Abstract
Drug development relies on high-throughput screening involving cell-based assays. Most of the assays are still based on cells grown in monolayer rather than in three-dimensional (3D) formats, although cells behave more in vivo–like in 3D. To exemplify the adoption of 3D techniques in drug development, this project investigated the automation of a hydrogel-based 3D cell culture system using a liquid-handling robot. The hydrogel technology used offers high flexibility of gel design due to a modular composition of a polymer network and bioactive components. The cell inert degradation of the gel at the end of the culture period guaranteed the harmless isolation of live cells for further downstream processing. Human colon carcinoma cells HCT-116 were encapsulated and grown in these dextran-based hydrogels, thereby forming 3D multicellular spheroids. Viability and DNA content of the cells were shown to be similar in automated and manually produced hydrogels. Furthermore, cell treatment with toxic Taxol concentrations (100 nM) had the same effect on HCT-116 cell viability in manually and automated hydrogel preparations. Finally, a fully automated dose-response curve with the reference compound Taxol showed the potential of this hydrogel-based 3D cell culture system in advanced drug development.
Introduction
It is well recognized that cells grown on two-dimensional (2D) flat and rigid substrates are facing an unnatural environment, leading to differential cellular behavior when compared with cells grown in their native three-dimensional (3D) in vivo environment.1,2 Therefore, researchers have begun to study cells in 3D to obtain more relevant results with in vitro cultures. In consequence, cell-based assays in which cells are grown in 3D are also increasingly gaining importance for drug development because the effects of drugs are reflected in such cultures much more closely to those found in the human body than they are in 2D cultures. 2 Thus, the advantages of such cultures are (1) drugs that have been identified as false-negatives would be identified as effective and (2) late drug failure and possibly even drug removal from the market could be avoided, and thus a lot of money could be saved. 3
Three-dimensional cell culture opens new dimensions in cell-based assays. 4 For screening drugs, 3D cell cultures must be automated to obtain appropriate throughput and must be well reproducible and standardizable, 5 the latter being also important to obtain approval by the authorities. In the past decades, many systems have been developed to enable cell cultivation in 3D. 6 In a recent review by Rimann and Graf-Hausner, 5 commercially available scaffold-free and scaffold-based 3D cell culture systems are listed and discussed in detail. Among the scaffold-based cell culture systems, hydrogels are a prominent group. Hydrogels, unlike rigid scaffolds, are providing a cellular environment resembling the native extracellular matrix (ECM), displaying high water content and different stiffnesses. 7 The most widely used hydrogel is an extract of the basement membrane secreted by Engelbreth Holm Swarm mouse tumor cells. 8 This mixture contains proteins, such as laminin, collagen IV, proteoglycans, and entactin, and different growth factors. Because of the natural origin of this material, cells that are encapsulated inside the hydrogel are able to adhere, spread, and migrate. Drawbacks of this natural product are the batch-to-batch variation exhibited by different concentrations of the components and a given composition of proteins that is not suitable for all cell types. 9 In addition, its handling can be problematic due to a temperature shift required for gelation to occur and thus the need of cooled equipment to keep the hydrogel in a liquid state. This is particularly important when considering high-throughput hydrogel preparation with liquid-handling robots. To overcome these problems, many synthetic hydrogels have been developed, mostly based on poly(ethylene glycol) (PEG) and chemical polymerization. 10 To more closely mimic the natural functions of an ECM, such hydrogels are covalently modified with cell adhesion peptides containing cell recognition motifs of ECM proteins and cell-degradable cross-linker molecules containing matrix-metalloproteinase–cleavable peptides. 11 Although adhesion sites can already fulfill certain functions of the ECM, both components are required for cells to spread and migrate within the gel. 12 The advantages of such artificial hydrogels are manifold: (1) the hydrogel is chemically defined and highly reproducible in composition, and (2) the basic material is cell inert; bioactive molecules such as cytokines, growth factors, and cell adhesion sites are absent and can be incorporated to be compatible with different cell-type requirements and cell culture regimes. In our study, we used a hydrogel based on the cross-linking of dextran with PEG. This dextran-based hydrogel system allows the incorporation of bioactive factors independent of hydrogel stiffness and can be degraded to release cells after culture without adverse effects on cells. Such hydrogel degradation is required for many assay readouts based on signals derived from the whole-cell population such as ATP and DNA content measurements. Because of the easy handling of liquids before gelation and the modular, highly flexible nature, this dextran-based hydrogel system is very well compatible for automation with a liquid-handling robot. In this report, we show the feasibility of an automated compound-screening approach using tumor cells embedded in this hydrogel thereby forming 3D multicellular spheroids.
