Abstract
A nanochannel-based electrochemical biosensor has been demonstrated for rapid and multiplexed detection of a panel of three biomarkers associated with rapid detection of sepsis. The label-free biosensor detected procalcitonin (PCT), lipoteichoic acid (LTA), and lipopolysaccharide (LPS) from human whole blood. The biosensor comprises a nanoporous nylon membrane integrated onto a microelectrode sensor platform for nanoconfinement effects. Charge perturbations due to biomarker binding are recorded as impedance changes using electrochemical impedance spectroscopy. The measured impedance change is used to quantitatively determine the concentration of the three biomarkers using antibody receptors from the tested sample. We were successful in detecting and quantifying the three biomarkers from whole blood. The limit of detection was 0.1 ng/mL for PCT and 1 µg/mL for LPS and LTA. The sensor was able to demonstrate a dynamic range of detection from 01.1 ng/mL to 10 µg/mL for PCT and from 1 µg/mL to 1000 µg/mL for LPS and LTA biomarkers. This novel technology has promising preliminary results toward the design of sensors for rapid and sensitive detection of the three panel biomarkers in whole blood toward diagnosis and classification of sepsis.
Introduction
Companion diagnostics are in-vitro, clinical, or take-home diagnostic assays that are designed to aid in the diagnosis and prognosis of various diseases. Until recently, most companion diagnostic systems approved by the Food and Drug Administration (FDA) have been designed along a linear paradigm of “one/single biomarker per test.” However, the limitations of this are becoming apparent as in vitro diagnostics are shifting toward multiplexed detection of a panel of biomarkers that can help understand disease origin, progression, and growth nature. This can help toward making better informed clinical decisions in treatment and drug administration. There is a need for companion diagnostics in infectious diseases where the focus is on detecting the host response to pathogen infection. This helps in identifying the nature of effect that the host is experiencing and can help in designing successful treatment therapies.
Sepsis is the leading cause of death in critically ill patients despite the use of modern antibiotics and resuscitation therapies. The septic response is an extremely complex chain of events involving inflammatory and anti-inflammatory processes. The diagnosis of sepsis and evaluation of its severity are complicated by the highly variable and nonspecific nature of the signs and symptoms of sepsis. 1 However, the early diagnosis and stratification of the severity of sepsis are very important, increasing the possibility of starting timely and specific treatment.2,3
Biomarkers can have an important place in this process because they can indicate the presence or absence or severity of sepsis3,4 and can differentiate bacterial from viral and fungal infection, as well as systemic sepsis from local infection. Other potential uses of biomarkers include roles in prognostication, guiding antibiotic therapy, evaluating the response to therapy and recovery from sepsis, differentiating gram-positive from gram-negative microorganisms as the cause of sepsis, and predicting sepsis complications and the development of organ dysfunction (heart, kidneys, liver, or multiple-organ dysfunction). Procalcitonin (PCT) has been proposed as a more specific5,6 and better prognostic 7 marker than C-reactive protein (CRP), although its value has also been challenged. 8 It remains difficult to differentiate sepsis from other noninfectious causes of systemic inflammatory response syndrome, 9 and there is a continuous search for better biomarkers of sepsis.
More than 100-different molecules have been suggested as useful biomarkers of sepsis.10,11 The International Sepsis Forum Colloquium on Biomarkers of Sepsis in 2005 sought to develop a systematic framework for the identification and validation of biomarkers of sepsis. 4 The report from the colloquium proposed that the use of biomarkers could have the potential to alter our view of sepsis, from one of a broad physiologic syndrome to a group of distinct biochemical disorders, and this would help in therapeutic decision making and ultimately improve the prognosis for septic patients and enable us toward designing companion diagnostics assays. Sepsis can be diagnosed or detected with a variety of biomarkers that include proteins, endotoxins, bacterial genetic material, cytokines, and interleukins. These biomarkers can stem from host responses or the bacterial agent directly. Responses from the host include biomarkers that relate to proinflammatory conditions such as CD11b 12 and CD64, 13 which are expressed on the surface of many leukocytes that include monocytes, neutrophils, natural killer cells, granulocytes, and macrophages. There also exist anti-inflammatory biomarkers as a function of immunodepression such as HLA-DR that are used toward early detection of risk for infectious complications. 14 In addition, binding proteins that recognize biomarkers on infectious agents such as lipopolysaccharide binding protein can be used toward sepsis diagnosis. 15 For direct disease correlation, endotoxins or cell wall materials from gram-negative and gram-positive bacteria can be used as biomarkers. 16 While there exists a wide variety of biomarkers, it should be noted that in order for biomarkers to be useful in guiding the treatment of sepsis, there needs to be better standardization of assay methodologies. 17 Current methodologies use immunoassay-based biomarker recognition followed by expensive and time-consuming assays such as enzyme-linked immunosorbent assay (ELISA), flow cytometry, and high-performance liquid chromatography (HPLC) for detection and quantification. New technologies that focus on molecular techniques such as the SespiTest and SeptiFast provide reliable detection windows but are limited by software developments that influence cutoff windows for detection. 17 Time of detection is critical and threshold of detection is important for studying infectious disease progression, and toward this end, we have performed a study in designing a standardized label-free assay for simultaneous and rapid detection of three biomarkers widely used in screening for sepsis and onset of subsequent infection. The long-term goal of this project is the design of platforms suitable for translation as point-of-care companion diagnostic assays.
