Abstract
Airborne pathogens affect both humans and animals and are often highly and rapidly transmittable. Many problematic airborne pathogens, both viral (influenza A/H1N1, Rubella, and avian influenza/H5N1) and bacterial (Mycobacterium tuberculosis, Streptococcus pneumoniae, and Bacillus anthracis), have huge impacts on health care and agricultural applications, and can potentially be used as bioterrorism agents. Many different laboratory-based methods have been introduced and are currently being used. However, such detection is generally limited by sample collection, including nasal swabs and blood analysis. Direct identification from air (specifically, aerosol samples) would be ideal, but such detection has not been very successful due to the difficulty in sample collection and the extremely low pathogen concentration found in aerosol samples. In this review, we will discuss the portable biosensors and/or micro total analysis systems (µTAS) that can be used for monitoring such airborne pathogens, similar to smoke detectors. Current laboratory-based methods will be reviewed, and possible solutions to convert these lab-based methods into µTAS biosensors will be discussed.
Background
Over the past 50 years, many advancements in the field of biosensors have been made. Traditionally, biosensors have been defined as devices or methods that are used to detect specific or nonspecific biological analytes (bioanalytes), either directly or indirectly, using chemical, electrical, or colorimetric means. The first biosensor, described in 1962, used an enzyme as a bioanalyte in corroboration with an electrochemical transducer to continuously monitor blood chemistry during surgery. More recently, these sensors have been designed to be far more sophisticated, easy to use, specific, and sensitive. 1 With the advancements in specificity and sensitivity came progress in modularity and multiplexing, allowing the end user to test for multiple targets in a single sample or even a single target in multiple samples.2–8 On a more complex scale, multiple targets may be tested in multiple samples simultaneously, and it is possible for feedback and automation mechanisms to be integrated within the unit. Overall, appropriate planning, design, and fabrication provide the manufacturer and user with the flexibility to easily change the targeted agents with simple manipulation. Although the altering of intended target is often a minute detail, different targets require different levels of sensitivity, thus complicating the issue of modularity.
With advances in medicine along with the growing concerns over antibiotic resistance, food safety, and bioterrorism, there has been a paradigm shift in biosensor technology to field-deployable diagnostics and point-of-care diagnostics.8–14 In particular, sensitivity and specificity have been targets of optimization and scrutiny in designing these sensors, with high attention to target type, native environment (e.g., the types of samples in which they are typically found), and testing conditions. Specifically, these tests have been enhanced to detect bioanalytes in a variety of conditions (e.g., wastewater, rinse water, soil, food, blood, tissue sample, and aerosol) despite the presence of interfering substances such as proteins, cells, lipids, and dust particles. However, a comprehensive biosensor that can easily be manipulated to test for various types of pathogens (viruses, bacteria, or parasites) in various samples with complete automated sampling, packaging, and feedback control has rarely been demonstrated. Paramount to the layout of a particular biosensor, especially portable biosensors, is the nature of the pathogen being detected as well as the nature of the sample in which it resides. For example, bacteria such as Escherichia coli, Salmonella enterica species, Enterococci species, Cryptosporidium parvum, and Campylobacter species are fecal coliforms and may be found in contaminated water supplies or fresh produce,15–18 whereas hepatitis, human immunodeficiency virus (HIV), filoviridae viruses, and malaria are commonly found in blood.19–22 Due to the diverse nature of these pathogens (e.g., bacteria cell walls, viral capsids, and cyst-forming units), it is very difficult to create a comprehensive protocol to test for broad-spectrum pathogens.
Although great strides have been made in detecting various pathogens in complex samples, little has been published regarding airborne pathogen detection, especially with realistic sampling procedures. In addition, virtually no commercially available biosensors exist that indicate airborne pathogens. Research that focuses on biosensors for diagnosing airborne bacteria and viruses typically does not include real-time air sampling and automation due to extreme complications, such as low pathogen content in aerosol particles, dust interference, and the problem of interfacing the captured pathogens with the actual sensing device. To truly test for highly infectious airborne pathogens such as arenavirus, influenzas, Neisseria species, severe acute respiratory syndrome (SARS), Mycoplasma species, and various fungi in the field, a complete biosensor accounting for the aforementioned parameters must be established. Proper design and flow will allow for field-deployable and stationary sensors to detect, in real time, airborne pathogens in hospitals and air vents, and even bioterrorism agents in public places such as subway stations and airplanes. With the advent of evolved airborne disease as well as emerging epidemic panzootic (can be transmitted to different species, including mammals) diseases such as Ebola, 23 there is an increasing need for these rapid, portable airborne sensors.
Common Airborne Pathogens
Viruses (as Shown in Table 1)
Influenza viruses are some of the most common highly infectious viruses worldwide and are spread through coughing and sneezing in the form of aerosols. 24 Influenza A/H1N1 is one of the most prominent groups of strains and leads to untold millions of cases per year in the United States. 24 Furthermore, due to the fact that symptoms are ambiguous and that diagnosis is slow and often inaccurate, many do not seek medical attention, thus leading to underreporting, especially in third world nations.25,26 Because of the relatively short incubation time (average of 2 days) 27 and the fact that the infectious period typically begins 12 h prior to the onset of symptoms, 28 influenza A is very difficult to control. Furthermore, seasonal influenza A strains generally mutate every year, making it challenging to predict accurate vaccines. 29 Although infected individuals usually recover within a week with only rest and possibly antivirals, those with compromised immune systems often require hospitalization. Due to these factors, influenza viruses lead to astronomical annual costs to the U.S. health care industry.30–32 A special case of influenza A was the 2009 H1N1 strain, which was considered pandemic from 2009 to 2010. 33 Since its discovery, this strain has led to more 60 million reported cases, more than 275,000 hospitalizations, more than 10,000 deaths, and billions of dollars in health care costs in the United States. 34 Worldwide, the total numbers have not been accurately reported but are expected to be much greater than the numbers reported for the United States. Although excellent collaborative efforts by the U.S. Centers for Disease Control (CDC) and World Health Organization (WHO) helped bring 2009 H1N1 to a more nonpandemic, controlled state, this instance illustrates how quickly viruses can mutate and become much more contagious and morbid. The high virulence, the ease of spread through aerosol particles, and the relatively short doubling time (2–3 days)35,36 make sensitive and rapid detection of such viruses paramount.
Common Airborne Viral Pathogens.
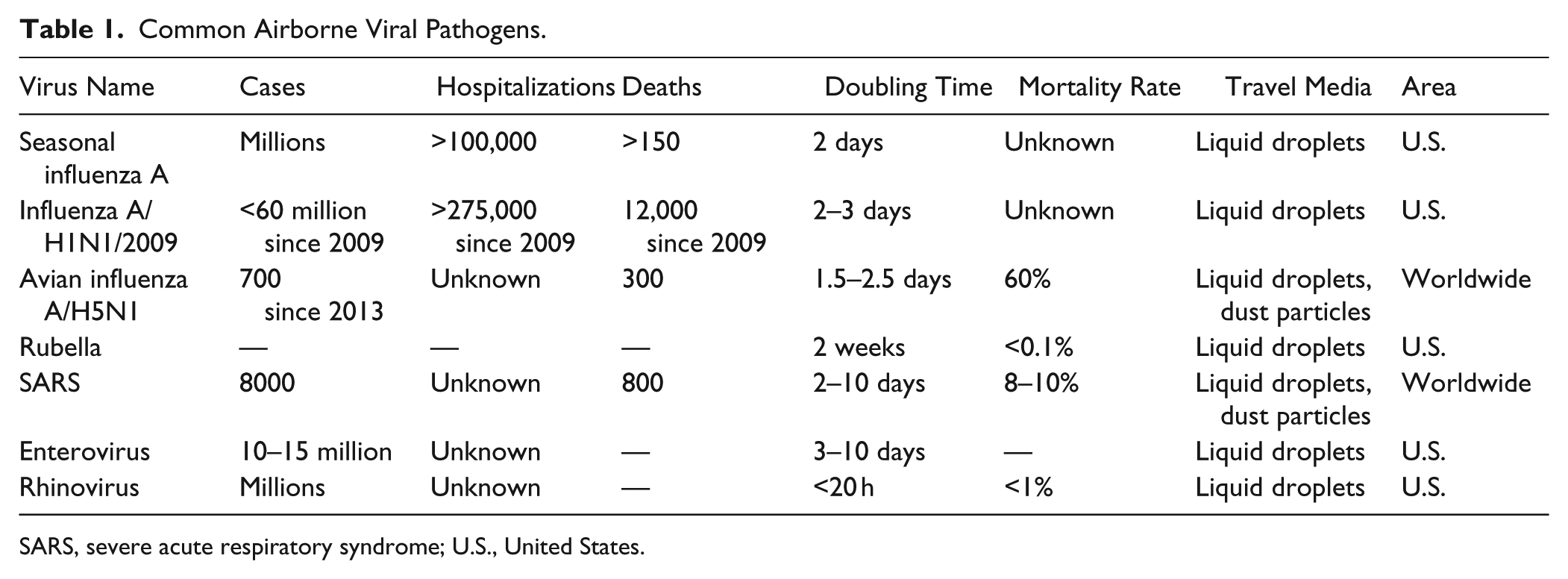
SARS, severe acute respiratory syndrome; U.S., United States.
