Abstract
Hyperactivity of the Myc oncogenic transcription factor dramatically reprograms gene expression to facilitate cellular proliferation and tumorigenesis. To elicit these effects, Myc coordinates the activation and repression of an extensive network of protein-coding genes and, as has recently been appreciated, noncoding RNAs including microRNAs (miRNAs). Consistent with their ability to potently influence cancer phenotypes, the regulation of miRNAs by Myc affects virtually all aspects of the Myc oncogenic program, including proliferation, survival, metabolism, angiogenesis, and metastasis. This review will summarize the current understanding of the mechanisms underlying Myc-dependent transcriptional and posttranscriptional control of miRNAs and the resultant effects on tumorigenesis. As miRNAs are integral nodes in the transcriptional network controlled by Myc, modulating their activity represents a promising new approach for cancer therapy.
The
miRNA Biogenesis and Function
Most miRNA-encoding genes structurally resemble protein coding genes. The primary miRNA transcripts (pri-miRNAs) are usually transcribed by RNA polymerase II (although a small number are produced by RNA polymerase III); they are generally several thousand nucleotides in length, and they are capped, polyadenylated, and frequently spliced. 4-8 Indeed, many protein-coding genes contain miRNAs within introns. miRNA processing is initiated in the nucleus with cleavage of pri-miRNAs by the microprocessor complex which includes as core components Drosha, an RNase-III endonuclease, and DGCR8, an RNA-binding protein. This cleavage releases precursor stem loop structures, known as pre-miRNAs, which are ~70 nucleotides in length. 9-11 Pre-miRNAs are then exported to the cytoplasm by exportin 5 in a RanGTP-dependent manner. 12,13 Within the cytoplasm, pre-miRNAs are further processed by the RNase-III enzyme Dicer, which removes the stem loop and produces a ~22 nucleotide RNA duplex. 10 One strand of the duplex, referred to as the guide strand, stably associates with an Argonaute (Ago) protein, while the opposite strand (referred to as the passenger strand or miRNA*) is discarded. 14-16 The resulting miRNA-Ago complex associates with target mRNAs through miRNA:mRNA base-pairing interactions, most often within 3′ UTRs. miRNAs bind to their targets with imperfect complementarity, although interaction between the 5′ end of the miRNA (especially nucleotides 2-7, known as the seed sequence) is usually required for efficient target regulation. Following recruitment of the miRNA-Ago complex, the target mRNA undergoes accelerated turnover and/or translational repression. Many excellent reviews providing further details on miRNA biogenesis and function can be found elsewhere. 4,17,18
Myc-Mediated Reprogramming of miRNA Expression
Prior to 2005, it was not clear that miRNAs could act as important downstream effectors of key signaling pathways that influence cancer phenotypes. Indeed, the principal mechanisms through which miRNA expression is regulated, including the identities of mammalian transcription factors that control miRNAs, were unknown. Nevertheless, emerging evidence pointed to an important role for miRNAs in cancer. Early on, miRNAs that regulate cellular differentiation, proliferation, and apoptosis had been discovered in
The first demonstration that miRNAs could indeed function as critical components of oncogenic pathways came with the discovery that Myc directly activates transcription of the polycistronic miR-17-92 cluster. 27 Induction of these miRNAs occurs through the canonical mechanism whereby Myc binds directly to an E-box within the first intron of the gene encoding the miR-17-92 primary transcript. This locus encodes 6 mature miRNAs: miR-17, miR-18a, miR-19a, miR-20a, miR-19b-1, and miR-92a-1. 27,28 The miR-17-92 cluster has been duplicated during vertebrate evolution, resulting in 2 other paralogs, the miR-106a-363 and miR-106b-25 clusters. Mice deficient in miR-17-92 have defective B-cell development and die shortly after birth due to lung hypoplasia and ventricular septal defects. 29 In contrast, deletion of the miR-106a-363 or miR-106b-25 clusters does not result in any overt phenotype. Both miR-17-92 and miR-106b-25 are broadly expressed and have some functional redundancy since their combined deletion results in embryonic lethality. Expression of the miR-106a-363 cluster is undetectable in most tissues, and its physiologic functions are unknown.
