Assay Volumes for the SET7/9 AlphaLISA Methyltransferase Assay.
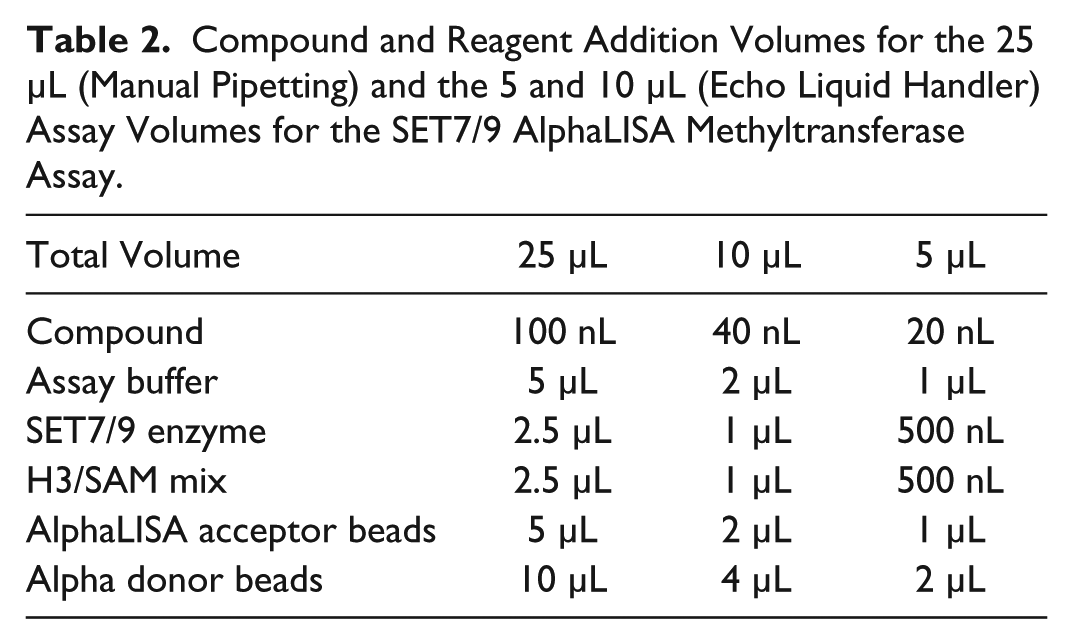
Assay Quality
For a comparison of signal-to-background (S/B) and Z′ factors, positive and negative controls were also run for the two assay platforms described at both miniaturized and full volumes to assess the quality and robustness of data generated in the assays miniaturized using the Echo liquid handler.
Results and Discussion
Epigeneous HTRF Methyltransferase Assays
To assess our ability to miniaturize the EPIgeneous HTRF assays, we used the Echo 555 liquid handler to optimize enzyme concentrations by running enzyme titration curves for G9a and SET7/9. From those results ( Fig. 4 ), an enzyme concentration of 40 nM for both G9a and SET7/9 was selected to run in subsequent