Avian influenza (including subtypes H5N1, H5N2, H3N2, and H7N9) is a group of highly infectious viruses that primarily affect poultry and have caused great economic burden to the livestock industries of many nations.37–46 Furthermore, mutations leading to panzootic strains have been occurring at an increasing rate, raising concern for health care industries worldwide. Since 2013, it has been reported that nearly 700 humans have been infected, and this led to more than 300 deaths.47–50 In addition, there is a growing threat of the possibility of highly pathogenic H5N1 being engineered for bioterrorism.51–53
The Rubella virus, which causes a rash and a range of symptoms and is termed German measles, is becoming a disease of interest in the United States after being rare for the past 40 years due to the development of a vaccine;54–57it remains an issue in developing nations.58–61 Although nearly 50% of Rubella cases are mild enough to not require treatment, serious complications often occur, including blindness and congenital Rubella syndrome, which causes pregnancy miscarriages in nearly 20% of cases.62–64 The average incubation time of Rubella is 2 weeks, and transmission typically occurs through liquid droplet secretions (and thus aerosols) from the respiratory system (especially sneezing). 65
Bacteria (as Shown in Table 2)
Mycobacterium tuberculosis (TB) is a Gram-positive bacterium that is responsible for tuberculosis infections and complications such as meningitis and pneumonia. 66 Good efforts, primarily through latent TB testing, have reduced the average number of annual cases in the United States to 11,000–20,000, with approximately 500 deaths.67–72 TB has an incubation time of 4–6 weeks (with a doubling time of 4–24 h)73–75 and may remain latent within an infected individual for years before the onset of an active infection. Although a latent infection is not communicable, the infected individual may develop an active pulmonary infection at any time, during which the infection may be easily spread.76–78 These bacteria are spread in aerosolized sputum and saliva droplets that are primarily generated through coughing, making it highly communicable. In addition, the very low infectious dose [infectious dose 50% (ID50) = <10 colony-forming units (CFUs)] contributes to the disease’s high communicability.79,80 Although the United States has been successful in minimizing the number of TB infections during the past 50 years, this disease has been a huge problem worldwide, with approximately 14 million persons affected each year and nearly 2 million deaths, currently rendering it one of the most common global bacterial infections. 81 Despite great strides in minimizing TB outbreaks, the disease is becoming more common once again, and this problem is compounded by newer antibiotic-resistant strains.82,83 Due to these factors, TB appears to be a good target for real-time airborne detection, along with the standard latent testing, to prevent future outbreaks.
Common Airborne Bacterial Pathogens.
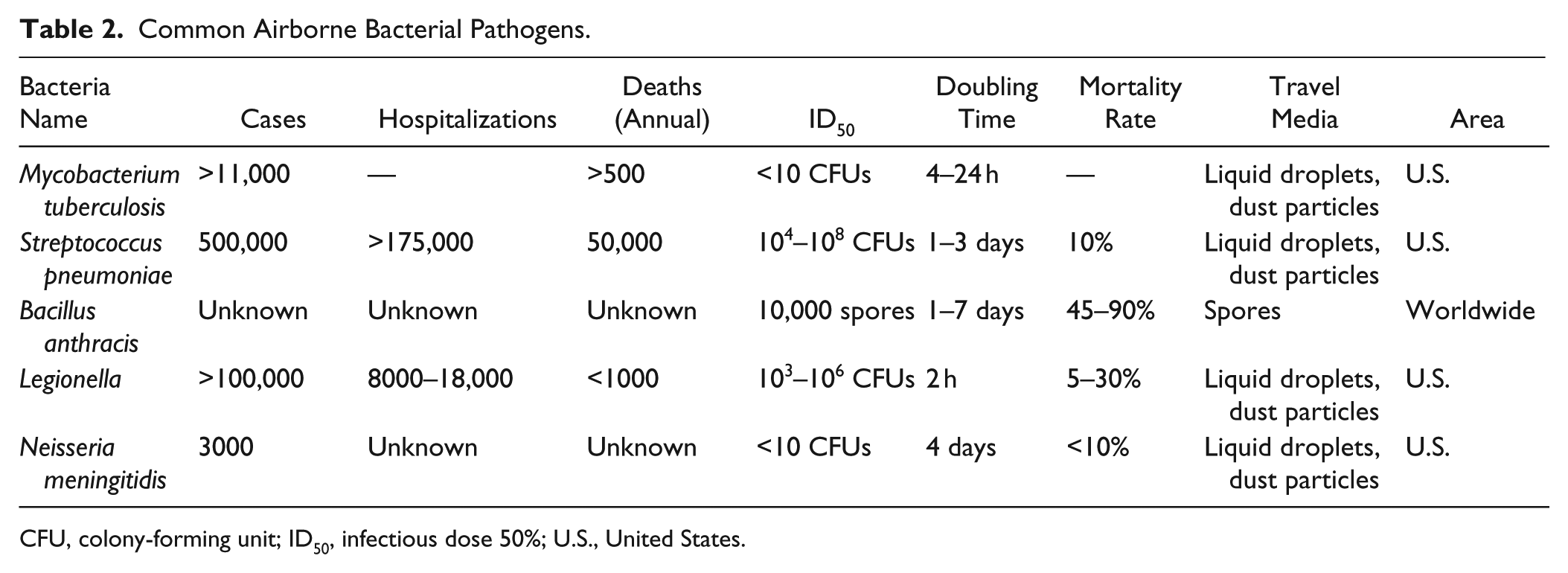
CFU, colony-forming unit; ID50, infectious dose 50%; U.S., United States.
Bacillus anthracis is a Gram-positive bacterium that causes various types of serious and often lethal infections. 84 Although anthrax is not contagious, the bacteria form endospores and lie dormant in harsh conditions for possibly long periods of time. Typically, humans contract anthrax through inhalation of spores (pulmonary type of anthrax, which is more serious) or through an open wound.85–87 Rarely, in developing nations, anthrax is contracted through the ingestion of infected meat.88–90 The infectious dose (ID50 as low as 10,000 spores) makes anthrax highly infectious, and this problem is complicated by stability of the spores. It has been reported that anthrax infection through inhalation occurred after a 70-year-old grave was disturbed. 91 Although many forms of anthrax have been highly lethal (with mortality rates of nearly 90%), antibiotic treatments have reduced the mortality rate to just over 45%.92–95 Although anthrax infections are rare, they may occur at any time due to the stability and prevalence of the spores. Furthermore, initial symptoms mimic a severe flu, with an incubation period of 1–7 days (and a fast doubling time of about 1 h),96–98 possibly causing the patient to not seek vital medical attention. Although there is no risk of transmission between humans,99,100 anthrax can be contracted very easily given the proper conditions, and it has been suggested and studied in the past for biological warfare and bioterrorism purposes.101–104 As recently as 2001, more than 20 people in the United States contracted variant forms of anthrax in what are thought to be rogue acts of terror, suggesting that this can be accomplished on a larger scale. Due to the robust nature of anthrax spores, the lethality of the infection, and the low infectious dose, anthrax should be closely monitored, especially for homeland security applications.
Streptococcus pneumoniae is a Gram-positive, facultative aerobic bacterium that is one of the leading causes of pneumonia, adult meningitis, and septicemia in immune-deficient patients. 105 Many known strains of these bacteria exist and lead to various minor infections such as rhinitis, sinusitis, and conjunctivitis. Although the bacteria reside in the nasopharynx of healthy carriers, the bacteria cause infection when the immune system is compromised, and healthy individuals may experience severe complications such as endocarditis, cellulitis, and even brain abscess.106–109 Even though more than 90 types have been isolated, 10 common serotypes account for more than 60% of the world’s infectious bacterial diseases.110,111 S. pneumoniae (or Strep) bacteria may manifest themselves in the nasopharynx, upper and lower respiratory systems, blood, or cerebral spinal fluid (CSF) of infected individuals, and transmission occurs through direct person-to-person contact or through aerosolized droplets from coughing or sneezing.112–115 Infections are highly transmittable (incubation period of 1–3 days, ID50 hypothesized between 104 and 108 CFUs, and low doubling time of 30 min),116,117 especially to the immune compromised, and are large problems in clinics, hospitals, and crowded spaces.118–120 It is estimated that nearly 500,000 Americans are affected by Strep each year, with 200,000 hospitalizations (primarily from pneumonia) and a mortality rate of nearly 10%. 105 Statistics also suggest that these infections cost the U.S. health care industry more than $1 billion per year. 121 In addition, it is estimated that, worldwide, pneumonia caused by this bacteria kills more than 1.2 million infants per year, especially in nations with a population at higher risk of HIV.122,123 Further compounding costs and complications is the developing prevalence of antibiotic-resistant strains.124,125 To reduce morbidity, mortality, and cost, it would be highly beneficial to detect low levels of Strep bacteria in hospitals and clinics to reduce the risk of transmission to immunocompromised patients.
