Abstract
Flow cytometry is a powerful tool providing multiparametric analysis of single cells or particles. The introduction of faster plate-based sampling technologies on flow cytometers has transformed the technology into one that has become attractive for higher throughput drug discovery screening. This article describes AstraZeneca’s perspectives on the deployment and application of high-throughput flow cytometry (HTFC) platforms for small-molecule high-throughput screening (HTS), structure–activity relationship (SAR) and phenotypic screening, and antibody screening. We describe the overarching HTFC workflow, including the associated automation and data analysis, along with a high-level overview of our HTFC assay portfolio. We go on to discuss the practical challenges encountered and solutions adopted in the course of our deployment of HTFC, as well as future enhancements and expansion of the technology to new areas of drug discovery.
Introduction
Flow cytometry is a flexible, rapid, and quantitative technology that enables multiparametric analysis of single cells or particles.1,2 Although well established as a cell biology technique, the relatively recent development of high-throughput flow cytometry (HTFC), enabled by the introduction of faster plate-based sampling, has transformed the technology into an attractive drug discovery platform.3–5 Various HTFC systems with fast autosampler devices aligned to traditional flow cytometers have been reported by different research groups3–9 (e.g., an autosampler aligned to the CyAn system from Beckman Coulter, Brea, CA; or to the FacScan, LSR II, and Accuri C6 from BD Bioscience, San Jose, CA); more recently, several fully integrated HTFC systems have become available5,8,9 (e.g., the iQue Screener from IntelliCyt, Albuquerque, NM; the MACSQuant from Miltenyi Biotec, Bergisch Gladbach, Germany; and the Guava from Merck Millipore, Darmstadt, Germany). In recent years, an increasing amount of reports have described diverse applications of different HTFC platforms using 96-, 384-, and 1536-well formats at various phases of drug discovery in pharma and biotech companies as well as in academic institutes.3,8,10–33 These reports demonstrate that HTFC technology has played, and will continue to play, an important role in early drug discovery. This article describes the deployment of HTFC platforms within AstraZeneca and MedImmune to support our early drug discovery process from target identification (TI), through hit identification (HI) and lead identification (LI), to lead optimization (LO).
Aligned to the relevant sample acquisition platform, successful exploitation of HTFC within drug discovery requires robust and reproducible upstream assay processes for sample preparation, such as cell staining or analyte capture, combined with appropriate downstream data analysis tools. Although there are numerous reports describing applications of HTFC technology, reports with details in terms of how samples are prepared prior to sample acquisition on HTFC platforms are very limited. To our knowledge, only a few research groups have reported the establishment and applications of automated robotic systems for HTFC sample preparation. The scientists at the University of New Mexico Center for Molecular Discovery established fully automated solutions with sample preparation equipment and an HTFC platform integrated on an Agilent BioCel System,16,34 and a group at the Genomics Institute of the Novartis Research Foundation (GNF) has developed one fully automated HTFC GNF system. 35 Use of separate workstations for sample preparation and sample acquisition have also been reported for performing HTFC.14–16,18,20 In this article, we describe AstraZeneca’s perspectives on our current and future-looking automation platforms that enable integration of HTFC assay development and screening, along with our current data analysis workflow. We also provide our insights into the practical challenges encountered during the course of HTFC deployment in our organization. In addition, our view on current best practices for HTFC screening, along with future enhancement and expansion of the technology applied to emerging areas of drug discovery, is discussed.
HTFC: High-Level Applications
Flow cytometry measures both physical and biological properties of cells or particles suspended in a liquid stream as they pass through a light beam. The technology’s high sensitivity enables multiparametric quantitative data capture of single cells or particles at rates up to 50,000 cells or particles per second. 1 It is broadly used in many areas of biological science, from basic research to the diagnosis of disease and immune-phenotyping of clinical samples, but traditional flow cytometry has been limited to low-throughput applications due to the relatively slow sampling speed and the requirement for large numbers of cells. The recent development and commercialization of the HyperCyt (IntelliCyt) sampler using air gap–delimited aspiration and sample feed, however, were a turning point for the platform, allowing flow cytometry to enter the high-throughput screening (HTS) arena.1,5
Compared to screening technologies with simple whole-well readouts, HTFC screens permit the simultaneous quantitative measurement of multiple parameters and can therefore be considered “high content” in nature. For example, flow cytometry allows quantification of secreted, cell surface, and intracellular proteins; specific protein modifications; DNA content; cell viability; cellular granularity; and cell size in a single acquisition step. This high-information content enables researchers to group (or gate) cellular subpopulations phenotypically based on one or more of the observed parameters, expanding the power of analysis beyond gross population measures. There are numerous and diverse recent examples of cell-based and bead-based applications of HTFC in preclinical drug discovery, many of which take advantage of the subpopulation ability that flow cytometry provides (e.g., phenotypic screening of cytotoxic T-lymphocyte lytic granule exocytosis in a 1536-well format, 19 phenotypic screening of human primary T cells,22,23 screening using fluorescent barcoded cells,11,24 and screening of Rho GTPases using a bead-based assay 12 ).
AstraZeneca HTFC Deployment: Historical Perspective and Business Drivers
As one might expect with a technology as established as flow cytometry, AstraZeneca has a wide range of traditional flow cytometers embedded within various core facilities throughout different research sites. As described earlier, however, these instruments generally have low-throughput, have long sample acquisition times (a requirement for large numbers of cells), and are often complicated to configure for use (e.g., the setup procedure includes manual detector voltage adjustment of each fluorescence channel to optimize signal resolution, and making a new compensation matrix every time the detector voltage settings have been changed). These combined attributes have long made such systems unsuitable for screening applications and have, until very recently, prevented wider adoption of the technology beyond specialist users.
With the advances in HTFC described earlier, AstraZeneca recognized the potential for this technology to meet the needs of our emerging portfolio, including the screening of soluble analytes or suspension cells, particularly when rare or expensive primary immune cells were needed. Although many flow cytometry systems can perform sample acquisition from 96-well plates, only a few can acquire directly from 384- or 1536-well plates. One option for acquisition from these higher density plate formats is the coupling of a HyperCyt autosampler onto the front end of a traditional flow cytometer. Such integrations have been reported previously (the CyAn system from Beckman Coulter, or the FacScan and Accuri C6 from BD Bioscience) as platforms enabling HTFC screens in 96-, 384-, and 1536-well formats.3,4,8,36–39 A fully integrated HTFC screening system comprising a HyperCyt sampler coupled to an Accuri C6 was also launched by IntelliCyt. 4
Although integration of a HyperCyt sampler onto a flow cytometer addressed part of the throughput concerns from a screening perspective, there were still a number of challenges associated with deploying these types of integration more widely. The physical size of the integrated platforms and the level of expertise required to properly operate them meant they were challenging for a wider user base. A few manufacturers looked to address these concerns by designing systems with built-in plate-based autosamplers (e.g., the MACSQuant from Miltenyi Biotec and the Guava from Merck Millipore). These designs brought the devices closer to a single reader platform that not only reduced the footprint but also attempted to reduce the system complexity of the user interface. Such fully integrated systems have become very attractive to laboratories looking to exploit the power of flow cytometry for plate-based screening. The only platform that combines the power of the HyperCyt sampler and the ease of use from a complete system in a box, however, is the iQue Screener system from IntelliCyt Corporation.21,40 This system is built on the original HyperCyt technology 9 and is commercialized by IntelliCyt Corporation.
