Abstract
Up to 6% of the global population is estimated to be affected by one of about 10,000 distinct rare diseases (RDs). RDs are, to this day, often not understood, and thus, patients are heavily underserved. Most RD studies are chronically underfunded, and research faces inherent difficulties in analyzing scarce data. Furthermore, the creation and analysis of representative datasets are often constrained by stringent data protection regulations, such as the EU General Data Protection Regulation. This review examines the potential of federated learning (FL) as a privacy-by-design approach to training machine learning on distributed datasets while ensuring data privacy by maintaining the local patient data and only sharing model parameters, which is particularly beneficial in the context of sensitive data that cannot be collected in a centralized manner. FL enhances model accuracy by leveraging diverse datasets without compromising data privacy. This is particularly relevant in rare diseases, where heterogeneity and small sample sizes impede the development of robust models. FL further has the potential to enable the discovery of novel biomarkers, enhance patient stratification, and facilitate the development of personalized treatment plans. This review illustrates how FL can facilitate large-scale, cross-institutional collaboration, thereby enabling the development of more accurate and generalizable models for improved diagnosis and treatment of rare diseases. However, challenges such as non-independently distributed data and significant computational and bandwidth requirements still need to be addressed. Future research must focus on applying FL technology for rare disease datasets while exploring standardized protocols for cross-border collaborations that can ultimately pave the way for a new era of privacy-preserving and distributed data-driven rare disease research.
Introduction
Rare diseases (RDs) are complex health conditions characterized by their low prevalence in the general population. RDs affect between 1 in 2000 to 1 in 200,000 people, with changing definitions depending on the region.1,2 A disease is classified as an ultra-rare disease (URD) if it affects less than 1 in 50,000 individuals. 3 Despite their low individual prevalence, collectively, RDs impact approximately 3.5% to 5.9% of the world's population.1,2,4 Their phenotypes are frequently chronic, progressive, and potentially life-threatening, 5 presenting unique challenges in diagnosis, treatment, and research compared to more common diseases. 6 This results in RD patients being chronically underserved due to limited advancements in RD management. For example, the accuracy of RD diagnosis, even when molecular data is considered, is only 50%. 7 Improvement is hampered by limited expertise, lack of research funding, and too small patient populations for clinical trials.8,9 However, RD research can reveal fundamental insights into biological pathways and genetic mutations, leading to broader scientific advancements that benefit rare and common conditions alike. 10
One inherent roadblock in RD research is the limited availability of comprehensive datasets that can facilitate the development of precise diagnostic tools and personalized treatment strategies. 11 To address this challenge, several crucial repositories for rare disease (RD) datasets have been established, including the Global Rare Diseases Registry and Repository, 12 Orphadata, 13 DECIPHER, 14 PhenomeCentral. 15 These siloed data sources can be obstacles to discovering various causes of considering the spectrum of overall RDs. To tackle this issue, the Matchmaker Exchange Application Program Interface 16 initiative has been developed, which uses a common data-sharing protocol to enable seamless searches and interactions across multiple databases while allowing each to maintain its own data organization schema. However, despite these valuable resources, the vast diversity of RDs necessitates even more comprehensive open-source datasets that include various genetic, phenotypic, and clinical information. The existing repositories, while important, do not contain all the estimated 10,000 RDs, especially newly discovered ones, covering around 6000–7000 each.1,17 Hence, collaboration among different hospitals or clinics to share their patient data for specific rare diseases is essential for advancing research in this field. It highlights the need for additional data sources to fill gaps and ensure robust research that can enhance accuracy and improve patient outcomes.