Complications of Airborne Pathogen Detection
Airborne pathogens are transmitted in a variety of ways, including (1) liquid droplets = aerosols (influenza A, Rubella, TB, and Strep.), (2) dust particles (influenza A, TB, anthrax, and Strep), (3) spores (anthrax), or (4) some combination of the above (influenza A, TB, anthrax, and Strep). Pathogens may be emitted in the air from an infected individual coughing, sneezing, breathing, or even speaking. In addition, some pathogens become aerosolized from other bodily fluids (e.g., blood or fecal matter). Many of these pathogens typically do not travel more than a few feet and may remain on a surface for a given time. 115 However, some of these bacteria and viruses may travel farther in smaller droplet forms, dust particles, or spores, rendering them more dangerous in highly populated areas with common ventilation sources. 126 The size of excreted aerosol particles varies with the mode of expulsion: 3.5–5 µm for speech, 127 0.3–20 µm for breathing, 1–40 µm for coughing, and 2–16 µm for sneezing. 128
To capture and analyze particulate matter for airborne pathogen detection, a variety of aerosol collectors has been developed. Primarily, these samplers, as shown in Figure 1 ,129–132 operate in two ways: (1) filtration 133 and (2) inertial/ gravitational. 134 Filtration collectors include button air samplers, porous-membrane filters, and particle-to-liquid samplers. Inertial/gravitational collectors include impactors, aerosol centrifuges, and impingers. Several variations of collectors may be implemented into real-time monitoring of bioaerosols, but the method of interfacing these samplers with the sensing platforms is a key to portable detection.
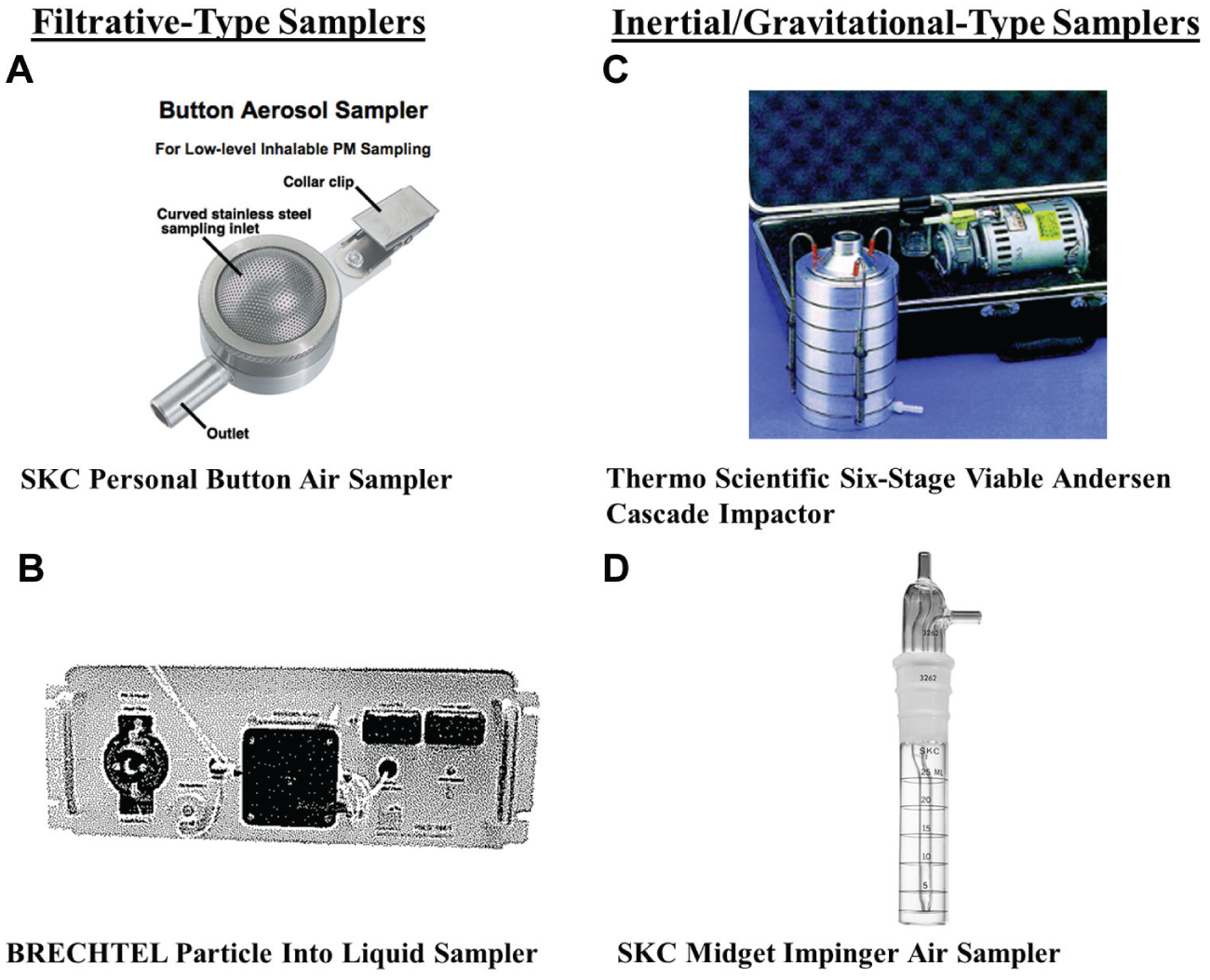
(
The near-real-time detection of infectious agents in air is difficult and is complicated by several factors. Perhaps the most difficult obstacle is the extremely low level of pathogens found in aerosols, making many biosensors insufficient for these applications due to poor sensitivity. For example, conventional PCR, enzyme-linked immunosorbent assay (ELISA), and conventional cell plating may have higher detection limits than the yield of pathogen content from the collected aerosol. In addition, many available air samplers are not very efficient collectors, thus leading to additional loss of captured pathogens. Furthermore, the presence of sample in the aerosol (i.e., saliva, sputum, etc.) may render pathogens difficult to detect. For these reasons, some concentration method is often required prior to the assay itself.
In addition to the problems of sensitivity and very low levels of pathogen present in aerosols, dust particle interference presents a major challenge in biosensing airborne pathogens. Dust particles, which are very polydisperse and heterogeneous in nature, may interfere with the aerosol collection and sensing methods, particularly in optical biosensors. For example, the presence of copious amounts of dust may clog air samplers and restrict pathogen collection from the captured aerosols.135,136 In addition, because dust particles range in size from 1 to 100 µm in diameter, they scatter light at different angles and different intensities, adding another level of complication to optical sensors. 137 Due to the abundance of dust in the air, choosing a proper sampling location is paramount to reduce the risk of interference. Furthermore, depending on the mode of sensing, certain parameters must be optimized to avoid this interference, including location of the air sampler, filter size, and any light-scattering parameters (for optical sensors).
Although the limited quantity of pathogens in aerosol particles, dust particle interference, and poor sensitivity of many sensors are major hindrances, selecting locations to sample air, particularly for stationary sensors, is often difficult. For sufficiently sensitive sensing, air samplers must be positioned to capture the maximum amount of aerosol possible for accurate sampling, but care must be taken to minimize the amount of dust present if at all possible.
Furthermore, the types of aerosol particles dictate where they will appear. For example, larger particles (d > 100 µm) settle, whereas particles significantly smaller than 100 µm will typically travel farther and will more likely carry through ventilation ductwork. 138 Tiny particles (d < 10 µm) often remain suspended in the air, may carry sizeable distances, and are the most likely to fuse with cells in host membranes, 139 rendering them the primary aerosol particles of concern. Furthermore, because human expiration creates aerosol droplets well smaller than 100 µm in diameter,127,128 exhaled pathogens in liquid droplets are likely to travel an appreciable distance. Consequently, airborne pathogens vary greatly in size, from hundreds of nanometers (many viruses) 140 to a few microns (many bacteria) 141 to tens of microns (endospores), 142 thus making choice of sampling location very important. Depending on the size of the aerosol particle, enough pathogens may be carried to where few particles are needed to transmit infections to host cells, further proving the necessity of diligent sampling and sensitive sensing. The nature of the pathogen and how it travels through air should be the primary factors in choosing sampling locations.
The final hurdle in airborne pathogen detection is the sample handling and automation. These biosensors should contain two main components: (1) the air-sampling portion and (2) the biosensor. Although both of these components have been fairly well established individually, few studies have joined the parts into a single, comprehensive unit. Currently, air sampling is typically accomplished by capturing particles on a filter or membrane using some type of sampler. From there, the sample is usually loaded into the actual sensor for the detection component, which requires manual labor. Because of the nature of aerosol particles drying on filters, an extra enrichment or resuspension step is required to achieve the liquid (colloidal) forms necessary for the biosensor. Although this process is effective in many cases, it is not practical for real-time detection because of the lack of continuous monitoring. Therefore, a continuous sampling system that automatically loads captured sample into the sensor (nondisposable or multiuse filters or membranes) should be established. In such a scenario, a sampling mechanism would be positioned, constantly collecting aerosol and dust particles. The captured sample would need to be automatically sent, at set time intervals, to a biosensor that would house some method (optical, electrochemical, etc.) to determine if targeted agents were present in the collected sample. Electrical transduction could be used to provide feedback to a remote computer and to possibly create a feedback loop within the entire system.
Current Standards in Airborne Pathogen Detection (as shown in Table 3)
Conventional Cell Culture and Colony Counting
Typically, pathogen detection of airborne substances is accomplished with separate air sampling and sensor steps. For example, Bacillus species and S. pneumoniae can be detected with conventional cell culture and colony counting. However, seldom do these methods include a sampling component, or they may contain an isolated sampling system prior to culturing. Airborne viruses such as influenza A/H1N1 and avian influenza A/H5N1 can be detected using similar methods yet most likely lack a single comprehensive system and require much manual labor. Furthermore, traditional culturing methods typically require >24 h, microscopic techniques, and highly trained personnel, thereby eliminating the primary benefits of a fully automated and rapid system. To counter these problems, Hsiao et al. 143 used CHROMagar culture media for bacterial enrichment to detect airborne antibiotic-resistant Enterococcus species collected using a system of nebulizers and air filters. Although this research successfully detected relevant levels of the bacteria (103 CFUs per mL) in the collected aerosolized samples and reduced total assay time (down to 24 h), the process remained unacceptably slow, and samples still had to be transferred and cultured, thus requiring sample treatment with no automation. To study the impact of relative humidity on aerosol collection and culture of Pantoea agglomerans, Rule et al. 144 investigated the efficiency of two aerosol collection systems. They found that a minimum relative humidity of 15% was necessary for minimal culture efficiency, which is important to consider in dry environments. The sampling process required less than 1 h, but the culture component still required 18 h. Furthermore, the entire system was not automated.