In 2012, we began using IntelliCyt’s original HTFC screening system 4 at MedImmune in the United Kingdom. Previously, flow cytometry was used to detect binding of a limited number of antibodies to cells expressing very low levels of cell surface antigens. These assays were also used to identify antibodies with weak affinity to their cell surface target, which could be detected using signal amplification such as biotinylated secondary antibodies and fluorophore-labeled streptavidin. To develop sensitive assays, numerous wash steps were required to reduce the background to enable detection of very low binding signals. These assays were carried out in 96-well plates and read with traditional flow cytometers, which led to long sample acquisition times and difficulty analyzing the data because the software packages available at the time struggled with plate-based data. Implementation of IntelliCyt’s HTFC screening system reduced the sample acquisition time significantly from one 96-well plate/h to one 96-well plate/10 min. The use of IntelliCyt’s ForeCyt software enabled analysis of flow cytometry data throughout a whole 96-well plate in a few steps and simplified gating of the data to determine levels of fluorescence for each sample tested. Since its installation, assays such as antibody internalization, antibody binding to monitor species cross-reactivity and selectivity, receptor ligand competition assays, and mechanism of action studies have all been run successfully for limited panels of antibodies. There is still a bottleneck in running high-throughput washed assays, and additional investment is required to support running these assays at a larger scale, such as automated plate washing and additional liquid handling. Implementation of the HTFC system has increased the number of flow cytometry users because of the ease of setup (e.g., the wide dynamic range and fixed voltages of the detectors make setup more straightforward compared to other cytometers, and therefore this system is more amenable to the nonspecialist).
In 2013, the HTS department of AstraZeneca’s Discovery Sciences IMED Biotech Unit retired its suite of FMAT readers, which had reached the end of their operational life and became unsupported by Applied Systems. The FMAT units had been successfully used in multiple receptor-binding and cytokine detection HTS campaigns. Recognizing the ongoing requirement for a technology suitable for HTS screening of such targets while observing the emerging need for HTS using suspension cells, the AstraZeneca HTS department explored various options and decided to invest in the iQue Screener HD platform (IntelliCyt). This HTFC screening system focuses on high throughput (384- and 1536-well plates), fast sampling speed (<20 min per 384 wells), and small sample volume (minimal sample aspiration volume as low as 1 μl per well), and it is configured with two lasers and six detection channels. As found with the earlier ForeCyt software on our MedImmune HyperCyt installation, the iQue HD ForeCyt software design allowed users with little flow cytometry expertise to operate the system (e.g., the ForeCyt software provides the flexibility to set up sample acquisition, probe rinsing, and sampler cleaning; provides plate-level annotation, multilevel analytics, data visualizations linked by well, real-time interconnected analysis, and result updates; and has optimized protocols that can be saved as templates and applied for new sample acquisition, with all desired parameters analyzed automatically).
The Discovery Sciences IMED Biotech Unit at AstraZeneca is a centralized function located on different AstraZeneca sites and encompasses all the capabilities required to support early drug discovery activities. As the HTFC technology became embedded within the HTS department and AstraZeneca’s inflammatory disease portfolio expanded, we also grew our HTFC capability across different sites. This enabled additional AstraZeneca sites to access the technology but also meant that assay development and iterative structure–activity relationship (SAR) profiling activities could be run concurrently with HTS campaigns. At this stage, we invested in the iQue Screener PLUS system, a HTFC system capable of 96- or 384-well sampling configured with three lasers and 15 detection channels, within our assay development and profiling groups in both Sweden and the United Kingdom. These aligned investments have enabled us to build a HTFC capability, which is able to support the early drug discovery process from the TI through HI and LI to LO phases within Discovery Sciences across three sites. We now have four iQue Screener systems deployed globally within AstraZeneca ( Fig. 1 ): two iQue HD systems within our Global HTS Centre in Cheshire, UK; one iQue PLUS system aligned with the local cardiovascular metabolic disease, and respiratory and inflammation disease, areas in Gothenburg, Sweden; and one iQue PLUS system aligned with the local oncology disease area in Cambridge, UK.
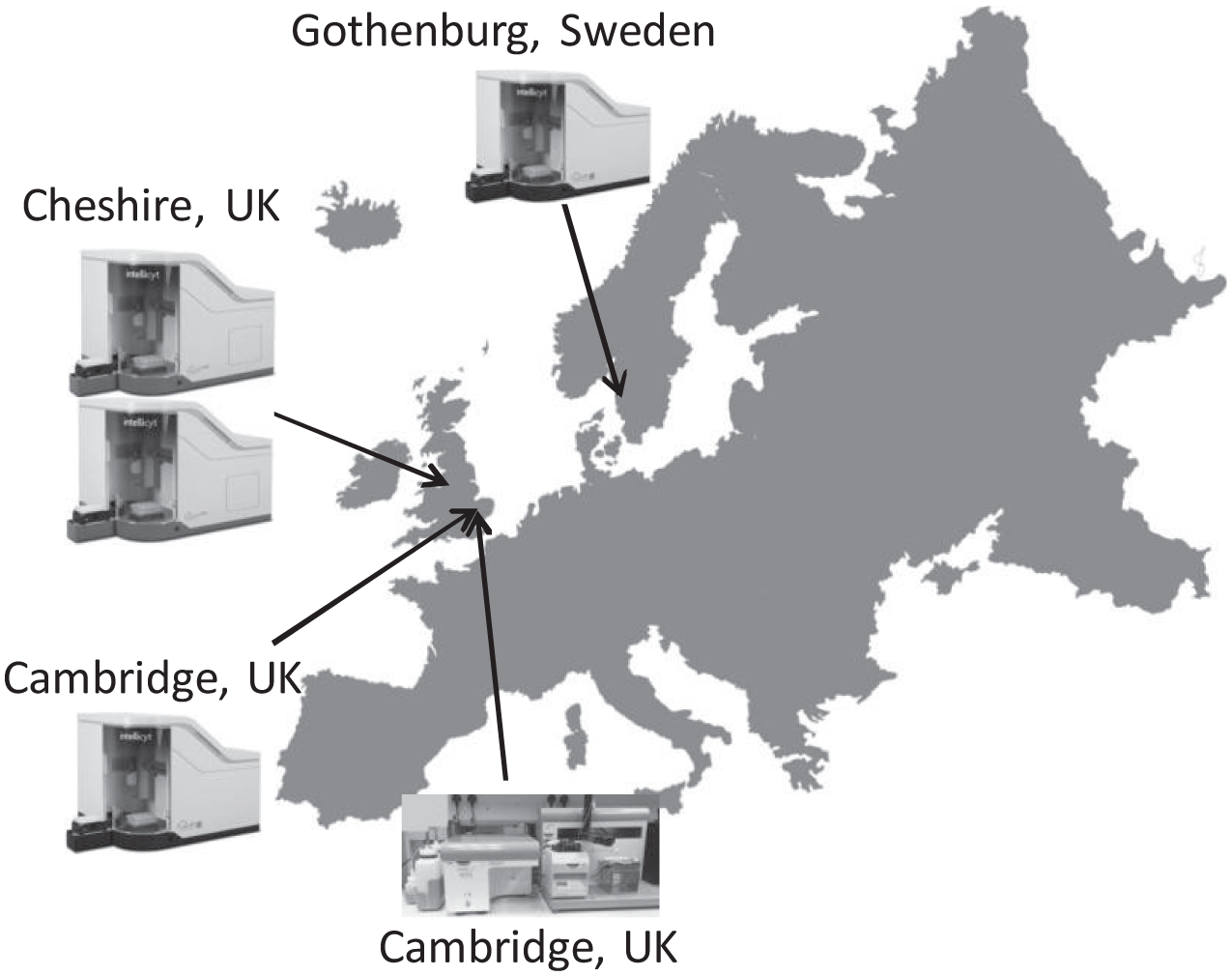
Deployment of high-throughput flow cytometry (HTFC) systems at AstraZeneca. Two iQue Screener HD systems at the HTS Centre in Cheshire, UK; one iQue Screener PLUS at Discovery Sciences in Gothenburg, Sweden; one iQue Screener PLUS at Discovery Sciences in Cambridge, UK; and one HTFC Screening System at MedImmune in Cambridge, UK.