Materials and Methods
Reagents
3-D-Life Dextran-PEG Hydrogel Kits (G90-1) were purchased from Cellendes GmbH (Reutlingen, Germany) containing Mal-Dextran, PEG-Link, 10× CBpH5.5, water, and dextranase. To improve the biocompatibility of the hydrogel with cell adhesion sites, RGD Peptide (P10-3, Cellendes GmbH) was incorporated covalently. Cell viability was determined with the CellTiter-Glo Luminescent Viability Assay (G7571; Promega, Madison, WI), and DNA content was assessed with the Quant-iT PicoGreen dsDNA reagent kit (P7589; Invitrogen, Carlsbad, CA). Taxol was purchased from Enzo Life Sciences (BML-T104; Farmingdale, NY) and stored at −20 °C in 1 mM aliquots dissolved in DMSO.
Consumables
Luminescence was measured in white half-area 96-well plates (Cat. No. 7.675075) and fluorescence in black 96-well plates (Cat. No. 7.655076), both purchased from Huber & Co AG (Switzerland).
Cell Line and Cultivation
The human colon carcinoma cell line HCT-116, kindly provided by InSphero AG (Switzerland), was cultivated in RPMI-1640 (R8758, Sigma, St. Louis, MO), supplemented with 10% fetal calf serum (Invitrogen, 10270) and 1% pen/strep. The cells were either cultivated in 2D in T-flasks and 48-well plates or in 3D dextran-based hydrogels in 96-well plates in a standard incubator at 37 °C and 5% CO2. Inoculation of cells in hydrogels and subsequent cell maintenance have been performed either manually or with a Freedom EVO liquid-handling robot from Tecan (Switzerland).
Instrumentation
The automated cell culture platform is based on the liquid-handling robot Freedom EVO 150, which was configured for cell culture requirements by adding the following four modules: (1) eight-channel liquid-handling arm with fixed tips, (2) controlled pump option for accurate media dispension, (3) orbital shaker (Te-Shake), and (4) clean air hood on top of the system causing vertical, low turbulence displacement flow (0.25 m/s ± 20%) to obtain purity level 4 requirements (VDI 2083) and class 1000 (U.S. Federal Standard 209).
The operating software Freedom EVOware (Tecan) was used to group pipetting commands and device commands into scripts and to create the specific experimental program used in this work.
Production of Cell-Containing Hydrogels (Manually and Automated)
Cell-containing hydrogels were produced manually according to the protocol provided by Cellendes. The protocol for the automated production of cell-containing hydrogels was adapted from the manual procedure. Cells were encapsulated in 50 µL hydrogel/well of a 96-well plate. In Figure 1 , a schematic outline of the different steps of hydrogel formation on the liquid-handling robot is shown. In all experiments, each hydrogel contained Mal-Dextran (2.5 mM maleimide groups), 0.5 mM RGD Peptide, PEG-Link (2 mM thiol groups), 1× CB buffer pH 5.5, and 1× 104 HCT-116 cells at the time of encapsulation. The freshly prepared hydrogel components (avoid freeze-thawing) and the cell suspension were supplied in 2 mL tubes, held in a tube carrier on the work table of the liquid-handling robot. A master mix of ddH2O, 10× CBpH5.5, Mal-Dextran, and RGD Peptide was prepared in a 15 mL tube and incubated for 10 min at room temperature (RT). During this time, the RGD Peptide was covalently coupled to the dextran by the Michael type addition of maleimide and thiol. Afterward, the cells were added and suspended in the master mix. As cells settle down during the course of mixing of hydrogel components, they must be well resuspended before pipetting into the master mix. Mixing must be slow in order not to shear the cells but must be efficient enough to result in a homogeneous cell suspension. This master mix with cells was then dispensed in 45 µL aliquots per well into 96-well plates previously supplied with 5 µL of PEG-Link (20 mM thiol groups) per well. Tips must be perfectly aligned within the middle of the wells in order to place the two components on top of each other. Careful mixing with no introduction of air bubbles was done, and gelling was induced by the reaction of the remaining maleimide groups on the dextran with thiols on the PEG-Link. The tips were washed thoroughly with ddH2O right after this mixing step to avoid gelling within the tip, which would cause a clogged tip. The hydrogel formation was completed after 10 min at RT. However, all hydrogels were prepared in the well plate before adding 200 µL media, leaving the first eight produced hydrogels exposed to air for approximately 45 min. During this time, the hydrogels did not dry out. To avoid extensive hydrogel air exposure, the software can be adjusted to add media to the hydrogels step by step directly after preparation. Two hours later, the medium was replaced manually. The cell-containing dextran hydrogels were cultivated in 200 µL growth medium for 8 days. The medium was exchanged every second or third day.
