Abstract
Background:
Bioinformatics and statistical analysis have been employed to develop a classification model to distinguish toxic and non-toxic molecules.
Aims:
The primary objective of this study is to enumerate the cut-off values of various physico-chemical (ligand-centric) and target interaction (receptor-centric) descriptors which forms the basis for classifying cardiotoxic and non-toxic molecules. We also sought correlation of molecular docking, absorption, distribution, metabolism, excretion, and toxicology (ADMET) parameters, Lipinski rules, physico-chemical parameters, etc. of human cardiotoxicity drugs.
Methods:
A training and test set of 91 compounds were applied to linear discriminant analysis (LDA) using 2D and 3D descriptors as discriminating variables representing various molecular modeling parameters to identify which function of descriptor type is responsible for cardiotoxicity. Internal validation was performed using the leave-one-out cross-validation methodology ensuing in good results, assuring the stability of the discriminant function (DF).
Results:
The values of the statistical parameters Fisher Discriminant Analysis (FDA) and Wilk’s λ for the DF showed reliable statistical significance, as long as the success rate in the prediction for both the training and the test set attained more than 93% accuracy, 87.50% sensitivity and 94.74% specificity.
Conclusion:
The predictive model was built using a hybrid approach using organ-specific targets for docking and ADMET properties for the FDA (Food and Drug Administration) approved and withdrawn drugs. Classifiers were developed by linear discriminant analysis and the cut-off was enumerated by receiver operating characteristic curve (ROC) analysis to achieve reliable specificity and sensitivity.
Keywords
Introduction
The National Cancer Institute defines cardiotoxicity as the toxicity that affects the heart. 1 It is associated with the deviation of cardiac electrical activity and contractile dysfunction leading to heart failure. 2,3 The development of competent medication is also challenging due to the high investment and research needed to pursue safe drugs with no potential of adverse side-effects. 4,5 Drug-induced cardiotoxicity is majorly caused by anthracyclines, arrhythmias, antidepressants, beta-blockers, among others. 6 –8 A high dose of cardiotoxic drugs is reported to cause apoptosis and deregulation of myo-contractility. 9,10 Also, these drugs induce cardiac muscle injury and modification in the normal functioning of the ion channels and pumps (voltage-gated sodium and potassium ion channel and Na+-K+ ATPase pump). 11 –13
Various drugs including antidepressants, antipsychotics, antihistamines, analgesics, opioids, beta-blockers, antiarrhythmics, ACE inhibitors, diuretics, etc. are known to contribute side-effects and toxicity by interacting with numerous receptors both centrally and peripherally resulting in cardiac deaths. 8,14 –16 For instance, psychotropic drugs directly affect cardiac repolarization contributing to heart muscle disease and high-risk mental disorder. 14 Dopaminergic, serotonergic, histaminergic, beta-adrenergic, and muscarinic receptors are the examples of psychotropic targets producing adverse cardiovascular side-effects such as orthostatic hypotension, syncope and related medical ailment. 17
A drug discovery cycle ranges from 10 years and more and their cost in research and development is estimated to $1,000 million for a pharmaceutical company.
18
The methods to forecast blocking orproduction of some existing drugs for the treatment of various diseases is threatened with discontinuation, because of limited markets and suspicion of long-term adverse effects.
19
Computational approaches reduce these risks by learning the available toxicity data and make informed decisions to pursue new chemical entities or optimized leads under
We have proposed a toxicity prediction model in this article for the prediction of probable cardiotoxic drugs using a multi-parametric approach combining drug status, calculated physico-chemical and ADMET properties (ligand-centric descriptors) and docking scores obtained from protein target interactions (receptor-centric descriptors). 20,25 All these properties were then used to decipher the discriminant function using Linear Discriminant Analysis (LDA) for distinguishing cardiotoxic and non-toxic compounds. Fisher Discriminant Analysis (FDA) classified the compound data in the form of two classes based on independent and dependent variables. 26 –28 The classification of drugs being toxic or non-toxic relied on the coefficients of linear discriminant function and selected targets from each group play a crucial role in probability generation. 29
The objective of LDA is to derive a function to classify cardiotoxic and non-toxic compounds by determining the “cut-off” values for each independent variables and the LDA classification performance was evaluated using various statistical measures of Receiver Operating Characteristic curve (ROC curve) including sensitivity, specificity and accuracy 30,31 (Figure 1).
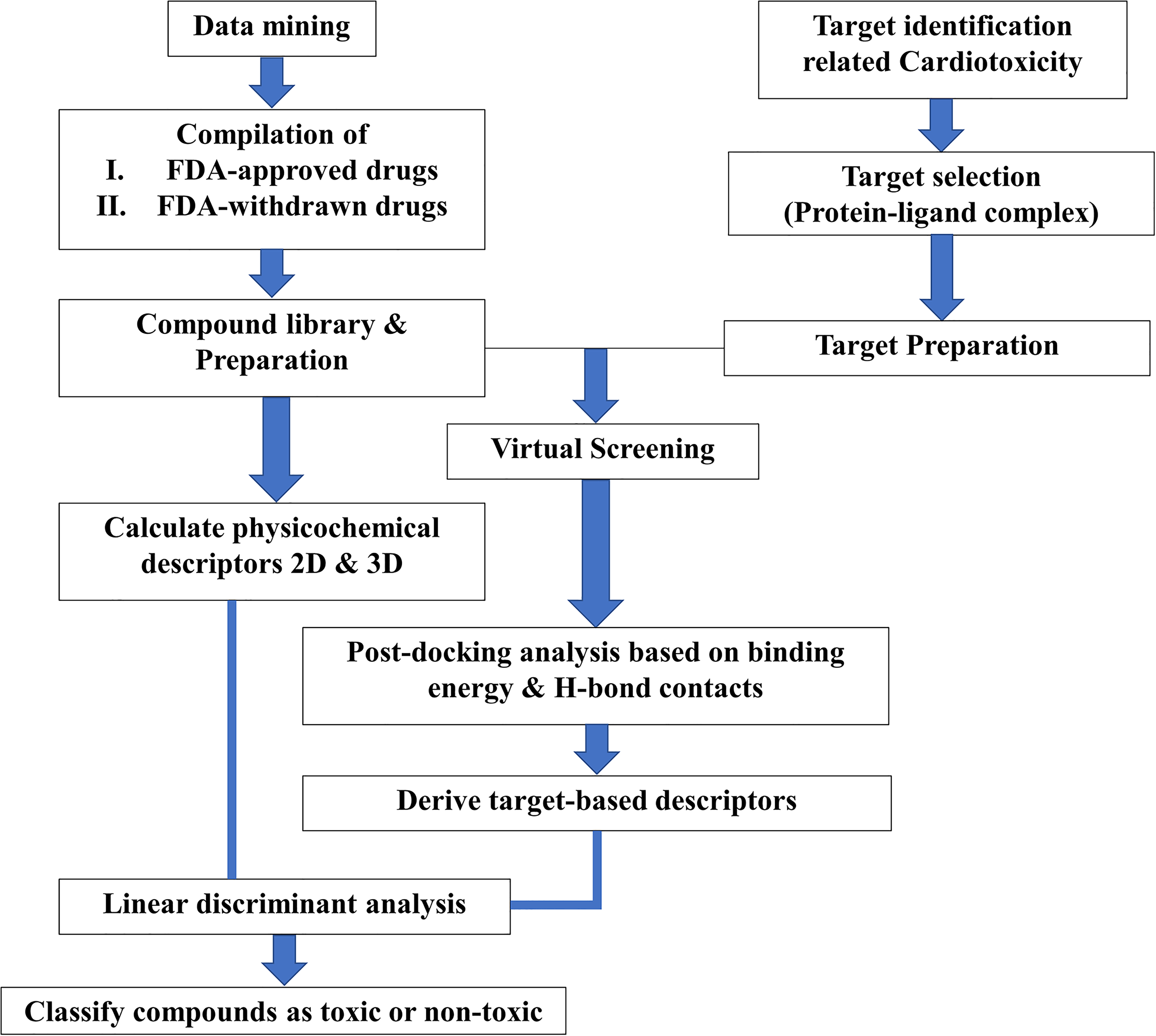
The flow chart of the proposed method combining ligand-centric and receptor-centric descriptors in LDA modeling.