The three biomarkers considered for this study are PCT, lipopolysaccharide (LPS), and lipoteichoic acid (LTA). PCT was first described for the diagnosis of sepsis in 1993. 18 Since then, it has been widely investigated as a potential biomarker for sepsis and has been used widely in Europe as a biomarker in the management of infection and sepsis. 11 In healthy individuals, serum concentrations of PCT are below 0.1 ng/mL. 19 Concentrations of PCT between 0.15 and 2 ng/mL indicate local infection arising from treatment or clinical procedures.20,21 In response to a bacterial infection, damage-associated molecular patterns and pathogen-associated molecular patterns stimulate cells to produce PCT, which results in a significant increase in serum concentrations. 19 Levels of PCT >2 ng/mL are highly suggestive of systemic bacterial infection/sepsis or severe localized bacterial infection, such as severe pneumonia, meningitis, or peritonitis. LPS and LTA are the main building blocks of the outer leaflets of bacterial cell wall membranes and as such contribute to and are essential for stability and growth. LTA is the gram-positive counterpart of LPS and has similar detection requirements as LPS and can be considered a biomarker for late-stage and severe sepsis. 22 Often, they are not directly exposed to the external environment because many naturally occurring gram-positive and gram-negative bacteria are fitted with a thick polysaccharide capsule. LPS-binding protein (LBP) is an acute-phase reactant that forms a complex with LPS. In human serum, LBP is constitutively present at a concentration of 5 to 10 µg/mL. During sepsis, LBP levels increase to median peak levels of 30 to 40 µg/mL within 24 h. However, it has been shown in many studies that at high concentrations, LBP effectively neutralizes LPS and may be less discriminating than other biomarkers with regard to classification of risk of sepsis and understanding its progression. 23 In addition, LBP binds to both LPS and LTA, which can affect the specificity of the assay designed for classification of the bacterial origin in sepsis. 24 Finally, it has also been shown that diagnosis and analysis of binding proteins for bacterial classification have been limited in success with patients who have undergone surgery recently and in patients who do not have any organ dysfunction from sepsis. 22 These limitations raise the need for direct detection of biomarkers and endotoxins expressed in very low to high concentrations relevant for clinical analysis and classification.
Several strategies have been developed for the detection of PCT, LPS, 19 and LTA in biological samples, such as capillary electrophoresis, HPLC, 25 radioimmunoassay (RIA), 26 fluoroimmunoassay, 27 and ELISA. 28 For LPS and LTA, the process typically requires a primary culture of bacteria from plasma or urine followed by an immunoassay detection. Recent developments have led to the use of Cambrex Limulus Amebocyte Lysate (LAL) tests (Lonza, Basel, Switzerland) and PyroDetect (EMD Millipore, Ballerica, MA, USA). However, with the exception of ELISA tests, these methods have many disadvantages: they employ expensive labeling detection methods, lack specificity, are time-consuming, and require qualified personnel and sophisticated instrumentation. Hence, there is a significant need for alternative methods that are rapid, reliable, and ultra-sensitive. Electrochemical biosensors are promising tools for point of-care testing due to low cost, ease of miniaturization, and possibility of integration with multiarray tools. 29 The use of electrochemical impedance spectroscopy (EIS) in biological applications has also apparently been reported since as early as 1925. 30 One of the first reported uses of EIS in these applications estimated the overall impedance of biological cells 31 and has more recently been used to study the activity of enzymes combining with target molecules.32,33 The technique is now widely reported in bioanalytical applications. EIS-based biosensing has been widely used for biomarker detection from a number if sensor substrates from both simple and complex buffers.
This article demonstrates the design development and testing of a cost-effective and robust, simple to use microdevice technology suitable for translation into clinical diagnostics assays, which can profile multiple biomarkers associated with gram-positive and gram-negative bacteria using EIS.
In this work, we report for the first time an EIS-based nanochannel sensor system for the detection of PCT, LPS, and LTA. To our knowledge, there are no reported electrochemical immunosensors for the simultaneous detection of these three biomarkers, although different biosensors have been developed for the detection of other markers of inflammation, sepsis, and cardiac pathological status. Herein, monoclonal antibodies were used to develop an electrochemical bioassay, coupling nylon polymer nanomembranes with printed circuit board electrodes. Monoclonal antibodies for the three biomarkers were coated individually on gold electrodes on printed circuit boards through the nanoporous nylon membranes. The antibody-modified nanomembrane-integrated gold electrodes were then tested with the three biomarkers in phosphate-buffered saline (PBS) and in human serum. Preliminary clinically relevant testing was performed on spiked patient serum samples to evaluate the performance of the sensor and validate the technology for feasibility toward diagnosis of sepsis or nonsepsis conditions.
Materials and Methods
Sensor Fabrication
Figure 1 shows the nanochannel sensor device formed by integrating a nanochannel polymeric substrate with gold electrodes fabricated on FR-4 substrate. The nanochannel polymeric substrate was aligned over each of the sensing regions as shown in Figure 1 . A microfluidic encapsulant was placed over the sensor, enabling pressure adhesion of the nanochannels over the sensing electrodes. Electrical contacts were interfaced with lead-free solder to connect to the companion electronic reader. The microfluidic channels were flushed with 0.15 M PBS followed by vacuum desiccation prior to introduction of the immunoassay reagents.