Summary of Studies.
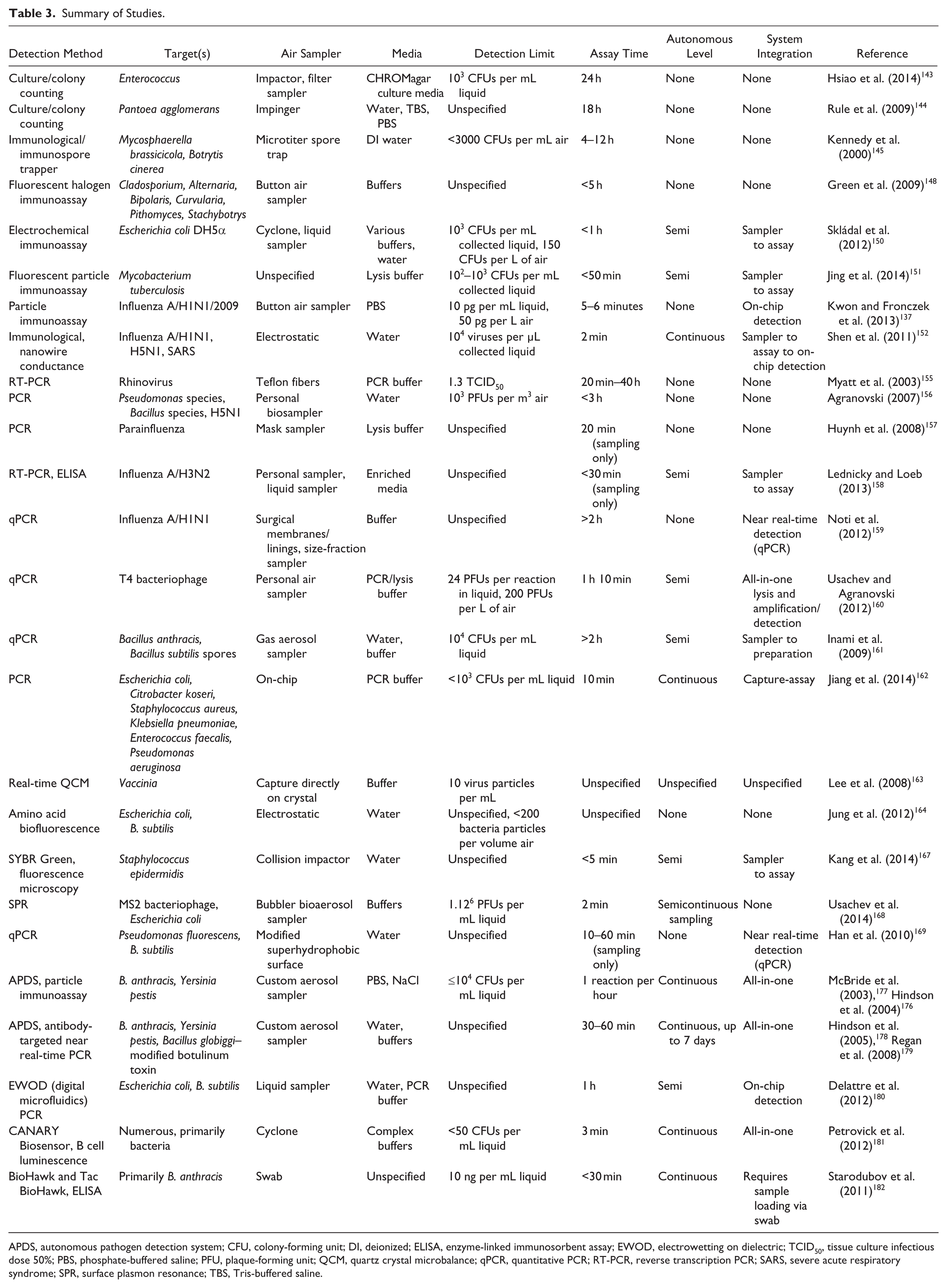
APDS, autonomous pathogen detection system; CFU, colony-forming unit; DI, deionized; ELISA, enzyme-linked immunosorbent assay; EWOD, electrowetting on dielectric; TCID50, tissue culture infectious dose 50%; PBS, phosphate-buffered saline; PFU, plaque-forming unit; QCM, quartz crystal microbalance; qPCR, quantitative PCR; RT-PCR, reverse transcription PCR; SARS, severe acute respiratory syndrome; SPR, surface plasmon resonance; TBS, Tris-buffered saline.
Immunological Detection Methods
Immunological methods ( Fig. 2 ), including radioimmunoassays, memory lymphocyte immunostimulation assay (MELISA), and enzyme-linked immunosorbent assay (ELISA), have been used more recently for such detection with much greater speed, specificity, and sensitivity. However, many of these techniques, especially ELISA, require several rinsing and pipetting steps and separate imaging techniques following sample collection from aerosols. Therefore, a total analysis system (TAS) is needed to create a portable or stationary sampling device. By building on existing technologies, a big push has been made to integrate these systems with bioaerosol samplers for complete detection and analysis. Kennedy et al. 145 demonstrated a portable microtiter immunospore trapping device (MTIS) for capturing Mycosphaerella brassicicola and Botrytis cinerea spores in microwells. Although this system was fairly efficient in capturing spores, was portable, and could be applied to stationary field-testing procedures, sampling was slow (required 4–12 h), shorter sampling times required separate culture enrichments (more than 10 h), and the actual detection component (ELISA) was completely separate, making the entire process insufficient for point-of-care diagnostics. In many cases, particularly when dealing with aerosolized allergens, skin scratch tests are the gold standards of detection.146,147 During the past decade, to reduce detection time and invasiveness, researchers have investigated methods to directly capture allergens from the air. One such method, the halogen immunoassay (HIA), has proven to be quite effective in detecting airborne agents of heterogeneous nature, such as fungal spores. Green et al. 148 developed a novel approach of using HIA via immunostaining to detect several aerosolized fungi. Even though the technique was successful and very specific, the whole process required more than 5 h, detection required microscopy, and the procedure contained several labor-intensive steps between the aerosolization and detection components. The process was later modified to optimize sensitivity through fluorescent HIA (fHIA), and although assay time was also reduced, microscopy steps were still required, and several intermediate steps remained. 149 To improve on assay portability time, Skládal et al. 150 developed an electrochemical immunoassay, coupled with an air sampler, for the detection of a strain of E. coli sometimes present in bioaerosols. This method was fairly sensitive (detection limit = 103 CFUs per mL collected liquid or 150 CFUs per L of air), showed a moderate linear detection range (103–106 CFUs per mL), was fairly rapid (total assay time less than 1 h), and was semiautomated, with a sampler output in a liquid form pumped to the assay platform. However, some pretreatment steps were required, and the device was not handheld. More recently, lab-on-a-chip techniques with implemented capture and immunoassay components have been developed to achieve more rapid and sensitive results. Jing et al. 151 described a novel microfluidic device containing components for the capture of M. tuberculosis, followed by stages of enrichment and immunodetection using secondary fluorescently labeled antibodies attached to antibody-bound microspheres. This method was somewhat rapid (less than 50 min), required little reagent and minimal sample handling, and was fairly specific (detection limit between 102 and 103 CFUs per mL of collected liquid). However, separate enrichment and cell lysis were required prior to the immunoassay, and detection was based on microscopy rather than some automated technique. Kwon and Fronczek et al. described a microfluidic particle immunoassay for the rapid detection of pandemic influenza A H1N1/2009 from captured aerosols, based on Mie scattering immunoagglutination assay using a smartphone. 137 A scaled-down mock classroom was fabricated, and dual fans were included to simulated realistic airflow. Filters were used to capture aerosol particles, which were then loaded onto a microfluidic chip for the immunoassay. Total assay time was just over 5 minutes, and detection was sensitive (detection limit = 10 pg per mL using a smartphone and 50 pg per L air). However, the captured sample was loaded onto the microfluidic chip manually, although the button air sampler included can, in theory, provide a means for total automation. To comply with CDC standards, viral antigen was used and compared to viral complementary DNA (cDNA) (as opposed to whole viruses), but the research does provide useful information about air sampling and viral capture. To take advantage of the specificity of antibody–antigen binding without the laborious washing procedures of ELISA, Shen et al. developed a nearly automated microfluidic device that captured airborne influenza A/H1N1, H3N2, and SARS viruses using antibody-coated silicon nanowires. 152 Unlike many optical detection methods, this sensor directly measured conductance fluctuations of the nanowires due to antibody–antigen binding. This system is nearly automated, requires minimal sampling handling, and is very rapid (total assay time of 2 min) and fairly sensitive (detection limit of 10 4 viruses per µL collected liquid, or about 1–5 pg per µL collected liquid).153,154
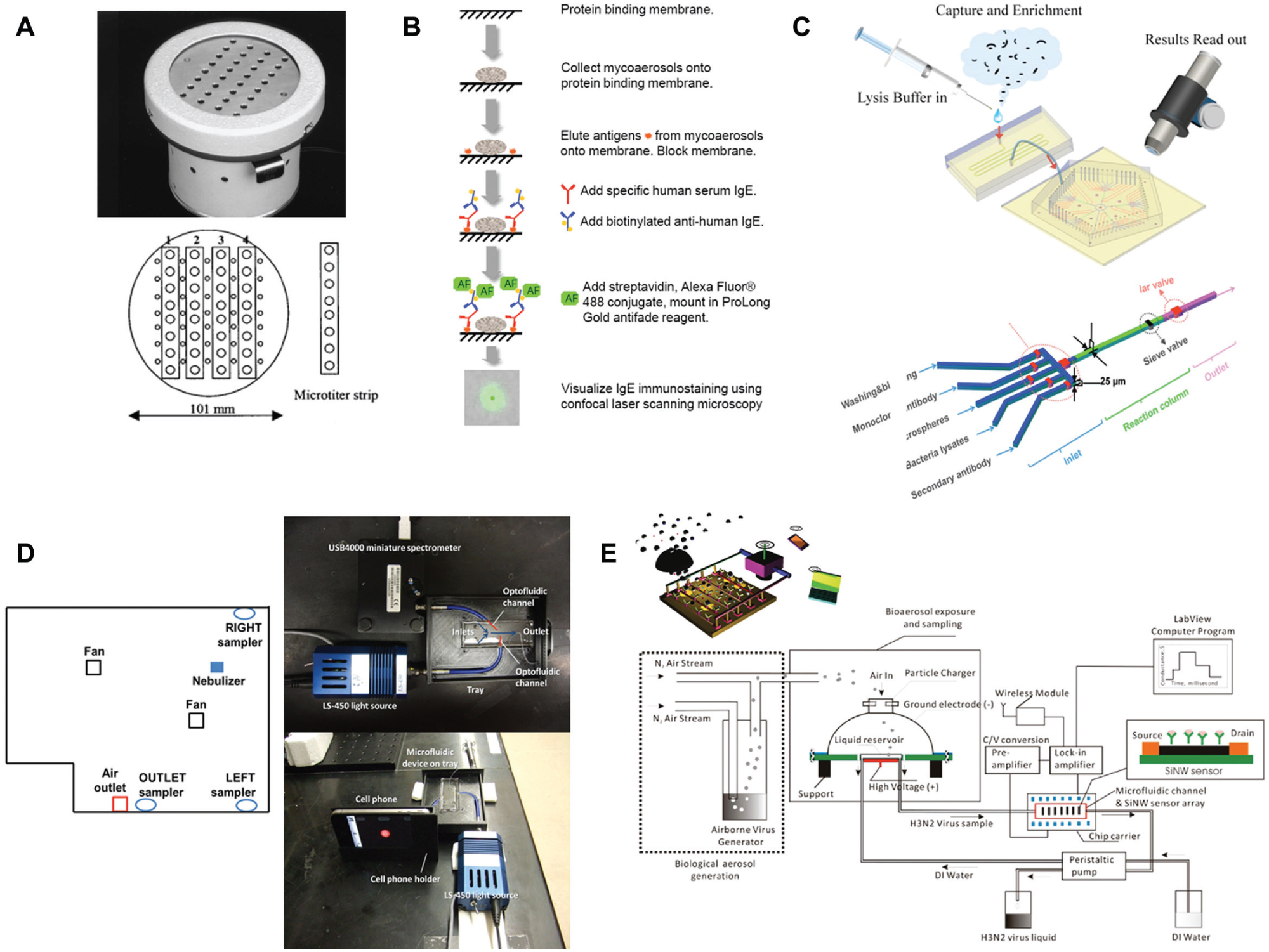
Examples of immunological biosensors. (
PCR and Nucleic Acid Detection