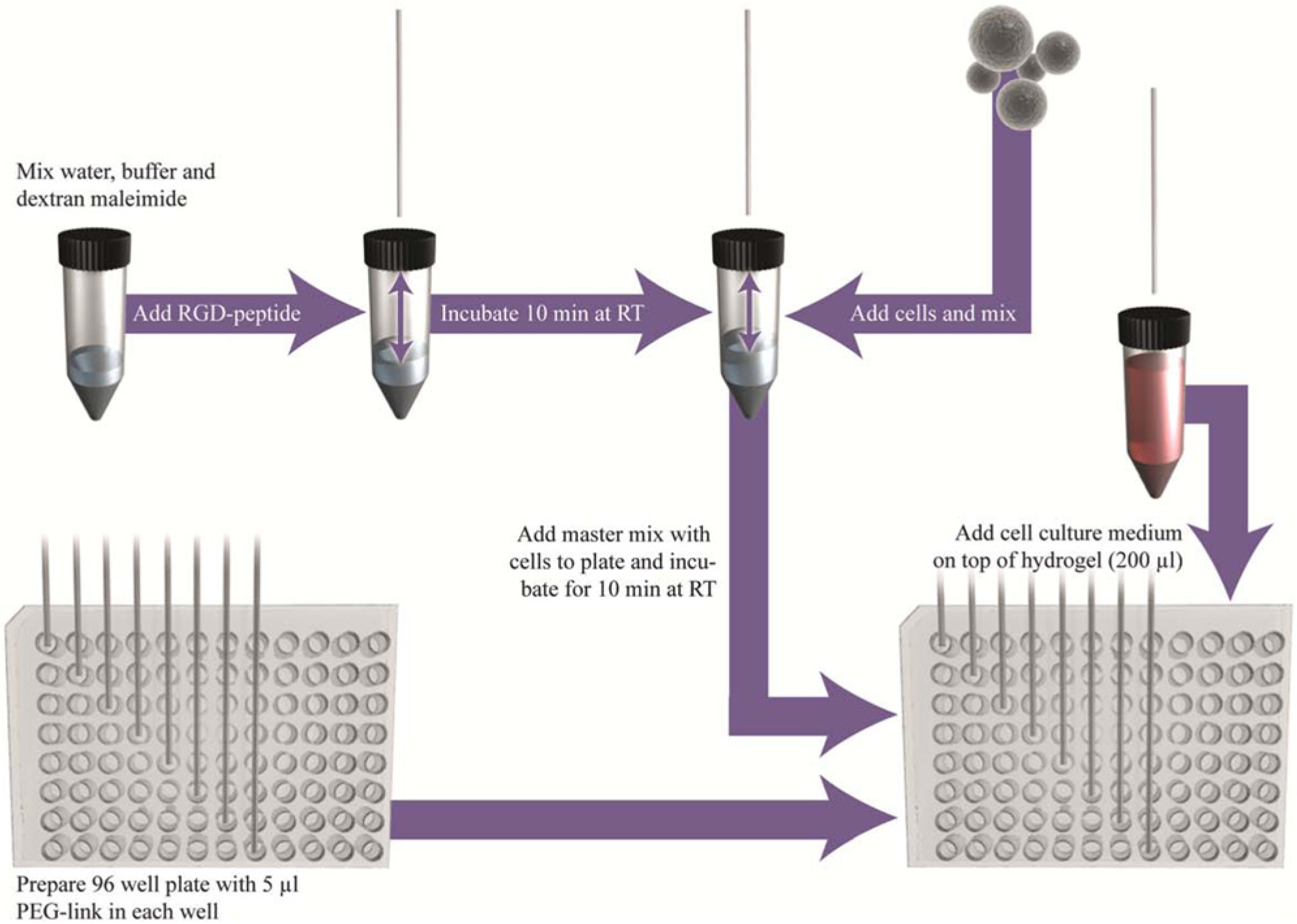
Fully automated preparation of a dextran-based hydrogel on a Freedom EVO liquid-handling robot.