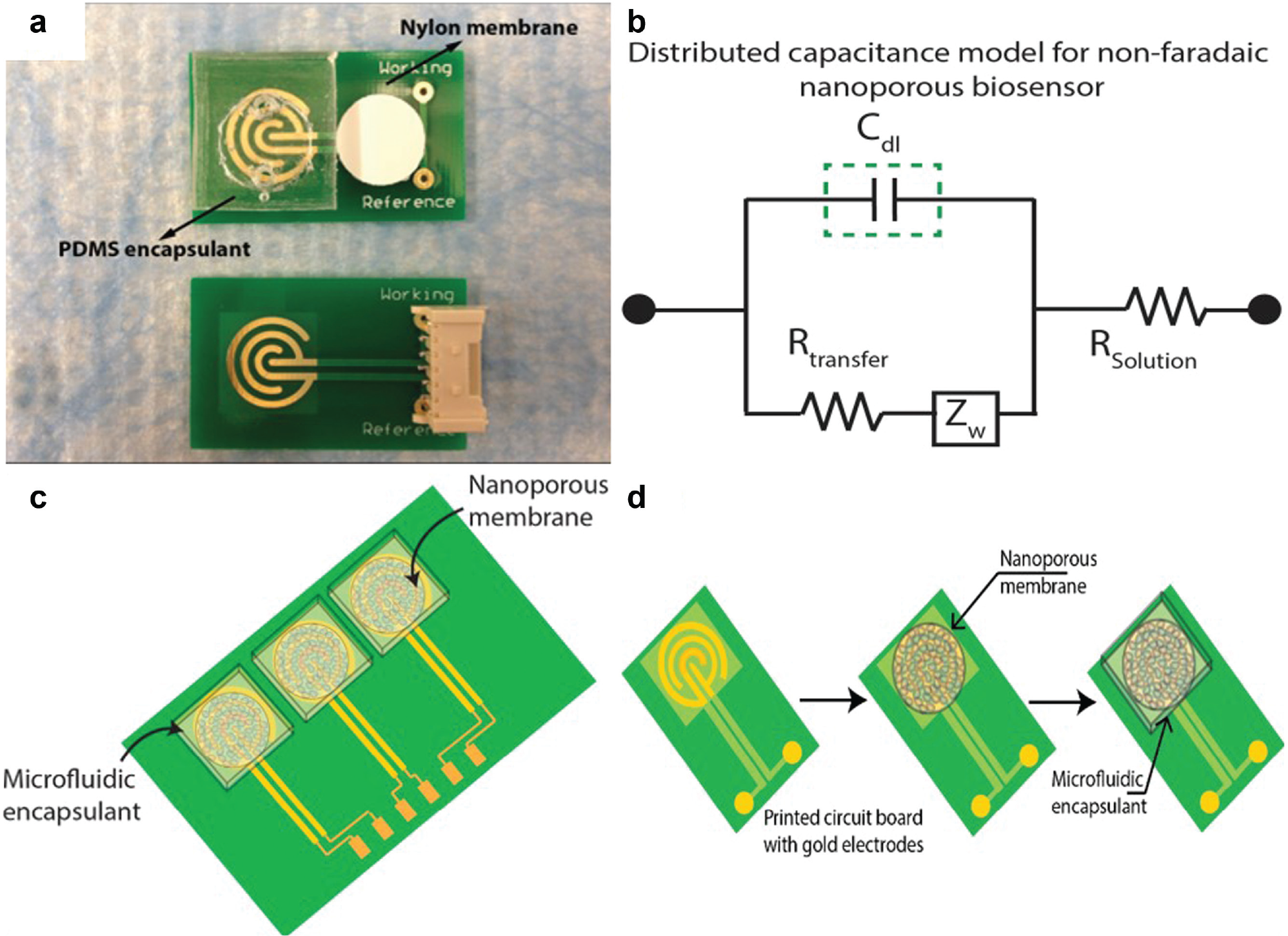
Electrical nanochannel biosensor for detection of biomarkers in a label-free manner: (
Baseline electrical performance of the fabricated sensor system was evaluated using impedance spectroscopy. Dithibois succinimidyl propionate (DSP; Thermo Scientific, Waltham, MA) was used to crosslink the capture antibody on the sensing electrodes. The electrodes were treated with 10 mM DSP dissolved in DMSO for 30 min to create a thiol linkage to gold and an open N-Hydroxy Succinimide (NHS) ester for immobilizing capture antibodies. Highly specific monoclonal antibodies to the three targets PCT, LTA, and LPS (Abcam, Cambridge, MA, USA) were added to the sensing regions. The electrodes were incubated with the antibody solutions for 30 min. After antibody immobilization, the sensing electrodes were treated with 1× SuperBlock (Thermo Scientific) blocking buffer solution to hydrolyze unbound NHS ester groups.
EIS for the Detection and Quantification of Target Biomolecules
The covalent conjugation of biomolecules and cross-linkers on the electrodes aids in the formation of an electrical double layer in the presence of an ionic electrolyte and applied voltage signal. To measure changes occurring in this electrical double layer due to biomolecular interactions (assay functionalization as well as target molecule sensing), we used EIS. The binding of molecules creates a coulombic potential that could be screened as impedance changes, leading to classification of binding effects (specific vs. nonspecific) on the sensing electrodes. Electrical double-layer capacitance is reflective of the columbic potentials developed due to biomolecular binding.
