Abstract

Mass Spectrometry Overview
Mass spectrometry is playing an increasingly important role in the molecular characterization of combinatorial libraries, natural products, and biopolymers.1-5 The development of matrix-assisted laser desorption/ionization (MALDI) and electrospray ionization (ESI) has significantly extended its application toward a wide variety of challenging problems in drug discovery and toward the identification of effective ligand-receptor binding, new catalysts, and enzyme inhibitors. Crucial to distinguishing the most active component or obtaining structure-activity relationships of compounds in a library is an efficient qualitative and quantitative assay. Toward this end, ESI and MALDI have been useful for the qualitative,6-8 and more recently, the quantitative screening of combinatorial libraries.9,10
In addition, mass spectrometry does not involve chromophores or radiolabelling, and thus provides a viable alternative to existing analytical techniques which typically require extensive sample preparation and optimization time, the disposal of radioactive waste, or require a significant amount of sample.
The utility of ESI lies in its ability to generate ions directly from the solution phase into the gas phase. The ions are created by applying a strong electric field to a very fine spray of the solution containing the analyte. The electric field creates highly charged droplets whose subsequent vaporization (or desolvation) results in the production of gaseous ions. The fact that ions are formed from solution has established the technique as a convenient mass detector for liquid chromatography and for automated sample analysis.
In addition, ESI-MS offers many advantages over other mass spectrometric methods including the ability to analyze low mass compounds, excellent quantitation and reproducibility, high sensitivity, simple sample preparation, amenability to automation, soft ionization, and the absence of matrix.1,11-13
In MALDI-MS, gas phase ions are generated by the laser vaporization of a solid matrix/analyte mixture. The matrix (usually a small crystalline organic compound) is necessary to absorb the laser radiation, in effect acting as a receptacle for energy deposition. This concentrated energy deposition results in the vaporization and ionization of both matrix and analyte ions. A useful attribute of MALDI is that it is especially well suited for the simultaneous analysis of multi-component mixtures. In the following review of some of our previous work, we have made use of the unique ionization qualities of both techniques for automation and quantitative analysis in combinatorial chemistry.
Automated ESI-MS as a Quantitative Assay for Enzyme Inhibitors
Using the advantages of ESI-MS listed above we have investigated inhibitor libraries designed to inhibit enzymatic glycosylation reactions. In this approach the entire enzymatic mixture (substrate, inhibitor, product, and internal standard) is introduced into the ESI mass spectrometer while analyzing for product formation as a function of the presence of inhibitor. Since only the product formation is quantitatively monitored, the effectiveness of the inhibitor can be readily determined.
We have applied this automated, quantitative ESI-MS approach (Figure 1) to glycoprocessing enzymes which are responsible for processing cell-surface carbohydrates. These carbohydrates in turn are associated with many specific cellular recognition and signaling events. Therefore, developing new assays and finding new inhibitors for glycoprocessing reactions is both medically relevant as well as scientifically interesting. To date, however, only limited success has been achieved in the design of these inhibitors since current assays require the separation of the compounds, the use of radiolabeled sugars, the performance of large scale reactions, or the use of additional enzymes.
