Abstract
Influenza A H1N1/2009 is a highly infectious, rapidly spreading airborne disease that needs to be monitored in near real time, preferably in a microfluidic format. However, such demonstration is difficult to find as H1N1 concentration in aerosol samples is extremely low, with interference from dust particles. In this work, we measured Mie scatter intensities from a microfluidic device with optical waveguide channels, where the antibody-conjugated latex beads immunoagglutinated with the target H1N1 antigens. Through careful optimizations of optical parameters, we were able to maximize the Mie scatter increase from the latex immunoagglutinations while minimizing the background scatter from the dust particles. The aerosol samples were collected from a 1:10 mock classroom using a button air sampler, where a nebulizer generated aerosols, simulating human coughing. The detection limits with real aerosol samples were 1 and 10 pg/mL, using a spectrometer or a cell phone camera as an optical detector, respectively. These are several orders of magnitudes more sensitive than the other methods. The microfluidic immunosensor readings are in concordance with the results of reverse transcription polymerase chain reaction. The assay time was 30 s for sampling and 5 min for the microfluidic assay.
Introduction
Influenza A H1N1/2009 is a highly infectious disease that initiated a global pandemic in 2009. 1 H1N1/2009 spreads through air from human respiration and/or coughing, and its spread is known to be very rapid. The use of a near-real-time, field-deployable biosensor device is the key to monitor the spread of H1N1/2009 (and other airborne viral pathogens as well). The current gold standard in detecting H1N1/2009 is reverse transcription polymerase chain reaction (RT-PCR) using the specimens collected directly from patients. Millions of copies of a specific genetic sequence from the virus can be produced with RT-PCR, ensuring a low detection limit and high specificity. Specimens are typically collected from the patients’ nose or throat (nasopharyngeal swab or oropharyngeal swab). 2 The RT-PCR assay may take up to 6 h, including sample preprocessing, gene extraction, reverse transcription, thermocycling, and gel imaging for product identification. This process still requires laboratory space, which adds additional time lag to sample delivery.
Field-deployable biosensors should ideally be portable, inexpensive, and near real time. The microfluidic platform is an optimum medium to meet these requirements. In fact, a couple of works have recently been published in detecting influenza A subtype H1N1 with RT-PCR in a microfluidic format. Ferguson et al. 3 used a specimen from a patient’s throat, collected the viruses in a microfluidic channel, and ran PCR thermocycling in a small heat chamber that was connected to a microfluidic channel. Since they did not use the microfluidic channel for thermocycling, known as “continuous-flow PCR,” 4 their total assay time was about the same as the laboratory-based conventional PCR time: 3.5 h. The lowest detection limit was 10 TCID50 (50% tissue culture infectious dose), which is typical of PCR assays. Kao et al., 5 on the other hand, did use the continuous-flow PCR method (but did not demonstrate the virus capture) and was able to finish the thermocycling in 75 min. Although 75 min is faster than 3.5 h, it did not include preprocessing and product identification time, and it is still too slow to be considered near real time. Their lowest detection limit was 20 to 30 pg/µL (= ng/mL) DNA molecules (they did not use the real virus samples). These microfluidic PCR methods do not meet the above-mentioned requirements, since they are not near real time and difficult to fabricate and operate.
Immunoassay-based (i.e., antibody-based) detection of H1N1 virus in a microfluidic format can provide much quicker assays, since they involve neither complicated gene extraction steps nor a lengthy thermocycling process. A couple of works have also been published recently, using antibody to H1 antigens. Unfortunately, their lowest detection limits are inferior to those of microfluidic PCR methods. Lee et al. 6 reported the detection limit of 1-µg/mL antigens using fluorescent-labeled anti-H1. Most commercial rapid kits that use membrane immunoassay (i.e., lateral-flow assay) show the detection limit of 103.3 to 104.7 TCID50/mL, 7 much inferior to the PCR methods. Li et al. 8 used nanoparticles to improve the detection limit down to 0.5 ng/mL antigens, but they used isolated antigens rather than patient sample.
None of the above microfluidic demonstrations used specimens collected from air. In fact, studies on H1N1 virus detection from air are very difficult to find, although it is a desired method in monitoring the spread of H1N1. Coughing from the infected patient generates aerosols that contain H1N1/2009 viruses. These aerosols can be captured with a commercial aerosol sampler, but the resulting specimen contains other proteins from the patient as well as dust particles inherently found in human environments. In addition, the H1N1 concentrations in the captured aerosol samples are considerably lower than those from the patients’ nose or throat.