Materials and methods
Compounds collection and descriptors calculations
We chose four different types of drug classes having implications in the cardiac system. These include antidepressants, antihistamines, antiarrhythmic and beta-blockers. A total of 91 drugs including approved and withdrawn composed of 72 non-cardiotoxic compounds (79%) and 19 toxic (21%) compounds. were compiled from different databases and resources such as Drugbank (https://www.drugbank.ca/), 32 Drugs.com (https://www.drugs.com/), 33 LiverTox (https://www.ncbi.nlm.nih.gov/books/NBK547852/), 34 etc (Table S1). The 3D structures of drugs were retrieved from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/) in structure data format (SDF) format. Subsequently, drug-likeness, Lipinski’s rule of five and physico-chemical properties were calculated using the drug-likeness tool (DrugLiTo) 35 and Osiris property explorer. 36 –41 Subsequently, we computed ADMET properties using ADMET Predictor (Simulation Plus Inc.) such as logP, topological surface area (TPSA Å2), ADMET_solubility, Perm_cornea, etc.
Virtual screening against selected protein targets
We selected protein targets by initially gathering the details regarding the primary targets of 91 drugs with evidence of pharmacological action and selected the most common four targets implicated in cardiotoxicity. For example, Timolol antagonizes the B1-adrenergic receptor. The targets were retrieved from Protein Data Bank (PDB) based on resolution and ligand-bound complexes: histamine H1 receptor (PDB ID: 3rze) 42 for antihistamines, cytochrome P450 2D6 (PDB ID: 3tbg) 43 for antidepressants and cytochrome P450 3A4 (PDB ID: 4d6z) 44 for antiarrhythmics. The homology model of the B1-adrenergic receptor for beta-blockers was developed using the Modeller program due to the unavailability of the crystal structure suitable for virtual screening. First, we retrieved the primary sequence of B1-adrenergic receptor from UniProtKB (P08588) 45 in Fasta format followed by template identification as a part of Modeller modeling protocol. 46 The best protein model was selected based on GA341, a composite score of compactness, statistical distance potential and sequence identity between target and template, and zDOPE score (z-statistic applied on Discrete Optimized Protein Energy), a distance-dependent potential native function of Modeller program. 47 The homology model was further evaluated for stereochemistry checks using Ramachandran plot. 48
All the four ligand-bound protein complexes (3 PDB protein-ligand complexes and 1 homology model with ligand-bound template) were prepared using YASARA Structure (academic license) 49 with AMBER03 force field. 50 Hydrogens were added and bond orders and hybridization states were assigned using Clean utility. 51 The ligand-bound site was defined as the docking site (simulation cell) for virtual screening using YASARA Structure. The simulation cell was placed by inspecting the centroid of the bound ligand and cell boundaries were defined in each direction by 8 Å distance. 25 We chose the YASARA Vina docking module for virtual screening in which Vina applies a stochastic-based dock pose search and implements the YASARA scoring function to score the docked pose in energy units (kcal/mol). According to YASARA conventions, positive energy score indicates high affinity in direct contrast to other known scoring functions where low negative energy values represent best binding pose. 52,53
Linear discriminant analysis of descriptors
The selection of descriptors was made by LDA in such a way to better classify cardiotoxic and non-toxic compounds and shed light on the mechanism of toxicity. The descriptors selection process was performed based on toxic and non-toxic compounds through linear discriminant function. 54 The discriminant function (DF) was defined as follows 54,55 :
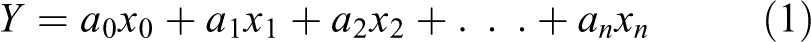
where
We utilized the rule of maximum likelihood picking up a DF with high importance while ensuring few descriptors to obtain a DF with statistical significance. Simultaneously, over-fitting as well as chance correlations among descriptors are avoided to create meaningful DF with better discriminating power. We assessed the following parameters for identifying the best DF: Wilk’s λ, Fisher degree (F) and the p-level (p) with 95% confidence limit and leave-one-out cross-validation (LOOCV). 56 –58 It was analyzed whether the subset of compound data has over the top effect over the created DF to examine over-fitting. We also examined the predicted classification in both classes (cardiotoxic and non-toxic). The LDA model consistency was also studied by evaluating the performance on test data (left-out data) upon selected DF which was not used for creating the DF.