Optical Observation of HCT-116 Cells in Dextran Hydrogels
To visually observe cell behavior and growth inside dextran-based hydrogels, the hydrogels were imaged with a light microscope from Zeiss (Axiovert 25).
Viability Determination (Manually)
To determine cell viability in dextran hydrogels, an ATP assay (CellTiter-Glo; Promega) according to the manufacturer’s protocol was used. The cell culture medium was completely replaced with 200 µL fresh medium, and 10 µL Dextranase (supplied with 3D-Life Dextran PEG Hydrogel Kit) was added and mixed. The hydrogels were incubated at 37 °C for 1 h while resuspending the medium containing the dextranase three times during the incubation. Then the wells were checked with a light microscope (Zeiss, Axiovert 25) for the dissolution of the gels and complete release of cells. The cells were then resuspended and transferred into a 1.5 mL tube containing 1 mL serum-free growth medium. The content was centrifuged at 200g for 5 min to pellet the cells. Afterward, the medium was removed, leaving 30 µL in the tube. Then, 30 µL of CellTiter-Glo Reagent (a combination of CellTiter-Glo Substrate with CellTiter-Glo Buffer) was added and mixed with the cells by pipetting up and down. After an incubation of 12 min at RT, 50 µL of that mixture was transferred into a white half-area 96-well plate, and luminescence was measured in a microplate reader (BMG Labtech, FLUOstar OPTIMA). The obtained signals were correlated to signals of known ATP concentrations (standard curve).
DNA Content Measurement (Manually)
DNA measurements were performed with the same solution used for luminescence measurements. To do so, 40 µL was withdrawn from the white half-area 96-well plates and was transferred into a black 96-well plate containing 60 µL phosphate-buffered saline. A total of 100 µL PicoGreen solution (Quant-iT PicoGreen dsDNA reagent diluted 1:200 in 1× TE buffer, supplied in the kit) was added into each well, and the solution was mixed by pipetting up and down. After a 5 min incubation at RT in the dark, fluorescence was measured in a microplate reader (BMG Labtech, FLUOstar OPTIMA) equipped with ex. 485 nm and em. 520 nm filters. The obtained signals were correlated to signals of known DNA concentrations (standard curve).
Dose-Response Experiment with Cells in 2D Including Viability Determination (Manually)
To get comparable values of the IC50 of Taxol with HCT-116 cells grown in 2D, the cells were subjected to different Taxol concentrations. To do so, 1 × 103 cells/well of a 48-well plate were seeded, and at day 3 of cultivation, different Taxol concentrations were applied. At day 6, medium was replaced containing the corresponding Taxol concentrations, and at day 7, ATP was measured as follows: media were removed from the wells, and 60 µL ATP reagent (50:50 of serum-free media and CellTiter-Glo Reagent) was added. Then, the well plate was shaken at 700 rpm for 15 min at RT, and 50 µL of the reaction solution was transferred into a white half-area 96-well plate, and luminescence was measured in the microplate reader (BMG Labtech, FLUOstar OPTIMA).