HTFC Automation Platforms at AstraZeneca
As with any screening technology, HTFC requires robust and reproducible assays. 41 For flow cytometry, sample preparation such as staining cells for surface or intracellular proteins is often required prior to sample acquisition. Staining of suspension cells for traditional flow cytometry is often performed in multistage manual processes (including sequential media removal, addition of staining solutions, centrifugation, and cell-washing steps), which can result in significant cell loss and large assay variability. Use of automation to facilitate HTFC assay development and screening is critical to minimize sample loss and intra- and interassay variability.
Various automation platforms have been deployed previously within Discovery Sciences to support different types of cell-based screening; however, the majority of these were designed for screening adherent cells and therefore do not have all the peripheral devices required for flow cytometry assays. This is particularly true for assays using suspension cells or beads in which medium exchange or cell washing requires the use of a plate centrifuge device to spin down suspension cells or beads, and liquid-handling equipment to remove supernatants.
Because sample acquisition times in flow cytometry (even on our HTFC units) can be comparatively lengthy, an ideal automation platform would be one offering the flexibility to perform full end-to-end screening (including assay build, sample preparation, and sample acquisition) or any subset of the full end-to-end process. Our current solution is to decouple the assay and data acquisition steps for our automated HTFC workflows. Figure 2 shows four of our existing platforms. Figure 2A shows an integrated assay platform primarily used for multistep suspension cell staining deployed for activities in the TI, LI, and LO phases of drug discovery. This platform is built around a 3 m ORCA rail, controlled via SAMI scheduling software (Beckman Coulter), and comprises a plate carousel (Beckman Coulter), a CytoMat2 incubator (Thermo Fischer Scientific, Waltham, MA), a Vspin centrifuge (Agilent Technologies, Santa Clara, CA), an ELx405 cell washer (BioTek, Winooski, VT), a Biomek NXP liquid handler (Beckman Coulter), and two Multidrop Combi dispensers (Thermo Fisher Scientific). Depending on the particular assay protocol, this automation configuration allows cells to be fixed, permeabilized, and stained in repeated cycles of liquid additions via Multidrop Combi, centrifugation by Vspin, and aspiration by the ELx405 cell washer. On this platform, cells can be incubated in the CytoMat2 prior to any staining procedure. The NXP liquid handler is used when protocols require transfer of sample between plates (e.g., transfer of detached adherent cells or transfer of cell supernatants). We have successfully used this automation platform to prepare samples for several HTFC phenotypic screens (up to 20,000 small-molecule compounds per screen, with a throughput of more than 3000 assay points per screening occasion), including one particularly complex screen to identify compounds affecting the stability of human regulatory T cells (Treg). 22 The Treg assay cell-staining protocol involves a total of 43 plate manipulation steps by the robotic arm, in which the cells are first stained with a fixable live/dead stain, followed by fixation, permeabilization, and staining for both CTLA4 and FOXP3. Human Tregs are a very rare cell population within the blood, and consequently running this screen was only possible through the use of a low cell number per well permitted by the HTFC technology, along with the minimal cell loss achieved through optimized staining protocols. Following sample preparation on the platform depicted in Figure 2A , we manually remove plates to a second automation platform dedicated to feeding the iQue PLUS instrument. This platform ( Fig. 2B ) is built around an ACell benchtop automation system, controlled via Cellario scheduling software (HighRes Biosolutions, Beverly, MA), incorporating a PicoServe plate carousel (HighRes Biosolutions), an ambient plate-nest, a CytoMat2 incubator (Thermo Fisher Scientific), and an iQue PLUS. In this setting, the incubator is used for cold storage of stained samples in plates, allowing maintenance of signal throughout time when large batches of plates are acquired. The extended period of storage time is assay specific, and it is determined based on signal-to-background and assay variability. In our hands, live cell assay storage time of stained samples can be up to 12 h prior to sample acquisition, whereas assays with fixed cells can be stored up to 72 h.
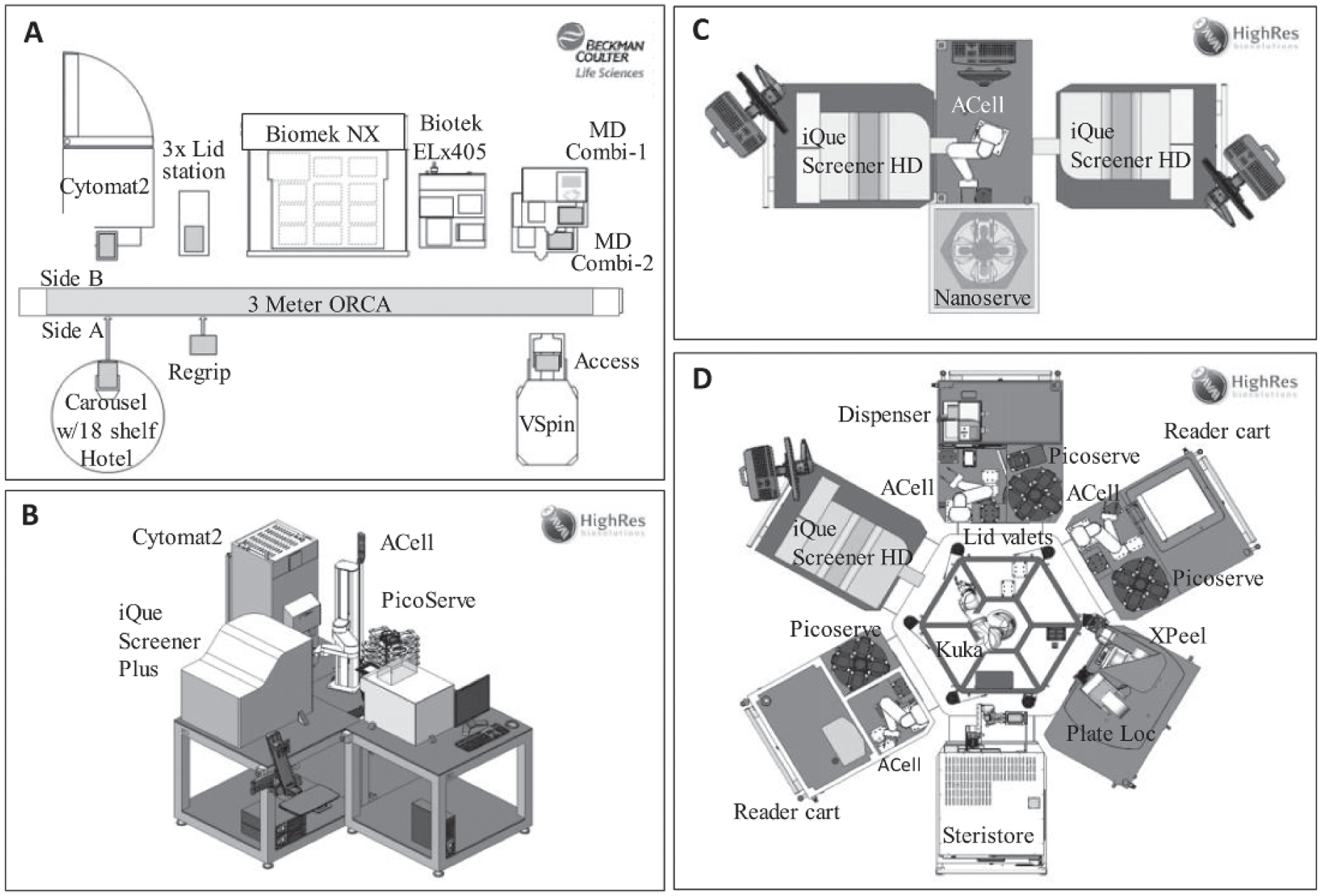
Examples of the current high-throughput flow cytometry (HTFC) equipment setup and automation systems. (