Calibration of Sensor System for Detection of Septicemia Biomarkers
The goal of the sensing system was to leverage the capture antibody-coated sensing electrodes toward the label-free detection of biomarkers in whole blood. Prior to testing with actual samples, the sensor was first calibrated by testing calibration standards of the target biomarkers in human serum solution. The standards for PCT were purchased from ProsPec Bio (Rehovot, Israel). LPS and LTA were purchased from Sigma-Aldrich (St. Louis, MO). PCT was prepared in the concentration range of 10 fg/mL to 100 ng/mL in 0.15 M PBS. LPS and LTA were prepared in the concentration range of 10 ng/mL to 100 µg/mL. The three biomarkers were calibrated for signal changes occurring due to their binding with capture antibody probes on the sensing electrodes. The biomarkers were calibrated individually with their respective capture antibodies on the different sensor systems. The calibration response was performed for a total of n = 5 replicates, and error was calculated as the standard deviation over mean. A polynomial fit was used to establish a correlation between varying concentrations of the calibration standards with respect to impedance measured. As a negative control, each of the sensor arms was also tested for nonspecific signal with the other biomarkers of interest.
Detection of Septicemia Biomarkers from Human Whole Blood
The sensor system was tested for feasibility and accuracy of detection of biomarkers in whole blood to show proof of feasibility for companion diagnostics. A total of n = 5 samples were tested, and the polynomial equation from the calibration study was used to determine the concentration of the biomarkers tested. Whole blood was directly procured from Carter BloodCare (Bedford, TX) in EDTA precoated tubes. The whole blood was tested negative for any diseases or conditions that could be considered harmful or unsafe for the personnel performing the experiments as detailed in this article.
Results
Electrical Circuit Model for Estimating Impedance Changes at the Electrode-Electrolyte Interface
The goal of using a label-free immunoassay is to transduce changes occurring at the electrode-electrolyte interface. The interaction of a biological fluid (PBS or human whole blood) with the electrode surface results in a charge distribution region that is perturbed with any subsequent functionalization as a part of the capture immunoassay. We used a Randles-equivalent circuit to identify the dominant parameter for changes occurring at the electrode-electrolyte interface. Figure 1b shows the Randles-equivalent circuit used in this study. To characterize the dominating circuit element, we measured impedance changes over the range of 1 Hz to 10 KHz. The use of nanochannels results in the formation of a structured double layer that can be modeled as a capacitance. From previous studies, we have identified that the covalent conjugation causes charge perturbations, which can be transduced in the frequency range of 1 Hz to 200 Hz. At frequencies greater than 200 Hz, the solution resistance dominates, which accounts for all molecules conjugated to the electrode as well as buffer constituents. This results in increased nonspecific noise and is not preferred to study specific biomolecular binding events.
Baseline Electrical Characterization of Sensor Surface
The polymeric substrate when integrated on the sensor surface forms nanochannels, inside which functionalization and detection occur. Impedance measurements were performed with 0.15 M PBS and pooled human blood to measure the baseline electrical properties of the electrodes. This measurement is used as a reference to validate cross-linking and immunoassay implementation. DSP cross-linker at a 10-mM concentration was incubated on the sensor for 30 min to achieve complete and effective functionalization. Impedance measurements were recorded and a maximum change in signal was identified at 10 Hz. Figure 2 shows the trend of impedance changes measured for DMSO (control, 168 kΩ) followed by an increase in impedance for DSP conjugation (365 kΩ). The data were collected and plotted for a total of n = 6 sensors. The increase in impedance was attributed to the highly electrical resistive nature of DSP.
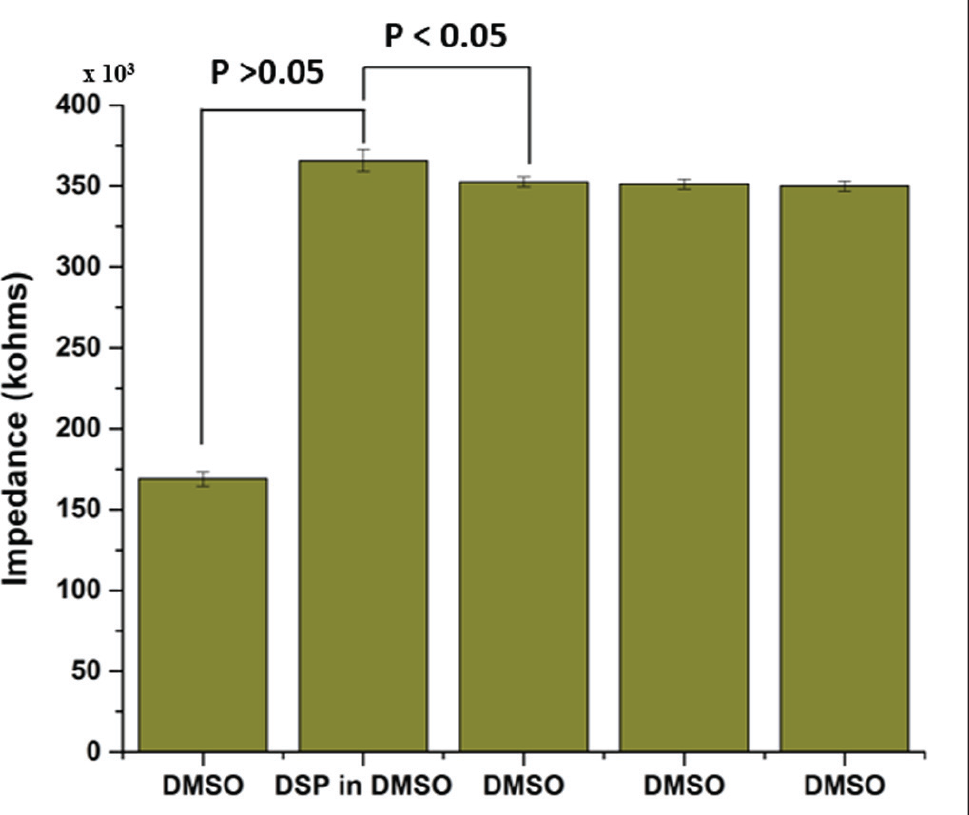
Validation study to assess linker binding. Impedance change indicates immobilization of resistive dithibois succinimidyl propionate (DSP) linker molecule. The 3× DMSO wash was performed to validate conjugation and stability.
