Abstract
Alterations in mRNA splicing play a critical role in driving the molecular heterogeneity of many cancers, including urothelial carcinoma, by contributing to disease progression, treatment response, and clinical outcomes. These splicing changes can arise from somatic mutations in core spliceosomal components or through alternative splicing events affecting cancer-associated genes. In this review, we examine how dysregulation of pre-mRNA splicing influences key aspects of urothelial carcinoma biology, including cell proliferation, invasion, metastasis, modulation of the immune microenvironment, metabolism, and therapeutic resistance. We highlight frequently observed splicing-factor mutations and discuss the impact of aberrant splicing and cancer-specific isoforms on prognosis. We also explore splicing alterations associated with susceptibility to urothelial carcinoma and review emerging therapeutic strategies, such as splice-switching oligonucleotides and small molecule spliceosome inhibitors, that offer promising avenues for precision medicine in this disease.
Keywords
Introduction
RNA sequencing of transcriptomes has broadened our understanding of the complexity of the pre-mRNA splicing process and its role in generating functionally diverse isoforms. Transcribed RNA must be processed before it becomes a mature messenger RNA (mRNA) that can direct the synthesis of protein. One key step in this processing is RNA splicing, which involves removing non-coding introns from pre-mRNA and joining exons, forming mature mRNA. If the introns are not removed, the RNA would be translated into a nonfunctional protein. Splicing occurs in the nucleus before the RNA migrates to the cytoplasm. Pre-mRNA splicing is regulated by a dynamic RNA-protein macromolecular complex composed of small nuclear ribonucleoproteins (snRNPs) and protein cofactors commonly referred to as the spliceosome (Figure 1A). Aberrant mRNA splicing is increasingly recognized as a hallmark of cancer, playing a critical role in disease progression and treatment resistance.
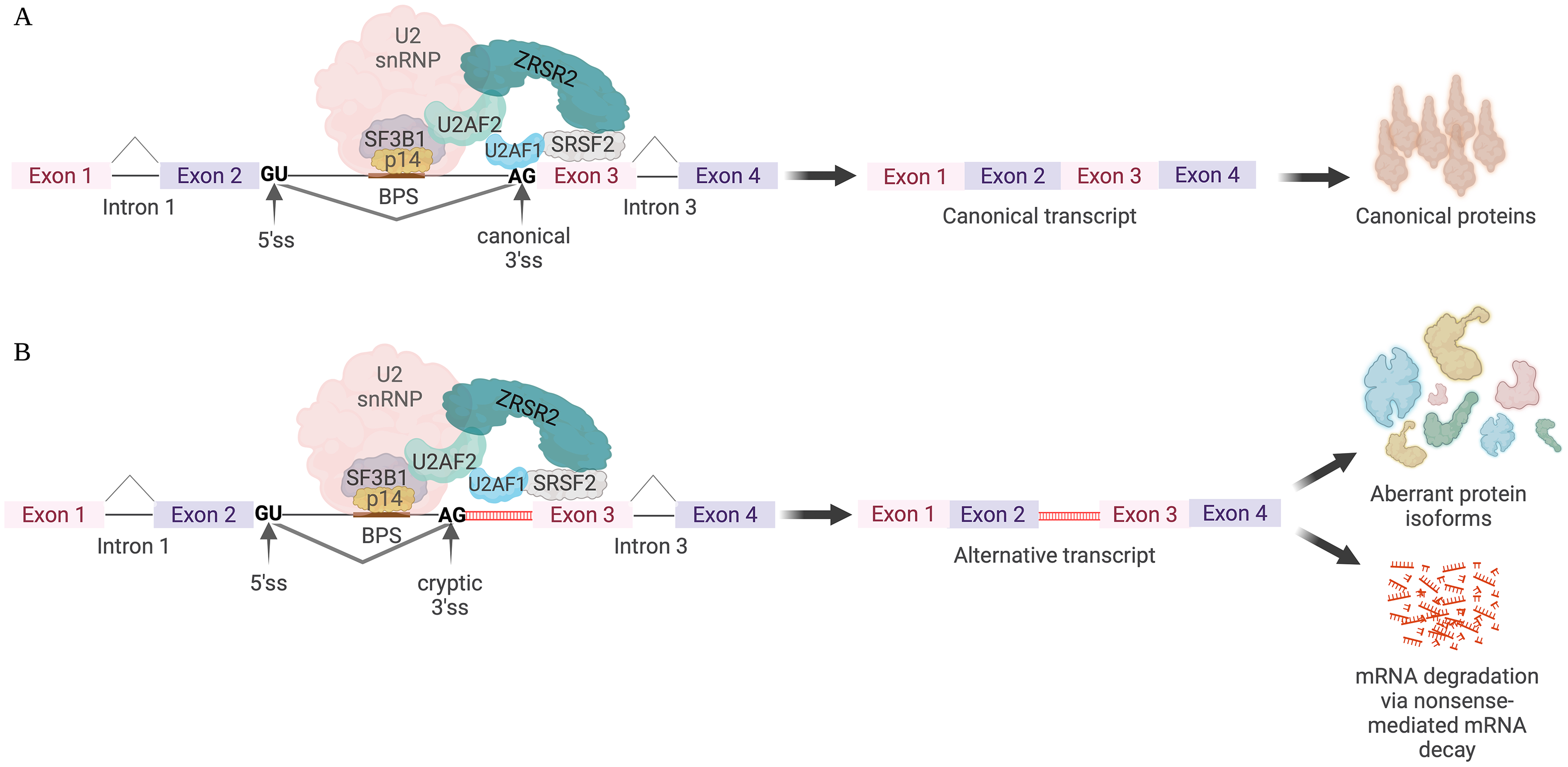
Normal (A) and aberrant (B) pre-mRNA splicing events mediated by the spliceosome.
Urothelial carcinoma is one of the most common cancers in the United States. 1 It is a heterogeneous disease that can be categorized as non-muscle-invasive cancer of the bladder and upper tract, muscle-invasive cancer of the bladder and upper tract, and advanced or metastatic disease. Treatment options and outcomes depend on the severity of the disease; however, many patients with advanced muscle-invasive or metastatic urothelial carcinoma progress with limited therapeutic options. Emerging evidence indicates that impaired mRNA splicing drives the molecular heterogeneity of urothelial carcinoma, contributing to disease progression, treatment responses, and clinical prognoses. Splice variants have also shown promise as potential therapeutic targets and predictive biomarkers. In this review, we summarize mRNA splicing alterations found in urothelial carcinoma and highlight their clinical and therapeutic implications for this disease, with an emphasis on potential applications in precision oncology and immunotherapy.
mRNA splicing alterations in urothelial carcinoma
Somatic alterations in the splicing machinery proteins in urothelial carcinoma
Pre-mRNA splicing is an intricate multiple-step process that is catalyzed by the spliceosome. Primary transcripts have introns removed by cleavage at conserved sequences called splice sites (ss). These sites are found at the 5′ (5′ss) and 3′ (3′ss) ends of introns and the branch site (Figure 1). The 5′ss is identified by the presence of a GU dinucleotide at the intron's 5′ end. The branch site typically has adenosine nucleotide; the 3′ss is composed of the polypyrimidine tract, a region that promotes the assembly of the spliceosome, and the AG dinucleotide at the intron's 3′ end. These consensus sequences are critical because altering one of the conserved nucleotides can lead to inhibition or missplicing. The spliceosome is composed of snRNPs and other protein factors. The major snRNPs are U1, U2, U4, U5, and U6; core splicing factors include SF1, U2AF, and SF3B1. The other major auxiliary classes of proteins that modulate splicing and selection of exons and introns include serine-arginine-rich splicing factor proteins (SRSFs) and heterogeneous nuclear ribonucleoproteins (hnRNPs). SRSFs and hnRNPs modulate splice-site selection through interactions with exonic and intronic splicing enhancer/silencer elements. All components of splicing machinery are essential in ensuring accurate and efficient removal of introns and the correct ligation of exons to generate diverse mRNA molecules with distinct cellular functions. The aberrant splicing can be caused by defects in both cis-/trans-factors of pre-mRNA splicing and contribute to pathogenesis (Figure 1B).