experiments. SAM + SAH standard curves were also prepared ( Fig. 5 ). We generated IC50 data for the G9a inhibitor UNC0646 and the SET7/9 inhibitor (R)-PFI 2 hydrochloride in the miniaturized assay volumes. UNC0646 had an IC50 value of 27 nM. IC50 values for (R)-PFI 2 hydrochloride were 341 nM at 2 µL in 384-well and 294 nM at 2 µL in 1536-well formats. A compound titration for (R)-PFI 2 hydrochloride was also done at a 20 µL assay volume in the 384-well format, using the Echo 555 liquid handler to dispense compound to the assay plate and manually pipetting the remaining reagents. At this volume, (R)-PFI 2 hydrochloride had an IC50 value of 227 nM, showing good agreement between the three assay formats ( Fig. 6 ). A published literature value of 2 nM was considerably more potent than what we observed; however, this was consistent with observations of the interactions between (R)-PFI 2 hydrochloride and SAM, and that less than saturating concentrations of SAM (<2 µM) and increasing concentrations of peptide resulted in increasing IC50 values. 21
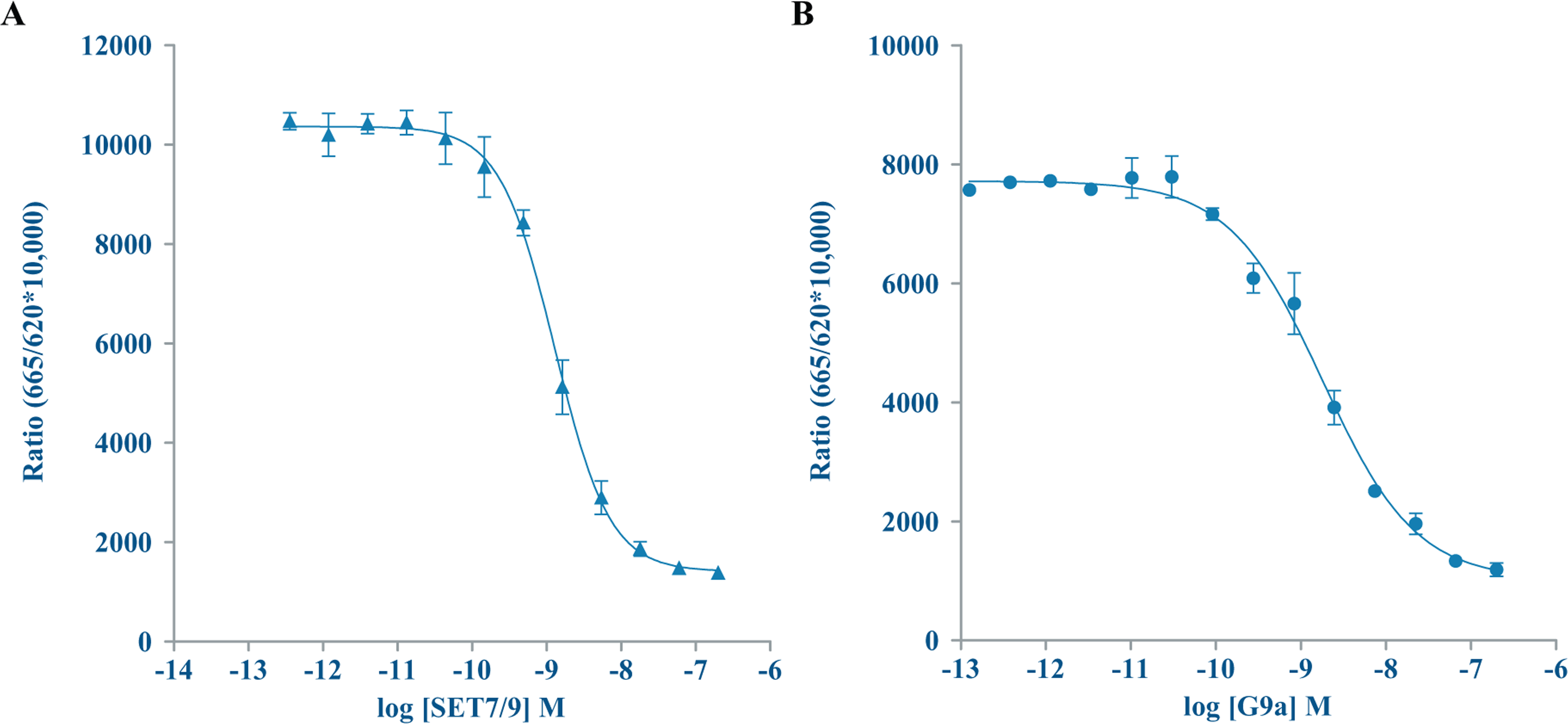
Enzyme titrations in 384-well, low-volume plates. (
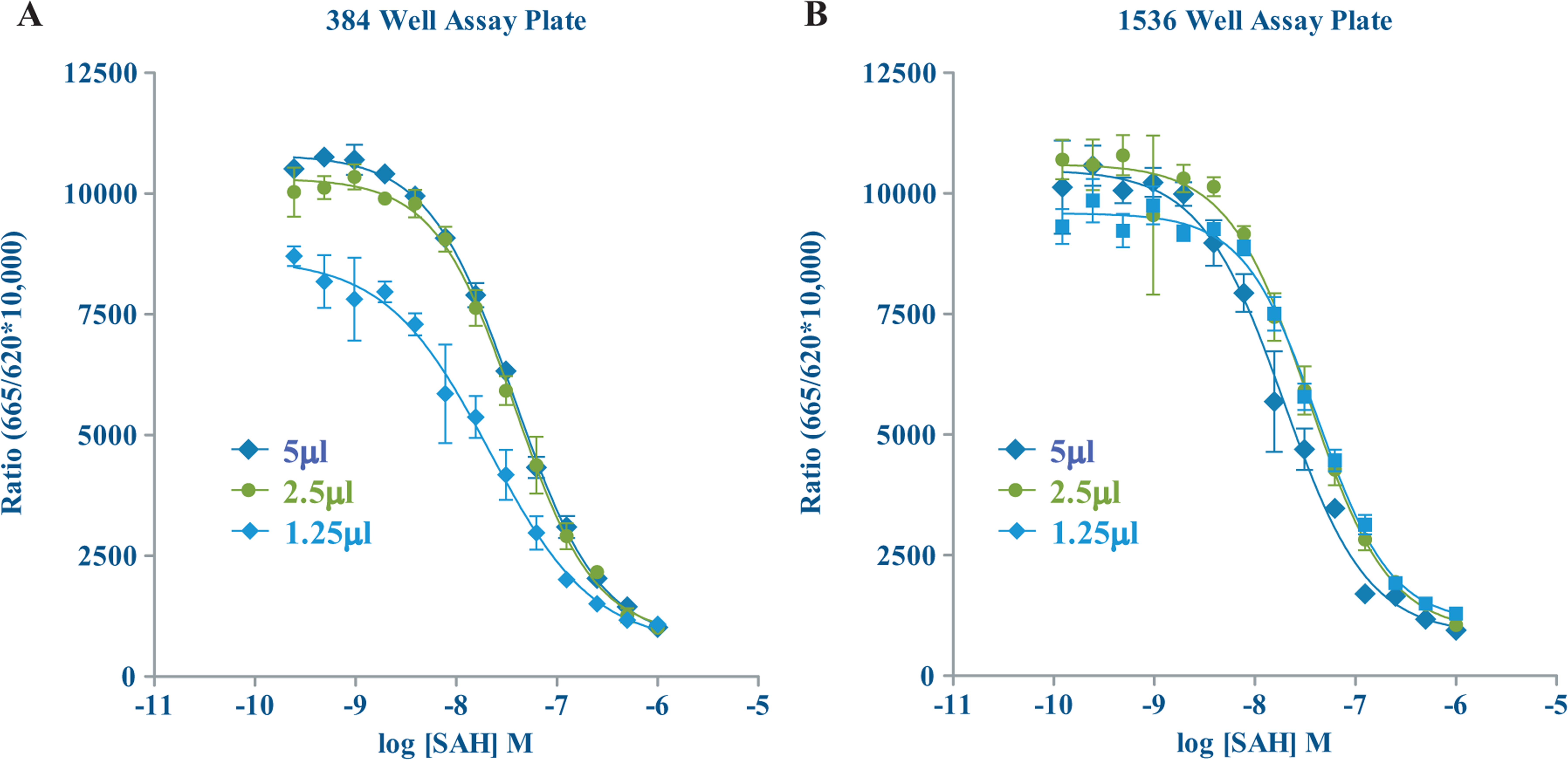
SAM/SAH standard curves in 384- and 1536-well assay plates. Total reaction volumes of 5, 2.5, and 1.25 µL. Standard curves were generated in enzymatic buffers to mimic methyltransferase activity resulting in SAH production. Various concentrations of SAH calibrator were mixed with varying concentrations of SAM while maintaining a constant 1 µM total concentration ([SAM] + [SAH]) throughout.
