Abstract
This article is a review of current and emerging methods used for prenatal detection of chromosomal aneuploidies. Chromosomal anomalies in the developing fetus can occur in any pregnancy and lead to death prior to or shortly after birth or to costly lifelong disabilities. Early detection of fetal chromosomal aneuploidies, an atypical number of certain chromosomes, can help parents evaluate their pregnancy options. Current diagnostic methods include maternal serum sampling or nuchal translucency testing, which are minimally invasive diagnostics, but lack sensitivity and specificity. The gold standard, karyotyping, requires amniocentesis or chorionic villus sampling, which are highly invasive and can cause abortions. In addition, many of these methods have long turnaround times, which can cause anxiety in mothers. Next-generation sequencing of fetal DNA in maternal blood enables minimally invasive, sensitive, and reasonably rapid analysis of fetal chromosomal anomalies and can be of clinical utility to parents. This review covers traditional methods and next-generation sequencing techniques for diagnosing aneuploidies in terms of clinical utility, technological characteristics, and market potential.
Clinical Utility
Chromosomal abnormalities in the fetus constitute one of the leading causes of stillbirth and births of infants with malformations. 1 One type of chromosomal abnormality is aneuploidy, which is defined as the fetus having an extra or missing chromosome. 2 Aneuploidies are caused by errors during the meiotic cellular division process. During normal meiosis, haploid sperm and egg cells containing one copy of each chromosome are formed from the diploid parent cells. Upon fertilization, a diploid zygote cell is formed, containing two copies of each chromosome, one from each parent. However, errors during meiosis can result in the fetus having an extra or missing chromosome ( Fig. 1 ).2–5 A number of risk factors can increase the probability of chromosomal aberrations in pregnancy, maternal age being the most important.2,3 Other risk factors are a history of infertility and a family history of chromosomal disorders. 1
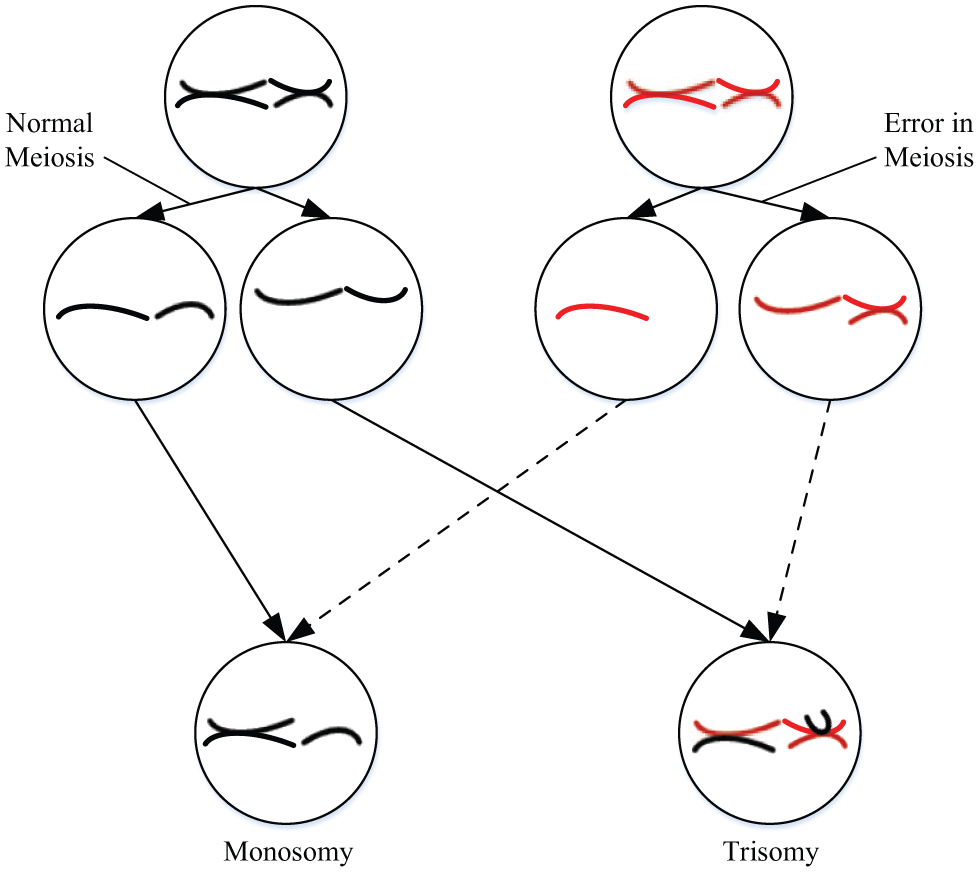
Chromosomal aberrations due to meiosis. Humans are diploid, with a total of 46 chromosomes, or 23 pairs of chromosomes per cell. During meiosis, gamete cells (sperm and egg cells) are formed that are haploid, each containing 23 single chromosomes per cell. Upon fertilization, the maternal and paternal haploid gamete cells combine, resulting again in a diploid cell with 23 pairs of chromosomes, or 46 chromosomes in total. Errors in the meiotic segregation process, as shown in the figure, lead to chromosomal aberrations.2–5
Down syndrome (DS), which is associated with mental retardation, is the most commonly occurring aneuploidy, with an estimate of 250,700 people living with DS, corresponding to a prevalence of 8.27 per 10,000 individuals. 6 DS is due to the presence of an extra chromosome 21 (trisomy 21). Chromosomal analysis of the fetus is the only way to definitively diagnose DS. 6 Trisomy 18, known as Edwards syndrome, is the next most common aneuploidy, occurring in 1 in 6000 to 8000 live births. 7 The probability of survival after birth with this aneuploidy is extremely low, with only about 10% of infants living more than a year.
Trisomy 13 or Patau syndrome also greatly reduces life expectancy and occurs 1 in 10,000 births. 8 Forty-four percent of children with this condition die within a month of birth. Turner syndrome (TS) is a serious sex chromosomal abnormality, in which most female patients are 45 X, meaning that all or part of an X chromosome is missing. Spontaneous abortion of TS fetuses is very common. Only 1% of TS pregnancies result in live birth, but malformations will make a normal life difficult. 6
In the 1960s, only women older than 35 years were considered a high-risk group for fetal chromosomal abnormalities. Links between maternal age and DS were made as early as 1933. 9 During the 1970s, studies established a correlation between maternal serum α-fetoprotein and pregnancies affected by DS. This allowed for prenatal screening of DS and was recommended to women who were at high risk for carrying such aberrations. In addition, this extended the screening age of women to those younger than 35 years. 10
It is crucial to detect fetal chromosomal aberrations early in pregnancy, so that the parents and medical team can discuss medical options. 6 If an abnormality is identified, the parents should seek genetic counseling to explore options that may include the decision to terminate the pregnancy. Due to the significant consequences of both false-positive and false-negative diagnosis, tests for fetal chromosomal abnormalities need to have close to 100% sensitivity and specificity. Chromosomal abnormalities need to be detected early on during pregnancy, since abortion is a time-sensitive issue.