Alternative splicing can also occur when exons from the same gene are joined in different combinations, resulting in distinct but often related mRNA transcripts that may encode proteins with different structures and functions. Several modes of splicing exist, including (1) constitutive splicing and alternative splicing events such as (2) mutually exclusive exons, (3) retained introns, (4) cassette exon skipping, and (5) alternative 3′ and (6) 5′ splice-site recognition (Figure 2).
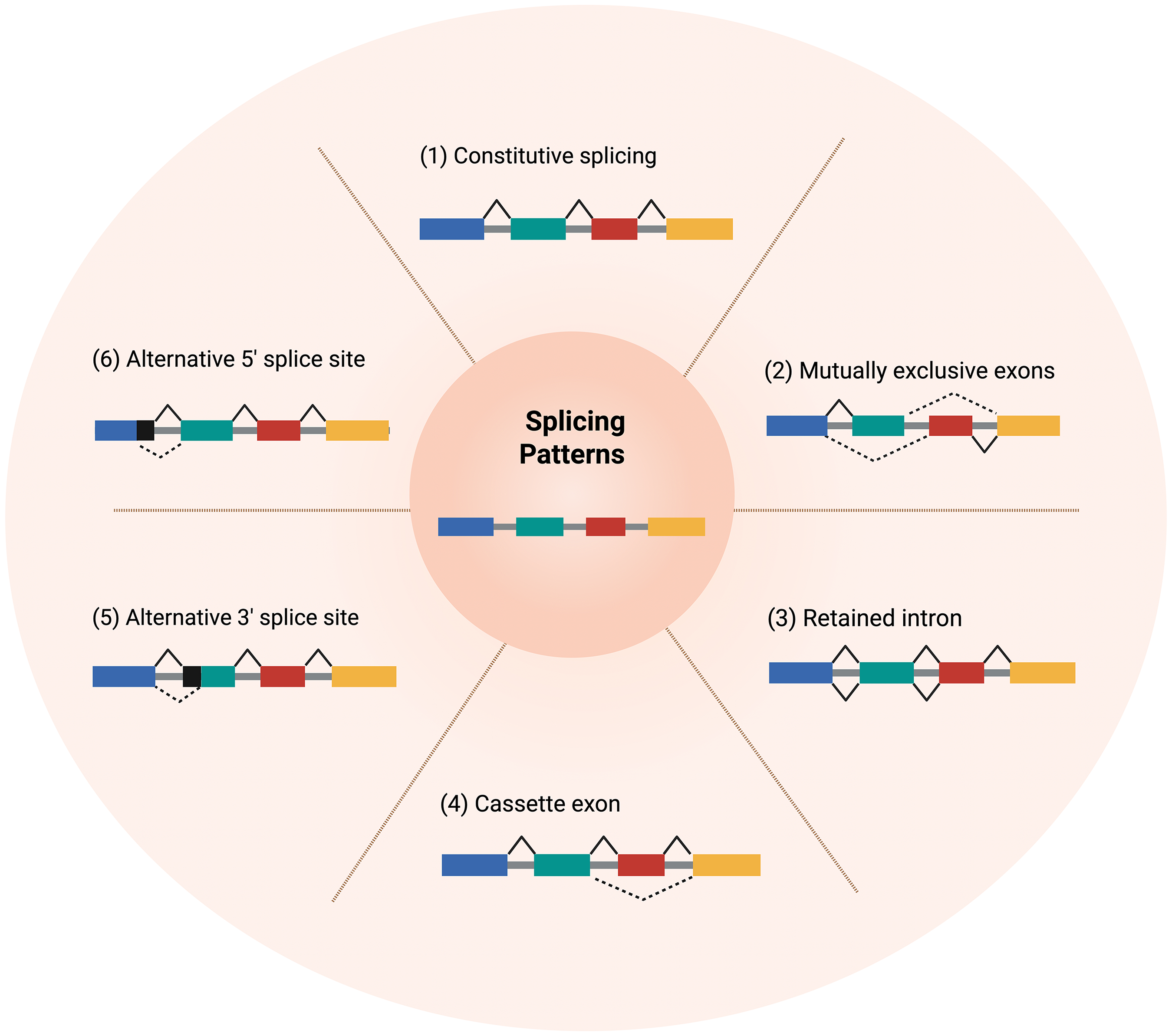
Overview of different splicing patterns. Constitutive splicing and the different types of alternative splicing: mutually exclusive exons, retained intron, cassette exon, alternative 3′ splice site, and alternative 5′ splice site. Exons are shown as boxes, introns as grey lines, and alternative splicing sites are indicated with dashed lines.
Although less common in solid tumors, somatic mutations in splicing factor-encoding genes are increasingly being recognized in cancer and were recurrently observed in approximately 13% of urothelial carcinoma cases based on TCGA data. 2 These heterozygous missense gain or alteration of function mutations are often mutually exclusive and recur at specific hotspots as implicated in various cancers.3–6 Splicing factor 3B subunit 1A (SF3B1) has the highest rate of alteration in all splicing-factor proteins found in urothelial carcinoma, which can lead to aberrant splicing. SF3B1 is a core component of the U2 snRNP, an important complex in spliceosome assembly and for branch-site recognition. The most common SF3B1 variant identified in urothelial carcinoma is E902 (Figure 3A).5,7 Although characterization of this variant is still lacking, it is known that missense mutations in the SF3B1 gene can lead to aberrant cryptic 3′ splice-site usage, leading to aberrant novel isoform or protein expression and/or nonsense-mediated mRNA decay affecting the proper function of several genes.8–10 Recent crystal structure studies of SF3B1 mutant proteins revealed that these alterations affect HEAT repeat domain conformations affecting SF3B1 interaction with other spliceosome proteins. 11 Additionally, alterations in the SRSF family proteins (SRSF1, SRSF2, and SRSF3), U2AF1, RNA-binding motif proteins (RBM 3, 4, 5, 6 and 10), heterogeneous nuclear RNPs (hnRNP A1, A3, A2/B1, F, L, K, and X), and Drosophila behavior/human splicing (DBHS) protein, NONO, have also been reported in urothelial carcinoma (Figure 3B–D).12–18 U2AF1 alterations were reported to cause exon inclusions with C- and G-rich 3′ss, while SRSF2 is known to cause genome-wide splicing alterations and aberrant mutation of different hnRNPs.19–21 The detailed mechanism and pathogenesis of these splicing-factor variants and the subsequent misspliced genes have not yet been described in urothelial carcinoma.