The objectives of this research are 4-fold. First, a microfluidic device should be fabricated and tested for detecting H1N1/2009 viruses from aerosol samples. Second, the lowest detection limit should be as low as possible to enable the detection from such aerosol samples. Third, such detections should be made in the presence of dust particles. Fourth, the total assay time should be made as near real time as possible, preferably less than 10 min.
Our group has used the Mie scattering measurement of latex immunoagglutination assay, since the greater intensity change with the growth of latex beads affords lower levels of detection. 9 In a latex immunoagglutination assay, antibodies are covalently conjugated to the submicron- or micron-sized latex beads. In the presence of target antigens, antibodies are able to bridge two or more particles together, effectively “agglutinating” the latex beads (i.e., the growth of latex beads). Quantifying Mie scatter is a perfect means to monitor this growth, as it is highly dependent on the size (and morphology) of latex beads and less dependent on the wavelength of incident light or the concentration of latex beads. Detailed theories and equations can be found in Heinze et al. 9 and You et al. 10 and will be described here briefly.
When incident light hits a latex bead, it scatters light to every possible direction. The scattering angle is defined as follows: the same direction of incident light = forward 0° scatter, perpendicular to it = 90° scatter, and back to the light source = 180° back scatter. The size of latex beads is typically submicron or micron, which is comparable to or greater than the wavelength of visible light (i.e., incident light can “see” or “recognize” an individual bead), making this particular situation fall in the Mie scatter regime. (If the bead size is sufficiently smaller than the wavelength of incident light, it becomes Rayleigh scatter.) Since the latex bead is large enough to be recognized by the incident light, it can scatter light from multiple points, and these different scattering centers can both constructively and destructively interfere with each other. This leads to nonsymmetric angular dependence of the scattered intensity for the latex beads.11–13 If the Mie scatter intensities are plotted against the scattering angle, they will fluctuate or oscillate over the angle (which will be shown in the Results and Discussion section). Due to the different refractive indices and sizes of latex beads and the sample matrices (especially dust particles in our case), such oscillation peaks may or may not overlap with each other. We can adjust the wavelength of incident light and the size of latex beads to deliberately make such peaks not overlap with each other. This effectively enables us to maximize the scatter from latex beads and minimize the scatter from dust particles.
Performing Mie scatter detection of latex immunoagglutination in a microfluidic device provides the additional advantage of “controllability” (i.e., the time-dependent immunoagglutination reaction can be precisely controlled under a strict laminar flow within a microfluidic channel).
In addition, the use of a cell phone camera as an optical detector (instead of a spectrometer) may further make the biosensor portable and easy to use. Cell phone cameras have recently gained popularity in chemical 14 and biological 15 sensing. For example, cancer cells were identified from an enzyme-linked immunosorbent assay (ELISA) plate using a cell phone camera. 15
In the present study, we sought to develop a microfluidic immunosensor using a miniature spectrometer or a cell phone camera as an optical detector. Mie light scattering from immunoagglutination of antibody-conjugated, submicron latex beads was used to quantify the amount of H1N1/2009 target in the aerosol particles captured by an air sampler. Aerosol samples were collected from a 1:10 scale mock classroom (with an air conditioning/ventilation system and a nebulizer that simulates human coughing). The ability to detect H1N1/2009 virus antigens in the presence of dust particles must be demonstrated, with an extremely low detection limit (substantially lower than 1 ng/mL, which has not yet been demonstrated so far). This assay will be compared with the RT-PCR assay, which is the current gold standard protocol. We will do this comparison only qualitatively, since quantitative comparison between the immunoagglutination assay (targeting proteins) and RT-PCR (targeting nuclei acids) would technically be incorrect.
Materials and Methods
Assay Reagents and Target Molecules
Antibody-conjugated latex bead suspensions were used to detect H1N1/2009 from the samples in a microfluidic device. Anti-H1 (R01419; Meridian Life Science, Saco, ME) was conjugated to the 920-nm highly carboxylated polystyrene beads (10.3 Å2 parking area per surface carboxyl group; Bangs Laboratories, Fishers, IN) by covalent binding, as described in previous research. 16 Anti-H1 specifically binds to the type 1 hemagglutinin protein on the surface of H1N1 virus capsid. We have also tested anti-N1 (type 1 neuraminidase protein) but found their binding strength (as determined by the slope of a standard curve) was inferior to that of anti-H1.
NATrol Influenza A H1N1 External Run Control stock (Zeptometrix Corporation, Buffalo, NY; cat. NATFLUAH1N1-ERCM), which includes modified, noninfectious virus particles from the influenza A/NY/02/2009 H1N1 strain, was used as a target.