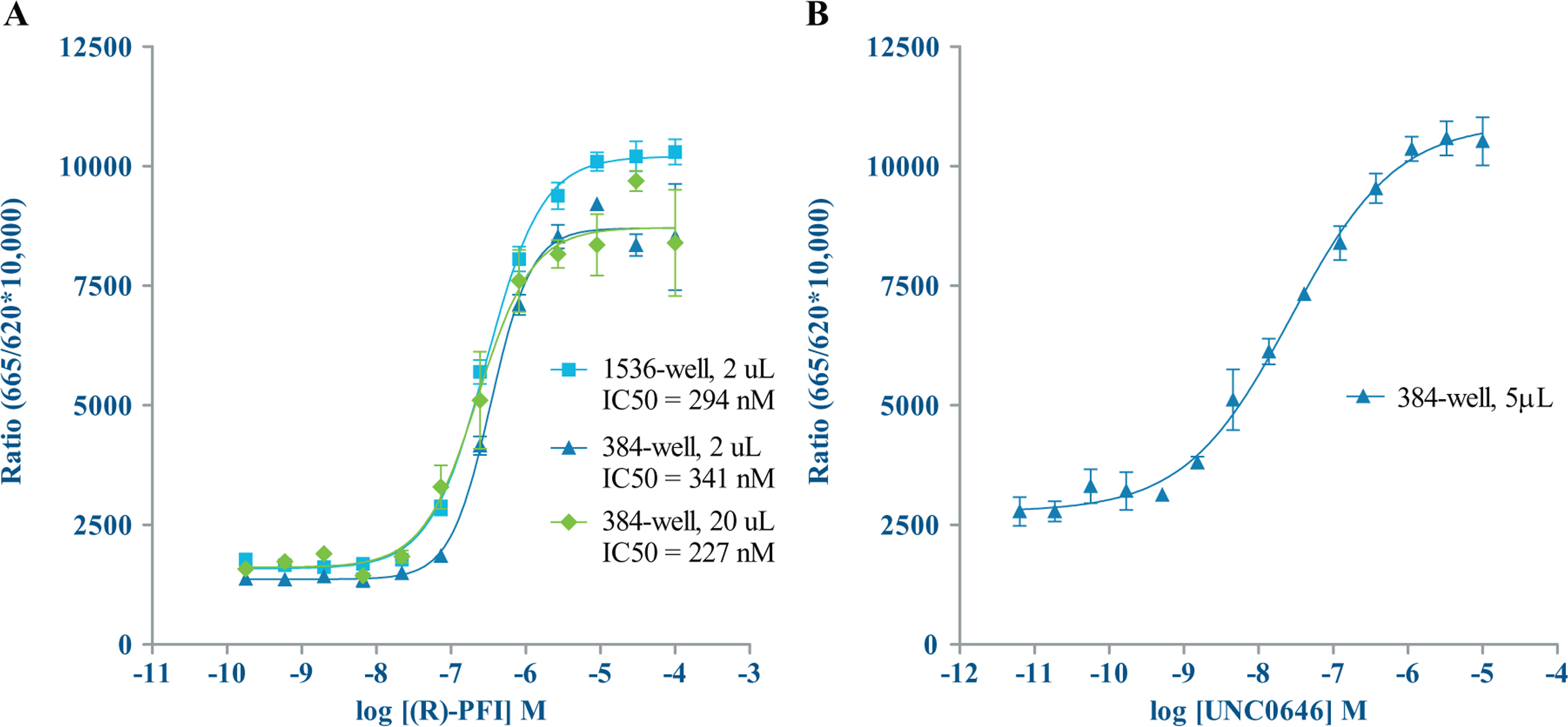
Inhibitor titrations—HTRF. (
Data quality in the miniaturized assays was excellent. The Z′ factors for the miniaturized assays were 0.88 and above, and S/B was 7.4 and above. These values were better compared to the manually pipetted standard 20 µL volume assay, which had a Z′ factor of 0.83 and S/B of 6.4 ( Table 3 ). The miniaturized assays assembled with the Echo liquid handler therefore not only had excellent data quality, but also showed improvement over the manually assembled, full-volume assays.