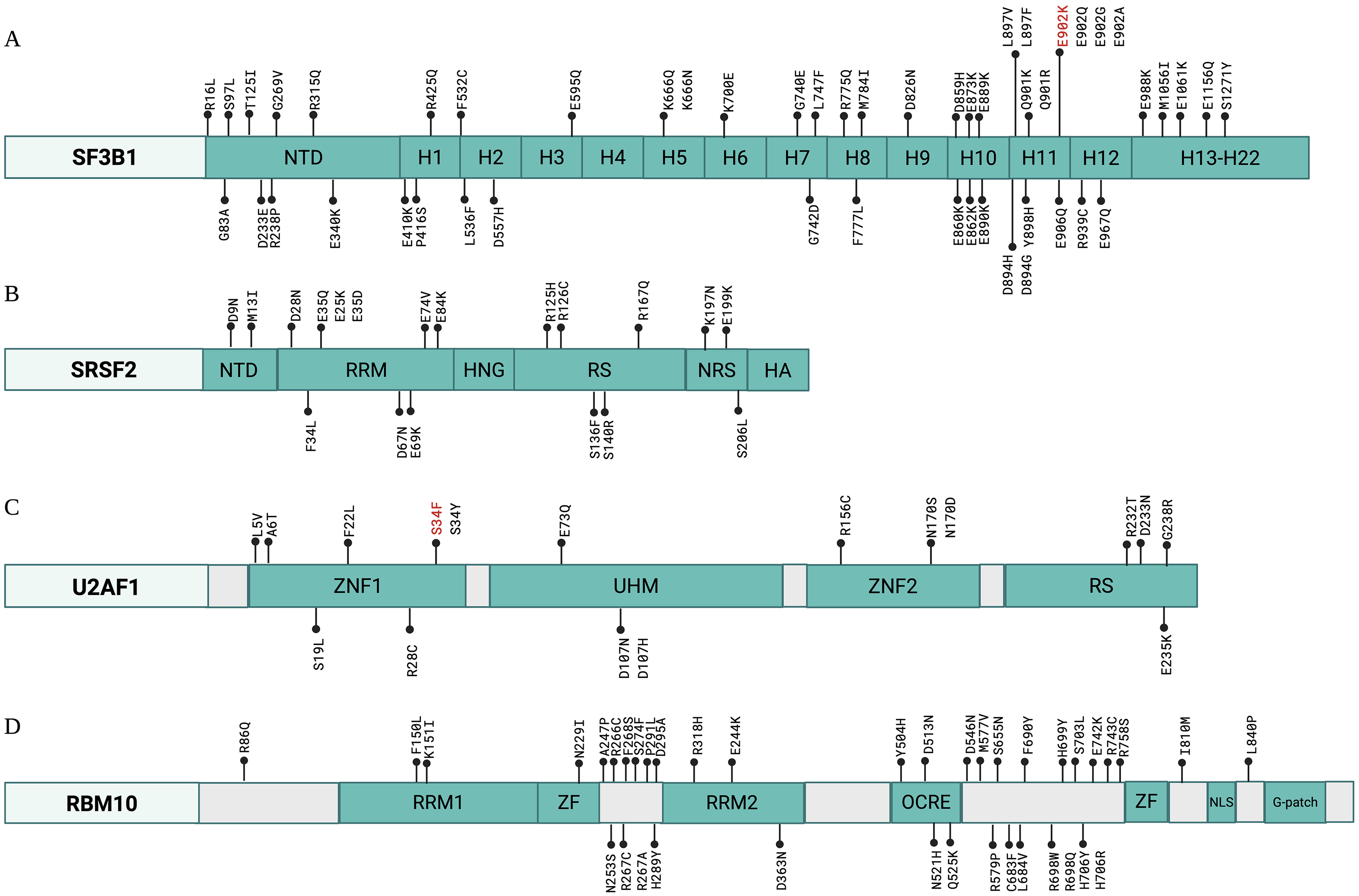
Comprehensive somatic mutation map found in spliceosome proteins reported predominantly in urothelial carcinoma. Somatic mutations found in splicing factors such as SF3B1 (A), SRSF2 (B), U2AF1 (C), and RBM10 (D) from TCGA data visualized and accessed via cBioPortal.
Aberrant alternative splicing in cancer-associated genes
While splicing factors can act as oncogenes in cancer by causing transcriptome-wide changes, genetic alterations in non-splicing factor genes—particularly known oncogenes and tumor suppressors—can also impact their isoform outcomes due to aberrant RNA splicing. These mutations often occur in splicing cis-elements, such as splice sites, splicing enhancers, and silencers. Aberrant splicing can drive carcinogenesis through the gain of oncogenic isoforms or the loss of tumor-suppressor isoforms. Such alterations resulting in missplicing in cancer-associated genes have been widely reported in urothelial carcinoma.
Oncogenes
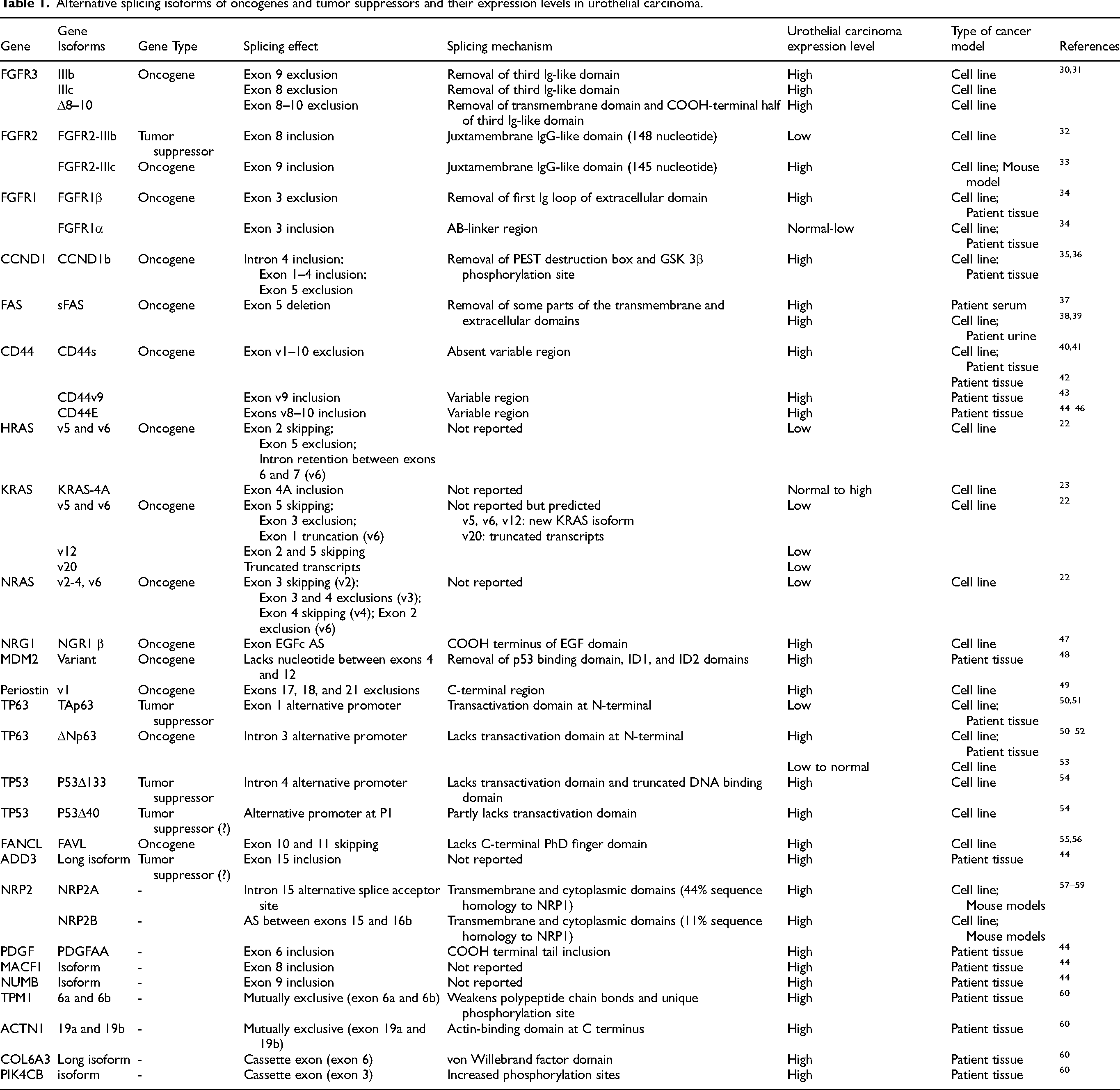
Proto-oncogenes are normal genes that regulate many vital biological processes, including cell growth, division, and differentiation. Mutated proto-oncogenes can become oncogenes that drive tumorigenesis by promoting cell survival and inhibiting apoptosis in various cancers. In urothelial carcinoma, major oncogenes such as the FGFR family and RAS family (HRAS, NRAS, and KRAS) undergo aberrant splicing, contributing to tumor development and progression (Table 1).22–29 Alternative splicing of HRAS intron D and splice-site mutations at the HRAS exon 5 lead to HRAS overexpression, while HRAS exon 2 skipping results in HRAS inactivation.61,62 Mutually exclusive splicing alterations in the HRAS, substitutions at exon 2, and small deletions at exon 5 are associated with young onset urothelial carcinoma. 63 Several other oncogenes also show isoform-specific overexpression in bladder tumors, including CD44, MDM2, FAS, NRG1, periostin, FANCL, and CCND1.35,39–41,43,44,46–49,56,64–67 A tumor-specific, O-glycoform spliced isoform of CD44, known as CD44s-sTn/Tn, is particularly associated with muscle-invasive urothelial carcinoma of the bladder. 40
Alternative splicing isoforms of oncogenes and tumor suppressors and their expression levels in urothelial carcinoma.