Following treatment with DSP, a stable impedance change that was statistically negligible (ΔZ <5%) was observed for consecutive DMSO washes that demonstrated the successful linker conjugation on the sensor surface. For immobilization of capture antibodies, the sensor surface was first washed with 1× PBS to remove any unbound linkers.
The sensor was next incubated with monoclonal antibodies for 30 min at room temperature. Impedance measurements were carried out before and after antibody treatment to identify success in immobilization. The three antibodies of interest were tested on individual sensors. Figure 3 shows the impedance changes observed at a frequency of 10 Hz for the PCT, LPS, and LTA antibody-coated sensors. The decrease in impedance from the PBS assay step to the addition of antibodies demonstrated the immobilization of the capture antibodies. Antibody complexes have been identified to be electrically conductive; hence, the immobilization resulted in the decrease in measured impedance. To further validate the immobilization, the sensor surface was washed 3× with PBS, and impedance measurements were repeated. There were no statistically significant differences between the three washes, p < 0.05.
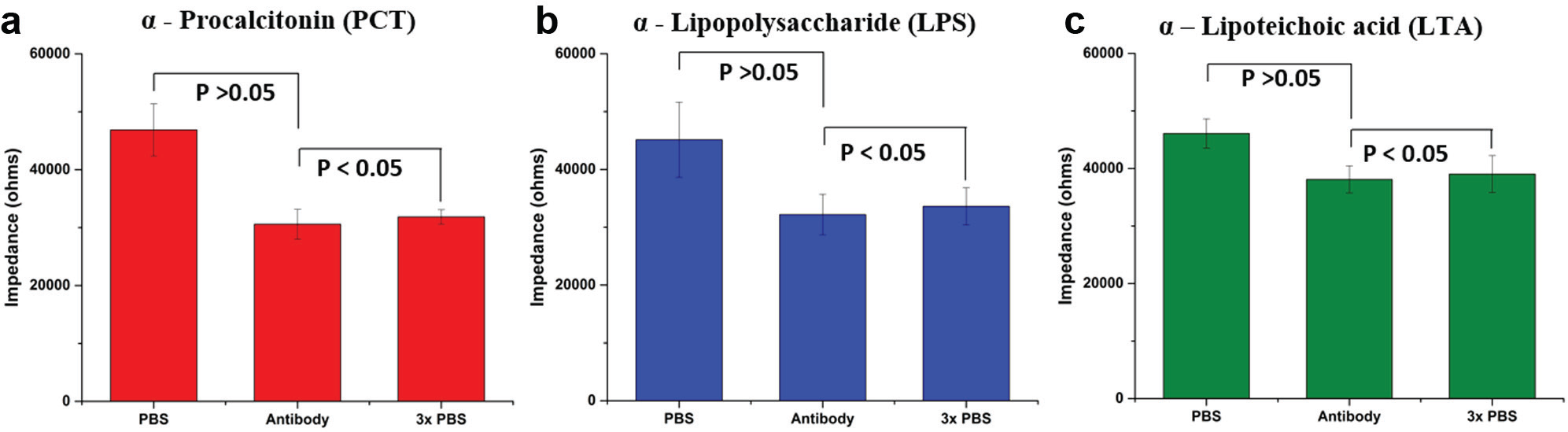
Immobilization of procalcitonin, lipopolysaccharide, and lipoteichoic acid antibodies to dithiobis succinimidyl propionate (DSP)–conjugated surface. The process is validated with impedance measurements at 10 Hz to identify charge perturbations in the electrical double layer as a function of antibody immobilization. Conductive nature of the antibodies conforms to a decrease in impedance observed. The 3× phosphate-buffered saline (PBS) wash was performed to validate stability of binding.
Noise Signal Estimation
To estimate the signal due to background buffer and nonspecific binding, we performed a noise signal classification study using human whole blood tested negative for the biomarkers of interest. Three individual sensors were immobilized with the different antibodies of interest and incubated with the control media (human whole blood) for 30 min. Impedance measurements were carried out for a total of n = 3 replicates. The change in impedance for noise signal was calculated as the difference (ΔZnoise) obtained by subtracting the impedance obtained at PBS wash after SuperBlock (control) from the impedance measured with human whole blood. The impedance measured at the control assay step (PBS wash after SuperBlock) and for the human whole blood for the three different antibodies is shown in Figure 4 . For a measured signal to be classified as specific signal, the specific signal threshold (SST) was defined as three times ΔZnoise (signal-to-noise ratio = 3) and hence estimated.