Comparison of Z′ Factors and S/B for Miniaturized (Echo Liquid Handler) and Full-Volume (Manual Pipetting) SET7/9 EPIgeneous HTRF Methyltransferase Assay.
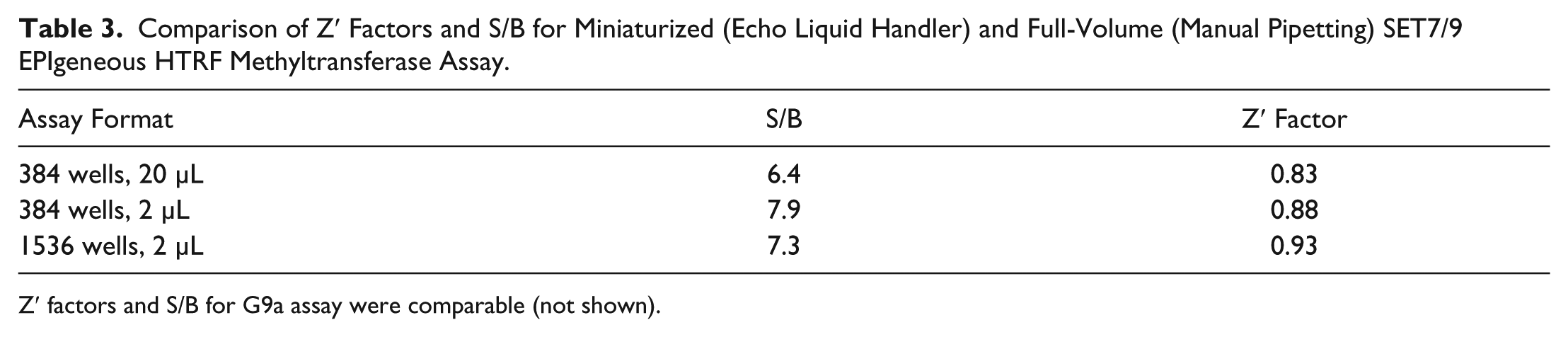
Z′ factors and S/B for G9a assay were comparable (not shown).
AlphaLISA Methyltransferase Assay
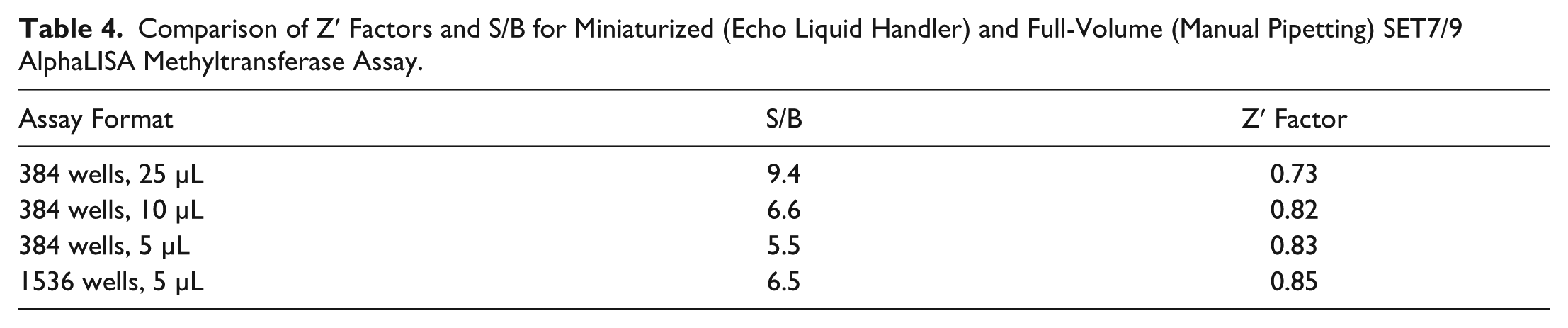
For the SET7/9 AlphaLISA assay, we generated IC50 data for SAH and Sinefungin and calculated Z′ factors at 5 µL assay volumes in 384- and 1536-well formats, as well as 10 µL assay volumes in the 384-well format. There was good agreement among the IC50 values for both compounds at the miniaturized assay volumes ( Fig. 7 ). IC50 values were also fairly consistent with literature values of 359 nM for sinefungin and 16 µM for SAH. 3 Both compounds showed slightly higher potency in our assay, which could be attributed to transfer of the compound via the Echo liquid handler, eliminating potential loss of compounds, particularly those with hydrophobic characteristics, from sticking to tips.22–25 Z′ factors for the miniaturized assays were excellent—0.82 and above in the miniaturized assay volumes, in both 384- and 1536-well formats, compared to 0.73 in the manually pipetted 25 µL 384-well assay. Assay window was smaller in the miniaturized assays—lowest at 5.5 S/B, compared to 9.4 S/B in the 25 µL assay. This could potentially be improved with further optimization at the miniaturized volumes ( Table 4 ).