Tumor suppressors
Tumor-suppressor genes normally function to prevent tumor development by regulating cell growth and division; however, their aberrant splicing can lead to the loss of tumor-suppressive activity. In urothelial carcinoma, aberrant splicing of tumor-suppressor genes such as TP53, which encodes the p53 protein, has been reported. For example, dysregulation of TP53 family isoforms (p53, p63, and p73) has been shown to drive tumorigenesis and influence therapy responses across various cancer types, including urothelial carcinoma.48,50,54,68–73 Impaired alternative splicing of STAG2 and FGFR2 tumor spuppresor variant IIIb has also been reported in urothelial carcinoma.32,44,74–76
Consequences of RNA splicing alterations in urothelial carcinoma
Pre-mRNA splicing alterations can affect multiple key aspects of urothelial carcinoma biology such as control of cell proliferation, metastasis and invasion, immune microenvironment modulation, metabolism, and therapeutic resistance (Figure 4).
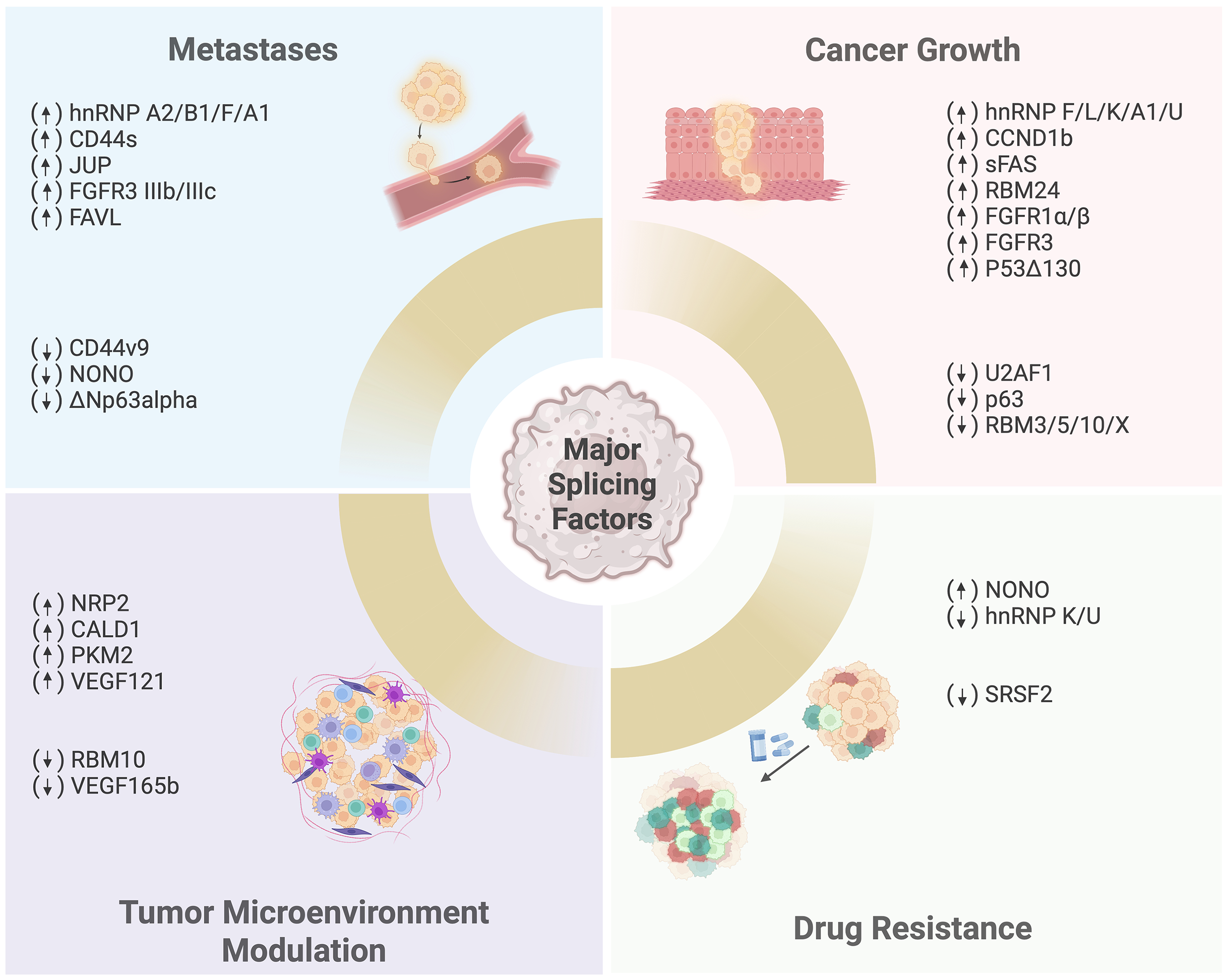
Major splicing factors reported to contribute to urothelial carcinoma oncogenesis. Dysregulation of expression of splicing factors and isoforms of oncogenes/tumor suppressors known to contribute to cancer growth, metastasis, tumor microenvironment modulation, and therapeutic resistance in urothelial carcinoma.