The Global HTS department provides HI capacity throughout all of AstraZeneca’s therapeutic areas, performing diverse HTS campaigns ranging in scale from 50,000 to 2,000,000 compounds. Screening 500,000 compounds (the largest HTFC screen run at AstraZeneca to date) in a 384-well format involves testing and analyzing samples from more than 1400 plates, and therefore requires well-validated and robust assays amenable to automation. 42 To enable the data acquisition throughput required while maintaining assay flexibility, the HTS department has also opted to decouple the sample preparation and sample acquisition steps. Figure 2C shows the current HTS HTFC implementation, comprising a pair of iQue Screener HD units linked to an ACell benchtop automation system. The system is controlled via Cellario scheduling software (HighRes Biosolutions) and further comprises a NanoServe plate carousel (HighRes Biosolutions) and an ambient plate-nest. Following the assay on upstream platforms, plates are manually loaded into the NanoServe carousel, then automatically fed into one of the iQue Screener HD units, then returned to the carousel once acquisition is complete. Even though the Cellario scheduler allows for the iQue devices to be used in a pooled (plates can go to the first free device) or round-robin (plates alternate between iQue #1 and iQue #2) fashion, we have found it important to define which plates are sent to each device. By controlling which plate set is acquired on which instrument, we can monitor each individual unit and follow the signal during the course of a screening batch. Because we know the signal stability of the assay, we can ensure that no signal change greater than 10% occurs, which is considered acceptable in our settings.
This flexible small-footprint automation platform allows unattended operation as well as automated cleaning of the cytometers at the end of each run. Although four HTS campaigns have been completed and more than 3 million data points have been collected using this simple platform, we are now in the process of integrating our iQue Screener HD units onto our larger HighRes platforms, through deployment onto CoLAB (collaborative laboratory) compatible carts. Figure 2D depicts a single iQue Screener HD cart docked into the CoLAB Flex HTS system. This modular automation platform can be configured to align different peripheral devices and therefore customized to suit many different assay requirements. The CoLab Flex Carts can work standalone, be partnered to other carts, or form a larger CoLab Microstar platform, as shown in Figure 2D . This ultimate flexibility will allow us to automate the full end-to-end process (or any subset of the process from assay build to sample acquisition), thus overcoming one of our current limitations (i.e., separation of HTFC sample acquisition vs. sample preparation) and moving us toward our idealized automation platform (i.e., a fully automated system with the sample preparation equipment and HTFC sample acquisition platform integrated).
Deployment of the HTFC Platform to Fulfill Business Needs
The global suite of iQue Screener systems integrated onto different automation platforms (as described earlier) along with the original HTFC screening system within MedImmune are continuing to deliver project-driving data to an ever-expanding portfolio of drug discovery projects at AstraZeneca. We not only are delivering an increased volume of data through our plate-based HTFC screening of antibody libraries, but also have shown impact across a broad range of our therapy areas and project phases ( Fig. 3 ) through delivery of the following: HTS campaigns of small-molecule libraries up to 500,000 compounds in size for the HI phase, 43 phenotypic screens for the TI phase, and mechanism-of-action studies and safety profiling for the LI and LO phases. 44 So far, HTFC as a technology has positively affected more than 15 discrete drug discovery projects within AstraZeneca since our initial 2014 investment in the IntelliCyt iQue platforms. Table 1 lists the various iQue screening applications within AstraZeneca to date. The vast majority of these have used suspension cell models, including various human primary immune cells and induced pluripotent stem cell (iPSC)-derived cells. Although it is more challenging to run adherent cell-based HTFC assays with an incubation longer than overnight due to the requirement of detaching cells from plates and making single-cell suspension before sample acquisition on HTFC, we have successfully delivered a 20,000-compound subset screen in 384-well plates against an adherent HEK293 cell model. The types of cellular assay endpoints and phenotypes that we have used can also be seen in Table 1 , and include both surface and intracellular protein expression, proliferation, apoptosis, cell differentiation, and viability, among others. It is this array of different possible endpoints that makes the technology such a powerful tool for drug discovery, enabling deep characterization of the assay system and the effect of modulators on that system.
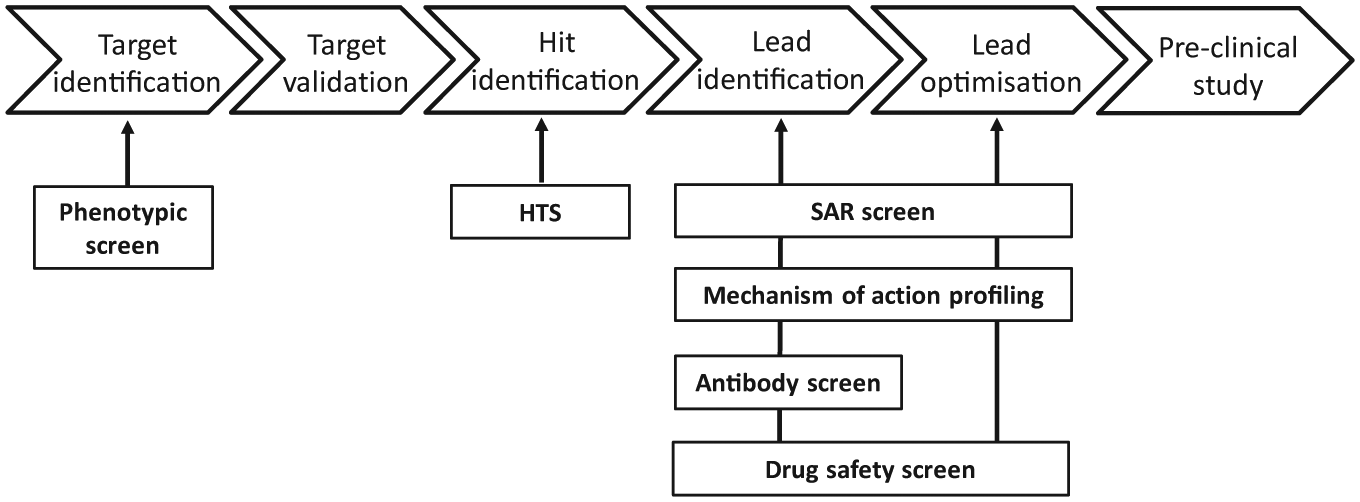
High-throughput flow cytometry (HTFC) applications at AstraZeneca in early drug discovery. The HTFC technology has been deployed to support numerous drug discovery projects: phenotypic screen for target identification, high-throughput screening (HTS) campaigns for hit identification, iterative structure–activity relationship (SAR) screening, mechanism-of-action profiling, antibody screening, and drug safety screening for lead identification and lead optimization.