Significant evidence now exists which convincingly implicates the miR-17-92 cluster as a
Despite the importance of activation of the miR-17-92 cluster, this represents only one aspect of a much broader Myc-regulated miRNA network. Further studies have demonstrated that Myc activity results in repression of numerous miRNAs, including many with documented tumor suppressor activity including let-7 family members, miR-15a/16-1, miR-26a, miR-29 family members, and miR-34a. 34 Each of these miRNAs have been demonstrated to exhibit antiproliferative, proapoptotic, and/or antitumorigenic activity in a variety of settings. 35-39 Accordingly, rescuing expression of several of these miRNAs in Myc-transformed B lymphoma cell lines dramatically inhibits tumorigenesis. 34 As expected, given their diverse targets, these repressed miRNAs broadly impact Myc-mediated phenotypes, as will be highlighted in greater detail below.
miRNA expression has been reported to be globally reduced in some tumor samples and cell lines,
40,41
and experimental inhibition of the miRNA biogenesis pathway accelerates tumorigenesis
Myc is also able to block the maturation of specific miRNAs without affecting transcription of the pri-miRNAs. Chang
Given the ability of individual miRNAs to regulate hundreds of targets, it is not surprising that the ability of Myc to broadly reprogram miRNA expression has dramatic and pleiotropic effects on cellular phenotypes. Although a full description of the known functions of all Myc-regulated miRNAs is beyond the scope of a single review, illustrative examples of how miRNA regulation has been integrated into the Myc oncogenic program will be provided in the ensuing sections (summarized in Figure 1 and Table 1).
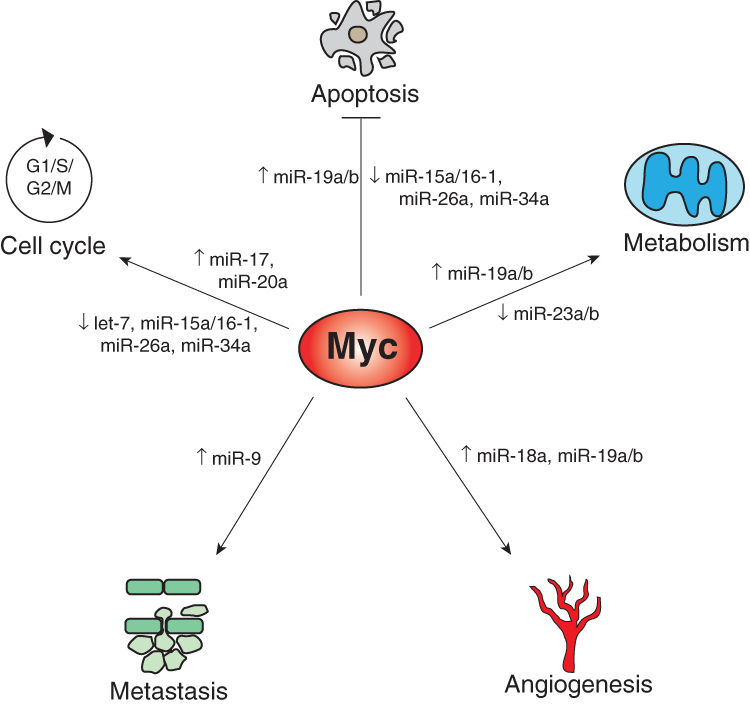
Myc-mediated induction or repression of miRNAs impinges upon multiple phenotypes that contribute to tumorigenesis, including cell-cycle progression, apoptosis, metabolic reprogramming, angiogenesis, and metastasis.
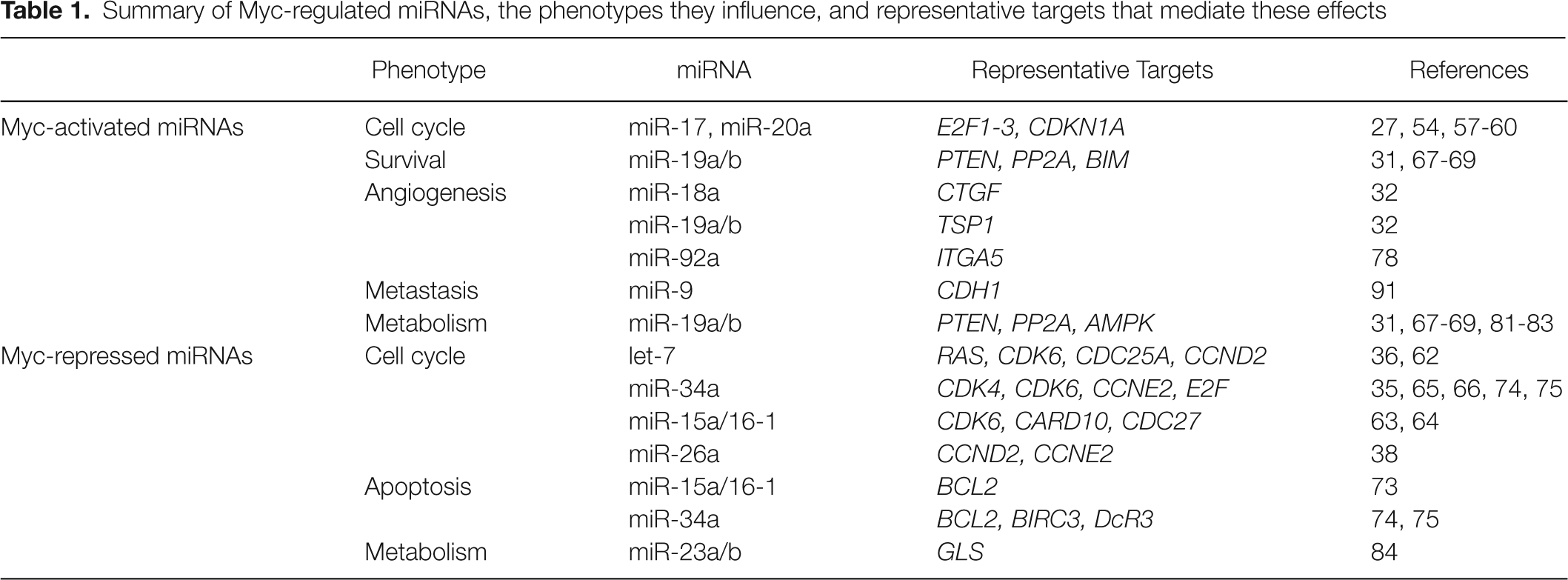
Summary of Myc-regulated miRNAs, the phenotypes they influence, and representative targets that mediate these effects
Cell-Cycle Control by Myc-Regulated miRNAs
Increased cellular proliferation is a major consequence of Myc activation in many cell types. Both induction and repression of specific miRNAs by Myc facilitates transit through the cell cycle through multiple mechanisms. For example, the initial studies which showed that miR-17-92 is directly induced by Myc also demonstrated that 2 components of the cluster, miR-17 and miR-20a, target the transcription factor E2F1.