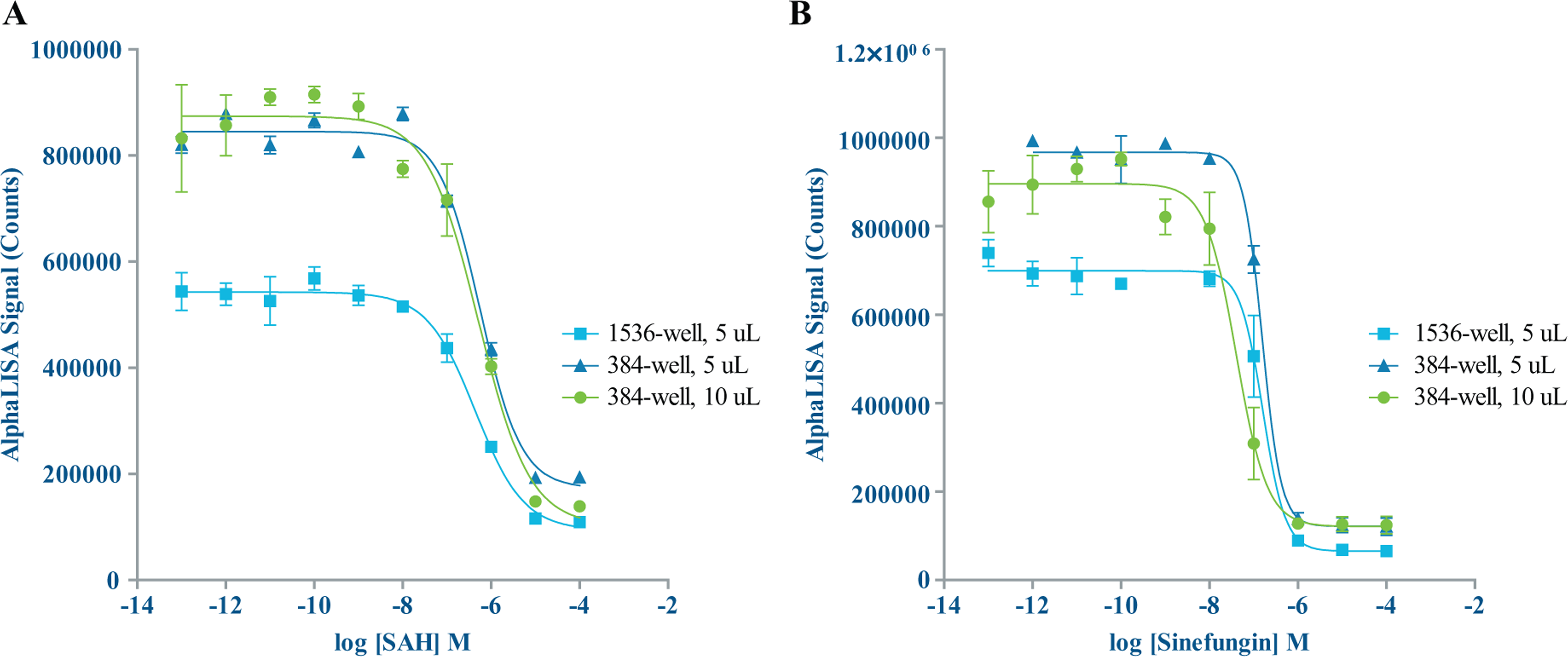
Inhibitor titrations—AlphaLISA. (
Comparison of Z′ Factors and S/B for Miniaturized (Echo Liquid Handler) and Full-Volume (Manual Pipetting) SET7/9 AlphaLISA Methyltransferase Assay.