However, the current process requires multiple tests with long turnaround times. After high-risk individuals are identified, a confirmatory diagnostic test is usually requested. Such confirmatory tests use sample acquisition methods that are invasive and may cause fetal harm. Once the sample is obtained, the turnaround time for the current gold standard, karyotyping, is at least 2 weeks. Given that these tests are typically performed during the second trimester, the duration of time to receiving results leads to potentially controversial decisions, such as abortion. Cytogenetic analysis of fetal cells conventionally requires amniotic fluid, chorionic villi, or fetal blood, which are sampled through amniocentesis (AMC), chorionic villus sampling (CVS), and cordocentesis, respectively. 11 These sample acquisition methods, described below, are invasive and pose a risk of miscarriage. 10
Amniocentesis
AMC is usually performed between 16 and 28 weeks of gestation. 12 A needle is used to aspirate approximately 20 mL of amniotic fluid under the guidance of ultrasound imaging. 12 The risk of abortion in AMC is 1% but increases considerably if performed prior to 16 weeks of gestation. 13 A clinical study showed a significant difference in total fetal losses (7.6% vs. 5.9%) as well as in amniotic fluid leakage (3%–5% vs. 1.5%) between the early AMC group (performed <13 weeks) and the mid-AMC group (performed >15 weeks). 14
Chorionic Villus Sampling
CVS sampling from the developing placenta is usually performed between 10 and 12 weeks of gestation. 15 There are two approaches for CVS, transabdominal and transcervical, which are proven to be equally safe for first-trimester diagnosis. 15 Tissue samples are obtained using a catheter attached to a syringe under the guidance of real-time ultrasonograms. 16 One advantage of CVS is that it enables an early and definitive chromosomal analysis. 17 While transabdominal CVS has been noted as being comparable to AMC, transcervical CVS poses a greater risk of fetal loss.13,17 CVS also has an operator-dependent learning curve, requiring about 100 to 400 CVSs before the procedure is considered mastered. 13
Cordocentesis
Cordocentesis sampling of fetal blood from the umbilical vein is usually performed between 16 and 37 weeks of gestation. 12 In cordocentesis, a spinal needle is inserted near the placental origin of the umbilical vein, and blood is drawn under the guidance of ultrasound. 18 Hemoglobin patterns are analyzed via Kleihauer testing, which measures the amount of hemoglobin transferred into the mother’s bloodstream from a fetus, and are performed to confirm the absence of maternal blood contamination. 12 While this procedure is more difficult to perform than AMC, it presents a decreased risk for miscarriage. 12
Novel approaches are needed that enable confirmatory diagnosis of fetal chromosomal aberrations early on in pregnancy (i.e., during the first trimester) based on minimally invasive sampling that does not introduce a risk of miscarriage.
Technical Product Analysis
A diverse range of screening and diagnostic tests exist for the detection of prenatal chromosomal aberrations. Maternal serum screening (MSS), nuchal translucency (NT), karyotyping, fluorescence in situ hybridization (FISH), and quantitative fluorescence polymerase chain reaction (QF-PCR), as described below, represent common methods for prenatal screening. In addition, the emergence of next-generation sequencing (NGS) has provided an additional platform for diagnosing prenatal chromosomal aberrations.
Maternal Serum Screening
Maternal serum screening for DS is generally offered by all obstetricians to their pregnant patients, irrespective of age. 19 It is a voluntary, minimally invasive blood test that measures specific biomarkers present in the serum of a pregnant woman to determine the risk of trisomy 21. 20 Since MSS only provides the patient with a risk assessment and does not provide a definitive diagnosis, 21 the patient may opt for more invasive screening methods, such as cytogenetic tests involving sample acquisition via CVS and AMC 19 to confirm a positive result. 21
The biomarkers measured during MSS are pregnancy-associated plasma protein A (PAPP-A), free β–human chorionic gonadotropin (β-hCG), α-fetoprotein (AFP), unconjugated estriol (uE3), and dimeric inhibin A (DIA) ( Table 1 ). 20 After the first and second trimesters, MSS is performed and the test results are combined and provide the patient with a risk assessment. 20
Biomarkers Measured in Maternal Serum Screening and Their Reference Values.
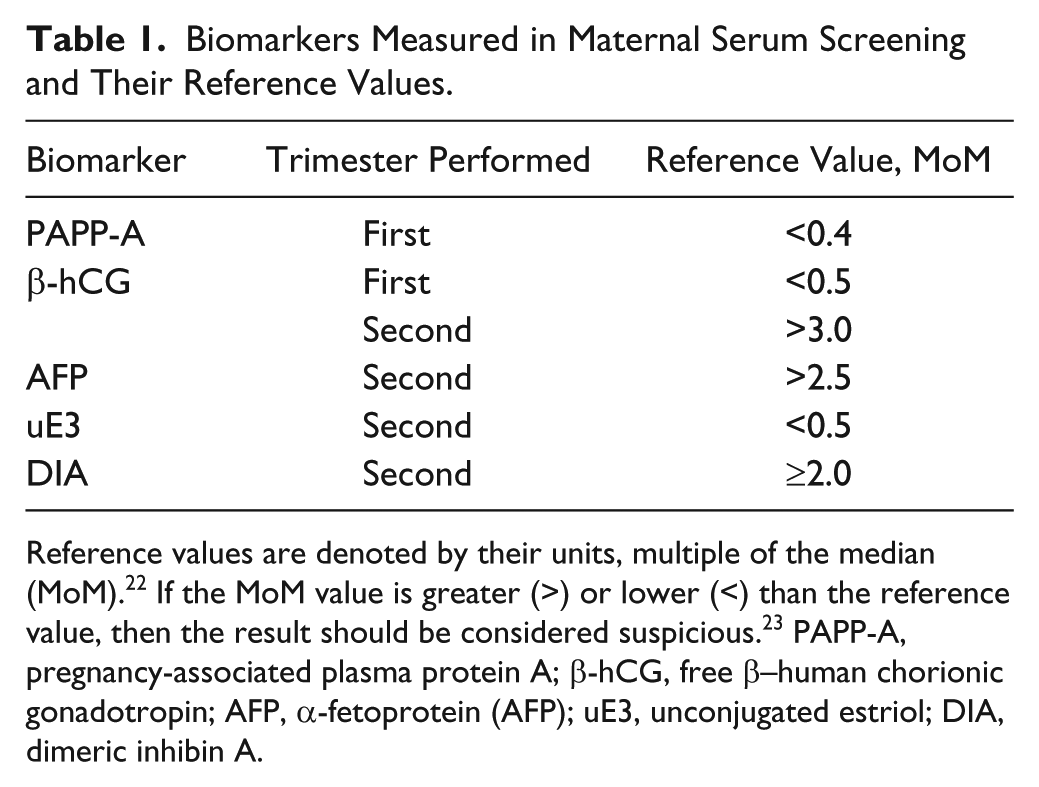
Reference values are denoted by their units, multiple of the median (MoM). 22 If the MoM value is greater (>) or lower (<) than the reference value, then the result should be considered suspicious. 23 PAPP-A, pregnancy-associated plasma protein A; β-hCG, free β–human chorionic gonadotropin; AFP, α-fetoprotein (AFP); uE3, unconjugated estriol; DIA, dimeric inhibin A.
Currently, three main types of MSS tests are offered ( Table 2 ), which measure different biomarker panels at different times during pregnancy. In general, in the second trimester of pregnancies with DS fetuses, AFP and uE3 levels are decreased, whereas β-hCG and DIA levels are increased.19,22 The diagnostic accuracy of these serum marker panels now reaches 90% sensitivity and 95% specificity. 22
Maternal Serum Screening Test Types.