Performance of the LDA model
To assess the performance of DF in better distinguishing between two classes of compounds, we utilized SPSS (Statistical Package for Social Sciences package) 59 statistical software to generate LDA classification models. After the model generation, ROC curve analysis was carried out to decipher cut-offs between two ROC parameters: sensitivity (true positives) and specificity (true negatives) values. 30,31,60 The choice of DF was made by assessing the area under the ROC curve (AUC) which is the widely used measure of discriminatory power of a diagnostic test 61 . We accepted a DF among various DFs which is superior in statistical significance (p < 0.01) 62 and Wilk’s λ with high sensitivity (most non-toxic compounds classified as non-toxic in LDA class prediction) and with best possible high specificity (few toxic compounds predicted as non-toxic exceptionally). In the regulative aspect of the study, the importance was given to the sensitivity rather than specificity which enabled the correct classification of non-toxic compounds while allowing few toxic compounds as non-toxic which slightly affect the accuracy of the LDA model. Various evaluative measures of ROC curve can be mathematically expressed 63 as follows:
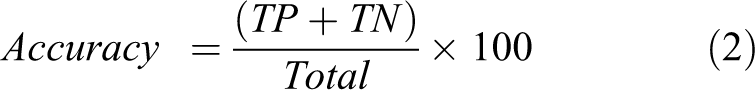
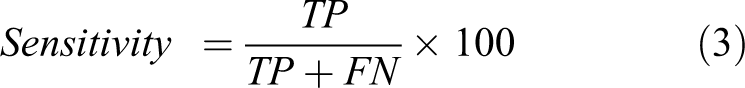
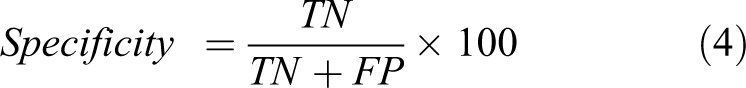
where TP = true positive, TN = true negative, FP = false positive, and FN = false negative.
Results and discussion
Target selection and virtual screening of compound set
We chose the most common targets of cardiovascular system to understand the interactions of compound sets viz. B1-adrenergic receptor, histamine H1 receptor, cytochrome P450 2D6 and cytochrome P450 3A4. The rationale behind selecting only four targets of cardiovascular system as the compound set used in the study possess binding capabilities to any one of the four targets. There are other interesting targets such as A2A adenosine receptor which have a myriad of clinical applications related to cardiovascular diseases. Further, the protein targets for the compound set was sought after the compilation of compound set which falls in four different classes, antidepressants, classified before high risk antihistamines, beta-blockers and antiarrhythmics. It will be desirable to build a comprehensive classification model by enhancing the compound set and target set.
Due to the unavailability of crystal structure of Human B1-adrenergic receptor, we modeled its structure using Modeller program. The template, Turkey (bird) B1-adrenergic receptor, PDB ID: 2y00 was chosen due to its best alignment with our query protein sequence: 1e-166 E value with 68% sequence identity and 73% sequence coverage (Figure S1). The template is complexed with dobutamine drug (a β1-agonist) which offered a reliable ligand-bound conformation to be modeled in query structure and subsequently used in virtual screening purposes. Moreover, the modeled protein obtained a GA341 score of 1.0 and zDOPE score of 1.03 indicating a reliable model for structure-based studies (Figure S2). Ramachandran plot showed 98% amino acid residues lie in favored region and 2% in allowed region without any outliers (Figure S3).
The remaining three targets were downloaded from PDB. The protein structures and compound set were prepared for virtual screening. The ligand-bound site in the structures were defined as simulation cell for docking with YASARA Structure (academic license). The binding energy of compound set (dock score; kcal/mol) with its respective protein target obtained from virtual screening is given in Table 1. The best binding pose of top five ranked compounds along with its crystal ligand across four targets is illustrated in Supplementary Figure S4 to S7. A variety of intermolecular interactions were observed including electrostatic bonds, hydrogen bonds, van der Waals and pi-sigma bonds, which boosted the dock score effectively. For example, Triprolidine secured a dock score of 10.808 kcal/mol (more positive means better affinity) with histamine H1 receptor. The aromatic moieties of Triprolidine is stacked between Tyr408 and Phe432 residues along with hydrophobic contacts by Asn19, Trp428, Phe435 and other pocket residues.
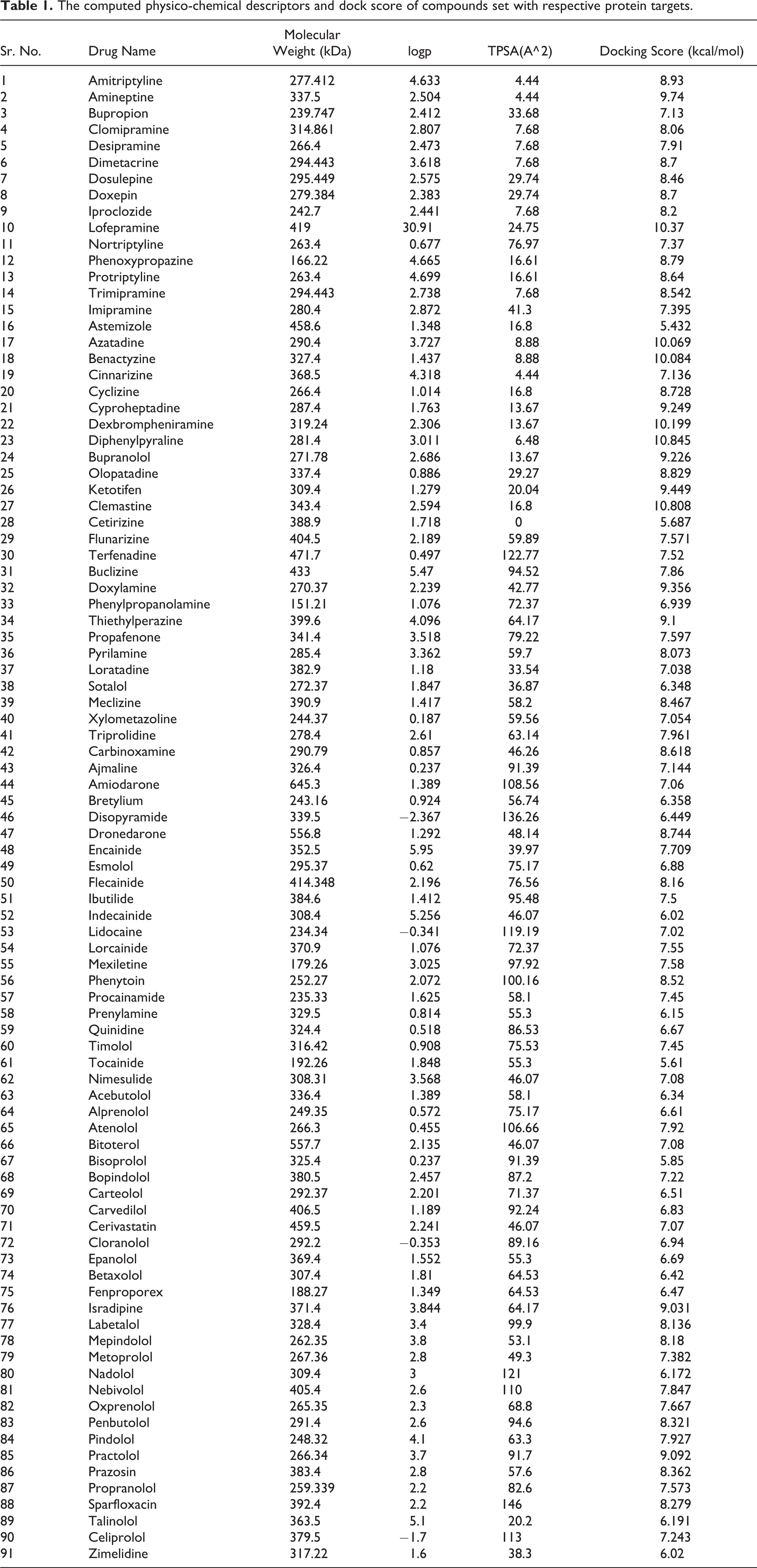
The computed physico-chemical descriptors and dock score of compounds set with respective protein targets.