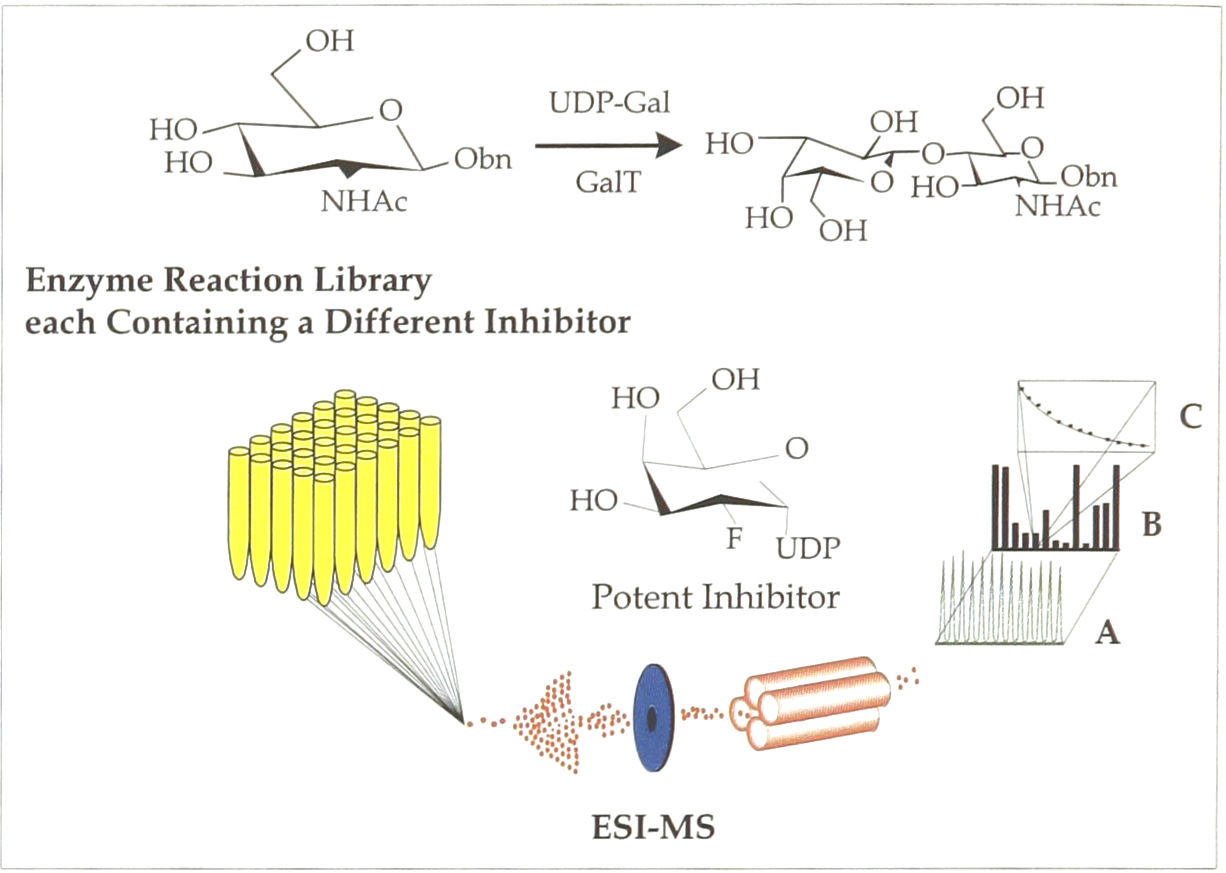
Quantitative determination of galactosyltransferase inhibition. (top) The enzyme reaction was monitored using an automated electrospray ionization mass spectrometry screening experiment. (bottom) Each solution contains the enzyme, reactants, an inhibitor (inset), and an internal standard. A) The total ion current is recorded on each injection into the mass spectrometer. B) Product formation is monitored with respect to the internal standard during each injection and C) if an inhibitor is found to be effective, IC50 data can be generated using the ESI-MS.
By monitoring one inhibitor at a time we could examine the effectiveness of a new inhibitor every two minutes. (Figure 2) In an effort to further increase the screening capacity, multiple inhibitors were screened against one enzyme simultaneously. The experiments were performed by pooling 5 potential inhibitors and the enzyme into one reaction vessel. The reactions that contained 5 inactive compounds showed no inhibition, whereas the reactions containing 4 inactive compounds and 1 active inhibitor (uridine diphosphate) showed potent inhibition. The compounds in the inhibited reaction were then individually analyzed to identify the actual inhibitor compound. Such strategies increased the screening capacity several fold such that 1 inhibitor/2 min allowed for 720 inhibitors/day, while 5 inhibitors/2 min allowed for 3600 inhibitors/day.