Fully Automated Dose-Response Experiment with Cells in 3D on Freedom EVO Liquid Handler Including Viability Determination
To show the versatility of the hydrogel system and its use for high-throughput screening in drug development, an automated procedure for dose-response curves was established. To do so, the whole process of hydrogel production, cell culture maintenance, Taxol serial dilution and addition, as well as ATP analysis was automated on the Freedom EVO liquid-handling robot. The automated production of cell-containing hydrogels was already functional (see the “Production of Cell-Containing Hydrogels” section). To perform an automated media exchange with the fixed tips only 150 µL of 200 µL medium was removed (to avoid hydrogel aspiration) and replaced with an equal volume of fresh medium two times in a row. The automated ATP assay was different from the manual procedure. After 150 µL medium was exchanged twice with fresh non–Taxol-containing medium, 10 µL Dextranase was added into each well, and the plate was shaken for 5 s with 1000 rpm on the Te-Shake. Then, the plate was incubated for 4× 15 min in the incubator at 37 °C and 5% CO2 and shaken in between in order to enhance the digestion of the dextran hydrogel and to release the encapsulated cells for further ATP analysis. After the digestion, the liberated cells were resuspended by pipetting 3× 150 µL up and down, and 30 µL of each well was transferred into a white half-area 96-well plate. Another 30 µL CellTiter-Glo Reagent from a 15 mL tube was added to the cell suspension with subsequent shaking (2 min, 1000 rpm). After a 10 min incubation at RT, the 96-well plate was shaken for another minute at 1000 rpm, and afterward, luminescence was measured in the microplate reader (BMG Labtech, FLUOstar OPTIMA).
Results and Discussion
Replacing the standard 2D with 3D cell cultures in high-throughput screening is a very promising approach to improve the drug development outcome. More relevant data are obtained due to the in vivo–like 3D environment cells are facing. 13 Although 3D cell cultures are well accepted and performed in research, the translation into automated processes is still in its infancy. In this project, we were using a highly modular dextran-based hydrogel system to translate 3D cell culture on an automation platform that is routinely used for cell-based assays. Gelation occurs via a Michael type addition of dextran macromeres harboring maleimide groups and thiol-bearing linear short PEGs. After mixing dextran and PEG molecules, the hydrogel formation starts instantaneously. This fast gelation has the advantage that cells seeded in the gel do not sink to the bottom but stay evenly distributed in the gel. We evaluated whether the automated pipetting and encapsulation of colon carcinoma cells HCT-116 led to the same quality of cell culture as the already well-established manual procedure. To examine cell behavior, cells were either microscopically observed or analyzed with relevant assays such as ATP and DNA content measurements generating a luminescence or a fluorescence signal, respectively. A total of 1 × 104 HCT-116 cells per 50 µL hydrogel per well of a 96-well plate were encapsulated manually or automated with the liquid-handling robot Freedom EVO and cultivated for 8 days in an incubator at 37 °C and 5% CO2. Microscopic images were taken at day 2, 5, and 8 of cultivation to compare cell growth visually ( Fig. 2 ). The cells were forming small multicellular spheroid-like structures over time in both preparation methods and showed no difference in cell growth or behavior. As a crucial issue, cell viability of HCT-116 cells in the dextran-based hydrogels was assessed. Cells were again seeded into hydrogels prepared either manually or automated. At days 2, 5, and 8, ATP content was determined. In this assay, ATP is used to propel the generation of a luminescence signal that is proportional to cell number. In Figure 3A , ATP content per hydrogel is shown over time of automated and manually produced hydrogels. HCT-116 cells show similar ATP values in both hydrogel preparation methods. To confirm this finding with another type of assay, the DNA content of the sample was monitored with PicoGreen. PicoGreen labels double-stranded DNA, generating a fluorescent signal that is measured with a microplate reader. The PicoGreen labeling was performed with the same sample used for ATP determination to show the multiplexed analysis of one sample with two readouts, one luminescent and the other fluorescent. In Figure 3B , the DNA content of automated versus manually produced 3D tissues is shown over the course of 8 days. DNA content steadily increased over time under both conditions, and the absolute values were similar ( Fig. 3B ). These findings clearly demonstrate the feasibility of analyzing cells grown in a dextran-based hydrogel with assays that require hydrogel solubilization and subsequent cell lysis. The results from automated versus manually produced hydrogels are comparable. Thus, hydrogel production is successfully translated to an automation platform that is suitable for high-throughput screening.