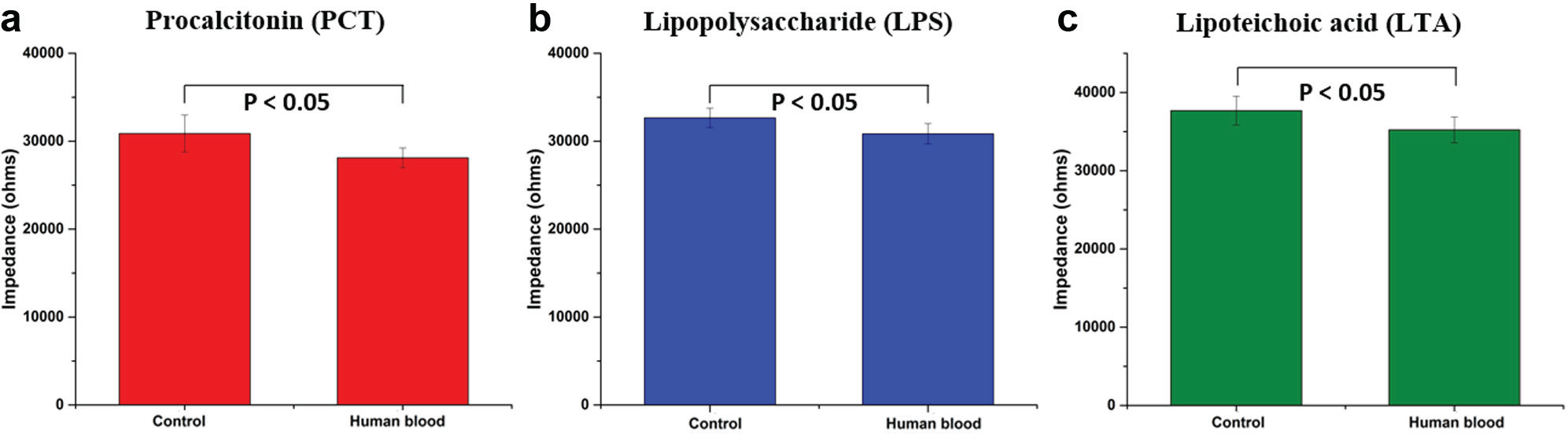
Performance of nanochannel biosensor. Noise signal was estimated as the change in impedance observed for an antibody-coated sensor (control) when treated with human blood.
Identifying the Impedance-Concentration Relation for Sepsis Biomarkers
A serial dilution of each target biomarker of interest was prepared in pooled human serum (Innovative Research, Novi, MI) over the range of 0.01 fg/mL to 100 ng/mL. The aliquots were prepared individually and used immediately for the calibration response study to identify the analytical parameters of the sensor device. The antibody-saturated sensor surface was treated with SuperBlock buffer to seal off any unbound linker sites. A PBS wash was performed following this, and the impedance measurements were recorded. The impedance measurement at this step was defined as the baseline/control value (ΔZBaseline).
All impedance changes for the subsequent testing with the target biomarker dilutions were calculated with reference to the baseline/control impedance. Figure 5 shows the calibration response for the different biomarkers over the concentration range of 0.01 fg/mL to 100 ng/mL. Error bars represent standard deviation over mean for a total of n = 3 replicates.
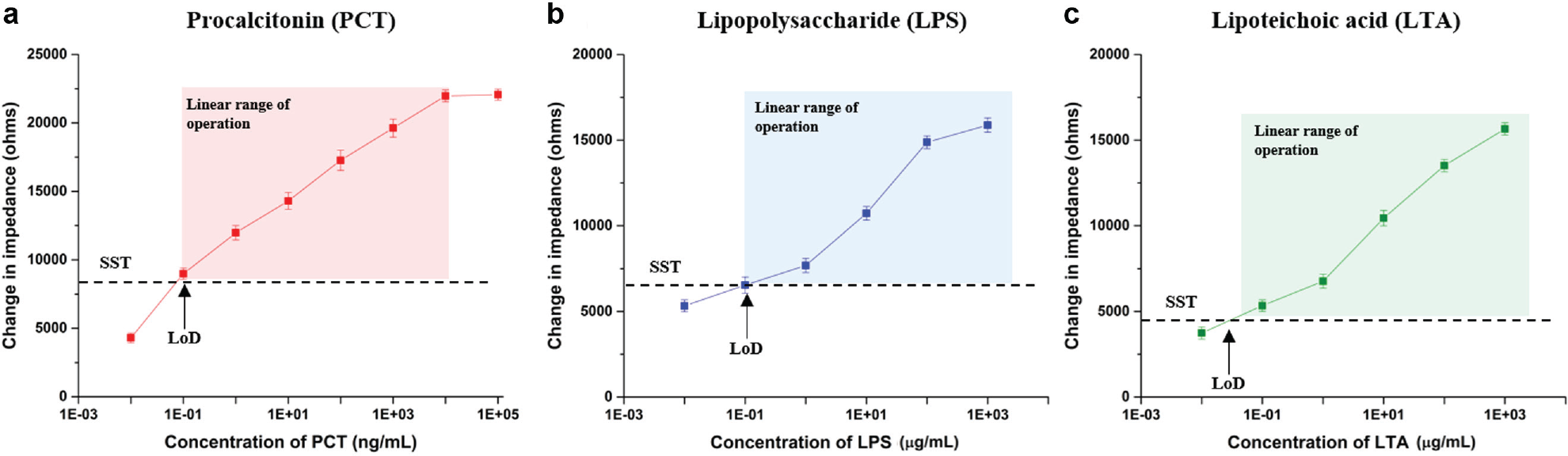
Calibration dose response correlating concentration of individual biomarkers with change in impedance observed at 10 Hz. Specific signal threshold (SST) was defined as three times the noise signal. LoD, limit of detection.