Mie Scattering Simulation
Mie scattering simulations were attempted to estimate the effect of dust particles on the microfluidic immunosensor readings. Online software by Prahl 17 was used with parameters of bead/particle diameter, including refractive indices of water, latex beads, and dust particles (1.33, 1.59, and 1.40, respectively), as described in previously published work. 10 The average size of dust particles was determined by analyzing the microscope images of aerosol samples using ImageJ software (National Institutes of Health, Bethesda, MD). The refractive index of 1.40 was used for the dust particles since most of the dust particles in the human environment were from human skin (i.e., collagens and elastins). 18
Fabrication of the Microfluidic Device
The microfluidic device (with optical waveguide side channels) used in this study was fabricated the same way as described previously. 9 The silicon master mold was single crystal silicon fabricated by photolithography and deep reactive ion etching (DRIE), where the fluidic channels were 1 mm wide and 100 µm high. To achieve a high-depth pattern without any sacrificial layer, thick photoresist, AZ 4620 (~6 µm thick), was used as a silicon etching mask. During the DRIE process, C4F8 (110 standard cubic centimeters per minute or sccm) and SF6 (130 sccm) with O2 (13 sccm) were used for the passivation and etching steps. Polydimethylsiloxane (PDMS) was then poured onto the master mold and cured in a convection oven for 1 h. The PDMS was then peeled off the master mold, exposing the y-channel, which was subsequently bonded to a thin PDMS cover slide via oxygen plasma treatment for 2 min (Plasma Preen Cleaner/Etcher; Terra Universal, Fullerton, CA). This oxygen plasma treatment causes the surface of the microchannel to become hydrophilic (water contact angle of 10°–20°). The fluidic sample channel is a simple y-channel design, containing two inlets and one outlet, with waveguide channels in “close proximity” (separated from fluid channels by 100-µm walls). The waveguide channels were prefilled with silicone oil (microscope immersion oil type A), where the silicone oil acts like a core (refractive index = 1.52) and the surrounding PDMS acts like a cladding (refractive index = 1.41) of an optical fiber. The critical angle of this optical waveguide is arcsin (1.41/1.52) = 68°. There is a reservoir at each inlet to load the antibody-conjugated bead suspension and the sample solution. Teflon tubing was then attached to the chip at the outlet channels to manually pump the bead suspension and sample into the y-channel by withdrawing action (at a flow rate of 10 µL/min).
Assembly of Immunosensor Devices
Two types of immunosensor devices were fabricated as shown in Figure 1 , one with a miniature spectrometer (USB4000; Ocean Optics, Dunedin, FL) and the other with a cell phone camera (iPhone 4; Apple, Inc., Cupertino, CA) as an optical detector. The disposable microfluidic device and optical fibers were placed on a “tray” as shown in Figure 1 . The tray achieved a friction fitting to the red light-emitting diode (red LED; light source) and a detector (a miniature spectrometer or a cell phone camera). Assays were performed as follows: first, the antibody-conjugated bead suspension and the sample solution were loaded to each inlet. Surfactant (Tween 80) was added to the sample solution to break off larger agglutinated beads, thus resulting in mostly doublets and triplets, where the scattering is maximum (see Results and Discussion).9,10 Then, these solutions were manually pumped using negative pressure created by a syringe (201000; Hamilton Company, Reno, NV) that was connected to the tubing attached at the outlet. After mixing, the 640-nm red LED light (LS-450; Ocean Optics) was directed to the center of the y-channel via the optical fiber and waveguide channel, which was filled with silicone oil. Forward light-scattering intensity by irradiated light was collected by a detector. In cases using a spectrometer, the collected light intensity signal was sent to a laptop computer and analyzed with SpectraSuite software (Ocean Optics). In cases using a cell phone camera, images were taken every 15 s for 3 min, and each image was analyzed using ImageJ software. The image was captured at 24 bit with 5-megapixel resolution, broken into three color components of red, green, and blue (RGB), each at 8 bit (0–255). Only the red intensities were considered. Any intensities lower than 20 were considered background noise and neglected. All other intensities higher than 20 (out of 255) were integrated for all 5 megapixels and used this value as the overall red intensity (I) for a single image. These values were normalized by dividing them with the integrated red intensities from the results of the phosphate-buffered saline (PBS) (I0).