The dock pose examination of top 5-ranked antidepressants against cytochrome P450 2D6 revealed that Lofepramine and Notriptyline exhibited a pi-pi stack with Phe120 residue similar to its co-crystal ligand, thioridazine derivative. In the case of histamine H1 receptor, doxepine, a compound in the set which is also the co-crystal ligand of target (PDB id: 3rze) secured a dock score of 8.4 kcal/mol with three pi-pi stacks established by residues Trp428, Phe432 and Phe435. Interestingly, all the top 5-ranked compounds had a score of ∼10.00 kcal/mol and created the three pi-pi stacks pattern of co-crystal ligand. The higher affinity of hits in contrast to co-crystal ligand is contributed to additional hydrophobic contacts.
For B1-adrenergic receptor, we first validated the ligand-bound pocket conformation of bird B1-adrenergic template modeled on the Human B1-adrenergic receptor by placing the template co-crystal ligand (dobutamine) in modeled one and re-docked using YASARA Vina. We obtained few best poses with root mean square deviation (RMSD) less than 2 A illustrating the reliability of docking protocol in generating crystal close conformations (results not shown). The best pose of top 5-ranked beta-blockers guided by high dock score did not generate common intramolecular interaction patterns. For example, bopindolol and esmolol established a pi-pi stack with Phe218 residue whereas prazosin produced a H bond with this residue. Although different types of contacts were developed, the dock score of top 5 beta-blockers secured less than 10 kcal/mol. Similar to B1-adrenergic receptor docking task, the top 5 antiarrhythmics obtained a dock score less than 10 kcal/mol. The pyridine-carbamate co-crystal ligand of cytochrome P450 3A4 (PDB id: 4d6z) produced pi-pi stacks with Phe108 and Phe304 residues. Although the same interaction pattern was not observed in the top 5-ranked compounds, the target pocket is enriched with phenylalanine residues enabling several pi-pi stack mode of interactions. The residues include Phe57, Phe108, Phe213, Phe215 and Phe304.
Descriptors calculations
We have used ligand-centric and receptor-centric descriptors for the development of the cardiotoxic LDA classification model. The former was calculated based on 2D and 3D geometry of ligands whereas the latter were taken from virtual screening of library molecules with four selected targets (protein) receptors. The Physico-chemical properties of all selected compounds were computed using Osiris property explorer and DrugLiTo programs. The computed 2D and 3D descriptors belong to major types of descriptors family including topology, fingerprints, fragment-based fingerprints and other properties such as clogP, molecular weight and topological surface area. The distribution of clogP values in the compound set was mostly in positive values (<5.0) with few negative values demonstrating most cardio system drugs are well absorbed and permeable. Also, calculations of fragment-based drug-likeness highlighted other important properties such as mutagenic, tumorigenic, irritant capacity and reproductive effect. The computed descriptors of physico-chemical types are listed in Table 1.
Nowadays, the prediction of toxicity risks can be performed parallelly and easily on the webservers. We used the ADMET predictor to calculate toxicity risk parameters such as ADMET_Solubility, ADMET_Solubility_Level, ADMET_AlogP98, CMR_Discriminant_Score, FRC_Discriminant_Score, among others. These properties can also be classified before high risk or low-risk compounds based on the distribution of different parameter values (Table S2). However, we chose to classify compound sets by supplying the computed descriptors (both Physico-chemical and ADMET properties) as input to LDA based on actual drug status (cardiotoxic or non-toxic) rather than interpreting the data distribution of different properties used to create descriptors by the software.