The sensor for PCT showed a linear range of impedance change between 8.9 kΩ and 22 kΩ for the concentration range of 0.1 ng/mL to 10 µg/mL. At doses <0.1 ng/mL PCT, the change observed was 4.3 kΩ (<SST), and the measured signal could not be classified as a distinguishable change due to binding events. The limit of detection for the PCT sensor was defined as 0.1 ng/mL. For concentrations >10 µg/mL, the impedance signal showed saturation and hence was defined as the upper limit of detection for PCT biomarker.
LPS and LTA are indicative of gram-negative and gram-positive bacteria and its respective role in infection. The sensor showed the ability to detect LPS and LTA in the concentration range of 1 µg/mL to 1000 µg/mL. For concentrations >1000 µg/mL, the impedance signal showed saturation and hence was defined as the upper limit of detection for LPS and LTA biomarkers. The impedance-concentration correlation was defined through a calibration equation that was used for the estimation of these biomarkers in human whole-blood test samples.
Detection of Biomarkers in Human Whole Blood
A blinded cohort of four samples was prepared with different concentrations of the biomarkers spiked in human whole blood. PCT was spiked in the clinically relevant concentration range of 5 to 50 ng/mL. LPS and LTA were spiked in the concentration range of 2 to 30 µg/mL. The sensors were assembled in a multiplexed configuration as shown in Figure 1c with each arm coated for one biomarker. The blood was added to the sensor and incubated for 15 min before impedance measurements were performed. The calibration equation from the response study was used to determine the concentrations of the different biomarkers in the sample. Figure 6 shows a plot of the concentrations estimated from the test samples.
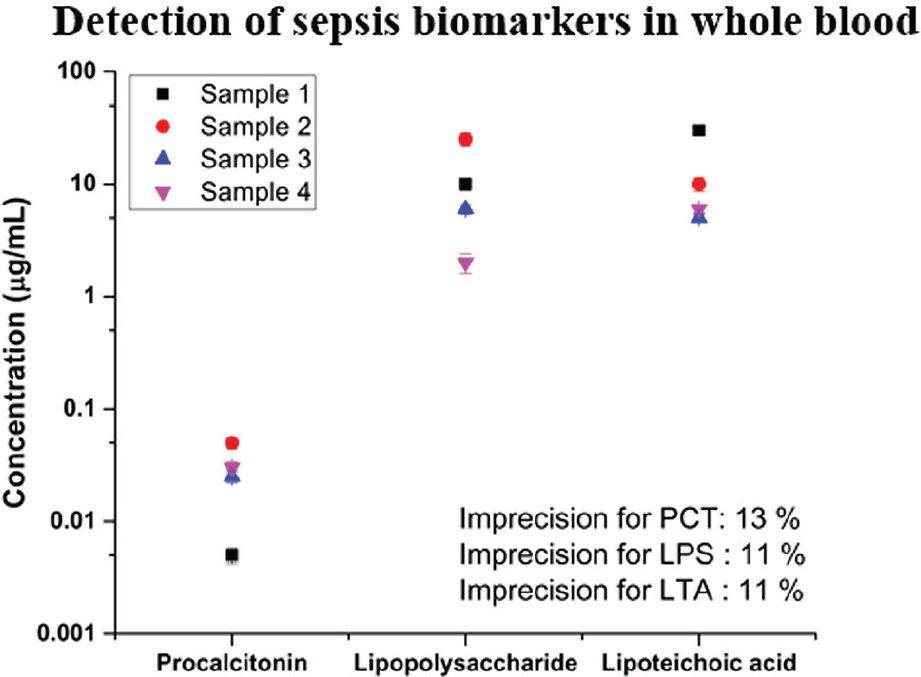
Quantification of sepsis biomarkers procalcitonin (PCT), lipopolysaccharide (LPS), and lipoteichoic acid (LTA) in human whole blood. Sensor demonstrated robust detection with imprecision estimated as error over mean from n = 3 replicates.
Discussion
The common approach for identifying and diagnosing infectious diseases and their primary cause depends heavily on optical immunoassays and cell cultures. These methods demand a laboratory setting and experienced personnel to initiate tests. The requirements that traditional methods pose limit the ability to screen every patient for risk of infection and diagnose it in real time at the time of occurrence. Companion diagnostics are highly important in settings where there are a number of patient scenarios who can present positive for infection. While there exist a number of standard immunoassay techniques and detection methods, there are larger schemes of issues in miniaturizing transduction methods as well as making them effective for point-of-care testing.
Quantification of biomarkers in clinical or laboratory settings currently depends on chemiluminescence assays. For rapid testing, the detection has to be performed directly in the sample of interest (urine, blood, serum, etc.) without the requirement for any purification or processing. Reporter labels for biomarkers are limited by nonspecific and interfering biomolecules in body fluids such as urine, saliva, and whole blood. Many alternative techniques to chemiluminescence assays have been reported. One such is EIS, which has been demonstrated as a rapid and robust technique for screening biomarkers based on affinity binding. Traditional EIS uses redox reporters and labels, which transduce the electrical signal based on specific biomolecular interactions. In summary, traditional EIS and ELISA use a label that critically limits the sensitivity and selectivity of the sensing system when testing with complex media samples, making it a challenge for implementation as companion or point-of-care diagnostics.
EIS based on electrical double-layer screening is a high-sensitivity and high-specificity method that can be used for label-free biomarker detection. The label-free operation is achieved by using a nonfaradaic detection modality that depends on electrical double-layer screening for detecting biomolecule binding events. In this article, we demonstrate the ability to detect biomarkers in pooled human serum and human whole-blood samples with enhanced sensitivity and selectivity. Our method provides a promising approach for screening a range of biomarkers directly in complex media in a rapid and robust manner. This can significantly help in the early detection and diagnosis of infectious diseases, which have sudden onset and rapid progression dynamics.