Conclusion
In this study, we set out to evaluate the miniaturization of two different epigenetic assay platforms, with two different enzymes, taking advantage of the ability of the Echo liquid handler to dispense nanoliter volumes of reagents with accuracy and precision. Both the HTRF and AlphaLISA technologies are robust, sensitive, no-wash assays that are amenable to being performed at lower volumes without negatively impacting data quality.
The ability to miniaturize these assays up to 10-fold represents a significant cost savings in reagents, enzymes, substrates, and compounds ( Table 5 ). Additionally, the ability to dose compounds at nanoliter volumes allows for direct dilution of compounds while keeping DMSO concentrations low, thereby eliminating the need for intermediate dilutions. We have shown here that we were able to miniaturize the EPIgeneous HTRF Assay for two methyltransferase enzymes, suggesting that this assay kit could be miniaturized across a range of methyltransferases. We also demonstrated the ability to acoustically dispense Alpha beads and miniaturize the SET7/9 methyltransferase assay to a 5 µL assay volume, a five-fold reduction from the standard assay volume. By using the Echo liquid handler to dispense all assay components and reagents, we were able to achieve excellent Z′ factors even while significantly reducing assay volumes.
Comparison of Costs between Standard Volume EPIgeneous HTRF Methyltransferase Assay and Miniaturized Volume Assembled with the Echo Liquid Handler.
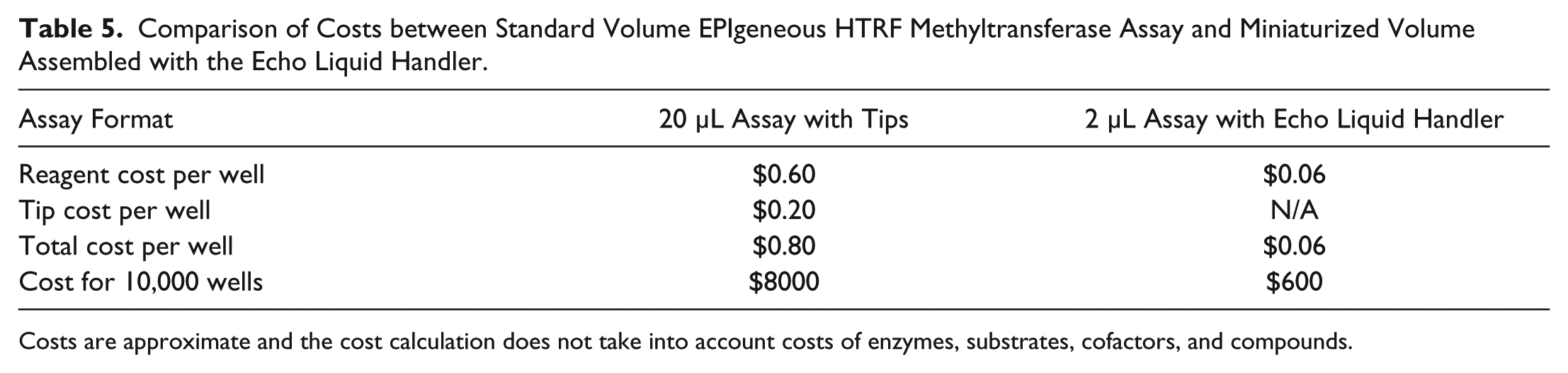

Costs are approximate and the cost calculation does not take into account costs of enzymes, substrates, cofactors, and compounds.
Footnotes
Acknowledgements
Bonnie Edwards and John Lesnick are employees of Labcyte Inc. Jing Wang was formerly an employee of Labcyte Inc. Nga Tang is an employee of Cisbio US. Carl Peters is an employee of BMG LABTECH.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