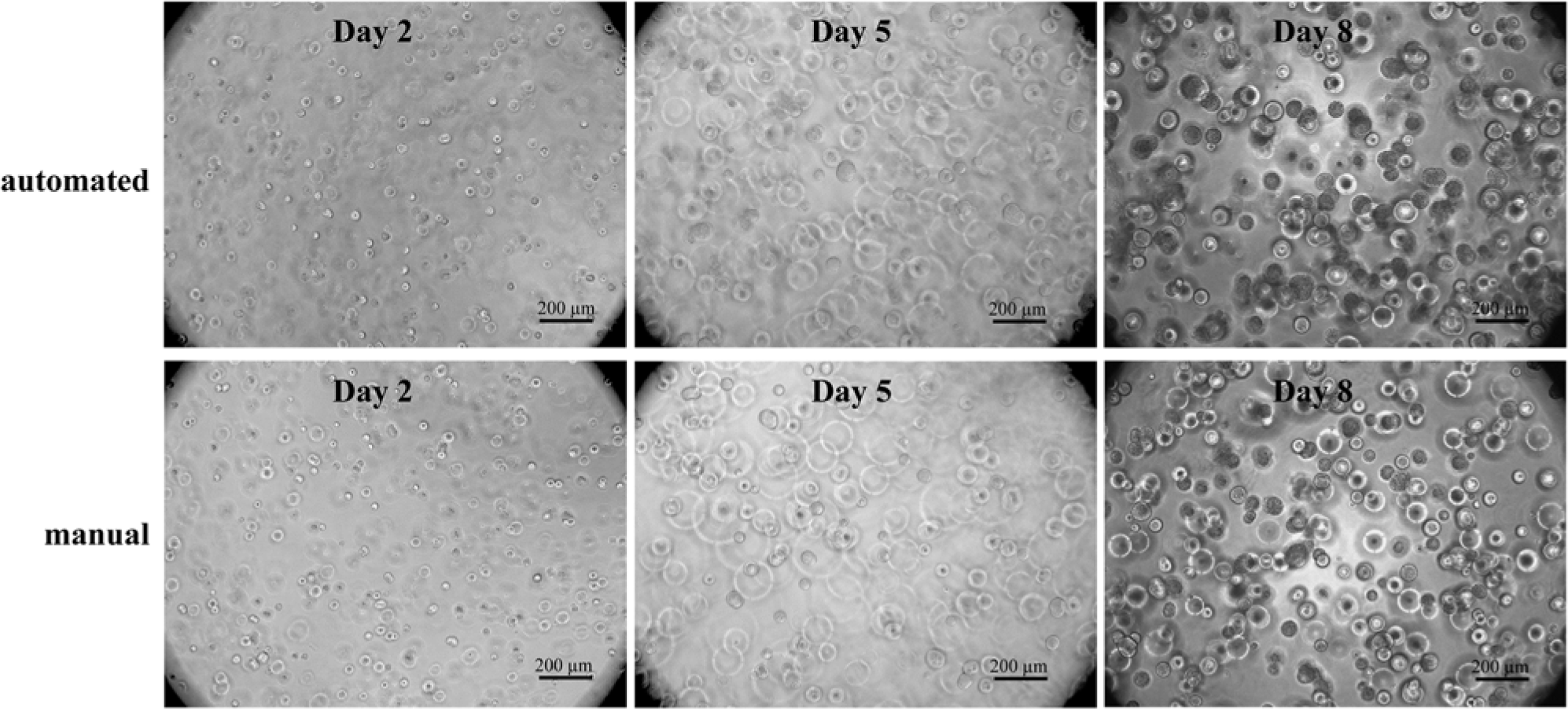
Light microscopical images of HCT-116 cells grown in dextran-based hydrogels over time. Cell encapsulation into hydrogels was performed automated using a liquid-handling robot. For comparison of the automated process, cell plating into hydrogels was done manually and is shown in the lower panel. Cell-seeding density was 1 × 104 cells per 50 µL hydrogel and well (96-well plate). Images were taken at days 2, 5, and 8 of cultivation.