Development of cardiotoxicity LDA classification model
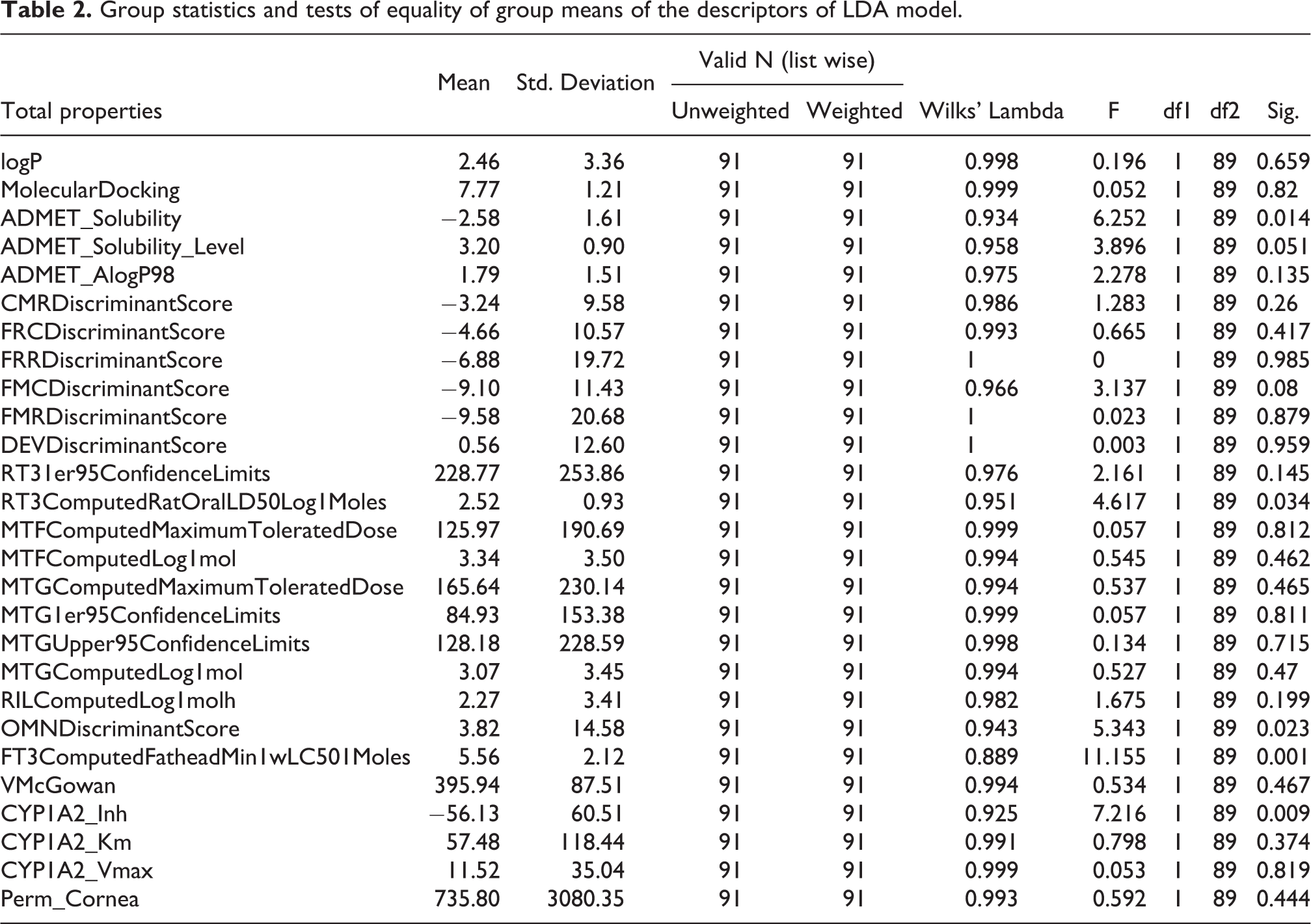
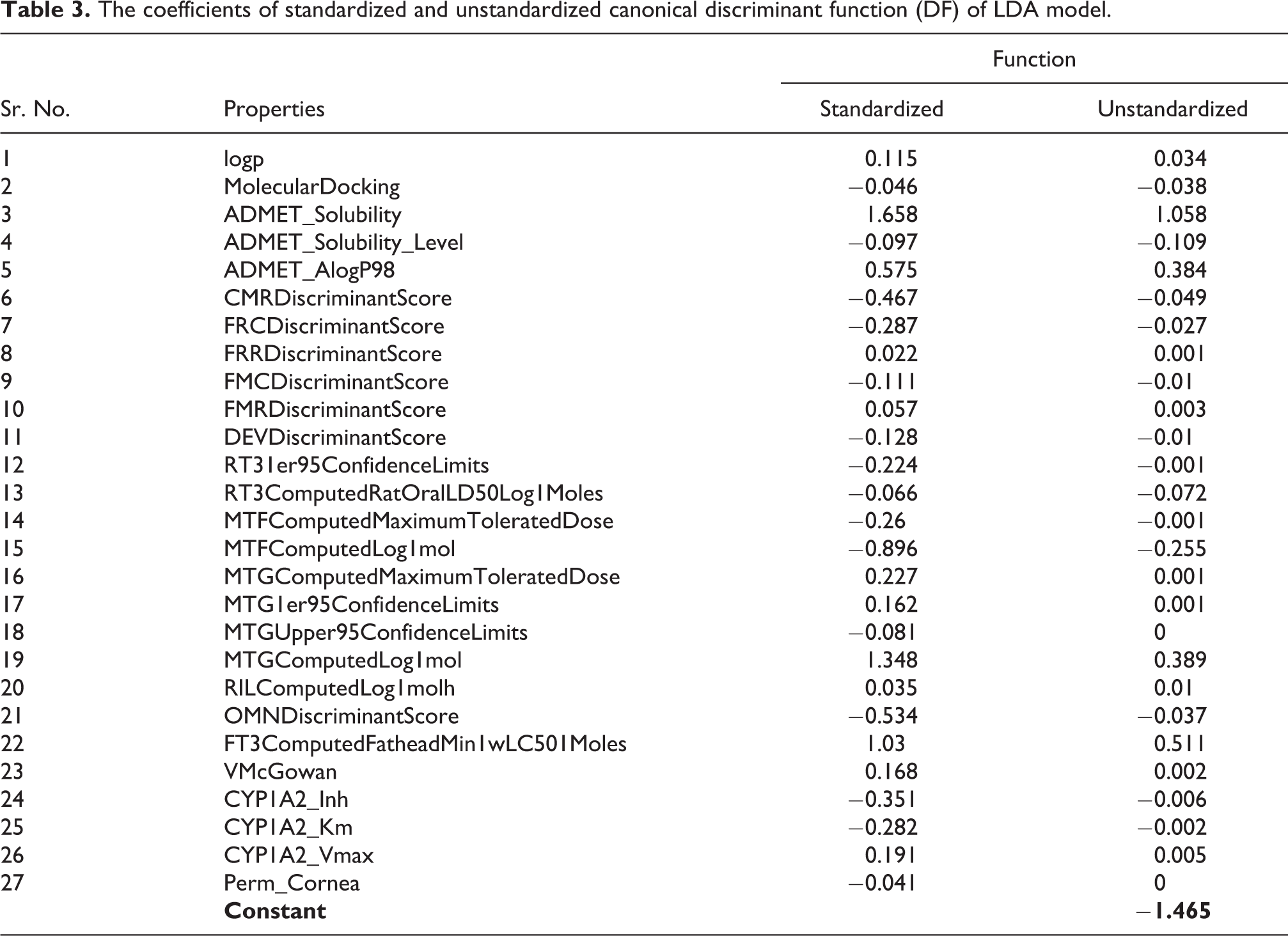
Similar to multiple linear regression, discriminant functions (DF) are generated by calculating differences between groups on each of the independent variables using group mean and one-way ANOVA results data. Table 2 lists the results of test of equality of group means. All the variables attained Wilk’s λ in the range of 0.889 to 1.000 demonstrating the entire 27 variables nearly have similar discriminating potential between groups. The standardized and unstandardized canonical DF coefficients for each variable is given in Table 3. Further, the pooled within-group matrices revealed the intercorrelations were low. The Eigen value of 1.094 accounting 100% variance showing the discriminating ability of DF function (function = 1) with a canonical correlation of 0.723 between variables and groupings (Table S3). We were able to generate one DF with 100% cumulative proportion which forbids the generation of other possible DFs. Hereafter, we discuss the DF function = 1 for its ability to distinguish cardiotoxic and non-toxic groups. Wilk’s λ multivariate statistic is 0.478 which is close to 0 highlighting strong discriminating power of DF (Table S4). Additionally, the p-value of 0.01 obtained from Chi-square statistic rejected the null hypothesis stating no discrimination between DF’s canonical correlation and all smaller canonical correlations.