Examples of iQue screening applications run at AstraZeneca.
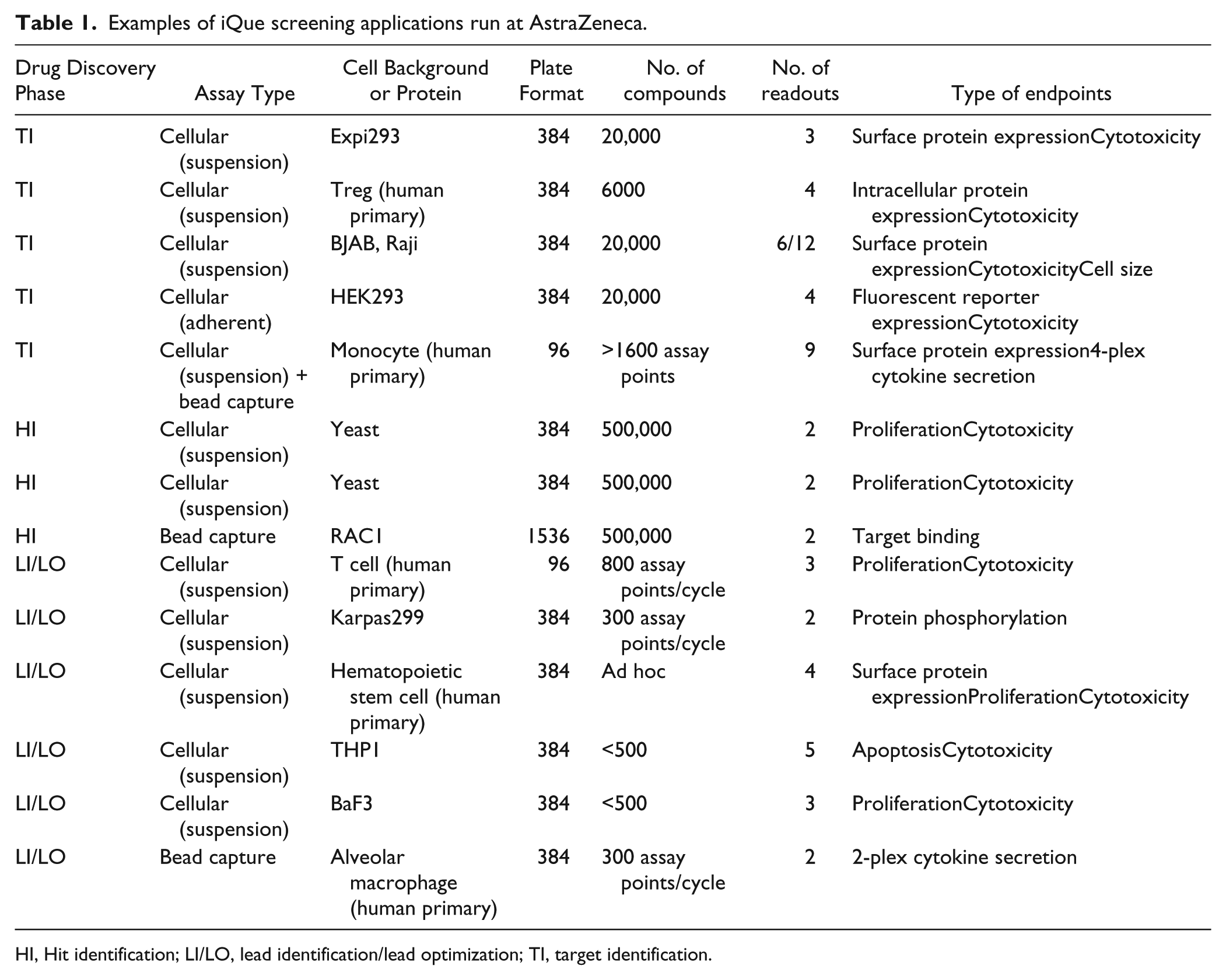
HI, Hit identification; LI/LO, lead identification/lead optimization; TI, target identification.
The nature of the single-cell analysis inherent in flow cytometry can be used to multiplex different cells together in a single well, and then subpopulate these at the data analysis phase. Provided there is a given measured feature to apply appropriate “gating” to the cellular objects identified, then quantification of different endpoints in any or all of those cell subpopulations can be obtained. We have exploited the multiplexing of different cell lines in a number of recent projects; for example, we have developed and validated an assay ( Fig. 4 ) using cell encoding to multiplex two B-cell lines (BJAB and Raji) within the same well by loading BJAB cells with an encoder dye (Cell encoder Red, FL4, and IntelliCyt) and mixing these labeled BJAB cells with unlabeled Raji cells. The multiplexed cells were subsequently dosed with test compounds, then stained for viability along with two surface markers. This assay has been successfully applied to the screening of a 20,000-compound subset, using the automation platforms shown in Figures 2A and 2B . The benefits of multiplexing two cell lines in one assay include reducing the number of physical plates that need to be processed, and therefore a resultant reduction in timelines and reagent costs for delivery of a data package across the different cell backgrounds. In addition, running a single physical screen but gathering data on multiple cell types ensures that compound dosing and treatment times are equivalent across the different cells tested (as they are within the exact same well of a microtiter plate).