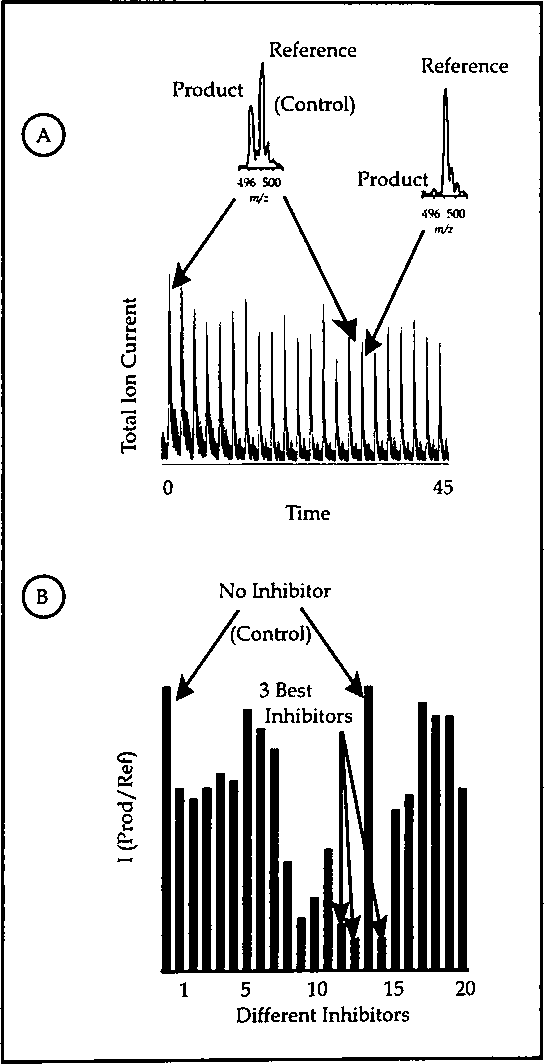
(A) The electrospray ionization total ion current observed for the 22 reactions (each having a different inhibitor). Inserts are the mass spectra for the reactions with no inhibitor (left, top) and an inhibitor found to be effective (right, top). (B) The bargraph of product ion/reference ion intensity ratio generated from the mass spectrum of each reaction containing potential inhibitors.
Automated Quantitation Studies using MALDI-MS
One of MALDI's greatest attributes is its ability to analyze complex hetergeneous mixtures. This ability has made the MALDI technique a valuable alternative to immunoassays and HPLC in the analysis of biological fluids. Immunoassays generally have low reproducibility and reliability, and provide little to no selectivity between a drug and its metabolites. This lack of specificity is a significant limitation since metabolites, although structurally similar to the parent compound, often have different biological activity. Mass spectrometric analysis, on the other hand, allows for co-extracted metabolites to be identified and quantitatively monitored (unless they have the same molecular weight). And although HPLC is relatively selective and accurate, the sensitivity is very compound-dependent and method development can be time-consuming. Thus, while both techniques are useful, they suffer when compared to the speed, sensitivity, and accuracy offered by MALDI mass spectrometry.15-17
Since some biofluids contain contaminants, it is often necessary to perform extractions for purification prior to sample analysis. In such instances the development of an efficient extraction assay can be very time-consuming. In an effort to create a more simplified and efficient extraction protocol, we have developed a combinatorial extraction method to be used with an automated MALDI mass spectrometry procedure. 15 In this example we demonstrate the approach to improve clinical analysis of the immunosuppressant drug cyclosporin A (CsA).