Nucleic acid assays ( Fig. 3 ), in particular PCR and real-time/quantitative PCR (qPCR), have been used to detect airborne pathogens from bioaerosols due to their very high degree of specificity and good sensitivity. Traditionally, PCR is time consuming and laborious and requires nucleic acid extraction and purification, reagent mixing prior to thermocycling, and separate imaging, typically by gel electrophoresis. This whole process may take up to 4 h. Furthermore, in the case of RNA genomes such as most viruses, cDNA templates must be synthesized prior to thermocycling [referred to as reverse transcription PCR (RT-PCR)], adding extra time and sample handling. Although great strides have been made in the PCR such as improvements to nucleic acid extraction kits, primer kits, real-time quantification of PCR amplification, and portable PCR, very little research has demonstrated integrated, rapid, and portable PCR with aerosol sampling and collection. However, several studies have attempted to interface rapid PCR with air sampling to some extent. Myatt et al. developed an RT-PCR biosensor to test aerosolized rhinovirus while attempting to distinguish UV-inactivated viruses from live viruses. 155 Although their detection system was fairly sensitive [detection limit 1.3 TCID50 (tissue culture infectious dose 50%)] and had a reasonably good viral capture limit compared to existing protocols (4–10% capture efficiency), no automation existed, thereby requiring several hands-on steps and thus possibly introducing error. Furthermore, although sampling could range from 20 min to 40 h depending on the desired goal, sample pretreatment was necessary as well as separate imaging through gel electrophoresis. Agranovski described a personal, portable air sampler that efficiently captured various pathogens, including Pseudomonas species, Bacillus species, and influenza A/H5N1. 156 This personal sampler was reported to capture bioaerosols with up to 94% efficiency in little time (ranging from 1 to 12.5 minutes) and was fairly sensitive [detection limit was 103 PFUs (plaque-forming units) per m3 air]. Furthermore, the total assay time, including air sampling, sample preparation, and PCR, required less than 3 h. However, this time is too long for practical in-field detection, and the protocol requires separate sample handling and laboratory diagnostics. Other groups have approached portable bioaerosol capture through simpler methods. Huynh et al. developed a mask designed to capture aerosolized rhinovirus, influenza A virus, and parainfluenza virus from patients following talking, breathing, and coughing. 157 Sampling itself was quite successful (>95% capture efficiency) after 20 min. However, detection was accomplished through conventional RT-PCR in a remote laboratory, adding additional time and labor. However, this approach has potential in clinical settings with coupled portable detection (rapid PCR, immunoassay, etc.) and can be modified to capture contagions or hazardous agents while protecting the tester. Lednicky and Loeb 158 described methods for rapid sampling of influenza A/H3N2 viruses using two samplers: (1) personal air samplers (less than 5 in. in length) and (2) media-containing air samplers (less than 1 ft. in length). Both types of sampler contained a system of valves for size exclusion, thus lending a high degree of control over the size of the particles captured. The media-containing sampler was not as portable as the personal sampler, but the media allowed for the possibility of culture enrichment and/or transfer to assay without separate sample handling. Enhanced RT-PCR and ELISA were used to analyze results. Although this protocol contained no automation and is not rapid (sampling typically <30 min, RT-PCR 2–4 h), it could be configured for total in-field detection. Noti et al. created a mock scaled-down surgical operating room with realistic conditions (notably, ventilation rates and temperature) and analyzed the capture efficiency and filter efficiency of surgical membranes, including surgical masks, respirators, and gloves. 159 The study noted that aerosolized seasonal influenza A/H1N1 virus particles were captured on these membranes and often passed through filters that were not fitted sufficiently, and the results were confirmed by qPCR. Although the total assay was not rapid (1 h collection time, RNA extraction, and more than 1 h for qPCR), this study demonstrates that, when equipped with appropriate on-site detection systems, pathogens can be efficiently collected in aerosol form using economical (nonexpensive) materials. Usachev and Agranovski developed a protocol to monitor T4 bacteriophage and recombinant M13 using personal air samplers interfacing with real-time PCR. 160 This study made an attempt at ease of use toward automation by using specialized PCR reagents requiring no separate nucleic acid extraction or purification. Furthermore, all necessary reagents were loaded into the collection liquid prior to qPCR. The total assay was more rapid (10 min collection time and 1 h qPCR time) and sensitive (detection limit of 24 PFUs per reaction). To solve some of the problems associated with this type of assay, especially time, portability, and ease of use, lab-on-a-chip nucleic acid assays have been developed to interface with aerosol sampling. Inami et al. developed a two-stage microfluidic chip system to capture and treat B. anthracis spores for qPCR. 161 The assay was semiautomated because the microfluidic chip output was bacterial DNA (5% collected from initial aerosol input), and detection required separate qPCR. The total assay time (excluding qPCR) was 2 h, and no separate sample handling was required. The limit of detection was 104 CFUs per mL. This work made major advancements toward portable air sampling with total analysis, which could be achieved through on-chip detection. Jiang et al. created a nearly fully automated continuous flow-through microfluidic apparatus that was capable of bioaerosol capture and detection of six separate aerosolized bacterial targets. 162 Including sampling, lysis, and on-chip nucleic acid amplification, the total assay time was 1 h 10 min, with a detection limit <103 CFUs per mL. Separate gel electrophoresis was required for detection, but existing methods can be implemented into this protocol for (relatively) rapid detection. With the progress made in nucleic acid assay, particularly in real-time PCR, final modifications must be made to bridge aerosolized pathogen capture with amplification and detection, whether it be for stationary rapid detection or portable detection. In addition, steps must be taken for automation to maximize ease of use.