27
Subsequent work also identified the related E2F2 and E2F3 transcription factors as additional targets of miR-17/20a.
54
E2F1-3, known as the activating E2Fs, positively regulate the transcription of genes required for DNA replication and are essential for G1-to-S phase progression.
55,56
However, their timing and levels of expression must be precisely controlled or, paradoxically, they can induce cell-cycle arrest or apoptosis. Interestingly, E2F1 is also known to be directly induced by Myc. Therefore, by simultaneously inducing E2F1 and its negative regulators, miR-17/20a, Myc may produce tightly controlled levels of E2F activity such that cell-cycle progression is promoted and apoptosis is avoided. Experimental support for this hypothesis was recently provided by Pickering
miR-17 and related family members (miR-20a, miR-93, and miR-106b) also promote cell-cycle progression by targeting the cyclin-dependent kinase inhibitor p21 (
Multiple miRNAs that are repressed by Myc, including let-7 family members, miR-15a/16-1, miR-26a, and miR-34a, have potent cell-cycle inhibitory activity.
38,61
Collectively, these miRNAs influence virtually all levels of the cell-cycle regulatory machinery. For example, key cell-cycle control factors regulated by let-7 include
Myc-Regulated miRNAs in Apoptosis
In the Eµ-
A major effector of the PI(3)K pathway is AKT, which has many downstream prosurvival activities, including inhibition of the proapoptotic protein Bim.
70
The ϵ isoform of protein phosphatase 2A (
Repression of miRNAs by Myc also contributes to cellular survival. For example, miR-15a/16-1, miR-34a, and miR-26a, which are repressed by Myc, 34 can each activate apoptosis in specific settings. The miR-15a/16-1 cluster directly downregulates the antiapoptotic protein Bcl-2, and expression of these miRNAs results in apoptosis in leukemic cell lines. 73 miR-34a is a direct transcriptional target of p53 and contributes to p53-dependent apoptosis. 74,75 In a mouse model of liver cancer, systemic delivery of miR-26a induced apoptosis specifically in tumor cells without affecting normal hepatocytes. 38 These observations highlight the broad impact of Myc-mediated miRNA reprogramming on cellular survival pathways.
Control of Angiogenesis by Myc-Regulated miRNAs
Myc promotes angiogenesis and vasculogenesis by upregulating the expression of proangiogenic factors such as vascular epidermal growth factor (VEGF) and angiopoietin 1/2, and by repressing antiangiogenic factors such as thrombospondin-1 (Tsp1) and connective tissue growth factor (CTGF).