Group statistics and tests of equality of group means of the descriptors of LDA model.
The coefficients of standardized and unstandardized canonical discriminant function (DF) of LDA model.

The regression equation of DF is expressed as:
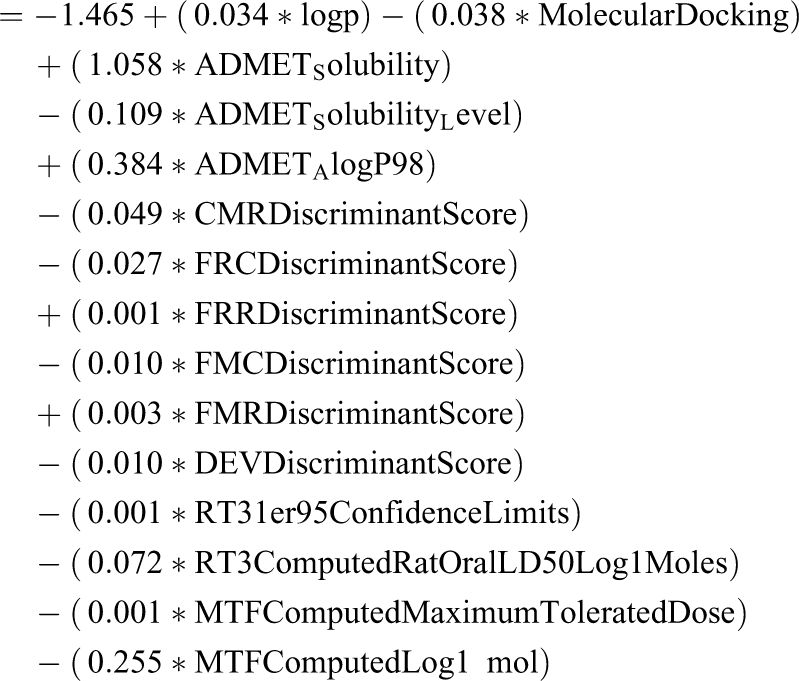

The group centroids table revealed that the difference between the means of the DF scores by non-cardiotoxic group (drug status = 1) and cardiotoxic group (drug status = 2) is −2.545 (Table S6) representing both the groups were satisfactory separated by DF and have high-discrimination ability. The DF of LDA model correctly classified 91.7% of non-toxic compounds and 84.2% of cardiotoxic compounds (Table 4). Cross-validation of hold-out cases revealed 73.6% grouped cases were correctly classified.
LDA classification of drug statuses of compound set.
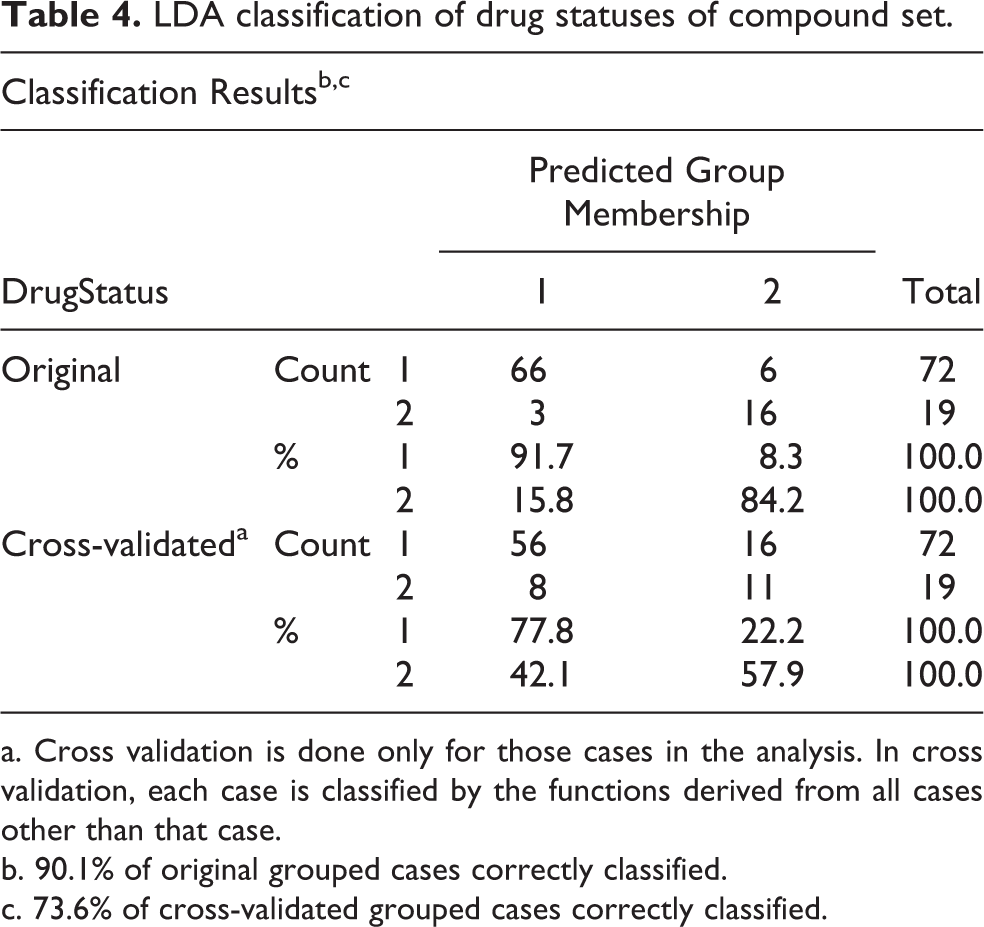
a. Cross validation is done only for those cases in the analysis. In cross validation, each case is classified by the functions derived from all cases other than that case.
b. 90.1% of original grouped cases correctly classified.
c. 73.6% of cross-validated grouped cases correctly classified.