Microfluidic immunosensors with a miniature spectrometer (USB4000 from Ocean Optics, Dunedin, FL; top) and a cell phone camera (iPhone 4 from Apple, Cupertino, CA; bottom). The optical fibers were partially inserted into the optical waveguide channels, all contained within a “tray.” The distal ends of optical fibers were connected to the red light-emitting diode (red LED; light source) and the detector (a miniature spectrometer or a cell phone camera) using friction fittings. Anti–H1-conjugated latex bead suspension and the diluted solution of collected air sample were separately introduced to two inlets of a y-shaped microfluidic channel and pumped through by manually withdrawing a disposable syringe from the channel outlet. The side channels were filled with silicone oil to act as liquid core optical waveguides.
Air Sampling from a Mock Classroom
A mock classroom was constructed with acrylic plates as shown in Figure 2 . The scale of the mock classroom is 1:10 of a real classroom at the University of Arizona. The area and volume of the real classroom is 37.96 m2 and 94.91 m3, respectively. Since two inlets with fans and one outlet were installed for ventilation in a real classroom, 1:10 scaled versions of inlets, fans, and an outlet were installed in a mock classroom modeled after the real ones. The fans were fabricated using the cooling fans for personal computers (35 × 35 mm), with the plastic enclosures designed by SolidWorks (Dassault Systemes SolidWorks Corp., Waltham, MA) and rapid prototyped with a 3D printer (Dimension uPrint SE; Stratasys, Inc., Eden Prairie, MN). The flow rates from these fans were controlled by the analog voltage output from an Arduino microcontroller (Boulder, CO). The actual flow rates from the fans were measured by traceable hot wire anemometers, installed within the plastic enclosures just underneath the fans. The fans introduced the naturally occurring dust particles from the laboratory into the mock classroom. The outlet was vented into a class II biosafety cabinet with UV sterilization capability (NuAire, Plymouth, MN) using 5-cm diameter plastic tubing. The upper casing of the classroom as well as the area around the base of the nebulizer was sealed for each experiment. The velocity of each fan was scaled down using the Archimedes number (Ar), as previously published, 19 using the experimentally measured velocity from the real fans. The ventilation rate of the mock classroom was 29.7 L/min or 1.05 cfm (cubic feet per minute), which is about seven times higher than the minimum requirement recommended by American Society of Heating, Refrigerating and Air-Conditioning Engineers (ASHRAE; 4.25 L/min = 0.15 cfm). 20 The minimum required ventilation rate (0.15 cfm) was also applied to compare the signal difference caused by the lower ventilation rate. No window was installed and the door was closed. A mock human was installed to spray aerosols that contained the modified H1N1/2009 viruses. The mock human was made of a commercial nebulizer (4650D-621; Sunrise Medical, Somerset, PA), and it was installed inside the mock classroom as shown in Figure 2 . The mean diameter of aerosols that contain viruses was 5 µm according to the specification of the nebulizer manufacturer.

A photograph (top) and a top view (bottom) of a 1:10 scale mock classroom. The locations of the air ventilation system (fans and outlet) and aerosol samplers (outlet, left, and right) are shown.
Sample Collections
The aerosolized sample injected by a nebulizer was collected with a button aerosol sampler (225-360; SKC, Eighty Four, PA). The aerosol particles were injected for 3 s by pressurized nitrogen gas. The velocity of injection was 11.0 m/s. After 3 s of injection, the aerosols were transported for 3 min by the ventilation system. Then, the button aerosol sampler collected the injected aerosol samples for 30 s at three different locations (outlet, left, and right, as shown in Fig. 2 ). This sampling time (30 s) was determined by the set of experiments that showed a sufficient difference in detectable signals over three different sampling locations. The aerosol samples were captured by a filter (1820-070; Whatman, Kent, UK) inside the sampler. The filter was soaked in 150 µL of PBS (pH 7.4; Sigma-Aldrich, St. Louis, MO) to dissolve virus particles into liquid and then squeezed by a syringe (S7510-3; National Scientific, Rockwood, TN). The button air sampler was cleaned with 70% ethanol and deionized water between each sample. A total of six sets of experiments were conducted, varying the concentrations of H1N1 virus particles inside the nebulizer (1–100 ng/mL), the ventilation conditions (29.7 and 4.25 L/min), and the three different sampling locations (outlet, left, and right), each of which was replicated three times. The sampler was installed at a scaled height of 0.15 m (a typical height of a human mouth, scaled down by 1:10) for all cases.