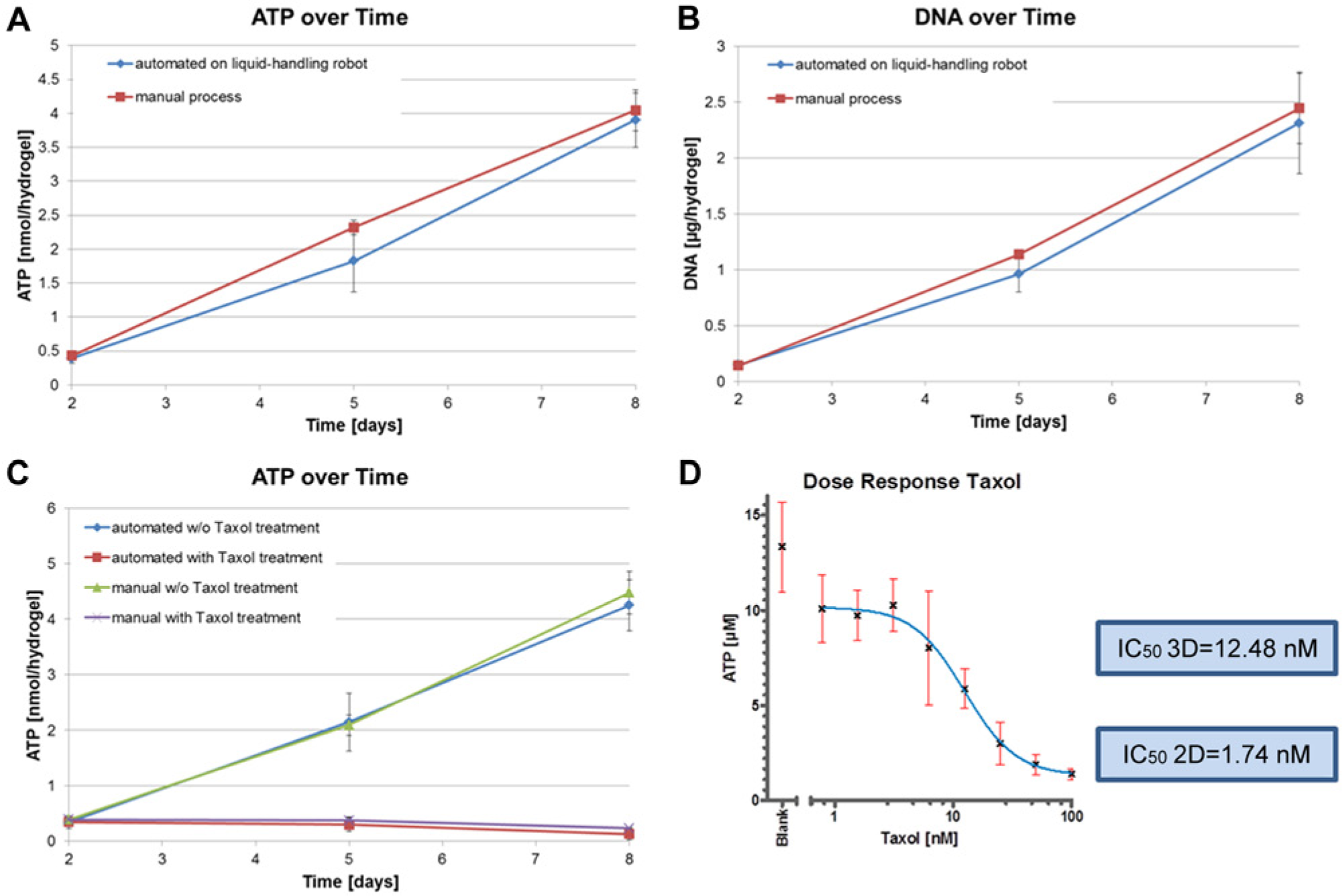
Viability and DNA measurements of HCT-116 cells encapsulated in 3D dextran-based hydrogels. (
To demonstrate the applicability of the automated hydrogel culture to drug development, the drug response of cells encapsulated in the dextran-based hydrogels was evaluated. Cytotoxic concentrations of the reference compound Taxol (100 nM) were applied on the encapsulated cells 2 days after hydrogel formation, and ATP content was analyzed at days 2, 5, and 8. Multicellular spheroids in manually and automated produced dextran gels were compared. Figure 3C shows the result of a representative experiment. First, it was shown that cells are susceptible to Taxol that was supplied in the growth medium, and second, the cells react in a similar manner irrespective of the hydrogel production mode.