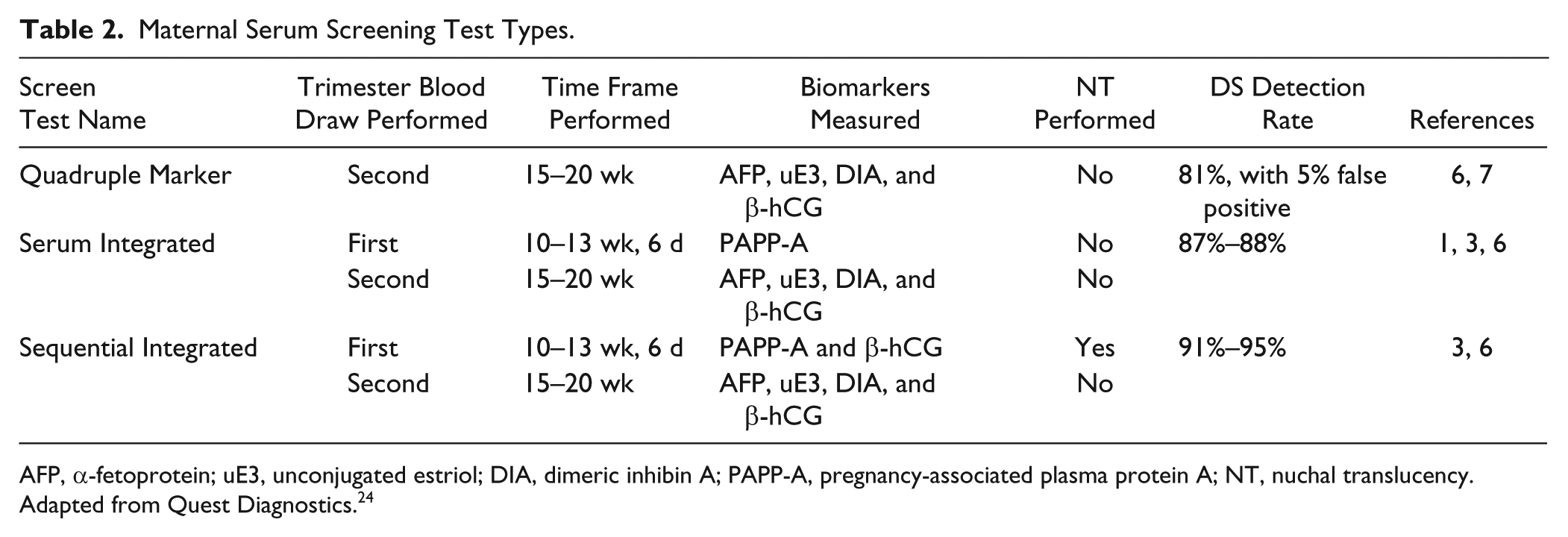
AFP, α-fetoprotein; uE3, unconjugated estriol; DIA, dimeric inhibin A; PAPP-A, pregnancy-associated plasma protein A; NT, nuchal translucency. Adapted from Quest Diagnostics. 24
Quest Diagnostics uses different testing principles to measure the respective biomarkers in MSS, including immuno-chemiluminometric assays (ICMAs) and enzyme immunoassays (EIAs). 24 Both types of immunoassays rely on antigen-antibody interactions. ICMAs are used to quantify the concentrations of AFP, PAPP-A, uE3, and hCG. 24 These assays use chemiluminescent substrates, such as amino-butyl-ethyl-isoluminol (ABEI), to label antibodies to specific biomarkers. 25 For example, rabbit anti-AFP IgG-ABEI is used as the antibody to quantify the amount of AFP in a serum sample. Dimeric inhibin A is the only biomarker measured by EIA. 24 Enzyme immunoassays use a pair of monoclonal antibodies against synthetic epitopes of the two subunits of DIA. 26
Nuchal Translucency Test
A nuchal translucency test is a noninvasive, inexpensive ultrasound test that measures the length of the fetal nuchal tissue, seen in a lateral view of an ultrasound, using a caliper function on the machine. 27 A fetus with a chromosomal abnormality will have an increased NT measurement. Among other chromosomal abnormalities, the NT measurement is useful for detecting DS in the late first trimester of pregnancy. 28 Between 11 and 14 weeks of pregnancy, the fetus is developed to the point at which a demarcated fluid-filled space behind the fetal neck is visible. This space is present in all fetuses, and by using the caliper function of the ultrasound, the space beneath the skin can be evaluated in comparison with the crown rump length, which is an indicator of fetal age.27,28 However, a chromosomal abnormality causes an abnormal amount of fluid to collect in this region of the fetus because the baby’s lymphatic system has not yet fully developed, thereby increasing the NT measurement. 29
The NT test is a diagnostic test that is useful in providing the mother and her physician with a better picture of possible future complications. Multicenter studies have shown that up to 80% of fetuses born with aneuploidy have increased NT measurements. The NT test is also useful for detecting chromosomal aberrations other than DS and can differentiate between chromosomal diseases based on morphological characteristics. 30
The probability of miscarriage increases from 1.6% for NT readings of 3.0 to 3.5 mm to 20% with an NT reading ≥6.5 mm.21,31 Subsequent to the test, counseling of the parents is required and other risk factors can be used to calculate the probability of an uncomplicated pregnancy. 29 In most cases, the mother will have an echocardiogram, AMC, and/or CVS test performed for a better idea of her diagnosis.31,32
Detection rates for a NT test are 70% with a 5% false-positive rate. 27 A false-positive rate means the percentage of cases in which the test shows a positive result even in the absence of aneuploidy. A perfect test result should have a 0% false-positive rate. The NT test has improved over recent years, which has enabled a 20% decrease of the false-positive rate. 33 Detection rates are improved when the NT test is used in combination with other prenatal tests, according to the National Society of Genetic Counselors’ guidelines of prenatal screening and diagnostic testing options. 21 Common combined tests include the combined first-trimester screening (FTS), which consists of an NT and CVS test, and the integrated combined screening (ICS), which consists of an NT and MSS test. 21 An ICS is usually performed for older mothers who have increased DS pregnancy risk, since the ISC has the highest prediction rates for DS (94%–96%) and trisomy 18 (91%–96%). 21 The mother must wait, however, until the second trimester for ICS results but can alternatively have a stepwise sequential or contingency test conducted during the first trimester with lower prediction rates. Stepwise sequential testing also combines an NT and MSS test and evaluates these results in comparison to the mother’s age. This allows physicians to determine a risk level, high risk or low risk, in which high-risk patients are offered diagnostic testing and low-risk patients are continually monitored through the second trimester. Similarly, contingency testing uses a process that creates a risk profile based on NT, MSS, and age to categorize mothers into high-, medium-, and low-risk groups. This differs from stepwise sequential testing in that the medium-risk group is evaluated through the second trimester, where high-risk patients are once again identified for diagnostic testing. 21
Karyotyping