Cancer growth
Urothelial carcinoma tumorigenesis and proliferation have been associated with differential expression of multiple splicing factors and cancer-associated gene isoforms. For instance, downregulation of splicing factors RBM5 and RBM10 were reported to cause tumor growth via apoptotic inhibition through β-catenin activation and miR-432-5p overexpression.77–81 Similarly, RBM24 overexpression enhanced bladder tumor proliferation by stabilizing RUNX1T1 mRNA and inhibiting miR-625-5p transcription. 82 HnRNPs were also reported to drive urothelial carcinoma proliferation. HnRNP-F regulates the expression of xenopus kinesin-like protein, while hnRNPs L and K suppresses apoptosis-related protein caspases and cyclin-D1 expression and upregulation of SOX2 mRNA, contributing to tumor growth and aggressiveness.83–88 Cancer-associated gene isoforms such as p63, CCND1b, P53Δ130, FGFR1, and FGFR3 have also been identified in urothelial carcinoma as having pro-tumorigenic properties.24,54,89–94 Constitutive expression of FGFR3 spliced variants caused by splicing factor hnRNPL and several microRNAs, including miR-99a, miR-100, and miR-133, enhances tumor-proliferative characteristics.95–97 Furthermore, ectopic expression of either FGFR1β or FGFR1α isoforms induces activation of MAPK and other signaling pathways in telomerase-immortalized human urothelial cells, leading to increased tumor proliferation. 34 Similarly, cancer-associated gene isoforms such as sFAS and HRAS have been shown to reduce apoptosis. 98
Not all mutations in splicing factors and cancer-associated isoforms lead to increased tumorigenesis. U2AF1 variants, and RBM3/RBMX overexpression, have been reported to reduce cell proliferation, attenuate tumor growth, and enhance apoptosis.99–103 Moreover, hnRNPA1 (PTBP1) and hnRNPU depletion can cause growth retardation and cell death in urothelial carcinoma cells.15,104,105
Metastasis and invasion
Muscle invasion and metastasis pose a challenge in cancer treatment as cancer cells increase their interaction with the microenvironment, fueling epithelial-mesenchymal transition. In urothelial carcinoma, hnRNP overexpression is linked to enhanced mesenchymal marker expression with a concomitant decrease in epithelial markers. 85 HnRNPF was reported to be essential in urothelial carcinoma epithelial-mesenchymal transition where it stabilizes its downstream target, Snail-1 mRNA, by directly binding to the 3′UTR. 106 Similarly, hnRNPA2/B1 and hnRNPA1 were linked to tumor invasion and metastasis through a VEGF-C-independent manner and HIF-1, respectively.107,108 Splice variants such as FGFR2 IIIb/IIIc, FGFR3 IIIb/IIIc, CD44 s, JUP, CD44v9, and ΔNp63alpha were also shown to promote urothelial carcinoma invasion.40,41,43,53,109–111 FGFR3 IIIb and IIIc isoforms induce aberrant autocrine and paracrine signaling upon binding to different FGF ligands.30,31,33,112,113 Loss of ΔNp63alpha promotes tumor progression through upregulation of N-cadherin, with subsequent activation of the extracellular signal-regulated kinase pathway. 53 Similarly, FAVL has been linked to urothelial carcinoma oncogenesis, chromosomal instability, invasiveness, and therapeutic resistance, especially to cisplatin.55,56,64,114 In contrast, NONO was reported to inhibit cell migration by its potential interaction with SFPQ, thereby regulating the splicing SET domain and mariner transposase fusion gene. 18
Tumor microenvironment modulation
Protumor modulation of the tumor microenvironment is increasingly recognized as a critical driver of oncogenesis in urothelial carcinoma. Loss of function mutation in RBM10 has been shown to exert immunosuppressive effects by reducing immune-cell infiltration, specifically through the disruption of RMB10-mediated transcriptome-wide cassette exon repression. 7 Overexpression of neuropilin-2, a VEGF-C receptor, has been negatively correlated with the expression of immune checkpoint gene sialic acid-binding Ig-like lectin 15, leading to CD4+ T-cell central memory depletion in urothelial carcinoma.115–117 CALD1, a gene expressed in cancer-associated fibroblasts, has been implicated in promoting urothelial carcinoma progression by remodeling the microenvironment. 118 Alternative splicing of CALD1 generates its long isoform via an alternative 5′ splicing event involving exons 5 and 6. This long CALD1 isoform has been reported to be reduced or absent in urothelial carcinoma tumors, although its functional significance remains unclear. 60
Tumor metabolism and angiogenesis
Recent studies have highlighted the role of mRNA splicing alterations in metabolic reprogramming and the promotion of angiogenesis in urothelial carcinoma. SRSF1 regulates alternative splicing of HIF1A through exon 14 inclusion, promoting HIF1A/BNIP3/mitophagy axis that drives metabolic adaptation, angiogenesis, and chemoresistance in urothelial cancer cells.119,120 PKM, a cancer cell metabolism gene, undergoes alternative splicing to produce PKM1 and PKM2 isoforms through selective inclusion of exon 9 or 10, regulated by hnRNPA1/A2/PTBP1 (exon 9 suppression) and SRSF3 (exon 10 promotion).103,121–123 PKM1 in stromal cells drives glycolysis and lactate production, fueling adjacent cancer cells via a reverse Warburg effect.124,125 PKM2 predominates in urothelial carcinoma where its dimeric form promotes the Warburg effect by diverting glycolytic flux toward anabolic pathways essential for proliferation.15,126 Additional metabolic control occurs through PCBP2, which affects serine metabolism and ferroptosis via interactions with phosphoglycerate dehydrogenase. 127
VEGF-A is key angiogenic regulator that drives tumor progression through alternative splicing of exon 8. The shorter, diffusible VEGF121 isoform (proximal splice site) is upregulated in muscle-invasive urothelial carcinoma, whereas downregulation of anti-angiogenic VEGF165b isoform (distal splice site) promotes aggressive tumor angiogenesis.128,129 Splicing factors such as SRSF1, SRSF6 and SRPK1 can specifically control this isoform balance. 130
Therapeutic resistance
Splicing factors also contribute to drug resistance in many cancers by generating protein isoforms that evade the effects of targeted therapies and promoting tumor survival mechanisms. For example, SRSF2 has been correlated with cisplatin, pirarubicin, paclitaxel, doxorubicin, and epirubicin hydrochloride resistance both in vitro and in vivo.12–14 The SRSF2 resistance mechanism is thought to act via activation of Notch and oxidative stress pathways. NONO has also been recognized as an SPHK1-induced cisplatin resistance regulator through activation of STAT3. 131 Similarly, reduced expression of hnRNPU and hnRNPK has been correlated with increased cisplatin sensitivity. 104 HnRNPK-induced chemoresistance is mediated through the regulation of ERCC1 and ERCC4. 86
Alternative splicing reprogramming by non-coding RNAs
Non-coding RNAs orchestrate alternative splicing in urothelial carcinoma through complex regulatory networks involving circular RNAs (circRNAs) and long non-coding RNAs (lncRNAs). CircRNAs, generated via back-splicing and exon circularization, exhibit dual regulatory function in urothelial carcinoma, as detailed by earlier reviews. 132 However, several studies have since uncovered novel circRNAs with defined mechanistic roles. Tumor-suppresive circRNAs include circLAMA3 (exons 45–47 of LAMA3), which directly binds and degrades MYCN mRNA to inhibit cell proliferation and migration, and circFAM13B (exons 8–10 of FAM13B), generated via HNRNPL-mediated back-splicing to bind IGF2BP1, destabilize PKM2 mRNA, reverse glycolytic acidification and enhance immunotherapy sensitivity.133–135 CircICMT (exons 2–3 of ICMT) similarly suppresses migration and proliferation through complex miRNAs and RNA-binding protein interactions. 136 Oncogenic circRNAs such as circNIPBL (exons 2–9 of NIPBL) activates Wnt/β-catenin signaling by sponging miR-16-2-3p to upregulate ZEB1, which promotes circNIPBL biogenesis by binding NIPBL flanking introns, creating a positive feedback loop driving epithelial-mesenchymal transition. 137 EIF4A3-induced circ0008399 (exons 2-5 of RBM3) enhances WTAP-METTL3-METTL14 complex formation, stabilizing TNFAIP3 mRNA through m6A modification and conferring cisplatin resistance. 138
LncRNAs also regulate alternative splicing through direct splicing factor interactions and chromatin remodeling. Tumor-suppresive BCLET promotes MSANTD2-004 isoform expression via MSANTD2 exon 1 alternative splicing. 139 Oncogenic lncRNAs such as LINC00839 promotes urothelial carcinoma EMT, invasion, and gemcitabine resistance through EGR1–LINC00839–miR-142–SOX5 regulatory loop while MALAT1 enhances metastasis by interacting with the splicing factor SF2/ASF to facilitate TGF-β-induced EMT.140–142 Beyond transcriptional control, UCA1 drives metabolic reprograming by recruiting hnRNP I/L to enhance GPT2-mediated glutamine anaplerosis (refilling) that supports glycolysis and tumor proliferation. 143 These non-coding RNA-splicing networks present promising biomarkers and therapeutic target for precision medicine approaches in urothelial carcinoma, thus more research is needed for their clinical utility.
Disease risk, clinical correlation, and splicing-based therapeutics
In this section, we describe the splicing alterations associated with urothelial carcinoma risk, clinical outcomes, and potential therapeutics.