To show the versatility of this 3D tissue format for a real drug-screening process in the pharmaceutical industry, hydrogel production, medium exchange, drug administration, and analysis of cell survival were automated. In the experiment, 72 dextran-based hydrogels were prepared with the liquid-handling robot. On day 3 after hydrogel formation, eight different Taxol concentrations were prepared by the robot and applied to the hydrogels. The hydrogel-encapsulated cells were treated with Taxol for 5 days. On the third day of Taxol treatment, medium was replaced with fresh Taxol-containing medium. The ATP assay was then conducted with the cells dissociated from the dextran hydrogels with dextranase. Afterward, the CellTiter-Glo Reagent was mixed with the cells and incubated for 13 min before the luminescence signal was measured with a microplate reader. In Figure 3D , the characteristic dose-response curve is shown. At low Taxol concentrations (0.78125, 1.5625, 3.125 nM), the HCT-116 cells showed the highest viability, whereas at intermediate Taxol concentrations, a drop in viability was observed (6.25, 12.5, 25, 50 nM). At the highest Taxol concentration (100 nM), cell viability was drastically reduced, shown by the low ATP levels ( Fig. 3D ). From this dose-response curve, the IC50 value of the 3D culture was calculated using GraphPad Prism5 software (GraphPad Software, La Jolla, CA). The obtained value of 12.48 nM was more than seven times higher than the value obtained with Taxol applied on standard 2D culture ( Fig. 3D ), which was in line with previously published 2D IC50 values. 14 Furthermore, Karlsson et al. 15 showed increased chemoresistance of 3D HCT-116 spheroids when standard cytotoxic drugs for treatment of colorectal cancer, such as irinotecan, oxaliplatin, and 5-fluoruracil were applied. These results reflect the differences of drug responses of cells cultivated in 2D versus 3D, respectively. 16
Conclusions
This study showed the easy implementation of automation of a novel dextran-based hydrogel system for 3D cell culture. The production and handling of the hydrogel-based 3D tissue was reproducible and reliable using a state-of-the-art liquid-handling robot. In this hydrogel, the HCT-116 cells showed a typical spheroid formation, as observed in other hydrogels. Furthermore, the presented data prove that cell viability and DNA content were similar in hydrogels that have been produced either manually or by automated methods. In an industry-relevant approach, we performed a fully automated dose-response experiment with Taxol. It could be shown that the manual procedure was easily adapted to the liquid-handling platform, making this dextran-based hydrogel system especially suitable to improve the drug development process. Novel 3D cell culture systems that are automation compatible and standardizable are urgently needed to replace state-of-the-art 2D screens to obtain more reliable data in drug development. The dextran-based hydrogel used here showed an excellent performance in automation due to the easy pipetting of liquid components and biocompatible gel formation and dissolution.
In this study, we showed the feasibility of automating a standard process that is used in the pharmaceutical industry in high-throughput screening with one compound, namely, Taxol. In further studies, focus will be on testing other compounds as well as the incorporation of other cell types in the hydrogels. The modular composition of the hydrogel allows for adapting the stiffness to mimic stiffnesses of particular tissues as well as modifying the gel with other cell adhesion motifs to create tissue models resembling the native properties of the respective in vivo tissue. The deposition of more than one gel layer can be used to create assays that require different 3D compartments, for example, for cell migration and invasion assays, or cultures mimicking tissues with different cell layers, such as epithelial barrier models. Accordingly, future tests will also include the automation of more complex 3D cell cultures, including heterogeneous cell types, to reflect the interacting behavior of cells in tissues.
Footnotes
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Brigitte Angres is an employee and co-owner of Cellendes GmbH, which produces and commercializes 3-D Life Reagents used in this work. Susanne Braum and Isabel Patocchi-Tenzer are employees of Tecan Schweiz AG, which manufactures and commercializes the liquid-handling robots Freedom EVO 150 used in this work.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was carried out within the framework of the competence center “Tissue Engineering for Drug Development” TEDD, which was provided with start-up financing by the Gebert-Rüf Foundation Switzerland [GRS-040/10].