Figure 3 shows the combinatorial-extraction approach followed by analysis, in which a MALDI mass spectrometer equipped with automated multi-sampling capabilities was used to facilitate data collection and analysis of cyclosporin A. The organic layer extracted from blood for each sample was placed on a MALDI sample plate (with a capacity for one hundred samples) and was then loaded into the mass spectrometer and analyzed using a computer-controlled algorithm. Extraction optimization was performed by generating an array of solvent systems and an automated analysis to identify successful extractions. The first generation of experiments revealed four binary solvent systems to be effective for cyclosporin extraction (hexane/EtOH, ACN/H2O, ACN/MeOH, hexane/CHCl3). A new array based on these solvent systems was generated and in a second iteration of these experiments, hexane/CHCl3 (70:30) was found to provide the most effective single-step extraction of these solvent systems for cyclosporin and its metabolites.
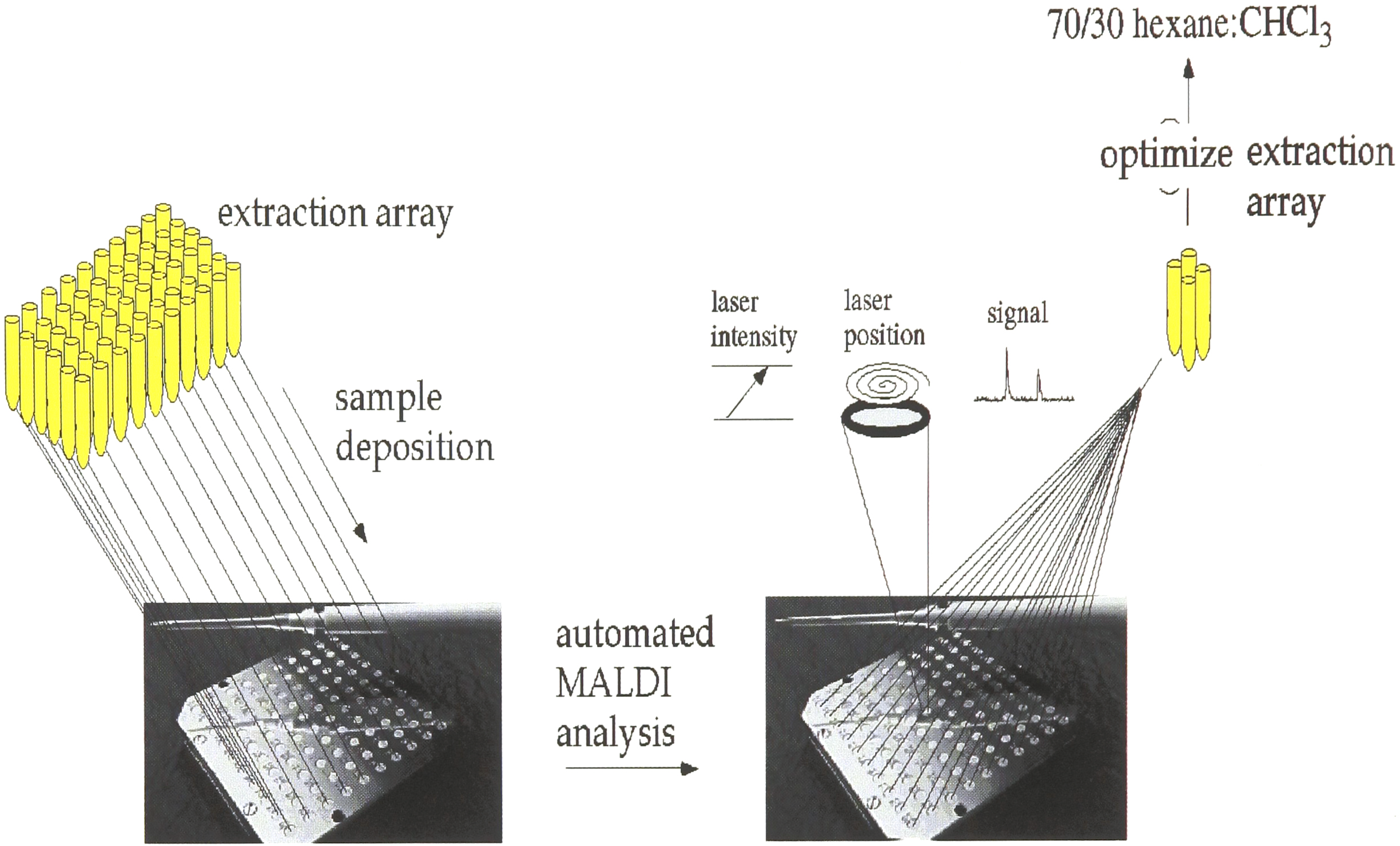
The automated MALDI mass spectrometry experiments were performed as a function of laser position (within each well) and laser intensity for each extracted CsA sample. The first iteration of these extraction experiments generated 4 useful solvent systems, and the second generation produced an effective 70/30 hexane:CHCl3 extraction solvent system.