Karyotyping is the current gold standard for detecting fetal chromosomal aberrations. The karyotype refers to the complete set of chromosomes in an organism. Fetal karyotyping is performed on the samples retrieved via CVS, AMC, or cordocentesis. In AMC and cordocentesis, fetal cells first need to be separated from the fluid by centrifugation. The fetal cells are then inoculated and cultured for 10 to 11 days at 37 °C, followed by addition of colchicine solution, a mitosis blocker to arrest cells in the metaphase of mitosis. After incubation and centrifugation, the medium is replaced with a hypo-osmotic solution, which leads to cell lysis. Subsequently, the sample is fixed using a methanol-acetic acid fixative, and a few drops of the suspension are placed on a microscope glass slide. The slide is dried, and the chromosomes are stained with Giemsa dye and then can be viewed under a light microscope for the cell’s karyotype. 16 This technique of producing a visible karyotype through staining chromosome is called Giemsa banding, or G-banding. Karyotyping yields accurate identification of fetal aneuploidy ( Fig. 2 ), structural rearrangements, and duplications and/or deletions of size greater than 5 Mb. 34
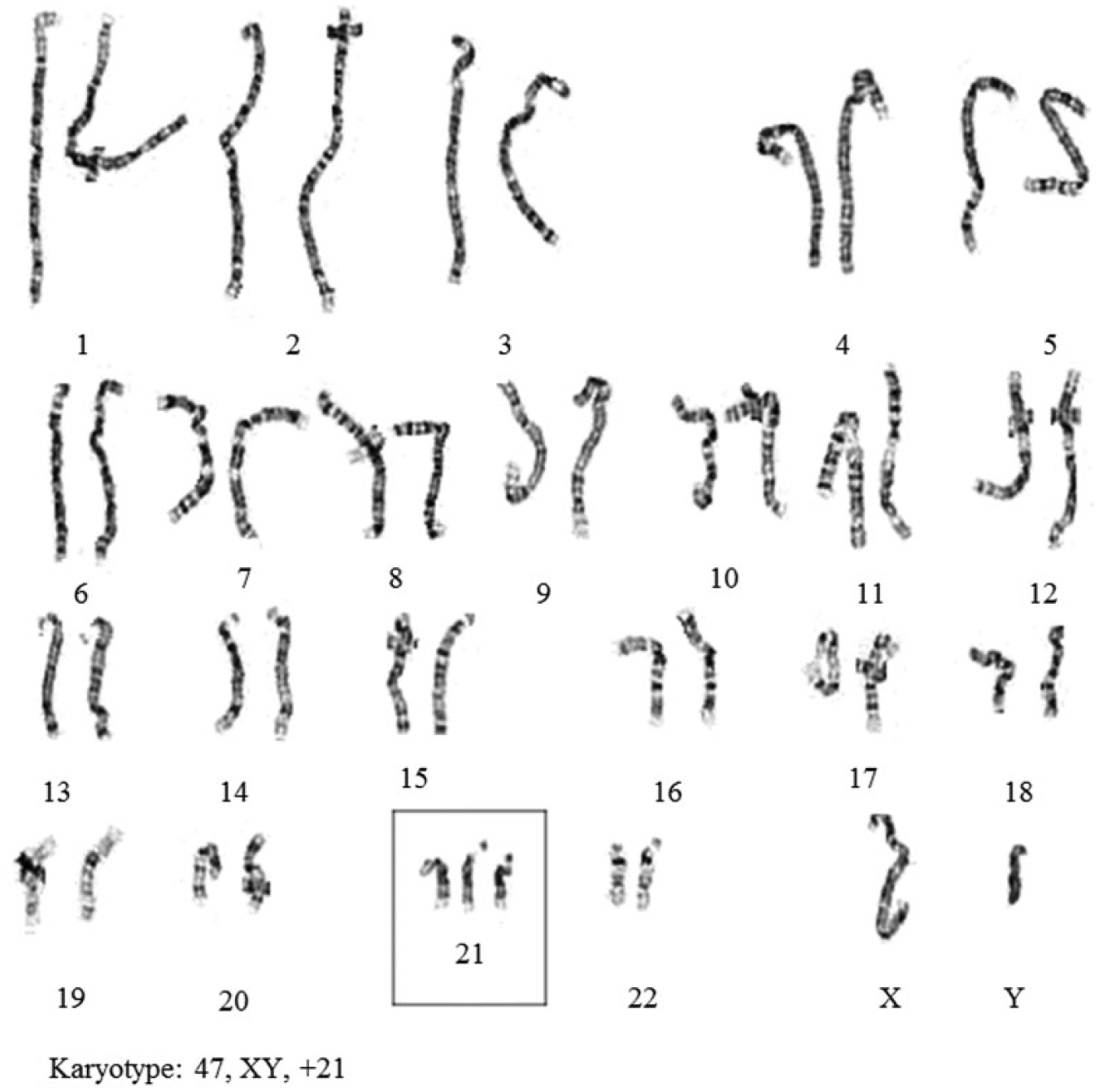
A karyotype of an individual with DS (Trisomy 21), with an additional copy of chromosome 21. 11
The cytogenetic analysis diagnosis rates for CVS and AMC are 97.8% and 99.4%, respectively. 17 In a retrospective study, 100% sensitivity was observed for cases using cordocentesis. 12 Karyotyping using amniotic fluid is considered the “gold standard” in fetal aneuploidy testing due to high sensitivity and relatively low risks. 34 However, the test requires fetal cells to be cultured, which causes some problems, including the low growth potential of the cytotrophoblast cells and the occurrence of maternal cell contamination (0.1%–0.2%).13,16 Promoting cell growth with hormone-supplemented medium or practicing caution to avoid contamination can help circumvent these problems. Another potential source of inaccurate results is mosaicism (prevalence of 0.1%–0.2%), which is the presence of a different genetic makeup of cells in one individual. Mosaicism is detected 10 times more frequently in CVS samples than in AMC samples. This could lead to both false-positive and false-negative results. 13
The major concern with chromosomal analysis is the invasive sampling method. Alternative sources of fetal samples, including fetal urine, pleural fluid, ascetic fluid, cystic hygroma, facial cyst, lung cyst, and pericardial fluid, have been shown to result in successful karyotyping in a single study. The study suggested that the “alternative” samples may be used in place of conventional fetal fluids when the recommended method is not feasible. 35
Chromosomal Microarray Analysis
Chromosomal microarray analysis (CMA) is a high-resolution array-based comparative genomic hybridization method that allows for identification and analysis of the chromosomes for a large number of genetic aberrations, including submicroscopic abnormalities that are below the resolution of conventional screening methods. CMA identifies copy number variations (CNVs), an imbalance of a number of chromosomes due to deletion or addition of sections of DNA. CMA is also known as cytogenetic microarray analysis that describes two other techniques, comparative genomic hybridization (aCGH) and single-nucleotide polymorphism (SNP) arrays. Both of these arrays use CNVs, but they identify different types of chromosomal aberrations. aCGH compares the genome of two individuals to identify genome-wide CNVs for relatively large deletions or duplications that are less than 100 kb, but it cannot detect triploidy. SNPs allows for genotyping based on allele frequency. SNPs provide even more genome coverage and better detection of CNVs compared with aCGH. These microarrays are also known to detect homozygosity, heterozygosity, triploidy, and uniparental disomy. 36
CMA has been investigated as a prenatal diagnostic tool that would use invasive methods to collect fetal cells. In December 2012, a team from Eunice Shriver National Institute of Child Health and Human Development published their findings of a cohort study comparing CMA and karyotyping. Study results have suggested that CMA could be used as an alternative to karyotyping as it offers several advantages: higher sensitivity, reduced turnaround time, less labor-intensive work, and use of standardized computerized analysis, unlike karyotyping, which uses microscopic examination of chromosomes. 36 The microarrays currently used in the prenatal setting are mostly targeted arrays used to reduce uncertain and insignificant results, thus reducing difficulty in making decisions and patient anxiety. 37
FISH
Developments in technical and laboratory equipment have led to improvements in chromosomal analysis. FISH has allowed for faster karyotyping of amniocytes (within 24–48 h), while maintaining sensitivities as high as 97.9% and specificities as high as 100%.13,38,39 FISH detects chromosomal abnormalities by fluorescently labeling specific chromosomes or chromosomal regions using labeled hybridization probes specific to the areas of interest. 40
FISH probes can be produced from vectors containing plasmids or other small inserts that are labeled either directly or indirectly. Direct labeling involves attaching fluorescent nucleotides to the complementary sequence of interest (the preferred method), whereas indirect labeling consists of a probe with biotin- or digoxigenin-conjugated nucleotides, which act as haptens that are recognized by fluorescently labeled antibodies. Indirect labeling allows for a variety of fluorophores to be used, but this process is more time intensive and has the potential of higher background noise.