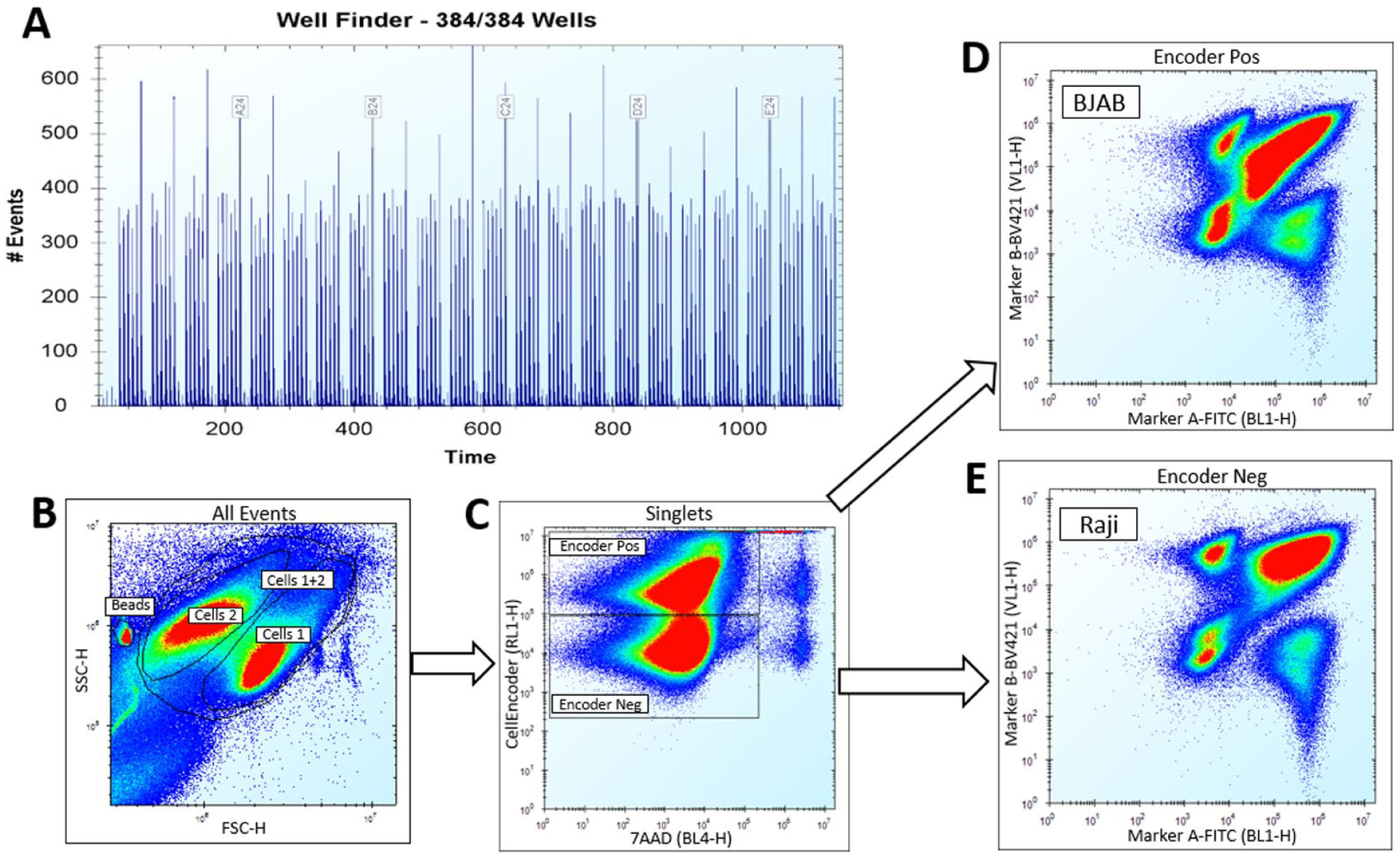
Gating strategy for the assay using multiplexing of two B-cell lines (BJAB and Raji). BJAB cells were loaded with cell encoder dye (Cell encoder Red, FL4) prior to being mixed with unlabeled Raji cells. Multiplexed cells received compound treatment followed by cell staining on automation. Marker beads were added for supporting automatic well identification before reading samples on an iQue Screener Plus. (
In addition to cellular assays, we have also successfully used iQue bead-based immunoassays for multiplexed cytokine quantification ( Table 1 ). We have been able to optimize and miniaturize such assays to 6 µl in volume (2 µl biological sample + 2 µl capture bead + 2 µl detection reagent) instead of 30 µl in volume using the standard protocol. For the cytokine immunoassays we have studied, the iQue Qbeads assays showed good assay correlation compared to other commonly used immunoassays (e.g., enzyme-linked immunosorbent assay [ELISA], mesoscale discovery [MSD], homogeneous time-resolved fluorescence [HTRF], and Luminex). For example, similar detection ranges and good correlation (with R2 values > 0.89) were observed between iQue and MSD assays for human cytokines interleukin-6 (IL6), IL10, IL12p70, and tumor necrosis factor-α (TNFα), and the iQue assay showed good correlation but a broader detection range than the ELISA assay for human Rantes (R2 = 0.7) and the HTRF assay for human TNFα (R2 = 0.92). The properties of multiplexing, a large detection range, high throughput, small sample consumption, and miniaturization opportunity of the iQue Qbead assays can result in cost savings and improved time efficiency for screening larger sample sets. Indeed, the possibility for large multiplexing of analyte detection using the IntelliCyt Qbead PlexScreen reagents or in-house customized capture beads against analytes of interest makes this an attractive alternative to MSD, Luminex, and ELISA formats. As an example, we compared a 2-plex iQue QBead assay (using the miniaturized protocol mentioned) versus MSD and HTRF quantification of human TNFα and IL10. Treatment of human alveolar macrophages or THP1 cells with a salt-inducible kinase inhibitor HG-9-91-01-induced concentration-dependent reduction in TNFα release. HG-9-91-01 treatment induced a similar effect on human TNFα release, with IC50 values at 34 nM and 28 nM, respectively, for iQue versus MSD in the human alveolar macrophages assay, and with IC50 values at 120 nM in both iQue and HTRF in the THP1 assay. IL10 concentrations in the supernatants from human alveolar macrophages and THP1 cells were very low and close to the detection limits in the iQue, MSD, and HTRF assays. With these assay comparison results, we ultimately selected the iQue approach as the most appropriate option for ongoing support of one LO project. This assay enabled iterative SAR screening for optimizing salt-inducible kinase inhibitors by testing compound effects on TNFα and IL10 release in a relevant human alveolar macrophages cell model. We have gone on to 22 use the above miniaturized protocol to screen for the release of cytokines or other secreted proteins from many other human primary cells and to screen for cytokines in biological fluids taken from murine in vivo studies, supporting several projects over the LI and LO phases.
Along with LI and LO support, Table 1 also shows a number of HTS campaigns delivered on the iQue Screener HD platforms, one of the first being a bead-based target-binding screen against the small GTPase RAC1. 43 This campaign was built around a previously published assay 45 in which multiple GTPase targets were multiplexed in a single assay, whereby it was miniaturized further to a 1536-well format to enable the screening of a 500,000 small-molecule compound library in less than 5 weeks. Conversion of the published 384-well assay to a 1536-well assay was undertaken as part of a strategic collaboration between AstraZeneca and IntelliCyt, whereby IntelliCyt provided their expertise and resources for the initial proof of concept, then AstraZeneca performed further optimization and validation to deliver or develop a robust HTS assay. A robust assay in our view should satisfy the following criteria: a robust Z’ factor of at least 0.5, a signal-to-background ratio greater than 3, a coefficient of variation (CV) across whole plates of less than 10%, and showing reproducibility when testing a defined set of compounds on two separate occasions. 42 The partnership worked very well, being valuable to both sides as a mechanism to embed the HTFC technology quickly within the HTS setting, while leaving AstraZeneca assay development resources free to focus on other portfolio deliveries.