Characterizing Compounds Directly from the Solid Phase
The use of both the automated ESI and MALDI-MS approaches described above represent only a few of the many ways in which mass spectrometry can be used in the field of combinatorial chemistry. In addition to the development of new automated quantitative methodologies, the qualitative evaluation of combinatorial libraries is essential for the characterization of active compounds and for monitoring both the progression and efficiency of reaction pathways. Because many libraries are synthesized on solid polymeric supports (e.g., resin beads), this chemistry necessitates assays which allow for the rapid characterization directly from the solid support. In the following section we utilized both MALDI-MS and photolabile linkers as a means of directly characterizing compounds from the solid phase.
Several reports have shown the utility of mass spectrometry in the characterization of compounds subsequent to their chemical cleavage from solid polymeric supports. Demonstrated here is a new MALDI approach which allows compounds covalently bound to a single polymeric bead (∼50 μm in diameter) or set of beads to be mass analyzed prior to their chemical cleavage from the resin.18,19 The scheme, outlined in Figure 4, permits the characterization of resin-bound analytes in a single step which requires no pretreatment of the sample to induce cleavage from the support. We have demonstrated that peptides covalently linked to a polymeric support through a photolabile linker can be directly analyzed by MALDI-MS. In addition, we have also shown that the technique is suitable for monitoring chemical reactions on the solid phase.
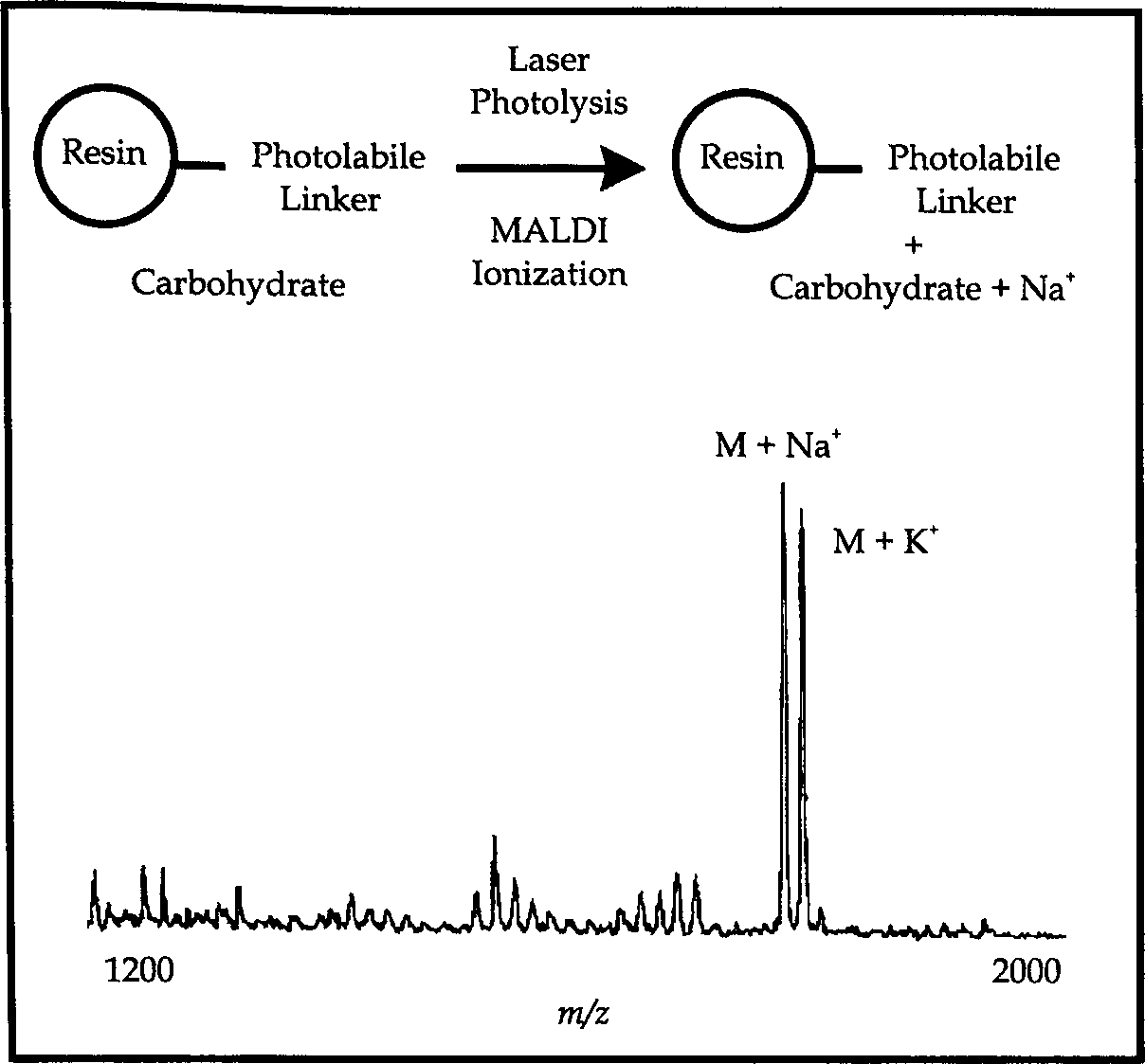
The MALDI technique facilitated the vaporization and ionization of analyte molecules by irradiating the sample/matrix mixture with a UV laser. (top) Schematic representation of the products generated upon laser photolysis (by MALDI) of a carbohydrate covalently attached to a solid phase resin through a photolabile linker. The mass spectrum of the carbohydrate was generated directly from the solid phase using a photolabile linker.