Urothelial carcinoma risk
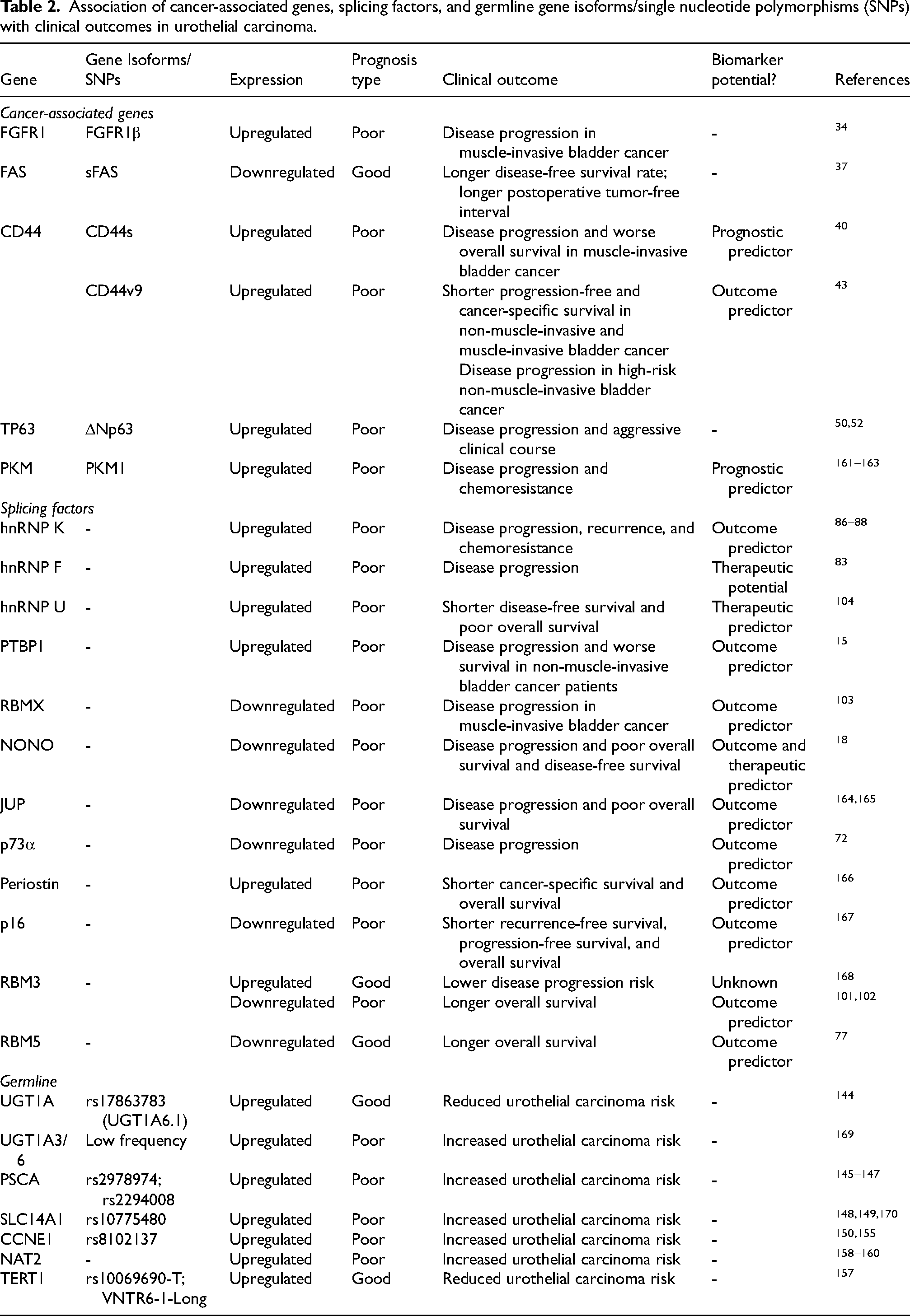
Genetic variants affecting genes such as UGT1A, PSCA, SLC14A1, NAT2, CCNE1, and TERT1 are associated with urothelial carcinoma risk (Table 2).144–160 For instance, the common T allele of the single nucleotide polymorphism (SNP) rs17863783 regulates expression of the UGT1A6.2 isoform, which is involved in the metabolism of aromatic amine in the bladder urothelium. 144 However, an alternative allele of this genetic variant produces UGT1A6 and UGT1A3 gene isoforms that increase the risk of urothelial carcinoma, especially in individuals with a history of smoking. 169 Increased expression of the UGT1A3 isoform has been linked to enhanced proliferation, migration, and invasion of urothelial carcinoma cells via LAMC2 activation. 171 The UGT1A genetic variant has also been associated with gender-specific differences in urothelial carcinoma incidence. 172 Overexpression of the T allele of SNP rs2294008 in the PSCA gene was reported to induce an alternative start site, leading to enhanced PSCA surface expression in high-grade bladder tumors.173,174 Another genetic variant, rs10775480, creates allele-specific isoforms of the SLC14A1 gene that are associated with low urine concentration and increased urothelial carcinoma risk. 170 In contrast, VNTR6-1-Long and the rs10069690-T variant produce the TERT-β isoform, which encodes a telomerase-nonfunctional TERT. This isoform extends cellular longevity of urothelial cells while reducing regenerative proliferation, thus reducing cancer risk. 157
Association of cancer-associated genes, splicing factors, and germline gene isoforms/single nucleotide polymorphisms (SNPs) with clinical outcomes in urothelial carcinoma.
Clinical outcomes
Expression of spliceosome proteins and cancer-associated genes has been linked to survival outcomes and tumor progression in multiple stages of urothelial carcinoma. SF3B1 overexpression in various cancers has been associated with tumor aggressiveness and decreased overall survival.175–178 Elevated SF3B1 expression has been specifically observed in non-muscle-invasive bladder cancer. 179 Overexpression of splicing regulators such as hnRNPF, hnRNPK, hnRNPU, and PTBP1 has been reported to be associated with disease progression, poor tumor prognosis, and worse survival in patients with non-muscle-invasive bladder cancer and muscle-invasive bladder cancer.15,83,86,87,104 Similarly, high expression of lncRNA AK023096, which recruits hnRNPK on target sites in pre-mRNA, was found to be associated with higher risk of recurrence following TURBT. 88 Downregulation of NONO, which also plays a role as a splicing factor, is associated with urothelial carcinoma lymph node metastasis, advanced tumor stage, and short overall and disease-free survival. 18
Overexpression of the cancer-associated splice variants FGFR1β, CD44 s, CD44v-10, JUP, periostin, p73α, PKM2, and ΔNp63 has been linked to more aggressive disease progression and higher recurrence rates of urothelial carcinoma.34,40,43,52,72,164–166,180 Decreased TAp63, p16, and p63 expression in urothelial carcinoma was correlated with higher tumor grades, tumor progression, and worse prognosis.50,89,90,167,181 MYC, a well-known oncogenic driver, can regulate alternative splicing through modulation of splicing-factor activity, contributing to cancer development and reduced overall survival. 182
Conversely, expression of certain RBM proteins, including RBM3, RBM5, and RBMX, has been associated with lower tumor grade, reduced metastasis risk, and improved survival.77,101,102,168 Low expression of soluble FAS (sFAS) isoforms in urothelial carcinoma was associated with improved survival rates and disease-free survival. 37 sFAS isoform expression has been proposed as a potential biomarker for diagnosing or monitoring primary or recurrent transitional cell carcinoma.38,39 HRAS gene isoforms associated with young-onset urothelial carcinoma have also been proposed as favorable prognostic markers.63,183 Additionally, highly expressed spliced variants of FGFR3, STAG2, ΔNp63, and P53Δ130 have been reported to correlate with low-grade tumors and better prognosis.54,94,181,184,185 Other gene isoforms correlated with tumor stage and survival outcomes include ACTN1, fibronectin ED-A, and MTAP/CDKN2A/2B.60,186,187
While these studies provide important insights into mRNA splicing dysregulation in urothelial carcinoma, most observations remain correlative and require further mechanistic investigation to establish the functional consequences of major splicing events, particularly in oncogenes and tumor suppressors. Moreover, the behavior of these RNA alterations in the context of targeted therapies remains largely unexplored. For instance, FGFR3-targeted therapies, such as erdafitinib, demonstrate clinical efficacy in urothelial carcinoma, yet it is unclear whether these agents differentially interact with specific FGFR3 splice variants across tumor subtypes, with potential implications for treatment outcomes.188–190
Potential splicing-based anticancer therapies
In preclinical studies, small molecule splicing inhibitors showed potential for therapeutic applications in many cancers. These agents typically disrupt spliceosome assembly by preventing the U2 snRNP from recognizing the branchpoint site, thereby blocking splicing. This leads to the accumulation of unspliced pre-mRNA, resulting in cell-cycle arrest and eventual cell death. Most small molecule inhibitors and their synthetic analogs target components of the SF3B complex, specifically SF3B1 (Table 3). Currently, there are no FDA-approved small molecule inhibitors of spliceosome proteins for urothelial carcinoma. Pladienolide B, a natural product, is an SF3B1 inhibitor that exhibits antitumor effects in vitro and in vivo in many solid and hematologic cancers.175–178,191 Despite the encouraging results from the phase I trial of a pladienolide synthetic analog E7107 in solid tumors, its efficacy was limited, and unexpected ocular toxicities were identified, leading to the early termination of its clinical development.205,206
Different splicing-based anticancer therapies.