Following hybridization of the probe to the sample, and after a series of washes, the sample is viewed under a fluorescence microscope, where the presence of a fluorescent marker is used as an indication of the probe-specific sequence ( Fig. 3 ).41,42 Fluorescent probes are particularly advantageous in diagnosing trisomies due to a more distinct fluorescent reading. However, FISH can also be used to label specific regions within individual chromosomes, to detect mutations and translocations.41,42 FISH probe production for fetal karyotyping is strictly regulated by the Food and Drug Administration (FDA) and must be validated before used. The first probes to be approved by the FDA for detection of aneuploidies included the AneuVysion sets, produced by Abbott Molecular (Abbott Park, IL), which use a multicolor probe set for chromosomes 13 and 21, in addition to a set of centromere probes specific to chromosomes 18, X, and Y. 41 Although FDA approval has been granted only for AMC, there is indication that CVS could yield comparable accuracy in FISH diagnosis. 41
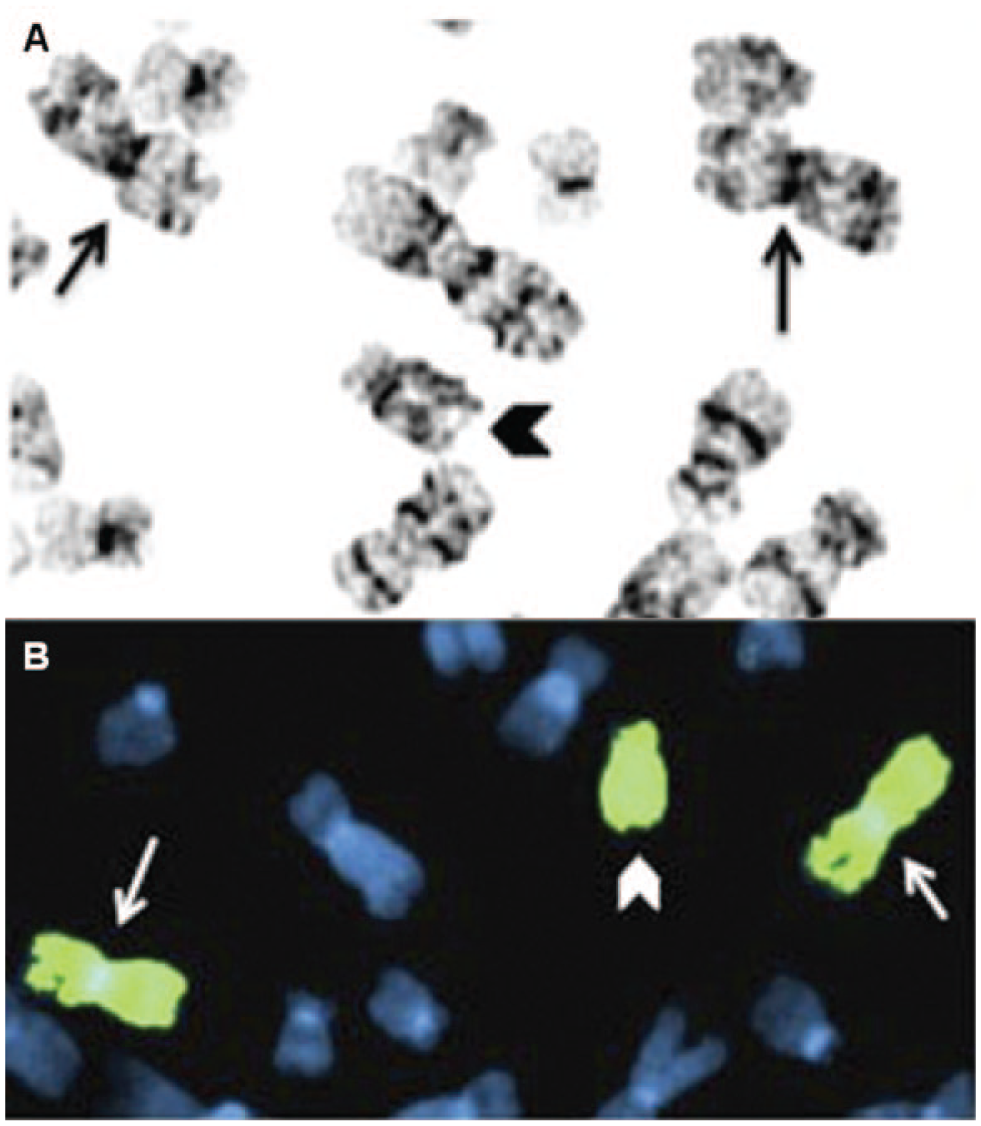
Automated FISH
As previously discussed, the standard of accuracy by manual FISH is high and provides high reliability and reproducibility compared with standard cytogenetic testing. However, since this technique is expensive and labor intensive, requiring subjective interpretation of its results, it has not been broadly adopted. Automated FISH may improve laboratory efficiency, reduce cost, increase throughput, and potentially improve the accuracy of results. 43 BioView Ltd. (Rehovot, Israel) launched automated FISH, Duet, into the market. This new system uses an optimized preparation protocol and commercially available probes to screen for aneuploidy in amniotic fluid cells. Major advantages of Duet are fast scanning of cells, high image quality, and high throughput. 43 When the efficacy of manual FISH was compared with automated FISH, it was found that analysis of samples is much less demanding in terms of technician time and shows high correlation (r = 0.9, p < 0.0001). An automated microscopy system is thus capable of providing similar or higher accurate results and rapid enumeration of FISH signals.43–45 The availability of such automated systems may enable its acceptance in broader patient populations for aneuploidy screening.