Artificial intelligence (AI) has demonstrated superior performance over traditional statistical methods in processing complex datasets essential for RD research. 18 A subcategory of AI is Machine learning (ML), which describes the ability of algorithms to extract correlations, distributions, probabilities, or other metrics from data that have predictive or classifying value. 19 AI models and its category of deep learning (DL) models turned out to be essential tools in enhancing our understanding of RDs by leveraging neural network structures to learn feature correlations, 20 integrating multimodal data sources, and enabling comprehensive analysis.21,22 These methodologies can facilitate personalized medicine development by identifying patterns that correlate with treatment success, which benefits RDs with limited treatment options and variable patient responses.23–25 AI-driven biomarker discovery already facilitates disease subtyping, patient stratification, and the identification of therapeutic targets.26,27 In this way, AI contributes to developing novel interventions and drug repurposing strategies for many diseases.28–30
While AI also offers transformative potential for RD research, the methods used to train these models present distinct challenges in balancing data privacy, model performance, and generalizability. Common approaches to model training come with specific limitations. Local training within one clinic (Figure 1A) is privacy-friendly but often results in models that don't generalize well due to limited and potentially biased datasets. 31 Centralized learning (Figure 1B) involves data collection from multiple sources, potentially balancing the data and improving model accuracy. However, it encounters significant administrative and patient privacy-related barriers as, in many cases, patient data cannot be freely shared between collaboration partners. This limitation arises from concerns regarding patient privacy, compliance with legal regulations, preventing unauthorized access, and maintaining data integrity. 32 In recent years alone, millions of clinical records have been affected by data breaches, 33 leading to patient stigmatization,34,35 even though data protection is always a critical concern when collecting medical data.36,37 Consequently, strict data protection regulations, like the EU's General Data Protection Regulation (GDPR), are crucial and indispensable to ensuring patient privacy. 38 However, data protection laws further increase the hurdles for collaborative research projects, rendering centralized data collection more challenging due to administrative overhead. Therefore, innovative approaches that balance the need for comprehensive datasets with stringent privacy requirements are crucial, especially in the context of RDs.28,39
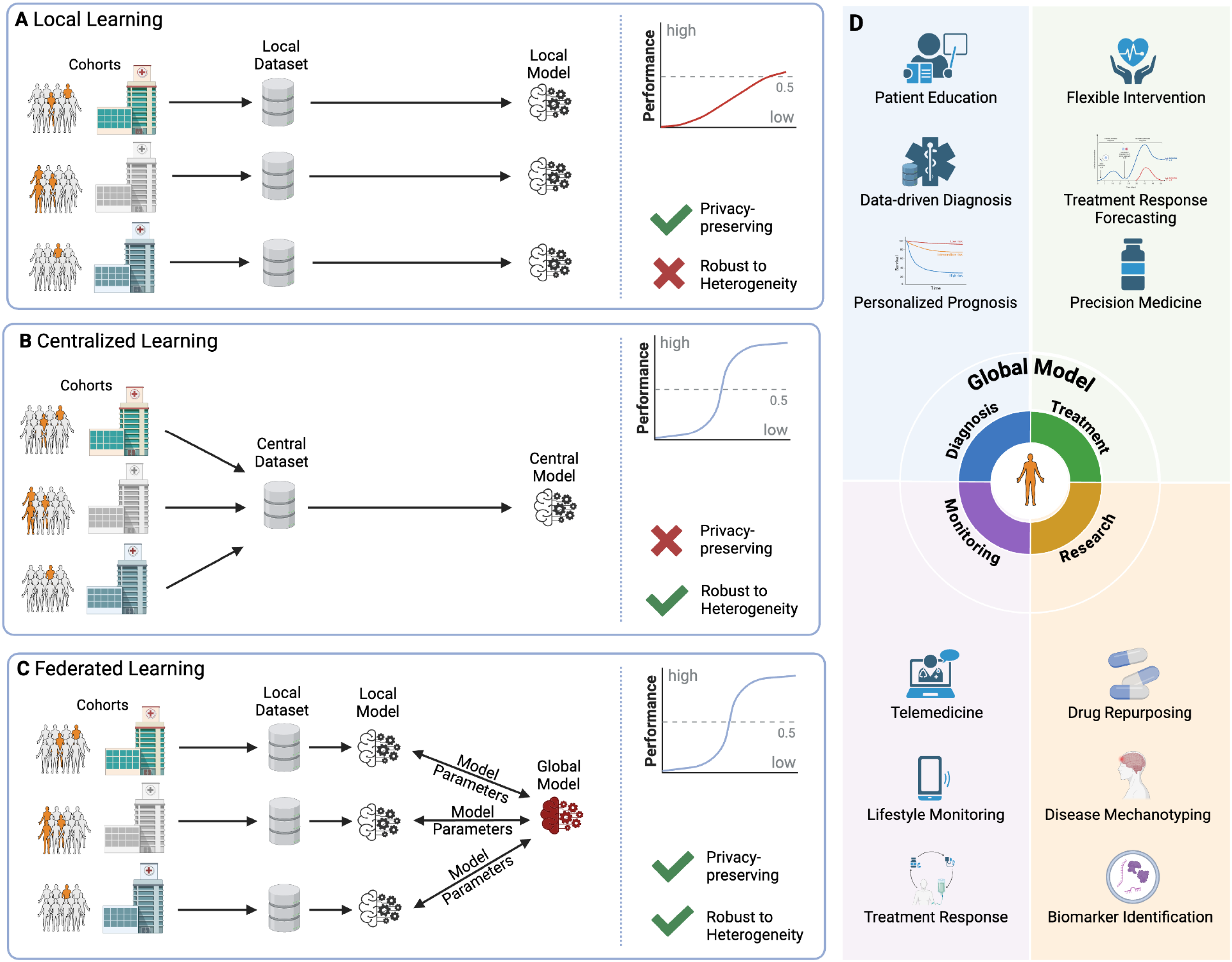
AI approaches for Rare Disease Research and Treatment—an overview of data processing methods in machine learning and specific applications and areas in healthcare. There are generally three approaches to analyzing patient data with AI techniques: local, centralized, and federated learning. Patients suffering from a rare disease are highlighted. In non-collaborative research scenarios, institutions limit their model training to locally available data (A). In comparison, centralized learning (B) collects data from all patient groups centrally. Federated learning (C) combines the advantages of the two previous approaches. Local datasets are used to create local models, and their parameters are then aggregated into a global model. (D) shows possible applications of the global model that offer improved performance while protecting patient data by applying the FL approach.