RT-PCR Assays
For verification of the immunosensor readings, RT-PCR was run on aerosol samples. In the immunosensor experiments, 1- to 100-ng/mL solutions of H1N1/2009 virus particles were aerosolized into the mock classroom for 3 s, and the resulting concentration after sampling was 1 to 100 pg/mL (see Results and Discussion). Although the Mie scatter-based microfluidic immunosensor was sufficiently sensitive to detect this low concentration, the concentration was too low to be detected with RT-PCR. Since the genome in H1N1/2009 virus particles comprised only 0.1% of the total weight, 1 to 100 fg/mL of the genome mass would appear in the collected aerosol samples.
Rather than increasing the concentration of virus particles for RT-PCR by 1000-fold, which would be very dangerous and in violation of Centers for Disease Control and Prevention (CDC) regulations, genetic material was extracted from the H1N1/2009 viruses and injected into the mock classroom at high concentrations. Zymo Research Viral RNA Extraction Kit (Zymo Research, Irvine, CA) was used for this extraction. Briefly, the modified viruses were denatured, and the genomic RNA was captured and purified using the filter columns provided in the kit. The final RNA elution volume was 20 µL per sample. Immediately, the RNA template was used to make complementary DNA (cDNA) using a High Capacity Reverse Transcriptase Kit (Invitrogen Life Technologies, Grand Island, NY). The RT program consisted of an initial 25 °C step for 10 min, a 37 °C step for 2 h, and a final 85 °C step for 2 min, and random hexamer primers were used for conversion. The resulting product was 20 µL cDNA per sample and was stored at −20 °C prior to mock classroom experiments.
Then, 10- to 50-ng/mL solutions of cDNA from H1N1/2009 virus particles were introduced into the mock classroom, under the identical conditions described in “Sample Collections,” and this yielded approximately 10- to 50-pg/mL cDNA concentrations in the collected aerosol samples.
Promega 2× Green Master Mix Kit (Promega Bio-Tek, Madison, WI) with DNA polymerase was used to amplify the cDNA sequence. The 2009 H1N1 M gene was targeted with the following specific primer sets: MP-39-67For (5′-CCMAGGTCGAAACGTAYGTTCTCTCTATC-3′) and MP-183-153Rev (5′-TGACAGRATYGGTCTTGT- CTTTAGCCAYTCCA-3′). 21 Up to 50 cycles were run in a conventional thermocycler (MJ Research, Watertown, MA, USA). To determine PCR product, standard gel electrophoresis was performed with fluorescent imaging. Briefly, 2% agarose gel (Sigma-Aldrich) was cured and immersed in 1× Tris-acetate-EDTA (TAE) buffer (Invitrogen Life Technologies), and a power supply was used to provide 120 V and 0.1 A for 45 min (Thermo Fischer Scientific, Waltham, MA, USA). Amplified samples were loaded into the gel, and a 1-kbp DNA ladder (Invitrogen Life Technologies) was loaded for size comparison. The gel was soaked in ethidium bromide solution (Sigma-Aldrich) for 20 min and imaged using a Gel Doc 1000 imaging system (Bio-Rad Laboratories, Hercules, CA). ImageJ software (National Institutes of Health) was used for image processing and band intensity determination in gray scale. Band intensity was determined by measuring pixel intensity, which was normalized to the gel background. These results were compared with those collected with the microfluidic immunosensor.
Results and Discussion
Ability to Detect the Latex Immunoagglutination in the Presence of Dust Particles
A series of Mie scattering simulations was performed for both immunoagglutinated latex beads and dust particles to optimize optical parameters—namely, the bead size d, the wavelength of incident light λ, and the angle θ where the scattering detection is made. Figure 3 shows the best result, indicating that θ = 45° maximizes the scattering from immunoagglutinated latex beads while minimizing that from dust particles, for d = 920 nm and λ = 640 nm. The size of dust particles in captured aerosol samples was measured to be on average 15 µm, evaluated from their microscope images. Mie scattering intensity of dust particles was an order magnitude lower than that of immunoagglutinated latex beads with θ > 40°. Since the size of dust particles in a real environment may vary, typically from 1 to 100 µm, 22 it is necessary to conduct a series of simulations for varying sizes of dust particles. For the dust particles larger than 15 µm, the scattering intensity became even smaller (results not shown), and hence these were not problematic. For the dust particles smaller than 15 µm (the result with 1.5 µm, shown in Fig. 3 ), the scattering intensity became substantially stronger for the angles smaller than 40°, while the scattering at θ > 40° remained relatively the same, with the minimum point also occurring at around 45°. This result indicates that Mie scattering light intensity of dust particles does not overlap with that of immunoagglutinated beads with the optimized scattering parameters. Therefore, it is possible to detect viral pathogens in the presence of dust particles.