QF-PCR
QF-PCR is a PCR assay producing fluorescently labeled amplicons with end-point detection via capillary electrophoresis. The most commonly used method for QF-PCR involves amplification of short tandem repeats (STRs), which are chromosome-specific short DNA sequences, normally 2 to 5 base pairs that are stable, and vary in length between subjects. 42 QF-PCR is performed on DNA extracted from amniotic fluid or processed chorionic villus samples. 46 Sample DNA is PCR amplified using fluorescently labeled primers. The primers are multiplexed so as to enable parallel detection of chromosome 13, 18, and 21 aneuploidies. 47 After amplification, the products are separated based on fragment length by capillary electrophoresis for quantitative analysis, by analyzing the peak areas of respective repeat lengths using a DNA sequencer.47,48
QF-PCR aneuploidy testing provides comparatively rapid diagnosis of chromosomal aneuploidies, with results available within a day or two since the fetal cells do not have to be cultured. 47 However, it requires an investment in equipment, infrastructure, and adequately trained staff. Cytogenetic analysis of the results also requires expertise for correct interpretation. 49
Clinical trials to validate this method in the United States showed 100% accuracy with no false negatives. 49 Karyotyping and QF-PCR testing on 7679 prenatal samples showed concordant results for 98.75% cases, and abnormal karyotyping was undetected in 0.05% of samples. 50 Thus, QF-PCR can provide clinically valid results, reducing the workload on cytogenetic laboratories and providing faster results for patients and physicians. 50
The major disadvantage of QF-PCR is its weakness to detect sex chromosome abnormalities. When STRs specific for chromosome X are used, QF-PCR may show homozygous patterns for normal XX females that are indistinguishable from those patterns produced by a single X chromosome. 49 However, this can be compensated by the use of highly polymorphic markers in the routine assay. 49 Keeping all the advantages of QF-PCR in mind, the process can be integrated into the normal workflow of prenatal diagnosis.
Diagnostic kits available in the market for performing QF-PCR are the Chromaquant Version2 (Cybergene AB, Solna, Sweden) and Aneufast (Genomed Ltd., Altendorf, Switzerland). Both kits have two multiplex QF-PCR sets that provide simultaneous analysis of four selected STRs located on chromosomes 21, 18, and 13 and pseudoautosomal STRs. 51 With Chromoquant, a high number of markers are analyzed in two capillaries. It is a ready-to-use kit, allowing 26 samples to be analyzed simultaneously with detection by capillary electrophoresis, using genetic analyzer software. 48 The Aneufast kit uses five fluorescent dyes for automated DNA analysis. 52 Multiplex amplification is possible, and electrophoresis of more than 15 loci can be performed simultaneously. Both kits have simple procedures, but the Aneufast kit also reduces the risk of contamination by combining all reagents into a single tube, which reduces the time associated with PCR setup, thereby reducing the opportunity for cross-contamination. 51
Next-Generation Sequencing
Another method of detecting genetic abnormalities is DNA sequencing, which determines the precise order of nucleotides in DNA. Early methods of DNA sequencing were based on chemically or enzymatically generating variable-length DNA fragments and separating them by gel electrophoresis. These methods were manually intensive and usually required confirmatory data. 53
In 1977, Sanger and Coulson 54 published an improved process, Sanger sequencing, using dideoxynucleotides (ddNTPs). Dideoxynucleotides do not contain a 3′ hydroxyl group and, when incorporated by DNA polymerase, terminate the growing chain. Primers used in this process were radioactively labeled with 32p. Four separate reactions, each containing all four dNTPs and one ddNTP, generated base-specific ladders of terminated chains. Since all four reactions used the same label, the fragments were separated, based on fragment length, by gel electrophoresis in four separate lanes to read the sequence. In 1986, Smith et al. 55 improved this process by using primers with different fluorescence labels, allowing terminated fragments to be combined and separated in a single lane and analyzed by fluorescence emission on an automated gel electrophoresis system. In 1987, Prober et al. 56 improved this process by using fluorescently labeled ddNTPs, allowing base-specific terminated fragments to be generated in a single reaction and analyzed by separation in single lanes by gel electrophoresis. This process in various embodiments, slab gel electrophoresis or capillary electrophoresis, was the primary technology used for the initial sequencing of the human genome.57,58
Continued advancement in DNA sequencing technology has led to the development of next-generation sequencing (NGS) techniques. Compared with traditional Sanger sequencing, which allows about 100 simultaneous samples per run, NGS allows for millions of DNA fragments to be sequenced simultaneously. NGS enables detection of fetal chromosomal abnormalities in a less invasive manner than traditional methods: maternal blood contains fragments of fetal DNA, which enter the mother’s bloodstream as cells from the placenta die and break down. 59 This DNA is cell free and makes up as much as 10% of the total cell-free DNA in the mother’s blood. This fetal DNA can be identified and quantified through NGS techniques. 60 AMC and CVS also contain fetal DNA but, as previously discussed, require a much more invasive sampling approach, which introduces the risk of miscarriage. NGS enables prenatal diagnosis of chromosomal aberrations from a routine maternal blood draw, which has little to no risk to the fetus in comparison to AMC or CVS. 59
Competition within the NGS market most notably comprises Illumina (San Diego, CA), Roche Diagnostics (Indianapolis, IN), and Life Technologies Corporation (Carlsbad, CA) with their HiSeq/MiSeq, 454, and SOLiD/Ion Torrent systems, respectively ( Table 3 ).59–66 Illumina offers a range of sequencers, with prices between $200,000 and $700,000. The HiSeq systems first appeared in 2009 and are geared for short-read applications, while the more recent MiSeq system, released in 2011, allows for rapid turnaround times on a benchtop system, but with reduced output. High-output systems can take up to 14 days for sequencing, whereas the short-run feature of newer models can produce results in 27 h. In addition, the variety of systems produced by Illumina can yield between 1 and 600 Gb per run. Illumina’s systems uses a “sequencing by synthesis” technique that adds fluorescently labeled reversible terminator nucleotides to halt a growing DNA chain after the addition of one base. 67 These fluorescent signals are recorded after each cycle of nucleotide addition, and then the reversible terminator is unblocked, allowing addition of the next reversible terminator to the growing chain. Illumina also uses a bridged amplification method, in which DNA strands are built upon the surface of a flow cell that holds a sequencing template. This procedure enables upwards of 10 million molecule clusters to accumulate within a square centimeter, given that bridge amplification occurs on the cell surface, in between bound primers. 68
Comparison of Sequencer Performance, Mid 2012.