Federated learning (FL) has emerged as a promising solution to overcome the challenges associated with centralized AI models (Figure 1C). 40 In FL, the local datasets of participating clients are used to create local AI models. After each local training round, the parameters of these models are aggregated to create a global model without the need to share sensitive data. 41 After multiple training rounds, this approach results in a model trained on all local datasets, not communicating any patient data, to reduce data bias and provide high-accuracy models. 42 Crucially, as the data remains within each institution, FL ensures privacy-preserving training of distributed datasets while maintaining data security.36,43 This characteristic is particularly relevant for RDs, where the number of individual cases in a single institution for a given condition is typically very small,44,45 and federated collaborative research opens the gates for previously unthinkable amounts of data without compromising privacy.28,46 This way, FL has the potential to advance RD research and treatment in an era of stringent data protection requirements.28,37,47
This review examines AI's contribution to healthcare to highlight its possibilities, with a specific focus on FL's role and potential in RD research. We assess AI's current state, limitations, and applications in healthcare, particularly for RDs, and discuss how FL can address these limitations, highlighting existing FL applications in healthcare. Additionally, we explore how FL can advance our understanding of RDs while addressing data privacy concerns.
Main
Artificial intelligence in healthcare
AI systems, specifically ML and DL approaches are revolutionizing healthcare, offering unprecedented opportunities to enhance patient care and clinical efficiency. From advanced diagnostic tools to operational automation, AI is reshaping how healthcare challenges are addressed. Integrating AI technologies promises significant improvements in early disease detection, personalized treatment plans, and overall patient management. As the healthcare sector continues to generate vast amounts of data, AI's role in processing and deriving actionable insights becomes increasingly crucial.
State of the art and current solutions
Approximately 10% of global healthcare expenditure is currently attributable to fraud and abuse, which can be reduced through AI. 48 Apart from means of protection, AI has demonstrated remarkable progress in healthcare applications, consistently outperforming traditional statistical methods in processing large and complex datasets.18,49 As computer vision is a highly advanced field within AI, medical imaging and radiology have been at the forefront of AI adoption in healthcare.50,51 New means of analysis were required, as the increasing speed and resolution of medical imaging devices, producing more and higher-quality output, has led to a significant rise in the workload for healthcare professionals, who are experiencing difficulties in keeping up with processing all the data.48–50 AI systems can process the growing volume and complexity of imaging data, providing faster and more precise analyses that complement or even outperform the work of healthcare professionals. The early detection of lung cancer is one area where computer vision has shown potential, facilitating the use of imaging techniques such as computed tomography or X-ray imaging. 52 On the other hand, analysis of molecular-level biological data enables more accurate disease diagnoses and deepens our understanding of disease mechanisms. 53 This is exemplified by using AI models to identify diagnostically and prognostically relevant correlations between genetic variants and cytomorphological changes in myelodysplastic syndrome, with the long-term goal of achieving disease classification only based on genomic data. 54 Recently, integrating multimodal data sources enables comprehensive analysis to enhance our understanding of diseases further, allowing the joint integration of genomics, imaging, and clinical records into one model.21,22,55,56 Such models are used in various contexts. For example, Unlearn.AI uses digital twin models that validate clinical studies on Alzheimer's disease and multiple sclerosis by simulating subjects with the same clinical attributes as the actual subjects.22,57,58 The used data include a variety of clinical and demographic covariates. 58 All these systems provide decision support to healthcare specialists, enabling them to analyze and interpret larger amounts of data in a significantly shorter time and more accurately.24,25
Even more possibilities arise from the recent emergence of large language models (LLMs) like OpenAI's GPT, Meta's LLama, or Google's Gemini.59–61 Their potential to enhance patient education and support, given a diagnosis or treatment, is already evident.62,63 Also, specialized LLMs for healthcare professionals, such as Med-PaLM, 64 have emerged, offering significant promise for further advancement. Possible application areas include LLM-driven assistants capable of comprehensive analysis and contextualization of a patient's medical history or electronic health records (EHRs), which could greatly benefit physicians. 64 In addition, EHRs and other unstructured data can now be analyzed more easily and categorized, expanding their role as information sources in healthcare applications. 64 This includes the extraction of medical relationships from doctoral notes in EHRs, such as the identification of drug-side effect relationships, as well as the resolution of medical questions. 64
In drug development and repurposing, AI allows for the identification of new target molecules, potentially leading to more effective drugs.22,23 By analyzing the molecular structure of existing drugs, drug discovery processes can be accelerated, particularly for immune-related genes, enabling the identification of off-targets or repurposing potential.26,27 For instance, in 2019, Insilico Medicine used AI to design and synthesize a novel drug candidate for fibrosis in 21 days and validate it in just 46 days, a process that traditionally takes years.65,66 Additionally, the introduction and improvement of DeepMind's AlphaFold has revolutionized protein structure prediction, a critical component of drug research, by achieving near-experimental accuracy in determining protein structures from amino acid sequences67,68
As the healthcare industry continues to shift towards more patient-centric and technology-driven care models, remote healthcare delivery has become an increasingly important aspect. The field of telemedicine, which provides clinical services remotely, is of increasing importance in healthcare due to its capacity to improve access to medical services and reduce costs for both patients and providers.69,70 Integrating AI into telemedicine has significantly enhanced its capabilities by improving diagnostic accuracy, enabling timely virtual triage, and providing personalized treatment recommendations. 70 These improvements are achieved by utilizing the knowledge and expertise distributed across the data on which the model is based. 71 It has also proven invaluable during public health crises and for individuals in underserved areas or with limited mobility by enabling remote consultations, monitoring, and treatment.69,72,73 A prime example of a successful telemedicine platform is Teladoc Health, which conducted up to 20,000 virtual visits per day during the COVID-19 crisis, 74 significantly reducing exposure risks and ensuring continuous care for patients, particularly in underserved regions, serving the demand for telemedical solutions at the time.75,76
These diverse applications demonstrate the broad impact of AI across healthcare domains, from molecular-level research to direct patient care. As AI technologies evolve, they promise to transform healthcare practices further, potentially leading to more precise, efficient, and personalized medical care.