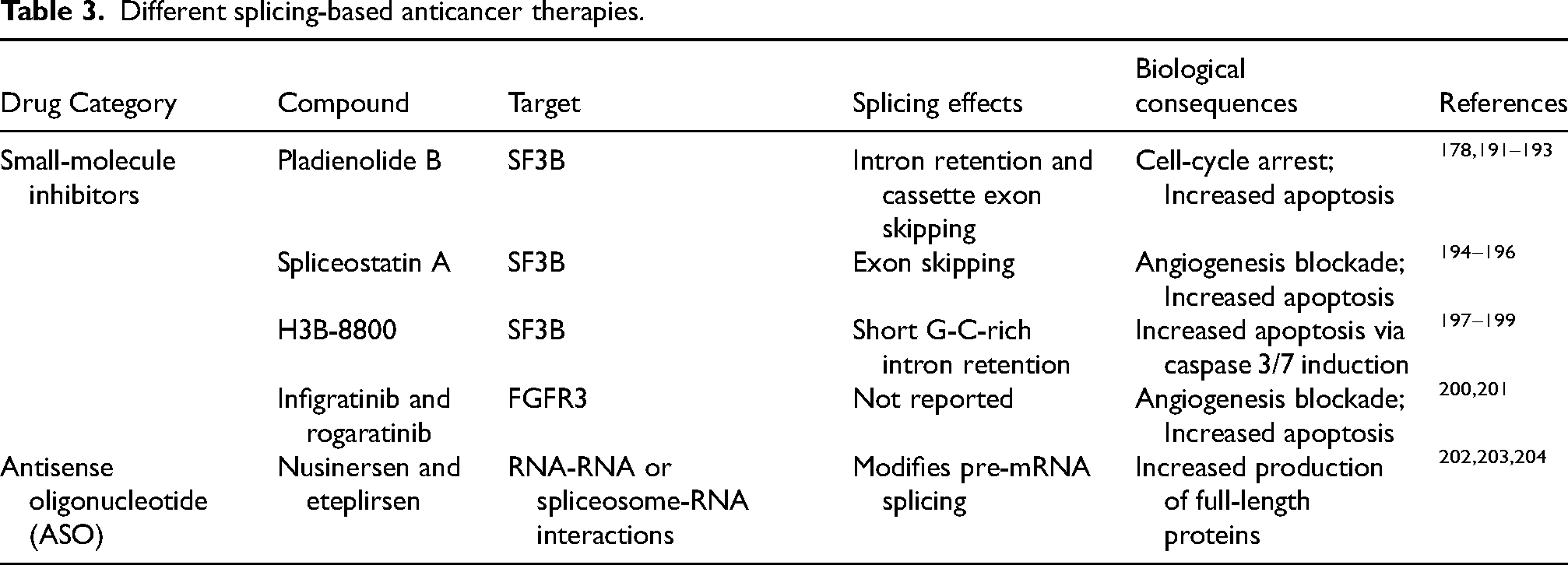

Another SF3B modulator, H3B-8800, has also shown encouraging antitumor effects in preclinical studies.7,198,199 H3B-8800 competitively binds to the wildtype and mutant SF3B1, inhibiting both canonical and aberrant splicing.7,207 A phase I trial of H3B-8800 to treat patients harboring SF3B1 mutations in hematologic disorders has been evaluated. 208 The drug was tolerated well; however, no objective complete or partial responses were observed. Further investigation of this drug in solid tumors is warranted. Other spliceosome inhibitors including spliceostatin A, herboxidenes, isoginkgetin, and sudemycin have exhibited antitumor effects by inducing apoptosis, inhibition of angiogenesis, and cell-cycle arrest in a variety of cancers.194–196,209–213 One limitation of spliceosome inhibitors is that they target both normal and altered spliceosome machinery, increasing the risk of life-altering side effects. However, combining splicing drugs with other existing chemo- and immunotherapies might offer unique strategies to treat specific tumor types. A preliminary in vitro study in urothelial carcinoma reported a synergistic effect of pladienolide B with cisplatin, though this remains to be tested in vivo.214,215 The combined use of spliceosome inhibitors and chemotherapies could potentially reduce side effects by allowing lower doses of each drug to target cancer-specific splicing modules. It may also be a potential strategy to overcome resistance to cisplatin and other effective therapies in urothelial carcinoma. This area of cancer therapeutics warrants further investigation, as it holds significant promise.
Antisense oligonucleotides (ASOs), which are small single-stranded DNA molecules that bind to specific mRNA targets and modify RNA splicing, have been used with clinical success in the treatment of non-cancer diseases such as spinal muscular atrophy and Duchenne muscular dystrophy (Table 3).202–204 However, ASO therapy faces several challenges in oncology, including delivery limitations and the molecular complexity of cancers, which warrants investigation of alternative ASO-based strategies for solid cancers. 109
Another promising therapeutic approach involves exploiting splice-derived neoantigens as novel targets for cancer immunotherapy, offering distinct advantages over conventional mutation-based strategies. Unlike single-nucleotide variants that alter only one amino acid, aberrant splicing events can induce frameshifts, generating extended novel protein sequences and highly immunogenic epitopes that are entirely foreign to the immune system (Figure 5).216–220 In normal cells, stringent quality-control mechanisms such as nonsense-mediated decay and the ubiquitin–proteasome system efficiently eliminate aberrant transcripts and proteins, resulting in minimal neoantigen presentation on MHC molecules. In cancer, however, elevated error rates in DNA replication, RNA splicing, and transcriptional fidelity overwhelm surveillance mechanisms, leading to the accumulation of frameshift transcripts, particularly from indel mutations and mis-splicing events. These aberrant proteins escape degradation, generating novel C-terminal sequences that are processed and presented on MHC class I and II molecules, where they are recognized as foreign by cytotoxic T cells. 221
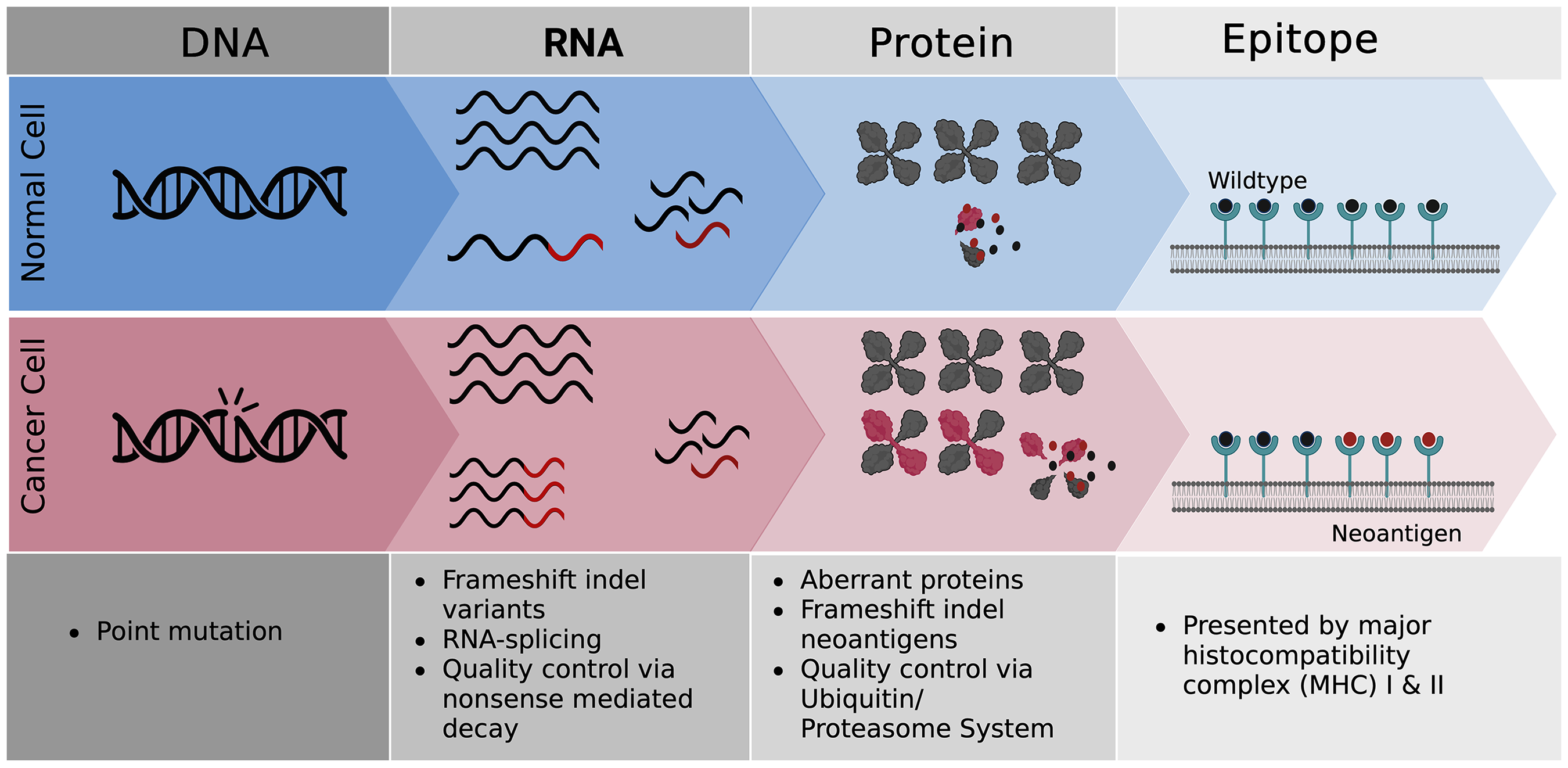
Neoantigen generation from frameshift events in normal versus cancer cells. In normal cells, nonsense-mediated decay, and the ubiquitin–proteasome system eliminate aberrant isoforms, limiting neoantigen presentation. In cancer, elevated replication errors, indels, and splicing defects produce abundant frameshift isoforms that escape surveillance, generating novel C-terminal sequences presented on MHC molecules and recognized by cytotoxic T cells.
Despite this potential, our current understanding of splice-derived neoantigens remains limited, especially in urothelial carcinoma, as most tumor datasets rely on short-read RNA sequencing, which identifies splice junctions but fails to resolve full-length isoforms and associated frameshifts 222 Long-read sequencing technologies (e.g., Oxford Nanopore, PacBio) now enable comprehensive characterization of aberrant transcripts, uncovering reservoirs of potentially targetable neoantigens.223,224 These insights can be translated into therapeutic applications, including splice-switching oligonucleotides designed to deliberately induce immunogenic splicing events, as well as the development of personalized neoantigen vaccines similar to NEO-PV-01—which has demonstrated enhanced efficacy when combined with immune checkpoint blockade. 225 Another promising strategy involves splicing-based modulation of tumor immunity through RNA splicing modulators. A recent preclinical study reported that such a modulator was able to enhance antitumor T-cell responses both in vivo and in vitro. 226 These and other methods of targeting RNA splicing appear promising as potential anticancer therapies but will need to be tested in preclinical studies to better understand their efficacy alone and in combination strategies.