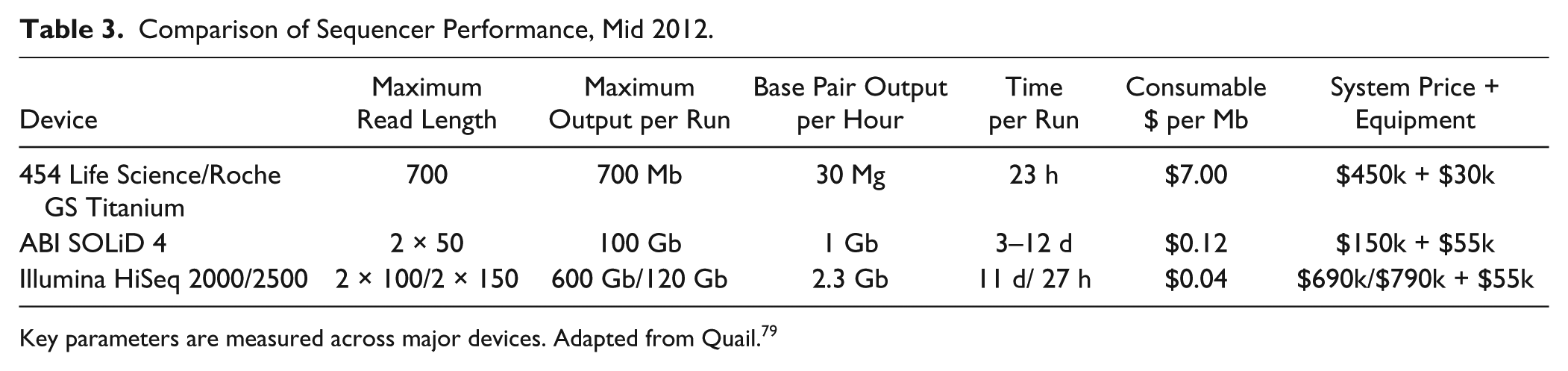
Key parameters are measured across major devices. Adapted from Quail. 79
The 454 Life Sciences (Roche Diagnostics) system is based on pyrosequencing technology and uses emulsion PCR for sample preparation. 69 During the preparation phase of this process, universal oligonucleotide priming sites are ligated to both ends of the fragmented DNA target and the sample is added to an oil-water emulsion containing Sepharose capture beads and PCR reagents. Conditions are optimized to have only one DNA fragment and at least one DNA capture bead in the majority of the aqueous micro-droplets. Amplification, emulsion PCR, then creates a clonal DNA library on the Sepharose beads in the aqueous micro-droplets of the emulsion. The beads are then transferred to a plate containing microscopic wells coupled to a fiber-optic array. Pyrosequencing technology identifies sequences through the release of a pyrophosphate when nucleotides are added to the growing chain. The sequential addition of dNTPs to the plate releases pyrophosphate into the wells containing complementary bases. The released pyrophosphate is enzymatically converted to a chemiluminescent signal that can be detected by a CCD camera. 67 These systems are able to produce longer reads than other NGS systems on the market due to the use of Sepharose DNA capture beads, which isolate specific sequences in the water-in-oil emulsion.
Life Technologies sequencers come from the SOLiD and Ion Torrent platforms. 67 The SOLiD system is unique in that it uses a technique for sequencing based on fluorescence stepwise ligation. This technique creates sequences that ligate priming sequences to that of chromosomal DNA, thereby creating sections of target DNA for amplification, which can be captured and immobilized for identification via fluorescent microscopy. The range for SOLiD systems can yield up to 150 Gb over the course of 8 days. 67 The Ion Torrent system, however, functions similarly to the 454 systems, yet instead of identifying a chemiluminescent signal, a semiconductor chip has the ability to identify protons from DNA synthesis without the use of a camera, laser, or fluorescent dye, yet through a spike in charge that identifies specific nucleotides. Also, similar to the 454 systems, the Ion Torrent systems yield approximately 1 Gb per run in as little as 2 h.57,67
NGS for Chromosomal Aneuploidies
The first step in NGS involves taking a sample of the mother’s blood, centrifugation to obtain plasma, and then extracting DNA from the plasma. Since the DNA is already fragmented, there is no need for further fragmentation. After ligating adapters to these fragments, clonal amplification of the fragments in the HiSeq platform is performed via bridge amplification. Finally, sequencing occurs by using fluorescently labeled, reversible terminator nucleotides in a massively parallel fashion on all DNA clusters within the flow cell.
NGS can be used to detect fetal chromosomal abnormalities by sequencing and analyzing the allelic ratio of the target locus. Fragments of DNA derived from allelic variants of a gene can be distinguished based on SNPs. Informative SNPs for this process identify genes that are homozygous in the mother and heterozygous in the fetus, meaning the mother has a genotype of AA and the fetus has a genotype of AB. Through NGS, it is possible to count the number of shared A alleles, which are derived from either mother or fetus, and the number of fetal-specific B alleles. However, one unknown parameter is the ratio of fetal to maternal DNA. Therefore, the allele frequency (i.e., the relative ratio of B to A alleles in the sample) from a specific chromosome known for chromosomal aneuploidy, such as chromosome 21, is compared with another chromosome, known as a reference. If the frequency of fetal-specific B alleles in chromosome 21 is twice as high as in the reference chromosome, then the fetus has two copies of the chromosome 21 bearing the B allele (i.e., is ABB) and thus has a trisomy 21, with leads to DS. 59
Deep sequencing techniques are used to enable quantitative analysis of the fetal chromosomal DNA present in the blood. Traditional sequencing has a depth or coverage of 5× to 10×, meaning approximately 5 to 10 independent reads are available for each nucleotide sequence present in the sample. Deep sequencing refers to a coverage >100×, which is possible due to the vastly higher throughput of NGS, which enables millions to billions of reads to be acquired simultaneously in one run, compared with 96 reads per run in high-end Sanger sequencing platforms. 59 The greatest benefit of deep sequencing is that it enables accurate quantification. Quantification is possible by analyzing how often a particular sequence was read relative to the total number of reads in a given run.
A large-scale validity study was performed using the Illumina platform against karyotyping as the gold standard. The study included 753 pregnant women at high risk for fetal trisomy 21 and 86 who were positively diagnosed with trisomy 21 using karyotyping. The results provided a 79.1% sensitivity and 98.9% specificity, with a positive predictive value of 91.9% and a negative predictive value of 96.9%. Therefore, if the referrals for the more invasive AMC or CVS were based on NGS test results, then about 98% of the invasive tests could be avoided. 61
In addition, a recent study in the New England Journal of Medicine has cited such noninvasive diagnostics as outperforming standard screening methods’ positive predictive value by a factor of 10 in low-risk populations, thereby suggesting that the analysis of fetal DNA in maternal blood may provide an effective, noninvasive screening method for parents. 70
Market/Competitor Analysis
Recent studies on the use of cell-free DNA (cfDNA) in high-risk women indicate that while the overall detection rate of fetal aneuploidies may be marginally effected by using NGS technology in prenatal testing, the false-positive rate may be dramatically reduced. 34 Bianchi et al. 70 demonstrated a false-positive rate of 0.3% in comparison to 3.6% for trisomy 21 in a study population of 1914 women (average age, 29.6 years), while detecting all cases of trisomy 21 (negative predictive value, 100%). This study, the Comparison of Aneuploidy Risk Evaluations (CARE), evaluates the use of cfDNA testing in high-risk pregnancies as well as the general obstetrical population. Whereas previous studies have focused primarily on the application of screening in high-risk populations, the CARE study finds the performance of sequencing (Illumina HiSeq 2000) cfDNA to be equivalent among the general and high-risk populations of women in the United States.34,71 The CARE study suggests that if all women were to undergo cfDNA testing as a primary measure, there would be an 89% reduction in the number of invasive procedures required to confirm a positive result since all women who test positive undergo posttesting counseling. Unlike many of the studies that focus on the use of cfDNA testing in high-risk women, the CARE study includes a diverse sample collection set, racial and ethnical diversity, complete follow-up, and the inclusion of standard screening methods. While the CARE study suggests consideration of cfDNA testing as a primary prenatal screening method, it does not evaluate the cost-effectiveness of this testing methodology.