Mie scattering simulation results for the immunoagglutinated latex beads and dust particles. θ = 45° maximizes the scattering from immunoagglutinated latex beads while minimizing that from dust particles, for d = 920 nm and λ = 640 nm. θ < 30° was not usable due to the interference of incident light, and θ > 60° was not usable due to the reflection at the fluidic-waveguide channel interface.
Standard Curves and Lowest Detection Limits
Standard curves were constructed with serially diluted H1N1/2009 solutions in PBS ( Fig. 4 ) using a microfluidic immunosensor with both optical detectors (spectrometer and cell phone camera). With a spectrometer, the light intensity readings (I) were recorded at 640 nm (the wavelength of incident red LED), which were divided with that of a PBS solution (I0). The standard curve with a spectrometer detector shows a good linear trend to the logarithm of H1N1 concentration, up to 100 pg/mL, with R2 = 0.9959. This linearity is significantly improved from the previous study, 9 through the use of optimized scattering parameters and the optimized use of surfactant (Tween 80). It seems that Tween 80 was able to break apart larger agglutinated beads and resulted mostly in doublet and triplet beads, as confirmed by microscope image analysis (data not shown). The strongest scattering increase was observed with triplet agglutination as confirmed with Mie scattering simulation; if the beads are immunoagglutinated into a very big clump, the effective number of beads in the microfluidic channel is reduced dramatically, negating the effect of bead size growth. The detection limit can be determined from a standard curve by using the three-sigma rule (i.e., the lower bounds of error bars multiplied by three times should be still above 1, to be statistically different from that of PBS). With this three-sigma rule, the detection limit was less than 1 pg/mL, significantly lower than most other studies (0.5 ng/mL to 1 µg/mL).6–8
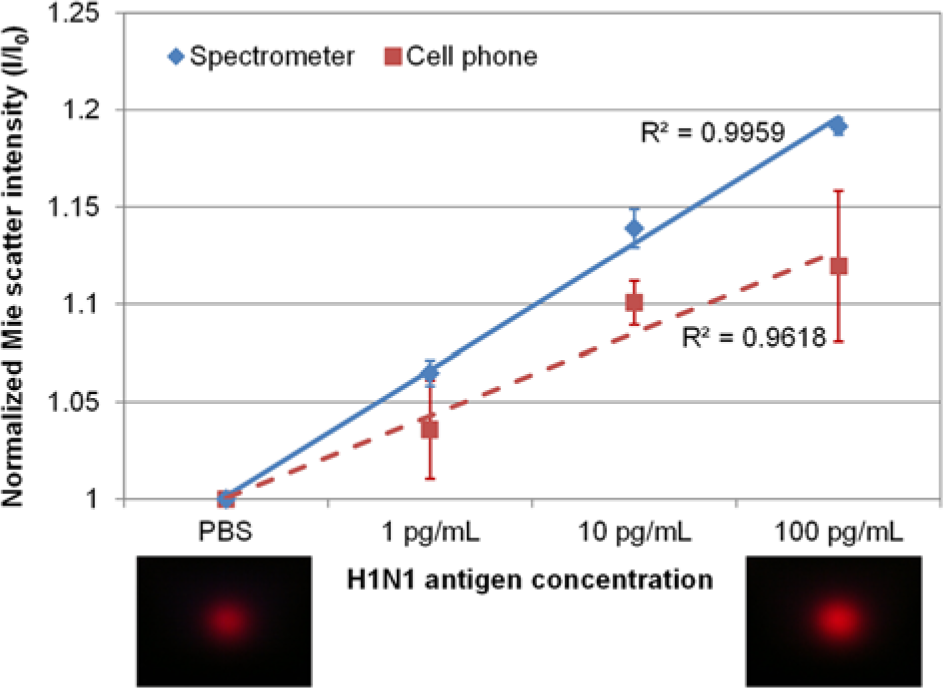
Standard curves constructed by microfluidic immunosensors. The detection limits were <1 pg/mL with a spectrometer and 10 pg/mL with a cell phone camera. All data points are averages of three different experiments. Error bars represent standard errors. Cell phone images from optical fibers are also shown at the bottom, for a negative control (phosphate-buffered saline [PBS]) and 100-pg/mL H1N1/2009 virus solutions, showing significant differences in brightness and area of the red spots.