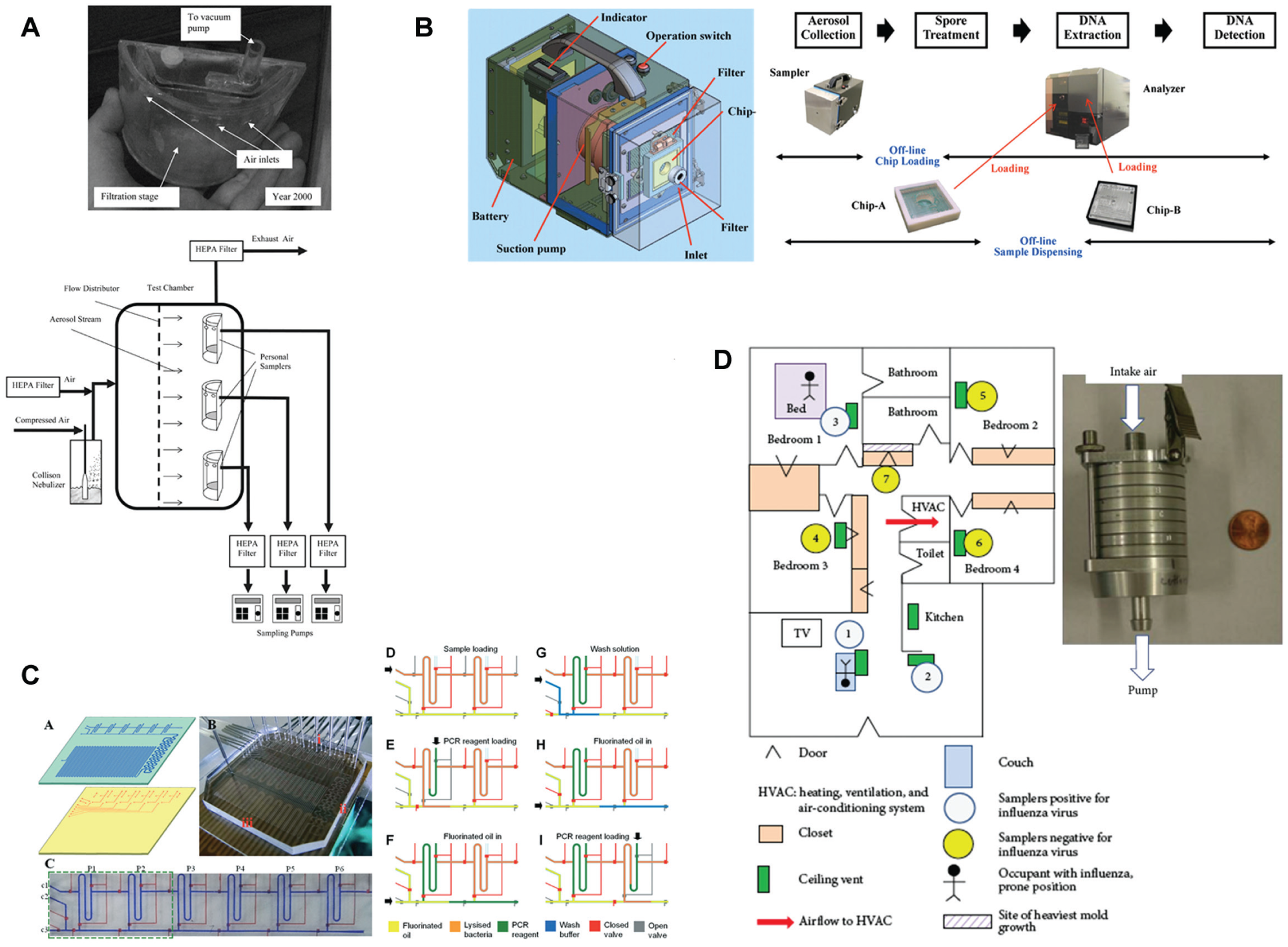
Examples of nucleic acid–based biosensors. (
Other Techniques
Besides conventional sensor techniques, other methods have been attempted for airborne pathogen detection ( Fig. 4 ). Lee et al. 163 created a real-time quartz crystal microbalance (QCM) to detect airborne Vaccinia virus particles. The aerosolized virus capture efficiency was moderately good (about 10%), lending a detection limit of 10 virus particles per milliliter. Although the results of this study were promising, QCM detection presents several problems, among them cost of materials, lack of portability, a larger required sample, and an unstable signal, thus leading to loss of sensitivity.164,165 Jung et al. took advantage of intrinsic amino acid biofluorescence to detect aerosolized E. coli and B. subtilis using a real-time aerosol fluorescence sensor (AFS) with dual fluorescence channels. 166 This protocol was fairly quick and allowed for the detection of <200 captured particles per volume of air, but the study was inconclusive regarding the assay sensitivity and specificity. Kang et al. developed an optical lab-on-a-chip biosensor for the real-time detection of captured aerosolized pathogens using a mini fluorescent microscope and SYBR green intercalating dye. 167 The group reported that the sensor was capable of distinguishing bacterial pathogens (in this case, Staphylococcus epidermidis) from interfering agents collected in aerosols due to the presence of intercalating dye but lacked true specificity (it typically only detected the presence of bacteria). Furthermore, the process, although rapid (total assay time of less than 5 min) and free of sample pretreatment, relies on microscopic techniques. However, with the appropriate modifications, such as image processing and microcontroller programming, this method can be made specific and more automated. Other groups have reported surface plasmon resonance (SPR)-based methods for the detection of airborne viruses, 168 but this technique is often laborious and expensive, and forces a tradeoff between time and sensitivity. Han et al. modified superhydrophobic surfaces to capture airborne Pseudomonas species and B. subtilis, and used an electrostatic precipitator to collect and concentrate liquid droplets for enhanced detection. 169 For these experiments, the concentrated droplets were prepared for qPCR, but this protocol is, in theory, suitable to be interfaced with more rapid and automated methods. Capture and concentration times ranged from 10 to 60 minutes, with capture efficiency ranging from 22% to 47%. Several groups have used mass spectroscopy techniques to study detection viability in various airborne pathogens.170–172 Although mass spectroscopy has been shown to be capable of such detection, several shortcomings have been noted;173–175 primarily, specificity is limited to clean samples because the spectra themselves do not necessarily provide succinct information on pathogen type. In addition, for larger pathogens (e.g., bacteria), consistent detection of multiple pathogens is possible only if their concentrations are roughly equivalent, rendering the method impractical for true pathogen detection. Finally, the protocols are expensive and require highly trained personnel. If these drawbacks could be overcome, automated detection would be a major hurdle due to the fact that, because of the experimental variation in spectra, a coherent algorithm would be nearly impossible to develop. Other methods have also been proposed and developed, ranging from lab-on-a-chip to flow cytometry technologies, and although most have not been optimized for portable airborne pathogen detection, modifications, in some cases only minor adjustments, can be made to create a more complete and desirable system.