Benn et al. 34 estimate that the cost of cfDNA testing ranges from $800 to $2000 in the United States and $500 to $1500 outside of the United States. On the basis of these costs, Cuckle et al. 71 conclude that universal prenatal testing using cfDNA does not provide a cost-effective replacement to standard diagnostic methods. While universal testing is not currently cost-effective, conditional use of cfDNA testing to standardize screen-positive women presents an option where the average lifetime cost associated with DS would be reduced through secondary testing. 34 Whereas the implementation of cfDNA testing is largely dependent on being cost-effective in comparison to standard methods, the adoption of use is heavily dictated by the willingness of insurance companies to reimburse the procedure. This presents a hurdle for cfDNA testing given the reimbursement issues that other novel prenatal testing procedures, such as CMA, have encountered with insurance companies. Although the American College of Obstetricians and Gynecologists (ACOG) has identified the increased sensitivity of CMA in comparison to karyotyping, the BlueCross BlueShield of Alabama’s medical coverage criteria only offer coverage on the condition of detection of a structural abnormality through ultrasound or magnetic resonance imaging. All other uses of CMA in prenatal testing are considered investigational and are not eligible for coverage. 37 Conversely, BlueChoice of South Carolina considers the use of CMA to be a “medical [necessity]” if abnormal anatomic findings are characteristic of a genetic abnormality, there is fetal demise with a congenital abnormality, patients with a structurally normal fetus are undergoing invasive diagnostic testing, or the individual is considered high risk due to being older than 35 years at time of delivery, having had a history of pregnancies with trisomies, or having had a positive standard biomarker screening test. 36 Such ambiguity in the reimbursement structure for procedures that insurance companies have identified as advantageous demonstrates the need for cfDNA to become cost-effective since adoption will be largely based on payer’s determination of the value at a given pricing point.
Prenatal diagnosis accounts for 69.3% of the genetic disease testing market and is expected to be the leading segment for the next 10 years. 72 Approximately 4 million women give birth in the United States each year.73 The estimated lifetime cost of having a child with DS ranges from $400,000 to $800,000, with an estimated baseline price of $677,000. 73 The purpose of screening for fetal aneuploidy is to reduce, or even prevent, the costs of disability and deaths associated with DS and similar disorders. 74
MSS is used throughout the United States, with only California and Illinois having an established prenatal screening program. 74 The program costs $162, regardless of which screening type—serum integrated, sequential, or quadruple—the patient chooses. 74 Fee coverage for the program depends on the individual insurance company and status of the patient’s yearly deductible. 74 The 5% false-positive rate is a potential weakness for MSS because it can lead to undue stress and the need for further follow-up testing. 73
The cost of an NT test ranges from $35 to $150, with a baseline estimate of $57 per ultrasound test. 75 NT is noninvasive, does not require any serum or tissues samples, and allows for the detection of approximately 1133 more cases of DS for each million pregnancies. 75
Karyotyping currently dominates the fetal genetic testing market. 72 The Mayo Clinic reported that approximately 200,000 cases of AMC are performed each year. 76 However, due to advances in genetic diagnostic technology, it is expected to be replaced by more modern technologies. 72
Current estimates indicate FISH tests account for 40% of the prenatal screening market ($72 million). FISH is currently the method of choice in small laboratories for prenatal diagnosis, given the reduced labor and time associated with FISH compared with karyotyping. The leaders in FISH diagnosis are Abbott/Vysis (Abbott Park, IL), Genzyme (Cambridge, MA), and Xenomics (New York, NY). 77 Most FISH-based prenatal tests are evaluated using an Applied Imaging (San Jose, CA) automated slide reader. 77 QF-PCR is becoming more common in large laboratories due to its reduced cost. The QF-PCR market is dominated by Applied Biosystems (Foster City, CA), with Roche Applied Science (Indianapolis, IN), and Qiagen (Valencia, CA) also having significant market share. 78 Large laboratories are increasingly adopting this method due to its low costs and fast results.
Karyotyping, QF-PCR, and FISH require sample acquisition via invasive AMC or CVS, whereas NGS enables minimally invasive testing from a blood sample and provides a reasonably sensitive method for detecting fetal chromosomal anomalies.
According to the 2013 Kalorama Information Market Intelligence Reports, the estimated market value for noninvasive prenatal screening services was $1 billion in 2012. 77 Sequenom, a San Diego–based company, developed a noninvasive prenatal test called MaterniT21 test, which screens for trisomy 21, 18, and 13 using massively parallel sequencing. Sequenom bills insurance companies $2900, offers the test to uninsured patients for $1700, and charges a $235 copay for insured patients while taking responsibility for securing the remaining costs from the payer. 77
As new technologies are introduced into the market, current methods will probably be outcompeted and newer technologies will complement the existing methods. Leading NGS companies comprise 80% of the $1468 genetic sequencing market (Illumina, $575 million; Life Technologies, $525 million; Roche 454, $75 million) ( Fig. 4 ). 67
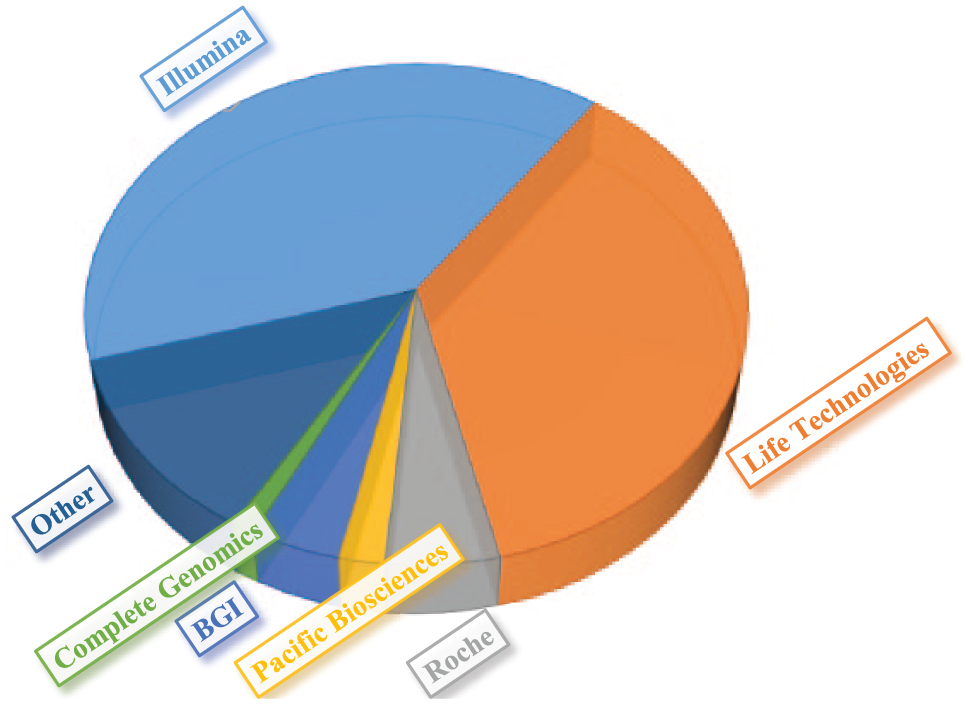
Market share for DNA sequencing. 67
The potential market for NGS is estimated to reach $1.5 to $2 billion in sales for 2015. 76 It is estimated that 10% of sequencing purchases will be for clinical applications, which is expected to increase to 16% to 20% by 2016. 67
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