Technological advances to capture full-length isoforms
Recent advances are reshaping the study of alternative splicing in cancers. Most current insights are derived from short-read RNA sequencing, which can identify splice junctions but cannot capture full-length transcripts or resolve complex isoform structures. Long-read platforms such as PacBio and Oxford Nanopore address this limitation by enabling complete transcript reconstruction, isoform quantification, and detection of RNA modifications—for example, mapping N6-methyladenosine (m6A) in muscle-invasive disease.227–229 These approaches can reveal the true diversity of isoform ratios and splicing switches that contribute to tumor heterogeneity and molecular subtypes. The field is now progressing toward single-cell long-read sequencing (e.g., PacBio Iso-Seq), which uncovers splicing complexity at cellular resolution. Efforts are also underway to integrate long-read sequencing with spatial transcriptomics, situating isoform heterogeneity within the tumor microenvironment.
Ongoing initiatives, including our lab, aim to generate such comprehensive splicing landscapes for urothelial carcinoma. These integrated approaches—spanning long-read, single-cell, and spatial profiling—will be critical for urothelial carcinoma biomarker discovery, therapeutic target identification, and the advancement of precision medicine.
Conclusion
In this review we discussed the functional consequences of different splicing alterations and cancer-associated gene isoforms in urothelial carcinoma, including their roles in cell proliferation and invasion, treatment resistance, metabolism, and modulation of the tumor microenvironment. We also elaborated on the implications of splicing-factor mutations commonly reported in urothelial carcinoma. Additionally, we highlighted how tumor-specific isoforms in oncogenes and tumor suppressors or specific alterations in splicing factors hold promise as potential biomarkers to aid diagnosis, prognosis, and monitoring of patients with urothelial carcinoma. However, realizing this potential requires the development of highly sensitive, specific, and cost-effective assays to detect alternatively spliced isoforms—tools that we currently lack. The described alternative splicing events found in urothelial carcinoma present new avenues for therapeutic targeting, including the use of small molecule inhibitors and splice-switching oligonucleotides. Despite growing reports of mRNA alterations in urothelial carcinoma, the biological roles and therapeutic applications of these changes have not been widely or systematically evaluated. Therefore, research priorities in this area should focus on systematic functional validation through gain- and loss-of-function studies, defining splice variant–specific therapeutic windows, and developing isoform-selective targeting strategies to unlock the full potential of splicing-based precision medicine in urothelial carcinoma.
Footnotes
Acknowledgments
This research was supported by the Intramural Research Program of the National Institutes of Health (NIH) and is subject to the NIH Public Access Policy. By accepting this federal funding, the authors grant the NIH the right to make the work publicly available in PubMed Central. The contributions of the NIH authors are considered works of the United States Government. The findings and conclusions presented in this paper are those of the authors and do not necessarily reflect the views of the NIH or the U.S. Department of Health and Human Services.
We thank Ms. Bonnie Casey, Clinical Editor at the Center for Cancer Research, National Cancer Institute, for her careful review of the manuscript and thoughtful editorial suggestions. All figures were created with BioRender.com.
Ethical considerations
As this manuscript is a literature review, it does not involve original research with human participants or animals and is therefore exempt from Institutional Review Board (IRB) approval.
Author contributions
RCAC, ABA, and ARB conceptualized the design and wrote the manuscript. RCAC prepared the figures. All authors read and approved the final manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors are funded by the Intramural Research Program of the National Cancer Institute, Center for Cancer Research. RCAC received funding from the NIH Medical Research Scholars Program, a public-private partnership supported jointly by the NIH and contributions to the Foundation for the NIH from the American Association for Dental Research and the Colgate-Palmolive Company.
Declaration of conflicting interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
Figure 3 is based on data from The Cancer Genome Atlas (TCGA) Bladder Cancer cohort, which is publicly available through cBioPortal (![]() ).
).