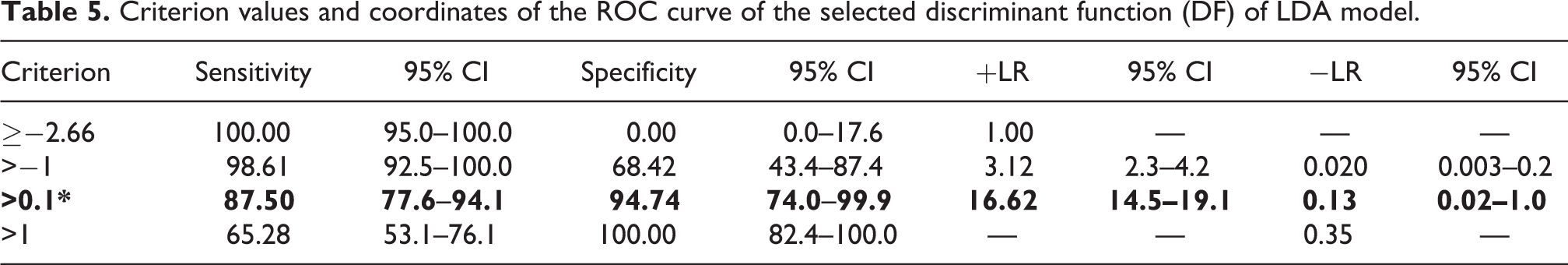
The cut-off points between two groups were predicted through the ROC curve. The DF calculated class along with the drug status (cardiotoxic/non-toxic) was supplied as input to ROC curve analysis. The true positive rate was defined as the correct identification of non-toxic compounds as non-toxic compounds by the LDA model based on the discriminant function (DF) values. Similarly, the true negative rate was defined as the correct identification of cardiotoxic compounds as cardiotoxic compounds (Table S6). We obtained the finest discriminatory ratio as area under the curve had attained 0.958 (Table S7) when the sensitivity reached 87.50% while specificity touched 94.74% in the ROC curve (cut-off value criterion: >0.1) (Table 5 and Figure 2). The positive likelihood rate (+LR) of 16.62 demonstrates very strong evidence in correctly classifying non-cardiotoxic compounds. According to this cut-off value, the greater class values showed more toxic while lesser value depicted non-toxic effects of any cardiotoxic drug from aforesaid targets (Table S8). We then explored whether there is any correlation between receptor-centric descriptor (Dock score) and ligand-centric descriptors (physico-chemical/ADMET properties). We found that the canonical correlation between Dock score and ADMET_AlogP (atom-based log P calculations) noticed in pooled-within groups table (not shown) was 0.461 while its counterpart ClogP (fragment-based log P calculations) secured 0.330. It was interesting to note both Dock score and AlogP are computed based on atom contributions which are additive in nature to come up with a single unique score. Both the descriptors provide positive values which are normally distributed and appears to provide apparent correlation.
Criterion values and coordinates of the ROC curve of the selected discriminant function (DF) of LDA model.
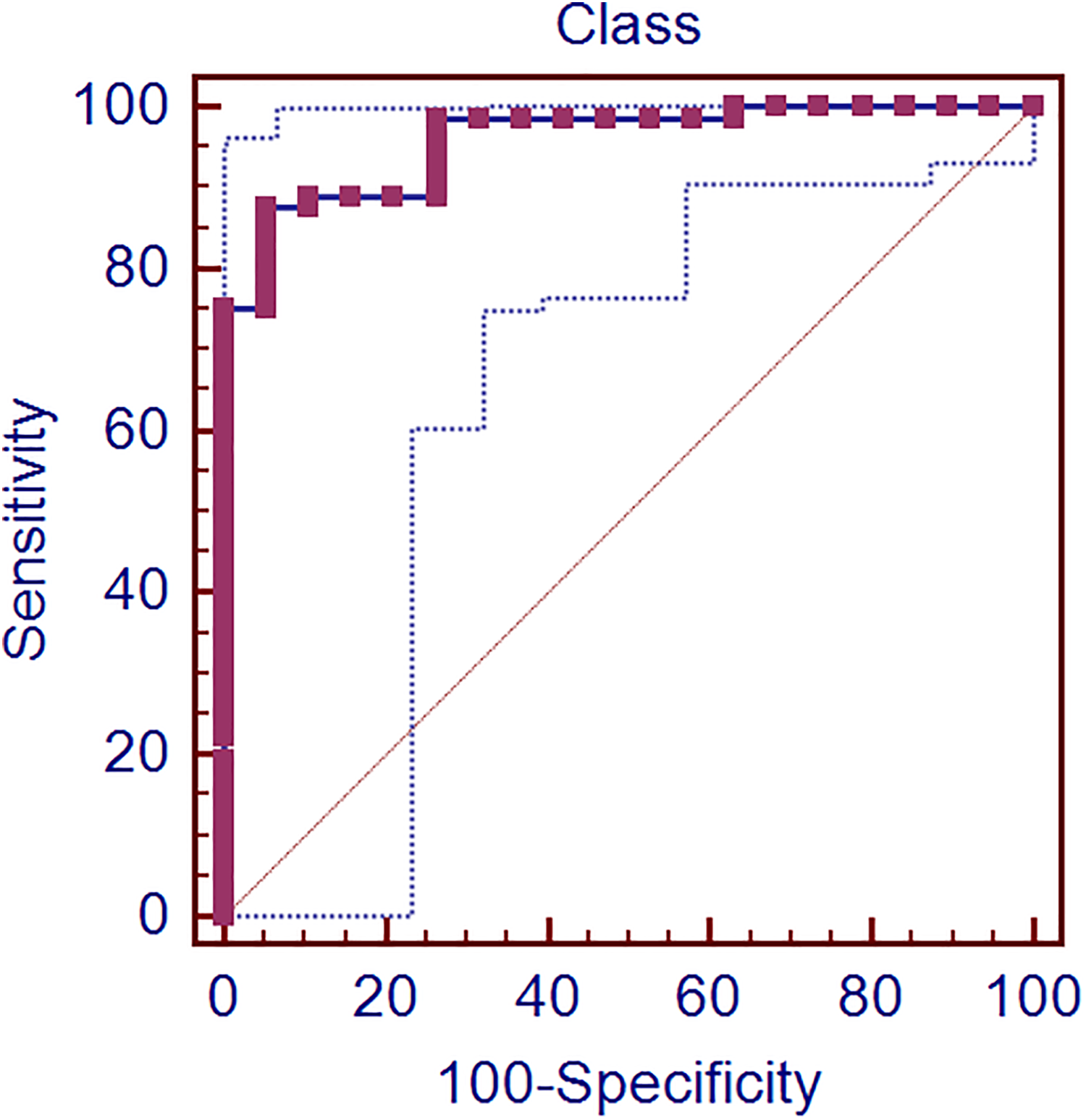
ROC curve analysis of the selected discriminant function (DF) of LDA model. The pink line represents the various data points related to sensitivity and specificity of the DF. The blue dotted lines indicate the cut-offs associated with each data point of the selected DF model. The straight diagonal line extending from the lower left corner to the upper right (dotted line) represents the no performance boundary.