Limitations of AI in medical applications
The advancement of precise AI models in the healthcare sector is confronted with many considerable obstacles, the most pressing being the need for extensive and uniform datasets. 77 As data harmonization is often challenging due to data scarcity and lack of homogeneity, the power of AI stays limited, particularly in the context of RDs, where data is scarce and fragmented.1,78 These issues led researchers to propose generating artificial time series data for electrocardiograms (ECG) and electroencephalograms due to the significant challenges posed by data scarcity and class imbalance in research. 79
On the other hand, a global trend towards data protection is evident, with more countries having enacted or preparing to enact relevant legislation, 80 all while RD patients are reported to be comparably open for data sharing yet scared of data misuse.81,82 Additionally, the EU AI Act introduces stringent regulations that could limit the application of AI in the medical field. It considers AI systems in healthcare as potentially high-risk as it can assess individuals’ health or profile patients. 83 This classification may lead to increased compliance requirements such as extensive risk management, data quality assurance, transparency measures, human oversight, and robustness checks, increasing the time, complexity, and overall cost of developing and deploying AI in healthcare. 83 However, to support organizations in preparing for these regulations, the European Commission has introduced the AI Pact, encouraging voluntary early compliance with the AI Act's requirements. 84
Conversely, even when comprehensive data protection measures are implemented, and centralized data aggregation is feasible, enforcing data quality and standardization remains a significant challenge. 85 This is due to the lack of consensus regarding evaluating various dimensions of data quality and its measures. 85 One source for quality issues is data entry errors that arise from the manual input of patient information. 86 Clinical records, but even more so, EHRs often include non-standardized questions, free-text patient responses, and personal comments from treating physicians, invoking substantial barriers to data quality, subjectivity, and standardization. 87 Similarly, in genomics research, issues like inconsistent naming conventions and varying experimental protocols are prevalent. 88 These and more quality and standardization obstacles inherently compromise the effectiveness of AI in medical applications by limiting the accuracy and reliability of the data used to train and operate AI systems.
AI in rare disease research
The potential for improving RD research and treatment has significantly increased with the advent of AI approaches. The developed models have been shown to outperform traditional statistical methods in processing large and complex datasets essential for studying RDs such as Huntington's Disease, 89 Hypophosphatasia, 90 Multiple osteochondromas, 91 Amyotrophic lateral sclerosis.18,92–94
From the initial stages of detection and diagnosis to the subsequent phases of treatment planning and drug development, AI models are transforming numerous aspects of RD management as they continue to evolve. For the detection and diagnosis of RD, AI models have shown remarkable effectiveness in phenotypic and genetic analysis. 95 By analyzing patient data encompassing symptom-related and genetic information, AI enables more expedient and accurate RD diagnosis, facilitating the identification of patterns that might otherwise be overlooked due to potential human error. For instance, the capacity to predict the progression of rare neurodegenerative diseases, such as multiple system atrophy, represents a promising avenue of research.93,96 Furthermore, AI models have exhibited promise in detecting RDs, such as Gaucher's disease and generalized pustular psoriasis, by detecting subtle patterns in clinical, genomic, and imaging data.78,95 Several studies use EHR data for the diagnosis of acute hepatic porphyria, thereby enabling a reduction in diagnostic delay with the potential to enhance patient outcomes.97,98 Another important advancement is the interpretability of results, one approach being a model that predicts infection risks in pediatric leukemia patients. 99 Using interpretable models ensures that the predictions are understandable to clinicians, enabling better clinical decision-making. By incorporating these sophisticated models into the decision-making process, clinicians can enhance their ability to select the most effective treatments, potentially improving patient outcomes and overall quality of care.24,25
Drug discovery and development are other highly relevant fields for RD research, and AI's ability to rapidly analyze vast amounts of scientific data is proving invaluable. It can facilitate the identification of novel therapeutic targets for various RDs by pinpointing specific molecular pathways and potential drug candidates with higher precision and efficiency. 23 A study has used various AI models to develop a new method for evaluating factors related to Metachromatic Leukodystrophy, a rare genetic disorder that destroys the protective fatty layers around nerves in the central and peripheral nervous systems with the accumulation of sulfatides. 100 This method helps understand the disease's pathogenesis, progression, and potential treatment options, identifying single and dual drug combinations as promising therapeutic targets.