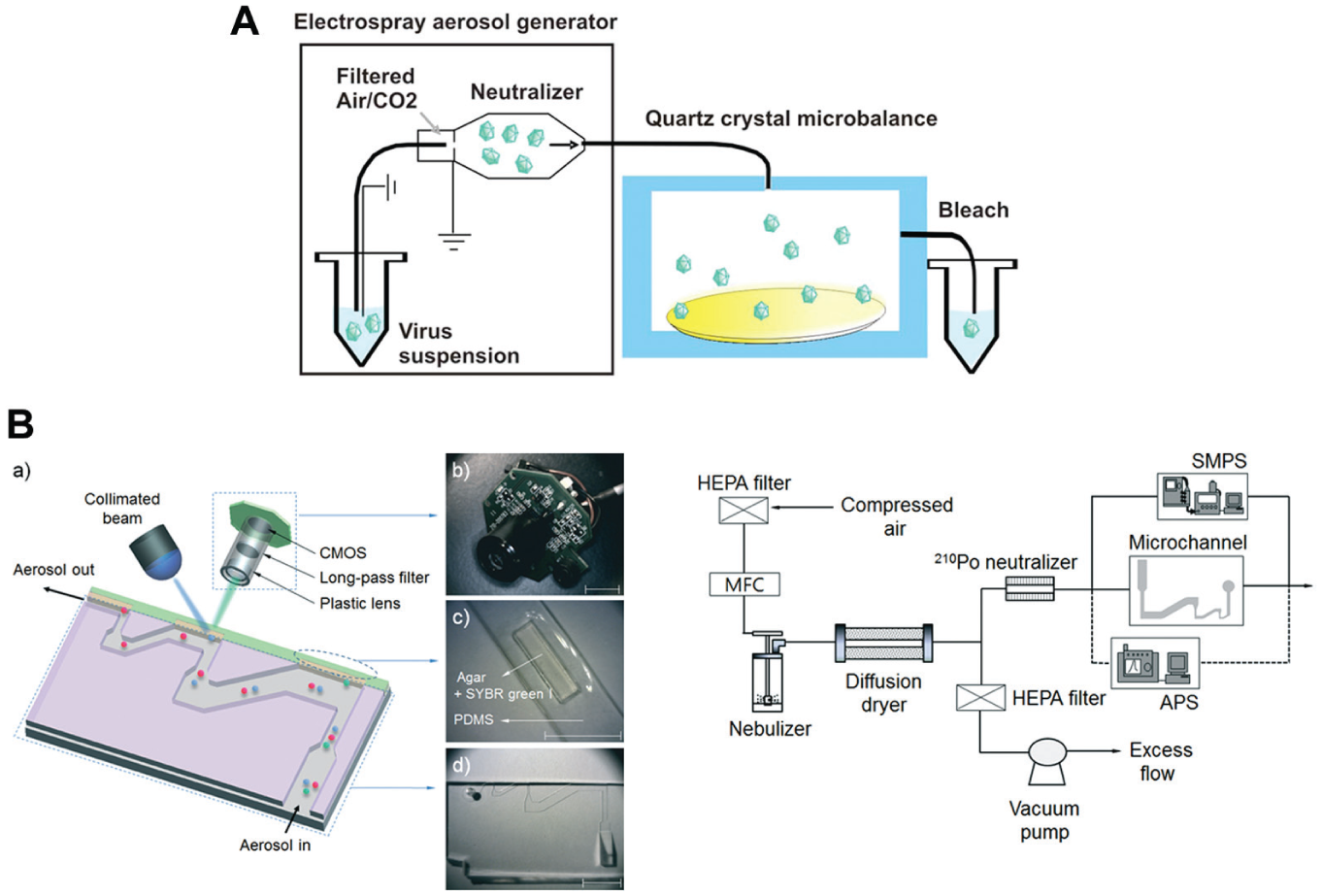
Other types of biosensors. (
Steps toward a Fully Automated Pathogen Detection System
Automated Pathogen Detection Systems for Academic Research
Although a vast majority of the described systems are not automated and contain discontinuous steps in the process chain, thereby introducing user error, methods exist that are nearly completely automated ( Fig. 5 ). Hindson et al. developed an automated system based on sequential injection analysis (SIA) that was capable of detecting aerosolized B. anthracis and Yersinia pestis.176,177 The entire system, which contained channels for aerosol capture, pathogen selection with antibody targeting, and fluorescent detection of labeled antibodies, is termed an autonomous pathogen detection system (APDS). Although the first generation of this system had high throughput and did not require separate user steps, it was not particularly rapid (one immunoassay result per hour) or very sensitive (≤104 CFUs per mL detection limit), and at times it lacked sensitivity due to nonspecific agglutination. In addition, APDS was bulky and only portable in an automobile. In response to these issues, Hindson et al.178,179 further built on APDS to simultaneously detect aerosolized B. anthracis, Y. pestis, Bacillus globiggi, and a modified botulinum toxin using a multiplexed antibody-based assay, with PCR confirmation. This second-generation protocol not only improved the assay time (30–60 min) but also decreased the number of false negatives by incorporating a system of controls and checks in the automation, including a full PCR control check run every 12 h. Furthermore, the system was established to be capable of running continuously for 7 days, allowing for continuous monitoring at locations such as medical clinics, airports, subways, and other areas at high risk for terrorist activity. However, this system has not been optimized for portable detection. In an attempt to create a miniaturized system, Delattre et al. created a lab-on-a-chip device for the simultaneous capture and detection of E. coli and B. subtilis using electrowetting-on-dielectric (EWOD) digital microfluidics for PCR. 180 This very nearly continuous protocol is capable of sorting and preparing aerosolized bacteria, using magnetic beads, for PCR and detection. The group did not report on the sensitivity of the system (which detected 104 CFUs per mL bacteria), but did state that one set of PCR data was reported every hour and that minimal sample input was required. However, the group did report that the aerosol capture component of the system needs to be fully integrated with the digital microfluidic portion. Also, although some reagents are preloaded and dried within the channels, other reagents must be replenished every so often. However, the platform is small, somewhat portable (25 kg total weight), and highly specific.
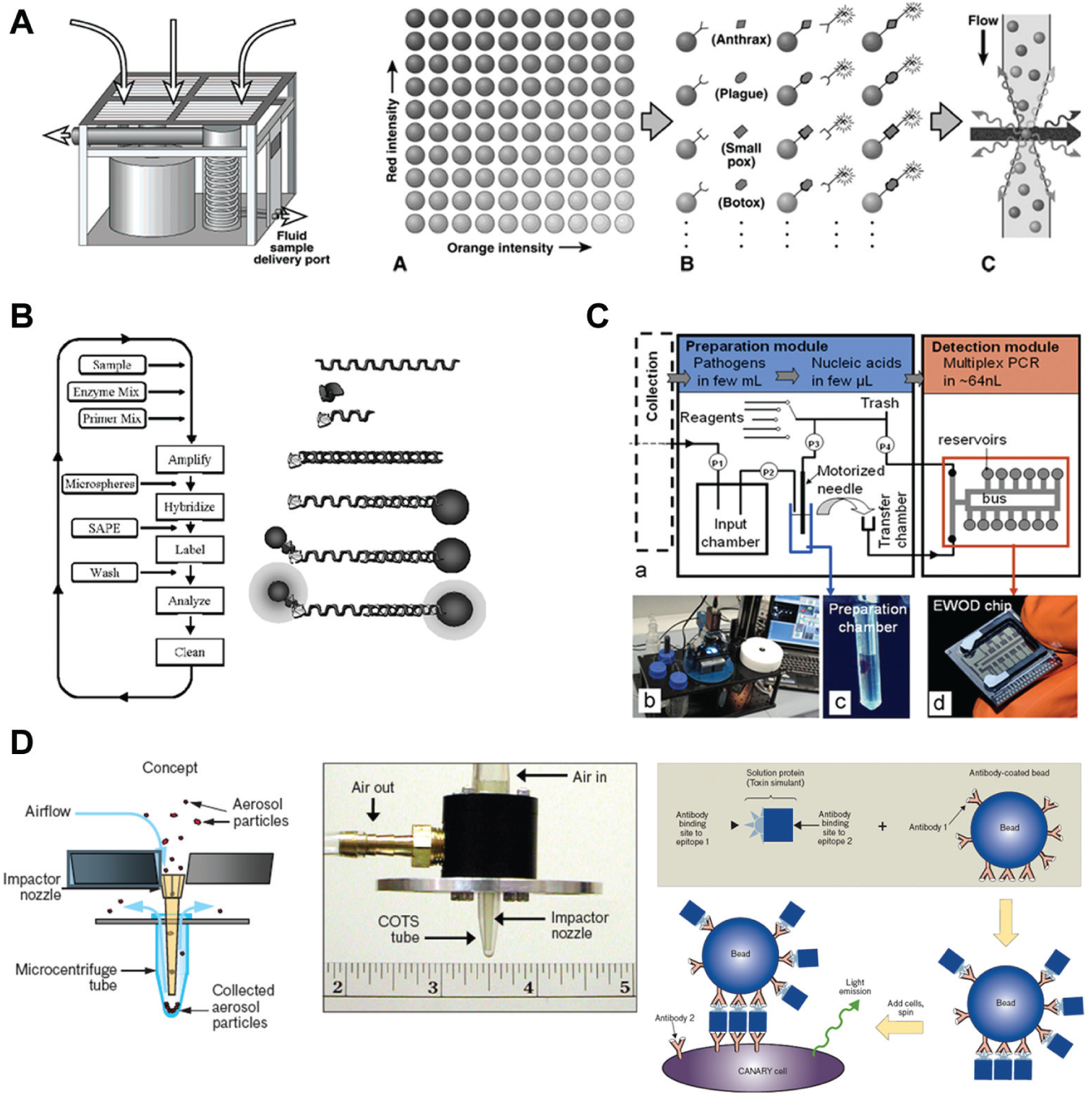
Fully automated biosensors. (
Automated Pathogen Detection Systems for Governmental and Commercial Uses
More recently, several companies have created APDSs for either commercial sales or use in military or homeland security applications ( Fig. 5 ). These platforms, including the CANARY Biosensor, 181 the BioHawk and Tac BioHawk, 182 and the Battelle Resource Effective Bio-Identification System (REBS), 183 vary in size, speed, portability, capture method, multiplexing, bioanalyte, and detection mode. Although these systems have set the tone for the future of counterterrorism biosensing technology, none of the existing systems, to our knowledge, can be modified easily to be used for portable detection or stationary detection (i.e., they are not modular enough to use). Furthermore, these systems are not cheap enough for most private individuals to own and are not currently interfaced with technology (e.g., smartphones) to be advantageous in field and clinical settings worldwide.
What Is Needed
With the advancements in airborne pathogen point-of-care diagnostics and field detection, more advancements are needed to create a complete, sensitive, and low-cost system that can be implemented into a stationary real-time monitoring system or personal sensor that can provide feedback to an individual anywhere in the world. For such a system to be created, it is necessary to optimize packaging, specificity, sensitivity, and feedback methods without sacrificing the detection limit or creating false positives and negatives. In providing feedback to provide the output levels of specific pathogens and verify the presence of false negatives, this new system could encompass multiple assay types (e.g., real-time PCR/nucleic acid detection and multiplexed immunoassays) in parallel to verify results and specificity. In addition, the system should provide some method for interfacing with standard laboratory benchtop analytical methods (e.g., exit valve with prepared sample for a standard agar culture) to verify the absence of false positives.