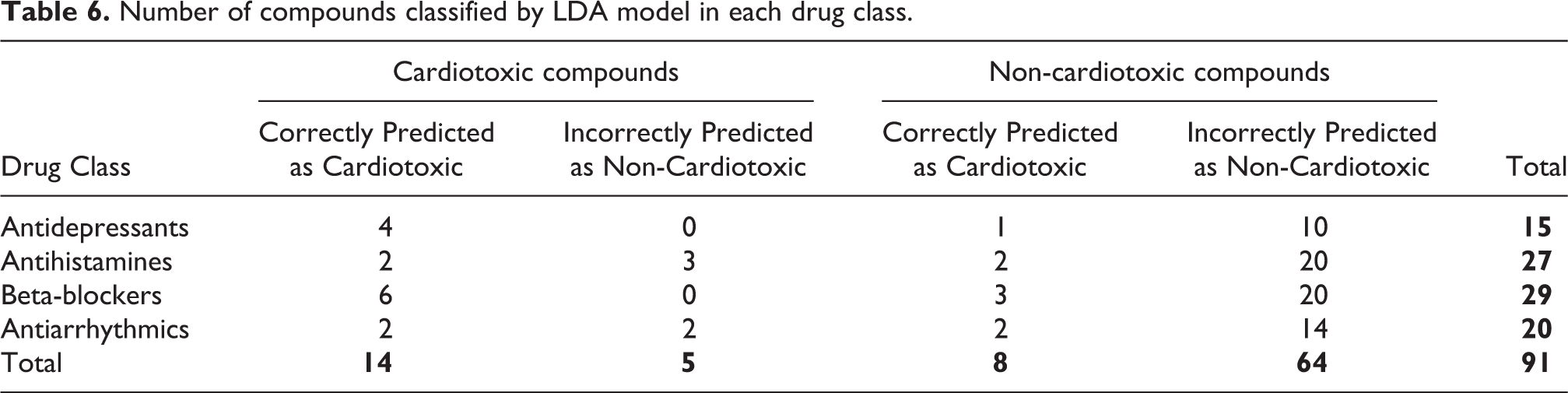
We further examined how the selected LDA model performed in each drug classes. The classification upon antidepressants class achieved best performance in correctly classifying correct drug status (10 non-toxic compounds predicted as non-toxic and all toxic compounds as toxic). In the case of antihistamines, 20 non-cardiotoxic compounds are correctly identified as non-toxic boosting the sensitivity. Only 2 cardiotoxic antihistamines were correctly predicted as compounds with cardiotoxic potentials. Similar to antidepressants, 20 non-cardiotoxic beta-blockers were predicted as non-cardiotoxic whereas all cardiotoxic compounds were correctly identified as cardiotoxic. The LDA model correctly predicted 14 non-toxic antiarrhythmics while sub-standard prediction was made in classifying cardiotoxic compounds as toxic (2 compounds) and as non-toxic (2 compounds). Collectively, the presented LDA model achieved better performance in the sensitivity across four drug classes (Table 6). The LDA model will be available upon request.
Number of compounds classified by LDA model in each drug class.
Conclusion
The presented LDA model strongly advocates the incorporation of both receptor-centric and ligand-centric descriptors in modeling cardiotoxicity. We obtained 90.1% correct classification of actual drug status and 73.6% correct classification in left-out samples during cross-validated. The ROC analysis showed AUC of 0.958 with best 87.50% sensitivity and 94.74% specificity. This good statistic upholds the need to extend this approach to multiple drug classes and its respective protein targets in LDA modeling. We found that both physico-chemical and ADMET properties contribute equally to discriminant function of LDA due to close values of Wilk’s lambda metric. Interestingly, Dock score of compound set achieved 46% correlation with AlogP property. The YASARA Vina scoring function partially distinguished dock poses of compounds with no common intermolecular interaction pattern with low scores, especially in the cases of beta-blockers and antiarrhythmics. The LDA model can be made more comprehensive by introducing more drug classes and targets and subsequently validate through
Supplemental material
6-10-20-Suppliment - Development of cardiotoxicity model using ligand-centric and receptor-centric descriptors
6-10-20-Suppliment for Development of cardiotoxicity model using ligand-centric and receptor-centric descriptors by Chirag N Patel, Sivakumar Prasanth Kumar, Rakesh M Rawal, Manishkumar B Thaker and Himanshu A Pandya in Toxicology Research and Application
Footnotes
Acknowledgments
The authors gratefully acknowledge the Department of Botany, Bioinformatics and Climate Change Impacts Management, Gujarat University for providing an opportunity to access the bioinformatics research facilities.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was financially supported by the Financial Assistance Programme – Gujarat State Biotechnology Mission, Gujarat, India [grant number GSBTM/FAP/1443] and Department of Science and Technology [grant number GSBTM/MD/JDR/1409/2017-18].
Supplemental material
Supplemental material for this article is available online.
Abbreviations
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