Figure 4 shows the mass spectrum of the protected, resin-bound carbohydrate. 19 MALDI analysis yielded a characteristic [M+Na]+ signal acquired from over 30 laser pulses on a single bead. The visualization system on the PerSeptive Voyager Elite instrument allowed irradiation of very specific regions of the sample plate. MALDI analysis of resin-bound peptide was also performed.
The direct analysis of resin-bound molecules by MALDI offers several important advantages, the first being the lack of an additional cleavage step prior to mass analysis, which is often required by other methods of characterization. Performing MALDI directly on the solid phase requires less sample handling and more efficient management since the resin-bound compound can be easily recovered for subsequent manipulations as less than femtomoles of material is consumed in typical MALDI analyses. The most significant advantage is that it can be used to monitor chemical reactions on the solid phase in real time, in much the same way that thin-layer chromatography is used to monitor reactions in solution. Furthermore, all analytes amenable to MALDI ionization should prove suitable for routine analysis by this procedure.
Conclusion
The rapid growth of combinatorial libraries has created a need for faster, more accurate, and more sensitive analytical techniques capable of large-scale (high-throughput) screening.
This mini-review of our previously published work 9,10,14,15,18,19 accentuates the numerous improvements in speed, sensitivity and accuracy of mass spectrometry, together with innovations in both automation and quantitation that make it an effective analytical alternative.
Overall, the strength of mass spectrometry lies in such versatility, placing it among the most powerful analytical techniques available today.