The same standard curve was constructed using a cell phone camera as an optical detector. The actual cell phone images were shown underneath the graph, for PBS and the 100-pg/mL H1N1 solution. It shows significant differences in brightness and area of the red spots. The red pixel intensities (above the background noise level) were integrated and used as the light intensity readings (I). These readings were again divided with that of a PBS solution (I0). The standard curve shows a linear trend up to the logarithm of H1N1 concentration, to 100 pg/mL, but with a smaller slope, bigger error bars, and with smaller R2 = 0.9618. The detection limit is 10 pg/mL, again using the same three-sigma rule. This can easily be explained with the dynamic range of analog-to-digital converters in two different systems. Despite the 24-bit image processing of a cell phone camera, it is divided among three different color components—red, green, and blue—an thus the effective resolution for red color is 8-bit (note that the system was exposed to a 640-nm red light source). The miniature spectrometer has a 16-bit resolution for all wavelengths, which is reflected in its greater sensitivity. Despite this, there is a stronger commercial potential in the cell phone camera usage as the miniature spectrometer has a price 10 times higher than that of a typical smartphone.
Immunosensor Readings From Collected Air Samples
We then detected H1N1/2009 virus antigens from the aerosol sample collected from a mock classroom. A nebulizer sprayed aerosols within a mock classroom, simulating a human coughing. The aerosols traveled through a mock classroom with the ventilation system, eventually escaped through the outlet, were trapped on the wall, or were captured by the air sampler. The button aerosol sampler collected a small portion of these aerosols at three different locations (outlet, left, and right). The total amount of injected H1N1/2009 viruses was 50 pg or 5 ng, coming from 1-ng/mL or 100-ng/mL solutions in a nebulizer, respectively. Two different ventilation conditions were considered, referenced as high (29.7 L/min) and minimum (4.25 L/min), where minimum refers to the minimum required ventilation rate suggested by ASHRAE, while high refers to the ventilation rate corresponding to the real classroom operation.
Figure 5 (top) shows the normalized light-scattering intensities for six different cases. For most cases, the cell phone camera detector showed a smaller signal compared with the spectrometer detector, consistent with the standard curves ( Fig. 4 ). Using the standard curves ( Fig. 4 ), all data were converted into the H1N1 concentrations, and the results are shown in Figure 5 (bottom). After the conversion, the estimated H1N1 concentrations with a cell phone camera are not very different from those with a spectrometer, mostly within one order of magnitude difference.
Top: microfluidic immunosensor readings for six different cases of experiments. All data points are averages of three different experiments. Error bars represent standard errors. Bottom: estimated H1N1 concentrations using the standard curves ( Fig. 4 ). Both types of immunosensors show similar concentrations mostly within one order of magnitude difference.
Under the minimum ventilation condition (4.25 L/min) and high H1N1 concentration in a nebulizer (100 ng/mL), the concentrations of captured H1N1 viruses are significantly different by three sampling locations: very high at outlet and right, while very low at left. These variances can easily be explained by the unequal distribution of air by the low (insufficient) ventilation condition. The H1N1 concentrations are 1 to 120 pg/mL with a spectrometer or 0.19 to 1000 pg/mL with a cell phone camera, which are higher than those with a high ventilation condition (29.7 L/min) and a low H1N1 concentration in a nebulizer (1 ng/mL).
Under the high ventilation condition (29.7 L/min) and low H1N1 concentration in a nebulizer (1 ng/mL), the concentrations of captured H1N1 viruses are 1.2 to 3.5 pg/mL with a spectrometer and in the order of outlet > left > right, although such differences are not very significant. In fact, these concentrations are at the detection limit of our system with a spectrometer, 1 pg/mL. The result with a cell phone camera is similar, again in the order of outlet > left > right, but with bigger variance, 1.9 to 10 pg/mL. This can easily be explained by the detection limit with a cell phone camera, 10 pg/mL. The total injected amount of H1N1/2009 virus was 50 pg for the 0.1-m3 mock classroom, indicating the detection limit of our microfluidic immunosensor would be 50 pg per 0.1-m3 volume.
Altogether, our microfluidic immunosensors are capable of correctly estimating the concentration of captured H1N1 viruses, with respect to the ventilation conditions and the H1N1 concentrations in a nebulizer. The assay time includes 30 s for sampling and 5 min for the microfluidic assay (i.e., total of 5 min 30 s).
RT-PCR Results from Collected Air Samples
As control experiments, cDNA solutions from H1N1/2009 viruses were aerosolized into a mock classroom and analyzed using RT-PCR. From Figure 5 , we determined that the concentrations of H1N1/2009 viruses in the captured aerosol samples were 1 to 100 pg/mL, which was about 0.1% of the H1N1/2009 concentration in the nebulizer (1–100 ng/mL). Since the genome is only 0.1% of the total virus weight, the genomic mass in the captured aerosol samples would be only 1 to 100 fg/mL (= 1–100 ag/µL), which would be impossible to analyze with RT-PCR. Therefore, we used 10- to 50-ng/mL cDNA solutions in the nebulizer, yielding 10 to 50 pg/mL (= 10–50 fg/µL) cDNA in the captured aerosol samples.