Advancements in gene and protein replacement therapies and stem cell and genome editing technologies have been driven by detailed studies of the biological pathways involved in rare genetic disorders. 101 For example, AI approaches have successfully identified key genetic variants and pathways associated with mitochondrial dysfunction in rare neurodegenerative disorders, which are crucial for developing targeted therapies. 102 Additionally, AI models were developed to examine gene expression patterns to predict and identify genes that could serve as early diagnostic or therapeutic markers, resulting in valuable findings for Chronic Nonbacterial Osteomyelitis, a rare autoinflammatory bone disease caused by abnormality in the immune system. 27 Another exciting approach is using AI to detect splicing defects in genetic data that are often overlooked in standard diagnostic workflows. This is advantageous in the context of RDs such as spinal muscular atrophy, in which splicing defects are known to play a significant role in the manifestation and progression of the disease. 103 From identifying genetic variants to uncovering specific pathways, the power of AI in advancing our understanding of RDs and paving the way for more effective treatments has been worth mentioning. 104 TRANSLATE NAMSE showed that AI can identify ultra-rare genetic disorders and novel gene-disease associations. 105 It identified 370 different genetic causes and discovered 34 new and 23 potential genotype-phenotype associations, mainly under the umbrella of neurodevelopmental disorders.
While AI shows great promise in improving RD research and treatment, 28 it's important to note that research demonstration is still relatively limited, often focusing on the more common RDs. 106 For URDs, AI applications remain scarce, but ongoing efforts are expanding AI's reach, as exemplified by the development of TRANSLATE NAMSE or AI-MARRVEL, a machine learning system designed to prioritize potentially causative variants for a broader range of Mendelian disorders, including ultra-rare cases. 107 Recently, AI has been employed to examine genetic data and discern novel genetic variants linked to a collection of ultra-rare metabolic disorders resulting from deficiencies in glycosylation. This process, which involves the attachment of sugars to proteins and lipids, is known as the congenital disorder of glycosylation. In general, the use of AI has enhanced diagnostic precision and facilitated a more complete comprehension of associated disease mechanisms. 105
Federated learning in healthcare
The potential of federated learning
FL emerges as a promising approach to overcome the challenges associated with centralized AI training in healthcare (Figure 1C) by effectively combining the benefits of centralized and local training. 40 This combination of local data utilization and collaborative learning allows institutions to benefit from training access to large-scale, heterogeneous datasets without being constrained by data privacy or security concerns.108,109 By limiting the transferred information to only model updates, FL mitigates the risk of data breaches, thereby addressing critical privacy and security concerns.33,43 At the same time, global models trained with FL on diverse datasets are usually more robust, thus less susceptible to bias, and can achieve high accuracy. 42 A study utilizing FL to diagnose patients with COVID-19 infection, based on chest radiographs from five international healthcare systems, demonstrated that local models trained on local data and integrated into a global model demonstrated greater accuracy when applied to previously unseen datasets. 110 Further, FL may significantly reduce communication costs and bandwidth usage, as the total size of communicated parameters is usually smaller than the raw data size.111,112 It has additional potential to be more cost-efficient than centralized learning, as it can utilize otherwise unused, locally available computing resources, in contrast to the need for investing in a central, high-performance computing system. 112 Since each clinic only requires sufficient computing power for its resources, FL provides a scalable solution for large-scale machine-learning applications.112,113
Continuous FL techniques can be employed to periodically train adaptable and optimized models, as data is constantly generated but not actively reshared at all times. This thereby reduces regulatory hurdles that might arise from direct data sharing.105,108,109,114 This approach enables quicker adaptation to new data, which is particularly beneficial in healthcare, where lots of data are generated, and data patterns frequently change. 115
Multiple instances of the utilization of FL capitalize on all the advantages. For instance, FL has been demonstrated to facilitate the development of more robust and accurate models by integrating data from diverse healthcare institutions and patient populations following privacy-compliant protocols, for example, for the computer-aided diagnosis of cancer. 114 Furthermore, using a federated framework validated the ability to predict the risk of sepsis and acute kidney injury in ICU patients from a range of clinical settings. 116 Moreover, FL offers potential for improvements in treatment decision support systems as new and more data, improved diagnostic procedures, and new medications become relevant to disease treatment and detection. 28 For non-small cell lung cancer (stage I-III), a two-year survival model has been trained on routine data from radiation oncology in a federated fashion. 117
Even a political incentive to establish FL infrastructures is evident from more than 28 ongoing EU-funded projects indexed by the Community Research and Development Information Service. 118 Notable projects include FLUTE, which applies FL techniques to prostate cancer research, 119 and dAIbetes which aims to create personalized type 2 diabetes treatment outcomes by developing a federated health data platform that consolidates data across multiple international cohorts. 120 Additionally, the extensive research project “Controls for Deep and Federated Learning” aims to develop new FL technologies with privacy preservation guarantees. 121 This extensive research initiative and other ongoing projects demonstrate the growing recognition of FL's potential, upcoming solutions, and investment in FL research within the European Union.