Infectious diseases are also well known for their ambiguity as the onset and development can be different organisms (bacterial and inflammation). Toward this, a multiplexed testing method would help in identifying the source of infection (gram positive or gram negative) as well as understand the response of the body to the infection (inflammation, immune response, organ dysfunction, etc.). Standard testing mechanisms are limited by the number of tests required to quantify different biomarkers as well as the sensitivity of few biomarkers expressed in low concentrations. Novel transduction techniques such as EIS, which can help quantify low-analyte concentrations with complex samples, can improve the throughput of analysis as well as provide information required for understanding the disease. The sensor system we have designed and developed focuses on multiplexed detection to quantify a panel of biomarkers, which can help understand disease dynamics and design a treatment strategy well suited for the specific case tested.
We have demonstrated the ability of the nanochannel sensor device to detect PCT over the concentration range of 0.1 ng/mL to 10 µg/mL, as well as LPS and LTA over the concentration range of 0.1 µg/mL to 1000 µg/mL. The sensitivity in detection is achieved through (1) use of nanochannels to overcome charge screening effects and (2) electrical double-layer screening for detecting specific binding of a target biomarker to specific antibodies. The integration of the insulating nanowells to the metal microelectrodes resulted in the formation of a charge screening system with reduced noise and interfering electrical signals from the bulk solution media. Hence, this helped in the measurement of electrical impedance changes with a high signal-to-noise ratio for ultra-sensitive quantification.
In the design of the nanochannel sensor, a polymeric membrane was embedded on the base metallic gold electrodes. This architecture resulted in the generation of a high-density array of nanoscale-confined spaces. These nanoconfined spaces have been engineered toward achieving macromolecular crowding-driven nanoconfinement. To support this, there is a compelling argument in the realm of biophysical understanding of various types of biomarkers and their structures. There exists a crowded environment inside a cell structure, the cytoplasm, which is typically different from the dilute solutions that are generally used in in vitro studies of proteins, and there exists a school of thought that states that this may significantly affect the behavior of the proteins. The high concentration of proteins, nucleic acids, and complex sugars in the cells has various energetic consequences and size constraints on the smaller molecules; the effect of these bigger structures is known as macromolecular crowding.34–36 The binding affinities and the rates of self-assembly can change by orders of magnitude as a result of this phenomenon. Crowding is therefore a very important factor when we are trying to perform an in vivo study in vitro. Thus, it is imperative to use this idea in a device that would be used to measure the accurate concentration of proteins and biomarkers as it is measured from direct body fluids.
The charge screening offered by the nanochannels helped in the implementation of specific electrical double-layer screening for the detection and quantification of protein and biomarker binding. The electrical immunoassay was implemented at the electrode-solution interface. By screening electrical impedance changes at this height from the electrode surface, specific binding events occurring due to target biomarker interaction with the capture probes (antibody) were detected. The modulus of impedance was used for subsequent conversion to meaningful protein and biomarker concentrations. From the circuit modeling, we identified that the dominant component of impedance was the double-layer capacitance ( Fig. 1b ). The changes in Cdl correlated to binding events occurring in the electrical immunoassay at the Debye length of interest. This confirmed the nonfaradic nature and the specific detection of binding events in the electrical Debye length.
The impedance change measured from a sample of human whole blood (tested negative for target biomarkers) was used as the noise signal of the nanoelectronic sensor device. As the sensor was an electrical system, a highly competitive signal-to-noise ratio of 3 (SNR = 3) was defined for the system. SST was defined as the impedance change that was three times the noise signal. The lowest dose concentration that the sensor detected was 0.1 ng/mL for PCT, 1 µg/mL for LPS, and 1 µg/mL for LTA. The sensor demonstrated a linear range of operation up to 10 µg/mL for PCT and up to 1000 µg/mL for LPS and LTA biomarkers.
To validate the applicability of the sensor for companion and point-of-care diagnostics, we spiked varied concentrations of the three target biomarkers in four samples of human whole blood. The concentration of the markers was estimated using the calibration response curve and repeated for a total of n = 3 replicates. This confirmed the ability of the sensor to perform robust detection in direct body fluids. Existing biochemical assays such as immunoassays, Limulus amebocyte lysate tests, and Western blotting are widely used in the diagnosis of sepsis. Quantification of the biomarkers of interest is further achieved through HPLC, ELISA, and so on, assisted by RNA amplification methods for improving sensitivity. The major drawback associated with existing methods are long preparation and incubation times followed by expensive techniques for quantification. In a number of scenarios, the cost of testing limits the need for rapid and continuous monitoring. Continuous monitoring can provide significant insights into the onset of sepsis that can lead to design of better treatment strategies, which is currently missing. The sensor system demonstrated in this article has the ability to quantify and report quantified biomarker concentrations with a total preparation time of less than 1 h, followed by immediate quantification and reporting (<15 min). The immediate clinical application of this technology is to detect various biomarkers in multiplexed configurations for a defined cohort of affected and healthy individuals to validate performance toward point-of-care testing and companion diagnostics.
Footnotes
Acknowledgements
We thank Dr. Thomas Barrett and Dr. Steven Kazmierczak for their inputs and feedback toward clinical relevance of this immunosensor assay. We also acknowledge Justin Wong, Bilal Quadri, Daniel Rodricks, Vikramshankar Kamakoti, and Arjun Menta for their contribution toward experiments and data compilation.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: financial support from UT Dallas.