Figure 6 shows the gel electrophoresis images after 50 cycles of RT-PCR taken at three different locations (outlet, left, and right), at two different ventilation conditions (high and minimum), and at various cDNA concentrations loaded in the nebulizer. Except for the cDNA concentrations, all other experimental conditions were identical to those of Figure 5 . Under minimum ventilation conditions, large amounts of cDNA were captured in the outlet and right samplers, while those captured in the left sampler were consistently smaller. Under high ventilation conditions, the amounts of cDNA captured in the samplers were as follows, from the highest to the lowest concentrations: outlet > left > right. Both RT-PCR results (under minimum and high ventilation conditions) show the same trend found in Figure 5 (with microfluidic immunosensors), demonstrating the capability of our microfluidic immunosensors in accurately quantifying the H1N1 viruses from the collected aerosol samples.
Gel electrophoresis images after 50 cycles of reverse transcription polymerase chain reaction (RT-PCR) for the aerosol samples collected from the mock classroom, under minimum and high ventilation conditions. The complementary DNA (cDNA) concentrations inside the nebulizer were 50, 25, and 10 ng/mL (from top to bottom), and the expected cDNA concentrations in the aerosol samples were 50, 25, and 10 pg/mL (= 50, 25, and 10 fg/µL). Experimental conditions are identical for each set of three gel bands. The intensities of gel bands were evaluated using ImageJ software (National Institutes of Health, Bethesda, MD), all normalized to the intensities of gel background.
For both the microfluidic immunosensor and RT-PCR, the outlet sample location showed the highest amounts of targets collected. This finding is evident as the outlet location is the nearest to the ventilation duct, so the bulk airflow movement in the mock classroom would carry a majority of the aerosols in that direction. The data also suggest that lower amounts of aerosols were collected at the left sample location, especially under minimum ventilation conditions. At the right sample location, however, there is a degree of randomness. Under minimum ventilation conditions, the immunosensor readings varied from experiment to experiment, resulting in larger error bars. They also varied due to the nature of the optical detectors (a spectrometer or a cell phone). The RT-PCR results were mixed at the right sampler location but consistently higher than at the other locations. This location is closest to the nebulizer (a mock human), despite the fact that the aerosols initially travel in the opposite direction, away from the right sampler. However, at high ventilation, collected sample concentrations were consistently lower than at the other locations, sometimes negligible, at the right location. This is because the aerosols travel toward the outlet for the duration, while they stay in the mock classroom substantially longer than at the minimum ventilation.
In conclusion, a microfluidic device was developed to detect and quantify H1N1 viruses in the captured aerosol samples from a mock classroom. A nebulizer generated aerosols within a mock classroom, simulating human coughing. A button aerosol sampler collected aerosols at three different locations. Mie scatter intensities were measured from a microfluidic device, through optical waveguide channels, where the antibody-conjugated latex beads immunoagglutinated with the target H1N1 virus antigens. Through careful optimizations of optical parameters, we were able to maximize the Mie scatter increase from the latex immunoagglutinations while minimizing the background scatter from the dust particles, inherently found in aerosol samples. Both a miniature spectrometer and a cell phone camera were used as optical detectors, and the detection limits with real air samples were 1 pg/mL with a spectrometer and 10 pg/mL with a cell phone camera, which are several orders of magnitude more sensitive than the other methods (which used nose or throat swab samples). The immunosensor with a cell phone camera has a higher commercial potential with a lower price tag than a miniature spectrometer. The microfluidic immunosensor readings are in good agreement with the results of RT-PCR, corresponding very well with the different ventilation conditions and the target concentrations in a nebulizer. Further work is needed to make the aerosol sampling completely automatic. A button sampler was used in this study, which is essentially a filter with a vacuum pump. A glass impinger may be used to collect aerosols into liquid, followed by automatic introduction into the microfluidic immunosensor. In addition, several different types of airborne viral pathogens need to be tested to evaluate the selectivity of the optofluidic immunosensor. The assay time includes 30 s for sampling and 5 min for the microfluidic assay (i.e., 5 min 30 s).
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Funding for this work was provided by Animal and Plant Quarantine Agency, South Korea, award number C-AD14-2006-11-0.