Challenges in implementing FL
Despite its potential, FL faces several significant challenges in implementation, particularly in healthcare settings. One of the most fundamental issues is the presence of non-independent and identically distributed (non-IID) data originating from different hospitals. 47 These significantly disparate statistical characteristics of the data, which can arise, for example, due to the different ethnic distribution of patients in the individual hospitals, can propagate biases and inconsistencies into the global model. 47 One potential solution to this issue is data augmentation, designed to expand and enrich local datasets. The objective is to achieve a more uniform data distribution. 122 Another proposed solution is the combination of FL with transfer learning (TL) methods. TL uses the knowledge acquired in one situation to improve performance in a related situation. The application of TL in FL is known as federated transfer learning (FTL) 123 and shares insights gained by a model at one location with models at other locations in a federated manner, potentially mitigating data scarcity issues and enhancing the model's capacity for generalization. 124 FTL techniques can counteract these problems by either pre-processing the data or trying to address the challenges in the model itself. 123 Another issue that can be addressed by FTL is the heterogeneity of labels. 123 This refers to the situation in which different labeling practices and criteria are employed in different hospitals, resulting in inconsistent and heterogeneous labels and meanings behind identical labels.125,126 Such inconsistencies can potentially introduce biases into the AI model. 127 Further, different hospitals may display disparate distributions of characteristics, including variations in units of measurement, scales, and the availability of specific characteristics.123,126 Such discrepancies can result in the model performing inadequately when generalizing across disparate sources, as it relies on characteristics that are not uniformly present or similarly distributed. To address this issue, feature normalization, standardization, and transformation techniques can be employed to align feature distributions.18,128,129 Furthermore, the temporal variability of data represents a significant challenge.114,123 Data collected over different time periods may exhibit temporal variability, such as changes in medical practice, diagnostic criteria, or patient demographics. Variability over time can cause a model to perform well with data from the past but poorly with more recent or future data, reducing its practical utility and sustainability. 130 To counteract this, the inclusion of temporal validation and time series analysis techniques is crucial. Continuous learning approaches that regularly update models with new data can also help to maintain their relevancy and accuracy over time. 123
While FL aims to protect privacy by keeping data where they are primarily generated or stored, the risk of reconstruction attacks remains a concern. Attackers listening to the communicated information in an FL network could infer original data from model updates, compromising patient confidentiality. 18 This risk necessitates implementing robust security measures, including access control, authentication, and verification mechanisms, as well as privacy-enhancing techniques like secure multi-party computation and differential privacy,131,132 which further increases the complexity and demands on the infrastructure. 36
Setting up a FL system also comes with its own issues. FL requires a robust technical infrastructure, presenting a significant obstacle for smaller or technically limited organizations or countries.40,47 Failure to participate in an ongoing FL training process due to setup-related issues could result in the generation of skewed or underrepresented models, as important health data could be missed. 37 Moreover, even if training on every FL node of a federated network is successful, FL often leads to longer training times due to increased communication and waiting times. 40 This increases the complexity of implementing models in comparison to classic centralized approaches. This issue is compounded by the limited availability of high-quality, labeled, and harmonized data in real-world healthcare settings, which poses a significant hurdle for effective model training. 124 Their complexity highlights the necessity for international collaboration within the scientific community. 28
Existing FL applications in healthcare
The potential of FL in healthcare is immense, but its practical implementations still need to be improved. A recent study found that only 5.2% of the results obtained from FL studies in healthcare are applied in practice, with the biggest part being feasibility studies. 43 Nevertheless, several noteworthy applications demonstrate FL's potential in various healthcare domains.
For example, in cardiovascular health, FL has been successfully used to detect hypertrophic cardiomyopathy, a condition where the heart muscle thickens abnormally, by combining ECG and echocardiogram data from multiple institutions. 133 Similarly, the ICU4Covid project developed a decision support system using FL that enhances the early identification of high-risk hypertensive patients while preserving data privacy, demonstrating its potential as a reliable predictive tool. 70 The ICU4Covid's telemedical network was used to propose an FL framework to significantly reduce communication costs and latency and enable small and medium-sized healthcare organizations to benefit from collective intelligence. 70 Finally, ADMarker shows that it is feasible to identify comprehensive multidimensional biomarkers, thereby facilitating the precise and early detection of the diverse manifestations of Alzheimer's disease. 134