32,76
Consistent with these activities, Myc-deficient mouse embryos exhibit vascular defects and die by embryonic day 10.5.
76
The miR-17-92 cluster plays an important role in Myc-induced angiogenesis. Dews
In contrast to the proangiogenic activities of miR-18a and miR-19a/b, recent work has uncovered an antiangiogenic role for miR-92a, another miR-17-92 component. 78 Systemic inhibition of miR-92a in mice using antagomiRs (modified antisense miRNA inhibitors 79 ) stimulates angiogenesis following multiple insults, including limb ischemia and myocardial infarction. The antiangiogenic activity of miR-92a is partially attributable to its ability to downregulate the angiogenesis promoting factor integrin α5. Precisely how the activities of the proangiogenic (miR-18a, miR-19a/b) and antiangiogenic (miR-92a) components of the miR-17-92 cluster are coordinated will be an important issue for future study. It is possible that in different cellular contexts, specific members of the cluster accumulate to different levels, resulting in distinct phenotypic outcomes. Additionally, differential expression of targets could certainly contribute to context-dependent effects of miR-17-92 expression on angiogenesis.
Metabolic Reprogramming by Myc-Regulated miRNAs
Glucose and glutamine are the 2 major carbon substrates utilized by highly proliferative cells.
80
As described in detail above, Myc-dependent activation of the miR-17-92 cluster potentiates signaling through the PI(3)K-AKT pathway.
31,67-69
miR-19 family members play a major role in this activity by virtue of their ability to downregulate the expression of PTEN, which antagonizes PI(3)K activity. PI(3)K-AKT signaling strongly promotes glucose metabolism and fatty acid synthesis through multiple mechanisms, including increasing glucose transporter surface expression and enhancing glycolytic enzyme activity.
81
AKT also activates ATP citrate lyase, an enzyme that is required for glucose dependent fatty acid synthesis and tumor growth
Glutamine metabolism is promoted by Myc through several mechanisms, including the direct transcriptional activation of glutamine transporters.
83
Myc activity also leads to increased expression of mitochondrial glutaminase (GLS), a key enzyme that converts glutamine to glutamate, which serves as a substrate in the TCA cycle for the production of ATP.
83,84
Both glutamine and GLS are required for Myc-mediated cancer cell survival and proliferation. Interestingly, Myc increases GLS activity by directly repressing transcription of 2 miRNAs that target the
Regulation of miRNAs by N-Myc
The
Targeting Myc-Driven Tumorigenesis with miRNA Therapeutics
The ability of miRNAs to potently influence cancer phenotypes suggests that strategies designed to inhibit or augment the activity of specific miRNAs in tumor cells might provide therapeutic benefit. Given the pleiotropic oncogenic functions of the miR-17-92 cluster and its importance in the Myc network, targeting these miRNAs with systemically delivered antisense reagents such as antagomiRs or locked nucleic acid oligonucleotides
79,93
represents a promising approach worthy of evaluation. Studies of therapeutic inhibition of miR-17-92 in tumor models
The ability of miRNA replacement therapies to suppress Myc-driven tumorigenesis
The effectiveness of miRNA replacement therapies is not limited to Myc-driven tumorigenesis. Delivery of let-7 has been shown by multiple laboratories to be an effective therapeutic approach in a well-established mouse model of non–small-cell lung cancer driven by oncogenic
As described in this review and elsewhere, the expression of many distinct miRNAs can inhibit cancer cell proliferation and survival in a large number of tumor types, underscoring the broad potential of miRNA replacement therapies. Although the lack of efficient nucleic acid delivery strategies for many tissue sites currently represents a major obstacle, the development of improved delivery methods is a major area of investigation in academic and pharmaceutical laboratories, offering hope for future implementation of this approach for many types of cancer.
Conclusion
Myc-mediated reprogramming of miRNA expression introduces a newly appreciated layer of complexity to the diverse physiologic and pathologic functions of this oncoprotein. The network of Myc-regulated miRNAs control key effectors of this oncogenic pathway that influence the cell cycle, apoptosis, metabolism, angiogenesis, and metastasis. Further study of the precise functions of these miRNAs will be essential to achieve a complete understanding of Myc-driven tumorigenesis and will likely reveal new diagnostic and therapeutic opportunities for cancer.
Footnotes
J.T.M. is an HHMI Early Career Scientist and a Rita Allen Foundation Scholar. The Mendell laboratory receives additional support from the NIH (R01CA120185 and P01CA143292).
The authors declare no conflicts of interest with respect to the publication of this article.