HTFC Data Analysis Workflow
HTFC generates large amounts of data. Raw data files for a single 384-well plate can easily exceed a gigabyte depending on the complexity and event count of the assay. The size and complexity of HTFC data as well as the need for standard flow cytometry data analysis procedures such as compensation, population gating, and generation of population metrics set specific minimum requirements for HTFC data analysis tools. ForeCyt is the software bundled with the iQue systems and, along with the direct control of the instrument, also provides functionality specific to the iQue technology (e.g., well identification through delimiting by air gap–delimited aspiration and sample feed, a proprietary technology by which the instrument autosampler separates well samples with an air bubble, allowing the software to determine what sample comes from what well and making the instrument a true high-throughput instrument). On sample acquisition via the iQue, one raw data file for the entire microtiter plate is generated, and data are then divided into that belonging to each individual test well on the plate. Once this annotation of data has been performed, it is then possible to extract relevant data files in Flow Cytometry Standard (FCS) format for downstream processing and visualization in third-party tools, or to use the in-built gating and visualization options directly within ForeCyt. We have found the ForeCyt software to be user-friendly and easy for applying flow cytometry–specific concepts such as compensation. The visualizations provided within the software are adequate for most needs, and we have found most users tend to perform all of their raw data analysis in this package. ForeCyt provides an easy mechanism for saving an entire assay setup (acquisition, gating, and visualizations) as a template, and we use this functionality frequently to ensure that once an assay is developed, the protocol can be “locked down,” thus ensuring consistent screening results each time the protocol is run.
Figure 5A outlines the end-to-end workflow for HTFC data analysis used at AstraZeneca today. In brief, the ForeCyt software is used for well identification, compensation, gating of populations, and generation of desired population metrics. We then load exported plate metrics into Genedata Screener (Genedata, Basel, Switzerland), a software used at AstraZeneca for analyzing screening data, to associate our HTFC data with identifiers for the test compounds and the specific plate-layouts (including the position of on-plate control wells). In this analysis workflow, each calculated feature or metric is aligned in Screener as a separate data layer. The downstream processing of assay data, including quality control (QC) through calculation of a robust Z’ factor 46 and other measures, data normalization, and finally dose–response curve fittings, is all performed in Screener. This aligns with our global process for data processing and ensures that the HTFC data are subject to the same strict business rules for data quality as any other screening technology used across AstraZeneca. Although the current workflow for HTFC data analysis has been used for several screening campaigns ( Table 1 ), it has disadvantages. Flexibility of analysis and traceability of results are restricted because raw data and analyzed data are stored in separate databases (the ForeCyt and Genedata Screener databases, respectively). If one observes that thresholds for a given population gate or gates should be adjusted at a later stage of data analysis, one needs to go back to the raw data in ForeCyt, implement any changes, and repeat the analysis process. Ideally, we would like to be able to take raw data (following well identification and compensation) directly into Genedata Screener and perform population gating within this application. We have been working with Genedata to help optimize the functionality already available within their Screener product for the manipulation and visualization of single-cell data in a flow cytometry context. The current tools available with Screener version 14 can be used for analyzing flow cytometry data using standard FCS files. We find, however, that the available visualizations are still insufficient to enable effective gating of cell populations under many circumstances. For example, histogram plots, density plots, and contour plots are currently not available, and it is only possible to view one plot at a time, making it difficult to track how modifications to one gate affect related gates in the hierarchy. We are continuing to work with Genedata to improve this. Figure 5B shows our idealized future workflow for HTFC data analysis when only the pre-processing steps of well identification and compensation are performed within ForeCyt. In such a workflow, raw data are stored together with gating strategy, ensuring consistency of gating throughout an entire screen, and providing both the flexibility of analysis at the assay optimization phase and traceability of results from end to end.
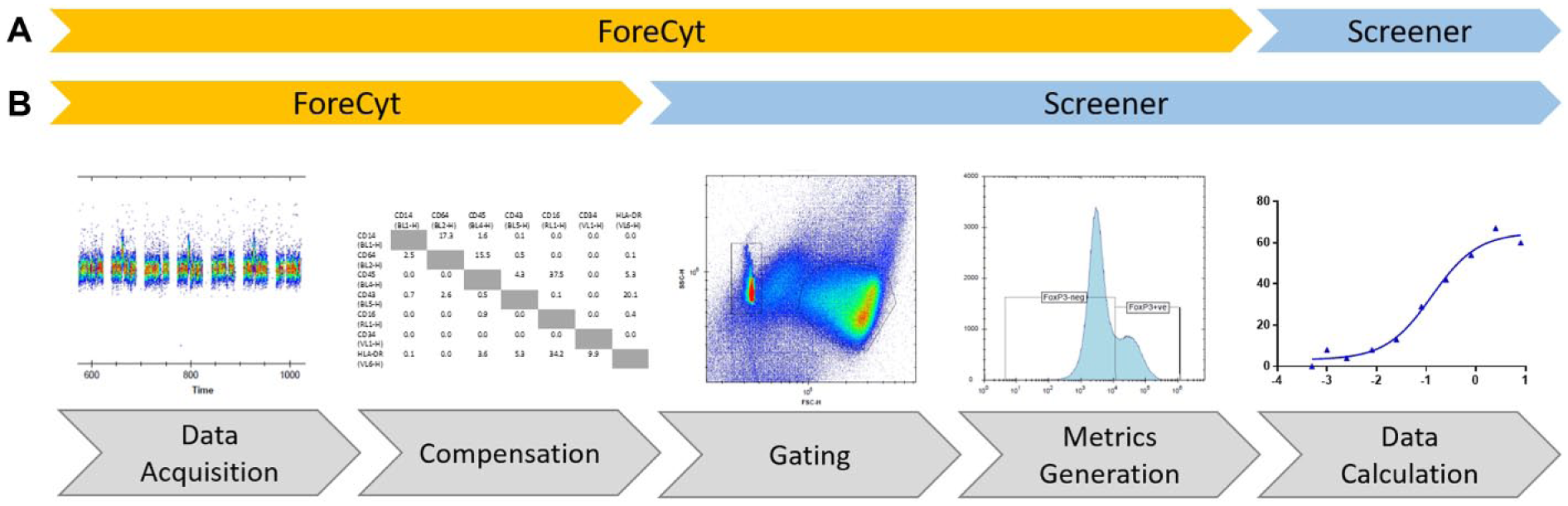
High-throughput flow cytometry (HTFC) data analysis workflow. (
Best Practice and Practical Challenges
As we have mentioned, flow cytometry is a very sensitive technology; however, data quality and reproducibility are directly related to instrument performance. Laser alignment and detection sensitivity are two key parameters that must be evaluated prior to running each experiment. In addition, ensuring the fluidics are free from debris is critical for maintaining a steady flow of particles through the flow cell. We have found that quickly establishing and ensuring all users follow specific HTFC best practice procedures for instrument QC, maintenance, and operation, as discussed in this section, are essential for optimum instrument performance (particularly as the user base for the technology expands). Due to the fact that AstraZeneca was an early adopter of the iQue platforms and one of the first to test these instruments in a true HTS setting, we encountered various practical challenges that IntelliCyt had not foreseen (discovering certain instrument limitations when run at scale, such as probes variability, valve board susceptibility to damage, etc.). During the past few years, we have worked together with IntelliCyt to find solutions and establish the appropriate best practice procedures for these instruments within our various settings.