Another significant area where FL has brought about substantial opportunities is health monitoring, particularly through the use of Internet of Medical Things (IoMT) devices that focus solely on Internet of Things (IoT) applications within the healthcare sector. IoT devices facilitate data exchange from physical devices connected to clinical management and patient well-being monitoring via the Internet. 135 The FedHome model highlights FL's ability to enable collaborative model training across multiple devices without sharing raw data. 136 The proposed approach effectively handles imbalanced data and reduces communication costs, common challenges in health monitoring scenarios. By accomplishing this, FedHome provides health monitoring in a residential setting through edge devices (end-devices), which facilitate the detection of falls, the monitoring of health-related activities, and the delivery of personalized health recommendations and alerts based on individual health data. 136 FL's capacity to enable personalized healthcare is evident in its ability to leverage the computational power of individual devices, allowing for the refinement of global models while maintaining data privacy, as exemplified by ClusterGAN for stress-level prediction using ECG signals. 137 This makes FL a compelling solution for health monitoring applications involving wearable devices. Smart monitoring systems that protect privacy and employ edge devices and sensors can alleviate pressure on healthcare systems and caregivers through continuous monitoring and early detection of potential health issues.138,139 Based on this, IoMT and FL can also be employed to construct decentralized networks that specialize, for instance, in neurological and metabolic diseases, thereby enhancing the collective analysis of sensor data for early detection and differential diagnosis. 140
Federated learning for rare diseases
FL has emerged as a powerful and highly considerable solution to address the limitations mentioned above and privacy concerns that limit the usability of general AI systems. 114 However, the scarcity of medical data for RDs poses significant challenges in training effective predictive models. This limitation stems from two main factors: the inherent rarity of these conditions and the heightened privacy concerns surrounding patient information. 82 As a result, the already limited number of available datasets can suffer from severe bias, for instance, patients of different socioeconomic statuses, 123 making it extremely difficult to develop accurate and generalizable models for disease prediction. 37
FL frameworks have the potential to enhance the accuracy of RD detection by drawing information from diverse data sources. 37 Furthermore, as the data does not leave the hospital, privacy-preserving training of distributed datasets is ensured, maintaining data security.36,43 This is particularly valuable for RDs as the number of individual cases in a single institution and for a given disease is usually very small.44,45
Recently, to explore its practicality, FL was applied to detect Tall Cell Morphology (TCM) in thyroid cancer, a rare but aggressive variant, from Whole Slide Images (WSIs). The federated training was simulated across the WSI datasets in three virtual clients to classify tissue patches as “tall” (expressing TCM) or “non-tall”. Model parameters from each client were then aggregated to ensure convergence. In various experiments, the FL models achieved accuracy comparable to that of centralized models, demonstrating the potential of FL to enhance diagnostic accuracy for rare but clinically significant detections. 141
Another noteworthy application case of FL in the detection of RDs is a model for glioblastoma, an aggressive form of brain tumor. 31 The comprehensive study utilized data from 71 geographically distinct sites across six continents and 6314 glioblastoma patients, representing one of the most extensive global FL studies. This approach enabled the creation of a robust and generalizable model for tumor boundary detection, showing improved performance for detecting different tumor regions, such as 27% for enhancing tumor (ET), 33% for tumor core (TC), and 16% for whole tumor (WT) against local validation data, and 15% for ET, 27% for TC, and 16% for WT against unseen data. 31
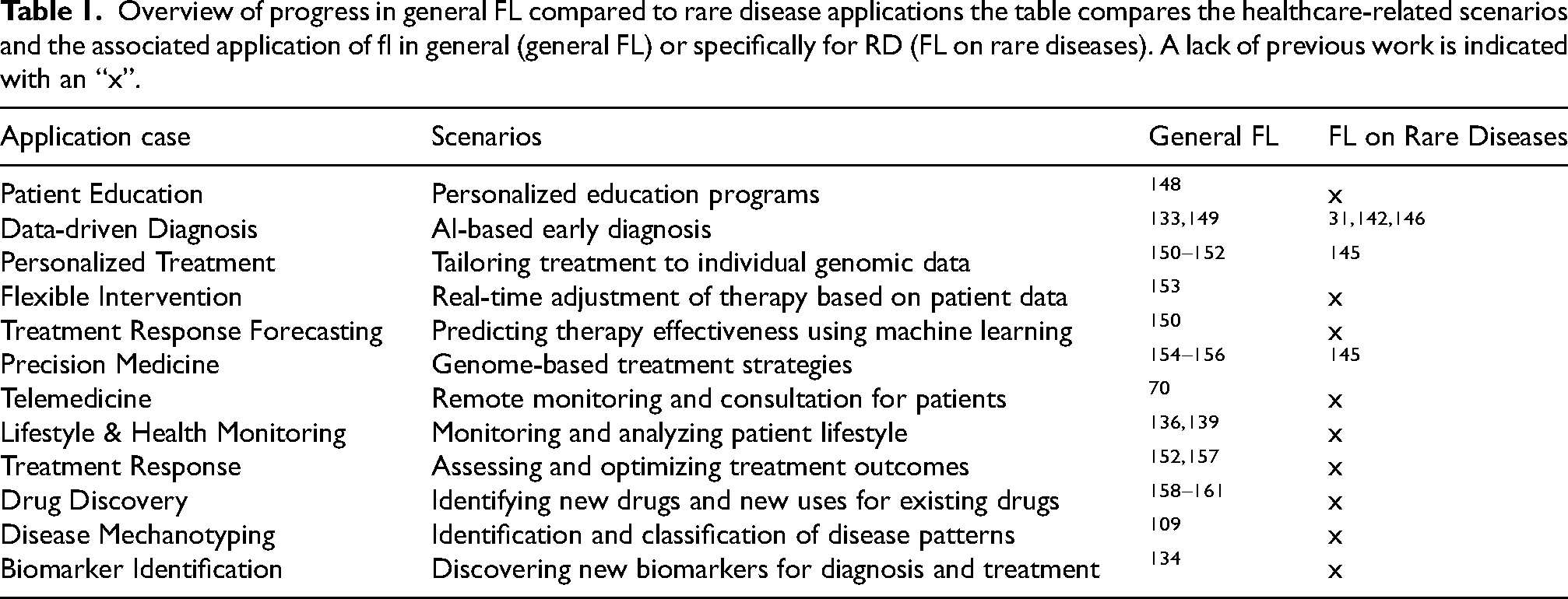
In the context of RD research, novel FL approaches have been developed. One notable contribution is the Dynamic Federated Meta-Learning approach that addresses not only the challenges of small sample sizes resulting in less usable models per data site but also focuses on optimizing the model's performance, where having very few positive samples makes training a robust model difficult, w.r.t. RD research. 142 Separately, FedIIC proposes a novel method specifically designed to address the issue of class imbalance in medical image classification, such as having far fewer examples of RDs compared to healthy cases. 