A Portable, Rapid, and Sensitive Multiplexed Sensor for Bioaerosol Sensing to Interface with a Smartphone
In light of the recent developments and the stated necessary improvements, we propose a concept of an autonomous, total analysis system ( Fig. 6 ) based on lab-on-a-chip technology and continuous air sampling. The total system will be (1) relatively inexpensive, (2) portable (small and lightweight), (3) rapid (processing n number of samples, yielding results in under 10 min), (4) sensitive, (5) multiplexed and capable of detecting multiple pathogen types simultaneously, (6) capable of interfacing with a smartphone, (7) capable of providing feedback, and (8) completely automated. Movement between assay segments will be driven by microfluidics. The device will contain several major segments: (1) sample loading with a membrane for aerosol capture; (2) sample preparation, including lysis, nucleic acid extraction and separation, and necessary buffer mixing; (3) analyte detection and signal augmentation, possibly in parallel; (4) detection to interface with a smartphone and/or some other detection apparatus; (5) multiple parallel channels for feedback; (6) programmed smartphone application to control lights and data collection; (7) a system of optical filters and fiber-optics to guide light appropriately; and (8) exit valves for sample outlet to interface with various other lab assays. This device will be intended for single use but capable of multiple readings in a single trail. To make this possible, components will have to be chosen carefully to minimize manufacturing costs. To make this prototype configurable with a more stationary apparatus, the device will be designed as a single cartridge, so that each new cartridge can be changed or replaced in a stationary system. Following collection, aerosol particles will be suspended in water, and microfluidic action will be driven by capillary action. At the entrance of the sample preparation stage, the channel will split in two: one channel for nucleic acid extraction and the other for particle-conjugated antibody capture. Nucleic acid extraction will be accomplished through paper microfluidics, 184 during which the nucleic acid will be tagged with preloaded fluorophores. The channel will split into several subchannels of preloaded nucleotides specific for a range of targets of interest (different targets in different channels), and a process of positive selection will be used to sort specific pathogens. Fluorescence excitation light will be incorporated at this point to excite fluorophores; fiber-optics will provide signal to the phone’s complementary metal-oxide semiconductor (CMOS). A separate negative control sample will be kept to reduce or eliminate background noise. Due to the presence of an exit valve, these pre-prepared samples can be off-loaded for conventional assays such as simple culture and PCR.
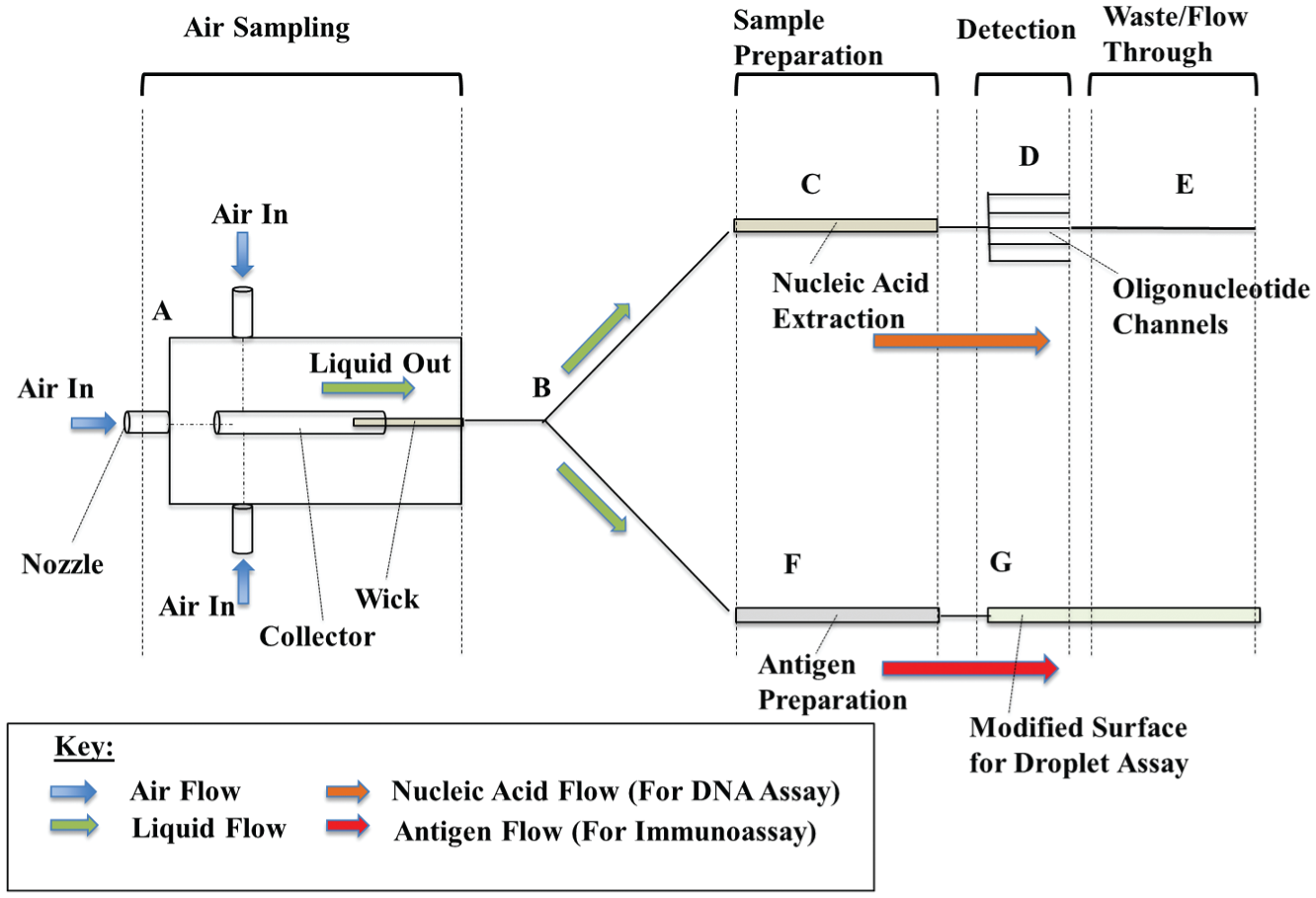
Proposed automated biosensor (not drawn to scale). Air samples will be collected using a modified miniature impinger sampler with multiple nozzles. The particles will be collected into liquid media and then filtered through paper microfluidics for pretreatment (
For the other detection channel, the traveling sample will diverge into several smaller channels and then encounter preloaded surfactant and antibody-conjugated particles of varying diameters and fluorescent properties, similar to those described in Nicolini et al. 2 The mixture will halt at specially fabricated surfaces that will cause them to bead up into highly defined droplets. At this point, the immunoassay will occur. A single source with necessary filters will be used to excite the various particles, causing multiple fluorescent shifts. Each type of particle will be conjugated with a different antibody to an antigen of interest, allowing for true dual detection. Mie scattering parameters will be established for signal readout. With proper orientation, a single smartphone can be implemented to simultaneously measure fluorescence from the PCR portion and immunoaggluttination scatter from the other. A single smartphone, including a flash light-emitting diode (LED) and CMOS camera, with a specialized application will be used to control the whole sensor.
Obviously, what we proposed in Figure 6 is just one example from multiple possibilities. It should be noted that, given the vast array of biosensor designs and their modularity, numerous detection methods may be implemented for this type of sensor, including rapid cell culture, bioluminescence, spectral imaging, ellipsometry, and electrochemical transduction. Although there are many possibilities, cost, ease of use, and level of specificity and sensitivity must be heavily considered.
Potential Problems versus Benefits
Although the sensor will be limited by the limitations of the existing technology (e.g., the weakness of fluorescence, potential lack of specificity of oligonucleotides, antibody cross-reactivity, limitations of CMOS, and shelf life of preloaded reagents), this schematic offers the specificity and sensitivity of technology previously reported with the added advantages of true multiplexing, incorporated negative controls, and true feedback using parallel assays. In addition, the whole device will be small, light, portable, disposable, and cheap (manufacturing cost <$10.00 per device).
Conclusions
Although much progress in the realm of biosensors has been made throughout the years, particularly in the areas of point-of-care diagnostics, environmental monitoring, and food safety, there is still much room for improvement when dealing with the rapid detection of airborne pathogens. Advancements in sensor technology as well as in aerosol collectors have enabled research into true bioaerosol sampling for continuous, automated biosensors. With more scientific breakthroughs and discoveries of drug-resistant pathogen strains and the looming threat of bioterrorism that may reach hospitals, schools, airports, and food supplies, there is a growing need for more portable, rapid, and sensitive biosensing technology. Despite numerous studies interfacing at least some air sampling with biosensor components, very few truly entail developed, comprehensive systems capable of point-of-care diagnostics or environmental monitoring. Furthermore, the systems that do encompass these features are typically large, not very rapid, expensive, and not accessible to private individuals, small clinics, or underdeveloped nations. The next major step in the process is the development of portable, disposable, and cost-effective biosensors that are more feasible for in-field use. With our proposed sensor, not only does the operator gain the sensitivity, specificity, and ease of use associated with the stationary sensors developed by the federal government, but they also obtain a method for ensuring that their specific environment is free from unwanted agents. On a larger scale, this type of comprehensive sensor can be implemented for soldiers overseas, in clinics, in counterterrorist applications, and for health care workers in the third world. Thus, this device, along with any improvements, has the potential to save lives, reduce morbidity, and save health care industries untold amounts of money.
Footnotes
Acknowledgements
The authors would like to express thanks to Ms. Ariana M. Nicolini for her kind help and support with this manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Funding for this research was provided by Seoul VioSys.