A daily QC routine is now performed before starting any experiment to validate the instrument performance for laser alignment, detector sensitivity against each fluorescence channel, and a clear fluidic path (achieved by running specific instrument validation beads supplied by IntelliCyt). For each of these tests, the fluorescence intensity and associated CV of the top peak in each fluorescence channel are measured and checked against expected values. Only once our instruments pass all the QC criteria do we proceed with the experiment. In our experience, the optics within the iQue instruments are very robust: we see little to no variability on the detected signals, with a typical CV lower than 2.5% every day. The fluidics components, however, require particular attention. The plastic tubing can become deformed in the peristaltic pump, affecting sample flow, and debris accumulation in the tubing can cause irregular sample patterns or complete blockage of the fluidic path. IntelliCyt recommends replacing components in the fluidics system every 1 to 2 months, depending on usage; but, in our experience, these recommendations are too general, because we have noted a large and unpredictable variation in lifetime between individual components. As a precaution against run failures, we change the probe every other day when running a screen that generates continuous reading on the iQue for up to 12 h nonstop. Furthermore, we found that issues with irregular sample patterns were not sufficiently captured by the initial manufacturer-recommended daily QC procedure. We therefore continuously monitor performance of the fluidics system during screening, using well identification within ForeCyt as a guide to performance. As described earlier, the iQue Screener has an autosampler that delivers samples in an air gap–delimited stream to the cytometer engine. This allows the software to determine which sample comes from which well, resulting in a well identification trace showing regularly spaced events’ peaks along a time x-axis. This does mean that, ideally, data collection needs to be monitored in real time (or as close as possible) to respond quickly in the event of fluidic instability.
To assist in accurate well identification and the early identification of fluidics issues, we routinely add in-well marker beads to our cell-based assays. We find this particularly useful for conditions in which compound toxicity causes low event counts or no event counts. Irregular patterns of marker bead count throughout time are a clear indication of fluidic instability and allow us to take early action to rescue subsequent sample wells by running additional ad hoc cleaning steps. If the automated ForeCyt procedure for well identification continues to fail and cannot be easily corrected by manual adjustment, the probe and entire tubing set are changed, the system is fully cleaned, and the plate rerun if sufficient sample remains. To minimize the need for such plate reruns, we take steps to prevent cell debris accumulation within the fluidic path during screening. Our current recommended procedure is to add 0.01% NP-40 to the buffer solution on the iQue liquid deck whenever possible. Standard protocols for iQue analysis include an initial step of priming the fluidic path with buffer and also plate shaking at regular intervals throughout the run. During shaking, the probe and tubing are washed with buffer, taking advantage of the enforced break in sample acquisition. In this way, the fluidic path is cleaned at several points during plate acquisition without adding additional runtime. In our experience, adding NP-40 to the buffer solution has the combined effect of reducing sample carryover and preventing protein buildup, resulting in more robust sampling and cleaner well identification. As a result, we have observed a reduced frequency of blockages and increased processing of samples without interruption.
For regular maintenance of the iQue instruments, we apply the cleaning solutions and procedures recommended by IntelliCyt, although we needed to adjust the recommended intervals for various steps to manage our sample load. For example, during HTS screening when the iQue Screener HD instruments are in constant use for 8 h a day, we change the probe and tubing every 2 days, and the filters and peristaltic tubing every 3 weeks. Finally, we have found that during HTS campaigns, it is necessary to run the extended enzymatic clean (a procedure involving overnight incubation with a protease solution to digest any accumulated cell debris in the fluidic path) at least twice weekly to maintain optimum performance.
Summary
Flow cytometry has been a staple of most cell biology laboratories for the past few decades, but it is only recently that the technology, with major advances in faster sampling, has started to become more widely exploited within early drug discovery. With the advent of integrated platforms allowing 384- and 1536-well plate-based sampling, HTFC is now beginning to catch up with other high-content screening paradigms such as cellular imaging. In just 4 years, HTFC coupled to flexible automation platforms facilitating assay-build and screening has become embedded as a core drug discovery technology within AstraZeneca. It has moved from the domain of bespoke mechanistic profiling experiments on small sample sets, to a technology that now spans the drug discovery cascade from TI through HI and LI to LO. Although initially brought into AstraZeneca to increase throughput within antibody profiling and later to address key gaps in target–ligand binding or screening of suspension cells at a high-throughput scale, we have now realized many gains in other settings. Building miniaturized assays in both 384- and 1536-well plate formats has enabled us to screen larger compound sets or screen rare cells in a number of drug discovery projects. The ability to perform assays with multiplexed heterogeneous cell populations has decreased cell requirements and shortened timelines. The introduction of multiplexed bead-based cytokine assays is a lower cost and higher throughput alternative to many commonly used immunoassays.
Although we have successfully applied HTFC technology to support our early drug discovery process, we have also experienced certain key challenges along the way. Flow cytometry assays are more demanding than technologies with simple whole-well readouts due to complex assay-build steps as well as the requirement for stable fluidics during sample acquisition. This is particularly evident in the HTS setting, where blockage of the fluidic path during an unattended run can result in a large-scale loss of sample. Although regular QC and standardized cleaning procedures can help, it is inevitable that flow cytometers will require more maintenance than many other acquisition systems to maintain optimal performance. Even with all precautions taken, real-time data monitoring and speedy manual intervention when a fluidics blockage occurs can be important to avoid costly waste of reagent and time.
Since our original 2012 investment in HTFC, we have seen the technology application expand rapidly to support many projects across a number of our therapeutic areas. More recently, building on the learning that AstraZeneca has amassed in applying HTFC technology for other disease areas, we are rapidly expanding our use of the technology to prosecute targets in the immuno-oncology space using a variety of primary immune and complex tumor-immune co-culture models.
Based on the overall success we have seen with our HTFC deployment, our view is that the technology looks set to expand further across the AstraZeneca drug discovery portfolio, not only allowing prosecution of new disease areas but also bringing deeper subpopulation understanding to existing cellular assays. We expect novel applications combining live cell imaging readouts with sampling of cell supernatants for cytokine profiling via HTFC to be something that will drive even more value from co-cultures of rare cell types and will be facilitated by integration onto flexible automation platforms enabling this combination of different modalities. As deeper mechanistic understanding of drug–target interactions is brought further forward in LI and LO phases of drug discovery, the necessity to understand subpopulations of cell responders becomes more important. Single-cell analysis is moving more toward the mainstream of early drug discovery, and flow cytometry, a technology that has always harnessed the power of single-cell analysis, now unleashed from its traditional throughput constraints, is well positioned to be one of the technologies at the forefront of this movement.
Footnotes
Acknowledgements
The authors would like to thank Erik Müllers and Johan Brengdahl for their contributions in reporting results of multiplexed B-cell lines, Melker Göransson for the MSD human TNFα and IL10 results, Nils-Olov Hermansson for the HTRF TNFα and IL10 results, Pia Berntsson for the human MSD 4-plex results, and Anders Carvallin for the ELISA results. We would also like to acknowledge the help and support of the wider AstraZeneca HTFC users. In addition, we would like to thank Mark Carter, Kim Luu, and the assay development team at IntelliCyt for their work in conversion of the Rac1 GTPase assay to 1536 format. Finally, we would like to thank Beckman Coulter and HighRes Biosolutions for their help in generation of the automation platform schematics.
Declaration of Conflicting Interests
The authors disclosed the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: All authors are employees of AstraZeneca or MedImmune.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