143 Another promising approach is feature-context-driven federated meta-learning, which employs dynamic weighting of clients based on the accuracy of each local model to enhance the prediction accuracy of RDs. 144 This approach has been demonstrated to be effective when applied to adapted datasets simulating RD data based on cases of cardiac arrhythmias and skin injuries. Another innovative initiative is GenoMed4All, which creates an FL platform to connect European clinical and biomedical datasets on RDs. It uses a common data model to maintain uniform client data standards. 145 In contrast, FedRare implements a technique called “contrastive learning”, facilitating more effective representation learning in the context of data heterogeneity. 146 The technique that is used allows a model to learn to group data points that are similar together and to keep those that are dissimilar apart. 147 As illustrated in Figure 1 (D), FL presents many potential applications in the context of RD. These are repeated in Table 1 with corresponding studies. It is noteworthy that there is a significant need for research into RDs, which is why the table also includes comparative studies that do not relate specifically to RD but are available for common diseases. While these models (“General FL”) may not have been explicitly designed for RDs, they nonetheless demonstrate potential for adaptation. With the requisite datasets for the desired RDs, these models could be constructed similarly.
Overview of progress in general FL compared to rare disease applications the table compares the healthcare-related scenarios and the associated application of fl in general (general FL) or specifically for RD (FL on rare diseases). A lack of previous work is indicated with an “x”.
The future of federated learning in rare diseases
The technological advancement of FL, coupled with its scalable and privacy-preserving architecture, presents a promising avenue for utilizing sensitive data in RD research while adhering to stringent data privacy regulations. Key advancements in FL, including edge learning techniques162,163 and continuous FL approaches,164,165 will significantly enhance the applicability and effectiveness of this technology in tomorrow's healthcare. It will particularly benefit RD research, where data and expertise are more widely distributed across multiple clinics.28,37 Innovations in the field of FL help to make it more accessible to a broader range of clinicians and researchers, potentially accelerating the detection and treatment of various RDs.
Building on these technological advancements, the increasing user-friendliness, scalability, and computing efficiency of FL frameworks41,146,166 have reduced the technical barriers to studying conditions that have traditionally been challenging due to limited data and resources, like RDs. Moreover, FTL has opened up new possibilities for cross-regional collaboration, addressing the persistent challenge of data heterogeneity in RD research. 123
Importantly, FL's distributed nature promotes more ethical use of patient data, 43 fostering greater trust and involvement from patients in the research process. This aspect is particularly crucial in RD research, where patient participation is often a prerequisite for successful research initiatives. 167
Looking to the future, applying these cutting-edge technologies promises to facilitate a profound enhancement of our comprehension and treatment of RDs. Large-scale collaboration will promote standardization as data need to be harmonized. Increased patient involvement could be hoped for with improved data privacy security, resulting in more data serving as a base for more promising research. These benefits could lead to a significant acceleration in the research and treatment of RDs. However, realizing the full potential of FL in RD research will require sustained efforts from the scientific community. There must be increased collaboration between healthcare institutions and technology developers to refine and implement FL technologies further. Additional research is required to address the remaining and pressing challenges, such as ensuring model fairness and robustness in the face of diverse and potentially biased datasets to not perpetuate existing inequalities and biases.
Conclusion
FL shows great potential to help patients with RDs identify fundamental causes, explore treatment options, and receive more accurate and faster diagnoses. However, this potential still must be realized by transferring expertise present in centralized AI development to federated RD research. With the increasing availability of general FL frameworks, incentives for cross-border collaborations must be established. This approach will facilitate the development of new RD-specific models based on existing AI or newly proposed paradigms to address the unique challenges of RDs. Simultaneously, the refinement and development of FL technologies can accelerate this transition to more FL-driven RD research, ultimately leading to improved patient outcomes worldwide. Future collaborations should focus on implementing data standardization protocols for impactful cross-border collaborations.
Footnotes
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The dAIbetes project has received funding from the European Union's Horizon research and innovation programme under the Grant Agreement no: 101136305. The information contained in this press release are however those of the author(s) only and do not necessarily reflect those of the European Union. This work was developed as part of the FeMAI project and is funded by the German Federal Ministry of Education and Research (BMBF) under grant number 01IS21079.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
